Abstract
Functions of microRNAs have been characterized in the embryologic, physiologic, and oncogenic processes, but the role of microRNAs in mediating tumor-specific organ metastasis was addressed only recently and still absent in gastric cancer peritoneal metastasis. Here, we used the microarray analysis to define the gastric cancer peritoneal metastasis–related microRNAs from highly peritoneal metastatic derivatives (GC-9811P cells) and the parental GC-9811 human gastric cancer cells. MiR-136 was found to be decreased in all peritoneal metastatic sublines when compared with that in the parental line. We further confirmed that miR-136 expression is frequently downregulated in gastric cancer peritoneal metastasis cells and tissues and its low expression is significantly associated with more peritoneal metastasis and worse prognosis. Moreover, restoring the expression of miR-136 could inhibit gastric cancer peritoneal metastasis in vitro and in vivo. Subsequent investigation characterized HOXC10 as a direct target of miR-136. In addition, knockdown of HOXC10 reduced GC-9811P cell migration and invasion, similar to the phenotype observed with miR-136 restoration in these cells, indicating that HOXC10 functions as a metastasis promoter in gastric cancer peritoneal metastasis. Upregulation of HOXC10 in parental GC-9811 cells resulted in a dramatic reduction of in vitro migration, invasion, and in vivo peritoneal metastasis. Furthermore, our results showed that ectopic expression of HOXC10 could reverse inhibition of metastasis by overexpressed miR-136 in GC-9811P cells. Our findings provide new insights into the role of miR-136 in the gastric cancer–specific peritoneal metastasis and implicate the potential application of miR-136 in gastric cancer peritoneal metastasis therapy.
Introduction
Metastasis, the spread and growth of tumor cells to distant organs, represents the most devastating attribute of cancer. A notable feature of this process is the variation in metastatic tissue tropism displayed by different types of cancer.1,2 In the case of gastric cancer, peritoneal metastasis is the most frequent pattern of recurrence in patients with gastric carcinoma.3,4 About half of patients with serosa-invasive gastric carcinoma develop peritoneal recurrence and die of this disease during the first 2 years, even if curative resection is performed.5,6 The 5-year survival rate of patients with positive peritoneal lavage cytology is only 2%. 7 An incomplete understanding of the molecular and cellular mechanisms underlying peritoneal metastasis hinders the development of effective therapies that would eliminate or ameliorate this condition.
MicroRNAs (miRNAs) are a class of small non-coding RNAs involved in a wide range of basic processes such as cell proliferation, development, apoptosis, and stress response. Recent reports even suggest that the expression profiling of miRNAs may be a more accurate method of classifying cancer subtype than using the expression profiles of protein-coding genes.8–14 For these reasons, we undertook to associate miRNAs with peritoneal metastasis of gastric cancer, with the hope that such associations might provide adequate information about the specific dynamics of tumor burden in these subjects during the course of their disease.
In this study, we have identified miR-136 that mediates gastric cancer metastasis to peritoneal and is clinically correlated with the development of peritoneal metastasis when expressed in primary gastric cancers. By means of in vitro and in vivo selection, microarray analysis, functional verification, and clinical validation, here we identify the biological functions and underlying molecular mechanism of miR-136 in gastric cancer–specific peritoneal metastasis. Restoring miR-136 could inhibit gastric cancer cell invasion and migration in vitro and peritoneal metastasis in vivo. Mechanistically, the inhibition of gastric cancer peritoneal metastasis of miR-136 can be explained to inhibit homeobox C10 (HOXC10). We also confirmed that gastric cancer patients with low expression of miR-136 were correlated with more peritoneal metastasis and deaths. Taken together, our findings indicate that miRNAs are involved in tumor-specific organ metastasis, and miR-136, which inhibits HOXC10, could be used as potential targets for gastric cancer peritoneal metastasis.
Materials and methods
Cell lines and cultures
Retroviral packaging cell line (HEK293T) and human gastric adenocarcinoma cells (MKN-45) were previously purchased from the American Type Culture Collection and preserved in our lab. GC-9811 cell line (also named XGC-9811, gastric cancer cell line with weakly peritoneal and liver metastatic potency) was established from tumor cells isolated from a human malignant ascites sample in our institute. 15 GC-9811P cell line and XGC-9811L cell line were derived from GC-9811 and established in our institute. GC-9811P cells can specific metastasis to peritoneal and XGC-9811L cells can specific metastasis to liver.15,16 All cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) containing 10% fetal bovine serum (FBS) supplemented with 100 U/mL penicillin and 100 mg/mL streptomycin at 37°C with 5% CO2. All transfections were done using Lipofectamine 2000 (Invitrogen). For bioluminescent tracking, cell lines were retrovirally infected with a triple-fusion protein reporter construct encoding herpes simplex virus thymidine kinase 1, green fluorescent protein (GFP), and firefly luciferase.17,18 GFP-positive cells were enriched by fluorescence-activated cell sorting.
RNA isolation and miRNA microarray hybridization
Total RNA from cultured cells, with efficient recovery of small RNAs, was isolated using the mirVana miRNA Isolation Kit (Ambion). RNA labeling and hybridization on miRNA microarray chips were performed as previously described. 14
RNA extraction and quantitative real-time polymerase chain reaction
Stem-loop reverse transcription (RT) for mature miRNA was done as described previously. 19 All reagents were obtained from Applied Biosystems. For miRNA detection, mature miR-136 was reverse-transcribed with specific RT primers, quantified with a TaqMan probe. The small nuclear U6 RNA was used as an internal control for normalization and quantification of the miRNA expression as previously reported. 19
Generation of retrovirus, miRNA expressing cells, and knockdown cells
The plasmid vectors encoding the miR-136 were constructed as previously described. 20 MiR-136 sponge and a control sponge containing tandem non-targeting binding sites were constructed as previously described by Ebert et al. 19 The production of viruses and infection of indicated cells were described previously. 21
Migration and invasion assay
For Transwell migration assays, 1 × 105 cells were plated in the top chamber with the non-coated membrane (24-well insert; 8-mm pore size, Corning Costar Corp). For invasion assays, 2 × 105 cells were plated in the top chamber with Matrigel-coated membrane (24-well insert; 8-mm pore size; Corning Costar Corp). Both assays were performed as pervious reported. 20
Luciferase reporter assay
Cells of 50% confluence in 24-well plates were transfected. Firefly luciferase reporter gene construct (200 ng) and 1 ng of the pRL-SV40 Renilla luciferase construct (for normalization) were cotransfected per well. Luciferase activity was measured 48 h after transfection using the dual-luciferase reporter assay system (Promega).
Western blot analysis
Cells were washed twice with Hank’s balanced salt solution and lysed directly in radioimmunoprecipitation assay (RIPA) buffer (50 mM Tris-HCl (pH 7.4), 1% (v/v) Triton X-100, 1 mM ethylenediaminetetraacetic acid (EDTA), 1 mM leupeptin, 1 mM phenylmethylsulfonyl fluoride, 10 mM NaF, 1 mM Na3VO4). The lysates were centrifuged at 14,000 rpm for 30 min at 4°C and the supernatants were collected. Cell lysate (60 µg) was separated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis, blotted onto nitrocellulose membrane, and incubated with a primary antibody: anti-HOXC10 (1:1000 dilution; catalog number: WH0003226M1; Sigma) or anti-β-actin (1:5000 dilution; catalog number: A5441; Sigma). After repeated washing, the membranes were incubated with horseradish-peroxidase-conjugated anti-mouse secondary antibody (1:2000 dilution; catalog number: sc-2005; Santa Cruz Biotechnology). The bands were visualized using the enhanced chemiluminescence (ECL) system (Amersham Pharmacia Biotech).
Animal studies
Briefly, 6 to 8-week-old BALB/C nude mice obtained from Shanghai Laboratory Animal Center of the Chinese Academy of Sciences were used for tumor growth, peritoneal colonization, and peritoneal metastasis xenograft assays. Primary tumor growth assays were performed as previously described. 15 Mice were anaesthetized and injected retro-orbitally with 1.5 mg of D-luciferin (15 mg/mL in phosphate-buffered saline (PBS)) and were imaged for luciferase activity immediately after injection to exclude any that were not successfully xenografted. Imaging was completed between 2 and 5 min after injection with a Xenogen IVIS system coupled to Living Image acquisition and analysis software (Xenogen). For bioluminescence imaging (BLI) plots, photon flux was calculated for each mouse using a rectangular region of interest encompassing the thorax of the mouse in a prone position. This value was scaled to a comparable background value (from a luciferin-injected mouse with no tumor cells) and then normalized to the value obtained immediately after xenografting (day 0), so that all mice had an arbitrary starting BLI signal of 100. Peritoneal metastatic colonization was monitored and quantified using non-invasive bioluminescence of normalized hindlimb photon flux.
Clinical sample validation
Formalin-fixed paraffin-embedded specimens of gastric cancer and information about the primary tumors and metastatic foci from the same patients in a sizable cohort of patients were collected from Department of General Gastrointestinal Surgery in Xijing Hospital and Tangdu Hospital, Xi’an, China. These patients were diagnosed and treated at Xijing Hospital and Tangdu Hospital from 1993 to 2000. All the patients had a well-documented clinical history and follow-up information and never underwent preoperative chemotherapy and/or radiation therapy. All of them had undergone gastrectomy with lymph node dissection and were also observed at two hospitals through the end of 2005. The protocols used in the study were approved by the Hospital’s Protection of Human Subjects Committee. Total RNA, with efficient recovery of small RNAs, was isolated from 20-mm sections from formalin-fixed, paraffin-embedded tissue blocks, using the RecoverAll Total Nucleic Acid Isolation Kit (Ambion).
Statistical analysis
All statistical analyses were conducted using SPSS version 11.05 software (SPSS). Student’s t-test was used to investigate the significance of the difference between the covariates. The Kaplan–Meier method was used to calculate the patients’ survival durations. Survival durations in the patient groups were calculated with the log-rank test; p
Results
Establishing a gastric cancer peritoneal metastasis miRNA signature
To identify patterns of miRNA expression associated with aggressive peritoneal metastatic behavior, we performed array-based miRNA profile of parental GC-9811 human gastric cancer cells and four derivative subpopulations 15 (GC-9811P1, GC-9811P2, GC-9811P3, and GC-9811P cell lines) with different peritoneal metastatic potentials in order to identify miRNAs that differ in their expression between the weakly and the highly peritoneal metastatic groups. The miRNA list obtained from a class comparison between parental and GC-9811P populations was filtered to exclude miRNAs that were expressed at low levels in a majority of samples and to ensure a three-fold or higher change in expression level between the two groups. A total of 27 miRNAs (out of 534 human miRNAs assayed) met these criteria and were differentially expressed in either one or more of the highly peritoneal metastatic sub-cell lines. Hierarchical clustering with the 27-miRNA list confirmed a robust relationship between this miRNA expression signature and the peritoneal metastatic activity of cell populations selected in vivo (Figure 1(a)). Out of a total of 27 selected miRNAs, 2 (miR-181b and miR-223) were found overexpressed and 1 (miR-136) was found decreased in all peritoneal metastatic sublines when compared with their expression in the parental line (Figure 1(a)).

MicroRNA-expression signature associated with peritoneal metastasis. (a) Hierarchical clustering of normalized miRNA expression levels of peritoneal metastatic GC-9811P populations with parental GC-9811 cells identifies 27 miRNAs that are significantly correlated with peritoneal metastatic activity. The red underlines indicate the specific three miRNAs that are differentially expressed across all peritoneal metastatic derivatives. (b) Solution hybridization detection of mature miRNA expression in GC-9811P sublines and GC-9811 cells. rRNA (5S) served as a loading control. (c) Real-time RT-PCR analysis of precursor miRNA expression in all peritoneal metastatic sublines and their parental lines. U6 served as a loading control (N = 3; error bars represent SEM). The p values are obtained using a one-sided Student’s t-test.
Furthermore, we used the solution hybridization detection method for mature miRNAs and real-time reverse transcription polymerase chain reaction (rt RT-PCR) analysis for precursor miRNAs to validate the results from microarray analysis. Our results showed that the mature and precursor forms of miR-181b and miR-223 were overexpressed in all peritoneal metastatic sublines when compared with their expression in the parental line; in contrast, miR-136 was decreased in all peritoneal metastatic sublines (Figure 1(b) and (c)). From the above results, we found that only miR-136 was decreased in all peritoneal metastatic sublines; therefore, we chose the moR-136 for our further studies.
MiR-136 inhibits gastric cancer cell peritoneal metastasis in vitro and in vivo
To determine whether miR-136 has a causal function in peritoneal metastasis, we used retrovirus of expressing miR-136 or its individual miRNA sponge to alter miR-136 expression level in GC-9811P cells and parental GC-9811 cells. We first used the miR-136 expressing retrovirus to infect GC-9811P cells. The miR-136 expression level was confirmed by rt RT-PCR. Our results showed that miR-136 expressing retrovirus could significantly upregulate the expression of miR-136 (rt RT-PCR; Figure 2(a)) in GC-9811P cells. We then silenced the expression miR-136 with retroviral miRNA sponge 19 in GC-9811 cells. We found that transfection of the miRNA sponge for miR-136 in GC9811 cells inhibited miR-136 function by >3.0-fold (Figure 2(b)).
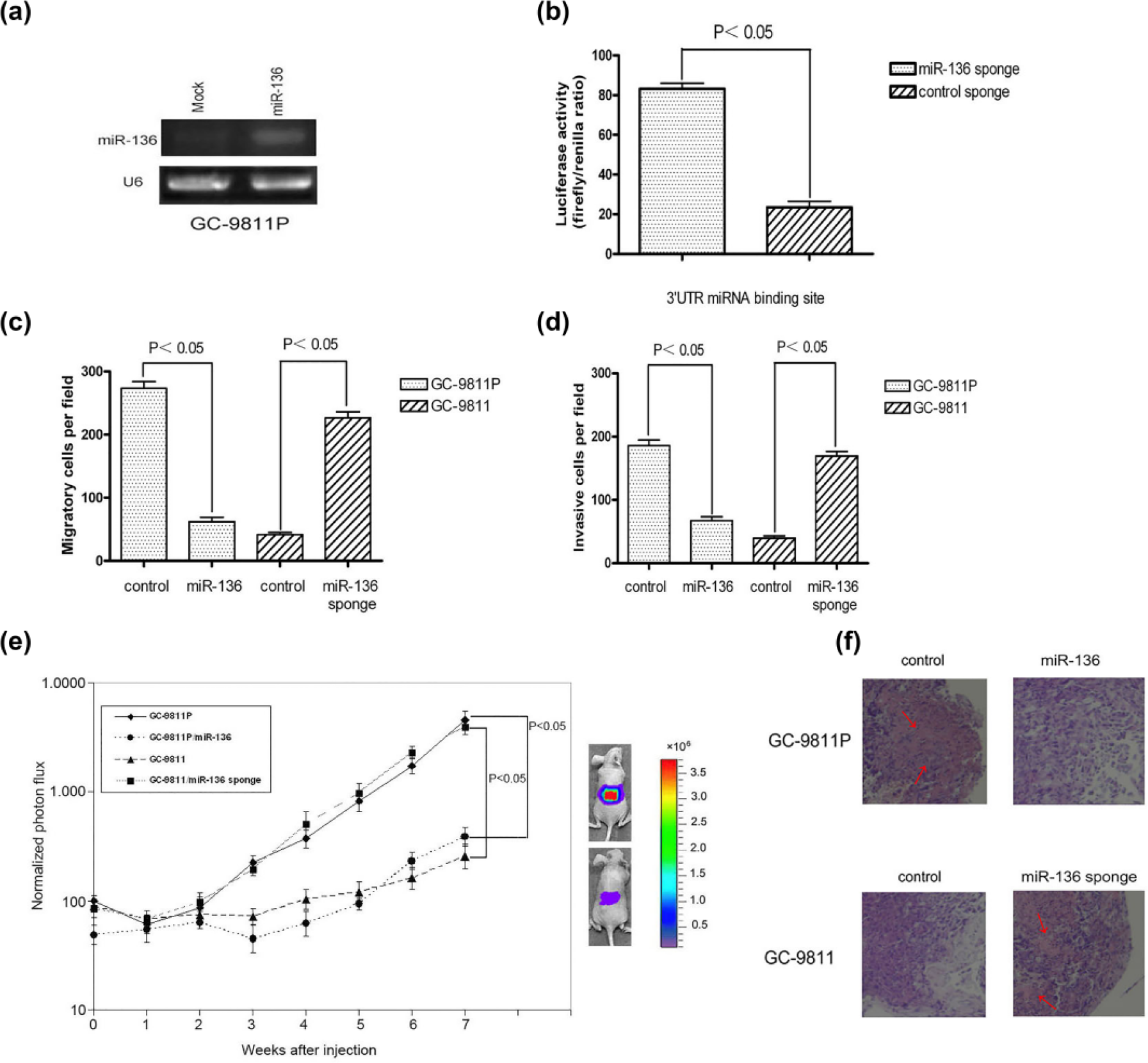
MiR-136 inhibits gastric cancer cell peritoneal metastasis in vitro and in vivo. (a) rt RT-PCR analysis of miR-136 in GC-9811P cells infected with miR-136 expressing retrovirus and control retrovirus. (b) MiRNAs inhibitor function for miR-136 was assayed in GC-9811 cells. Luciferase assays in GC-9811 cells were stable transfection of miR-136 sponge or controls. Then, indicated cells were co-transfected with a Renilla luciferase reporter containing an indicated miRNA motif in its 3′-UTR and a control firefly luciferase reporter; 24 h after transfection with the reporters, cell lysates were harvested and luciferase activity was quantitated (N = 3; error bars indicate SEM). The p values are based on a one-sided Student’s t-test. “+” labels mean cells infected of miRNAs retroviral sponges and “−” labels mean cells infected of miRNA control retroviral sponges. (c) Transwell migration assay and (d) Matrigel invasion assay of GC-9811P cells or GC-9811 cells transfected with miR-136 sponge or miR-136 expressing retrovirus (N = 3; error bars represent SEM). The p values are obtained using a one-sided Student’s t-test. (e) Bioluminescence imaging of peritoneal metastasis by GC-9811P cells with restored expression of miR-136 and GC-9811 cells with knockdown of miR-136. Cells were inoculated and injected into immunodeficient mice. Shown are representative mice corresponding to the GC-9811P set (top) and the GC-9811 set (bottom) at week 7. Peritoneal colonization was measured by bioluminescence and quantified. (N = 8–10; error bars represent SEM; p values based on a one-sided Student’s t-test.) (f) Representative whole-field magnification images of hematoxylin- and eosin-stained extract from peritoneum in representative mice 7 weeks after injection of indicated cancer cells. Red arrows in those four pictures indicate the specific metastasis in peritoneal tissues.
To verify the hypothesis that miR-136 may inhibit the peritoneal metastasis ability of GC9811-P cells and specifically confers the peritoneal metastatic ability of GC9811 cells, metastatic activity was assayed through in vitro and in vivo experiments. Cell proliferation, survival, and migration are among the common functions required by tumor cells for metastatic progression in target microenvironments. Following, we examined the cell proliferation rates in miR-136-restored GC-9811P cells and found that restoring the expression of miR-136 did not alter the proliferation or apoptotic rates of GC9811P cells in vitro or in vivo (data not shown), suggesting that miR-136 function may be required for motility but not for viability of these metastatic cells. Transwell migration assay and Matrigel invasion assay showed that restoring miR-136 led to significant reduction in the motility and invasive properties of GC-9811P cells (Figure 2(c) and (d)). To further confirm our above-mentioned results, we also examined these metastasis-relevant traits in miR-136-silenced GC-9811 cells and got the very similar results (Figure 2(a)–(d)). In short, these observations highlight cell-autonomous mechanisms of cell migration and invasion through which specific miR-136 may mediate metastasis.
We further used BLI of luciferase-transduced cells as well as gross examination of the peritoneal at necropsy. Indeed, the restoration of miR-136 expression in GC-981P cells significantly decreased peritoneal metastasis formation, and silencing miR-136 in GC-9811 cells showed the increased activity of forming peritoneal metastases, as assessed by BLI and histological analysis of micrometastatic nodules in the peritoneal cavity subsequent to the injection of GC-9811P or GC-9811 cells into stomach (Figure 2(e) and (f)). Collectively, these results show that miR-136 is not only a marker but also a functional mediator of gastric cancer–specific peritoneal metastasis.
MiR-136 post-transcriptionally downregulates HOXC10 expression by directly targeting its 3′-untranslated region
To understand the mechanisms by which three miRNAs mediate tumor invasion and metastasis, the target prediction program TargetScan or PicTar was used to search for predicted direct target genes of miR-136. Among the hundreds of the three miRNA targets, two genes (HOXC10 22 and insulin-like growth factor binding protein 5 (IGFBP5) 23 ) were previously implicated in promoting cancer cell migration and invasion. Bioinformatics analysis of the 3′-untranslated regions (UTRs) of two genes (HOXC10 and IGFBP5) revealed that HOXC10 and IGFBP5 were predicted to have at least one potential binding site at their 3′-UTRs for miR-136 (Figure 3(a)). Previously, we separately identified genes for which expression levels were changed across GC-9811P and GC-9811 cell lines. On the basis of available gene expression data sets, we identified 218 genes for which expression levels were increased (Supplementary Table 1) by at least two-fold in GC-9811P cell line compared with GC-9811 cells. We also identified 30 genes for which expression levels were decreased by at least half in GC-9811P cell line compared with parental line GC-9811 (Supplementary Table 1). The overlap between these two lists yielded a set of one gene (HOXC10) for which expression is changed in metastatic cells and regulated by miR-136.

MiR-136 post-transcriptionally downregulates HOXC10 expression by directly targeting its 3′-UTR. (a) HOXC10 and IGFBP5 were predicted to have at least one potential binding site at their 3′-UTRs for miR-136. (b) UTR reporter assays of miR-136 targeted genes in GC-9811 and GC-9811P cells. Reporter constructs consisting of the luciferase sequence fused to the 3′-UTRs of HOXC10 gene as well as the control gene CIAPIN1 were transfected into the GC-9811P and parental GC-9811 cell lines. Luciferase activity of cells was assayed at 32 h after transfection and the values were normalized to the GC-9811 cell line. (N = 3; error bars represent SEM; p<0.05; the p values are obtained using a one-sided Student’s t-test.) (c) miR-136 seed sequence in H0XC10 UTR or the miR-136 seed target site mutant were subjected to UTR reporter assays in GC-9811 cells with inhibited miR-136 expression. Luciferase activity of cells was also assayed in the presence or absence of the miRNAs inhibitors. (N = 3; error bars represent SEM; p values are obtained using a one-sided Student’s t-test.) (d) H0XC10 expression in GC-9811 and GC-9811P cells was obtained through real-time qRT-PCR in cells expressing miR-136 or silencing miR-136. (N = 3; error bars represent SEM; p values are derived using a one-sided Student’s t-test.) (e) H0XC10 protein level in GC-9811 and GC-9811P cells was obtained through western blot in cells expressing miR-136 or silencing miR-136. β-actin was used as an internal control.
In order to explore whether the altered expression of HOXC10 in metastatic cells is, in part, mediated through its 3′-UTRs, we cloned the 3′-UTR of HOXC10 downstream of a luciferase gene as a reporter and assayed their expression in GC-9811 cells (high miR-136) and GC-9811P cells (low miR-136). The expression from UTR reporter corresponding to HOXC10 was significantly lower in GC-9811 cells relative to GC-9811P cells, but not that of the control gene CIAPIN1 lacking the miR-136 target sequence (Figure 3(b)). In reporter assays, mutation of the miR-136 seed sequence in HOXC10 UTR as well as miR-136 inhibition with its sponge in GC-9811 cells significantly increased reporter expression (Figure 3(c)). In order to demonstrate that overexpression of miR-136 has a direct effect on targets, we assessed HOXC10 expression levels after miR-136 modulation by rt RT-PCR (Figure 3(d)) and western blot (Figure 3(e)). Our results showed that miR-136 directly regulates HOXC10, as indicated by the bioinformatics analysis.
MiR-136 inhibits gastric cancer peritoneal metastasis through downregulating HOXC10
Previous reports indicated that the expression of HOXC10 is overexpressed during the metastasis progression of cervical squamous cell carcinomas. 22 However, the biopathological significance of HOXC10 in gastric cancer peritoneal metastasis is still unknown. To better understand the potential role of HOXC10 in miR-136-mediated gastric cancer peritoneal metastasis, we examined the biological behaviour of HOXC10 on tumor cells in vitro and in vivo. We found knockdown of HOXC10 in GC-9811P cells with its siRNA reduced cell migration (Figure 4(a)) and invasion (Figure 4(b)) in vitro and significantly abolished metastasis by GC-9811P cells in peritoneal colonization assays (Figure 4(c)), similar to the phenotype observed with miR-136 restoration in these cells.

MiR-136 inhibits gastric cancer peritoneal metastasis through downregulating HOXC10. (a and b) Knockdown of HOXC10 in GC-9811P cells reduced cell (a) migration and (b) invasion and ectopic expression of HOXC10 reversed inhibition of (a) migration and (b) invasion by overexpressed miR-136 in GC-9811P cells. (N = 3; error bars represent SEM; p values are derived using a one-sided Student’s t-test.) (c) Knockdown of HOXC10 in GC-9811P cells significantly abolished metastasis by GC-9811P cells in peritoneal colonization assays. And ectopic expression of HOXC10 reversed inhibition of metastasis by overexpressed miR-136 in the highly metastatic cell line GC-9811P cells by peritoneal colonization assays. (N = 3; error bars represent SEM; p values are derived using a one-sided Student’s t-test.)
Considered miR-136 can post-transcriptionally regulate the expression of HOXC10 at the messenger RNA (mRNA) and protein levels by directly binding to its 3′-UTR, we hypothesized that upregulation of HOXC10 could directly reverse miR-136-restored inhibited gastric cancer peritoneal metastasis. We then re-expressed HOXC10 in miR-136-restored GC-9811P cells. Our results showed that ectopic expression of HOXC10 could reverse inhibition of peritoneal metastasis by overexpressed miR-136 in GC-9811P cells (Figure 4(a)–(c)). Thus, miR-136 inhibits gastric cancer peritoneal metastasis through suppression of HOXC10.
MiR-136 is downregulated in gastric cancer peritoneal metastasis tissues and associated with more peritoneal metastasis and worse prognosis
In order to confirm the cell-level experimental results, we further examined the miR-136 expression in 40 gastric cancer patients with (20 patients) or without (20 patients) peritoneal dissemination with quantitative rt RT-PCR. We found that miR-136 expression level was decreased in primary gastric cancer tissues from the patients with peritoneal dissemination, compared with that from the patients without peritoneal dissemination (Figure 5(a)). In stomach tissues from normal donors, we also observed a high expression level of the miR-136.

MiR-136 is downregulated in gastric cancer peritoneal metastasis tissues and associated with more peritoneal metastasis and worse prognosis. (a) Real-time qRT-PCR was performed with the clinical samples. Stomach tissues from normal donors were also examined. All of the PCRs were run in triplicate, and miRNA expression, relative to small nuclear RNA (U6), was calculated. (N = 3; error bars represent SEM; p values are derived using a one-sided Student’s t-test.) Patients in the (b) training set and (c) testing set were classified using a linear combination of miR-136. MiR-136 was weighted by its estimated Cox model regression coefficient for peritoneal metastasis to classify patients into a clinical low-risk group or a high-risk group. Shown are survival curves for peritoneal metastasis-free survival p values.
A biologically meaningful and clinically relevant miRNA that mediates peritoneal metastasis might be expressed uniquely by a subgroup of patients who suffered peritoneal relapse, and it should be associated with the clinical outcome. To test this, 84 gastric cancer patients treated at our institution were randomly assigned to a training set (n = 42) or a testing set (n = 42). Only the training data set is used for detection of miR-136 expression level. The training set was used in a univariate Cox proportional hazards model to relate the expression level of miR-136 with clinical outcome. We used miR-136 expression level to construct a signature by the risk score method. To investigate the effectiveness of miR-136 for peritoneal metastasis and clinical outcome prediction, a mathematical formula for this prediction was constructed, taking into account both the strength and the negative association of miR-136 with metastasis and survival. More specifically, we assigned each patient a risk score according to a linear combination of the expression level of miR-136, weighted by the regression coefficients derived from the aforementioned univariate Cox regression analyses. 21 The risk score for each patient was calculated as follows: risk score = (−0.174 expression level of hsa-miR-136). Patients having higher risk score are expected to have more risk to get peritoneal metastasis and poor survival outcomes.
We used the risk-score formula and calculated the risk scores for all patients in the training set. We ranked patients in the training set according to their risk scores and divided them into a high-risk group or low-risk group for developing peritoneal metastasis using the median risk score as the cutoff point. This risk-score formula distinguished patients who have high risk for developing peritoneal metastasis and low risk for developing peritoneal metastasis (5-year peritoneal-metastasis-free survival of 52.38% vs 76.19%, median survival of 37 months vs not reached, p = 0.036; Figure 5(b)).
We then used the same risk score formula obtained from the training set and calculated the risk scores for each of the 42 patients in the testing set. We classify them into the high-risk group or low-risk group using the same cutoff point as in the training set. We carried out the same peritoneal metastasis comparison procedures as in the training set. Similar to the findings from the training set, patients with high-risk miRNA signature had higher risk for developing peritoneal metastasis than patients with low-risk miRNA signature (5-year peritoneal-metastasis-free survival of 42.86% vs 71.43%, median survival of 33 months vs not reached, p = 0.0389; Figure 5(c)). Patients with high risk scores had more peritoneal metastasis and deaths than low-risk-score patients. These results suggested that low expression of miR-136 is closely related to the increase in gastric cancer peritoneal metastasis and may play an important role in the pathological process.
Discussion
Metastasis, the spread and growth of tumor cells to distant organs, represents the most devastating attribute of cancer. A notable feature of this process is the variation in metastatic tissue tropism displayed by different types of cancer. 1 In the case of gastric cancer, metastasis is the most frequent pattern of recurrence in patients with gastric carcinoma. 2 The prognosis of gastric cancer invading as deep as the gastric serosa remains poor, with a 5-year survival of <35%. 24 Peritoneal dissemination is reported to be the most frequent type of recurrence after curative resection in such cases. 25 Although coordination of multiple genes is involved in metastasis, only one or a few genes have been characterized in most previous reports examining this process. Differences in metastatic potential are expected to result from differences in expression in a combination of genes related to cell adhesion, apoptosis, drug metabolism, signal transduction, 26 and other processes. Yet, details of the molecular basis of peritoneal metastasis remain unclear.
Recently, miRNAs have been shown to be potential tools for cancer progression, diagnosis, and prognosis. 8 With the advent of miRNA expression profiles, significant efforts have been made to correlate miRNA expressions with tumor development.11–13 Since single miRNA can regulate hundreds of downstream genes, the information gained from miRNAs may be complementary to that from the expression profiling of protein-coding genes. Thus, identification of the actions of miRNAs could add new layers of complexity to our understanding of cancer dynamic.
In this study, we searched three gastric cancer peritoneal metastasis–related miRNAs by array-based miRNA profile. Among three miRNAs, we focused on miR-136, which was found decreased in all peritoneal metastatic sublines when compared with their expression in the parental line (Figure 1(a)). We further confirmed that miR-136 is frequently decreased in all GC-9811P subpopulations that metastasize to the peritoneal, regardless of whether these cells were randomly picked from the parental cell line or selected in vivo. When tested by change in miR-136 expression pattern in poorly metastatic cells or highly metastatic cells, miR-136 functions as a specific mediator of peritoneal metastasis in gastric cancer in vitro and in vivo. We also found that miR-136 expression level is tightly correlated with peritoneal metastatic activity. Furthermore, in the cohort of human gastric cancer primary tumors examined, patients having low expression of miR-136 had a significantly poorer peritoneal-metastasis-free survival. A direct and functional target of miR-136 was also identified. The target gene, HOXC10, is frequently repressed and functions as a metastasis promoter in several different tumor types. Taken together, these findings indicate that miR-136 plays a fundamental role in gastric carcinogenesis, especially in the process of gastric cancer–specific peritoneal metastasis.
Recent findings have shown that miR-136 is involved in various biological and pathological processes. Zhang et al. 27 reported that miR-136 plays an important role during transforming growth factor (TGF)-β1-induced proliferation arrest by targeting PPP2R2A in keratinocytes and could be used as a potential target for improving skin wound healing. Zhang et al. 28 reported that miR-136 could be used as a potential approach for the treatment of atherosclerosis through suppressing the extracellular signal–regulated protein kinases 1/2 (ERK1/2) pathway by targeting PPP2R2A. Chen et al. 29 reported that the low-level expression of miR-136 is significantly associated with a more aggressive and/or poor prognostic phenotype of patients with gliomas. But Shen et al. 30 reported that miR-136 acts as an oncogene by targeting PPP2R2A in human non-small-cell lung cancer. The existing studies have shown the complexity of miR-136 in different types of cancer. We postulate that this might be a consequence of different cancer heterogeneity, and different tumor types have its own specific genetic background. However, little is known about the role and underlying molecular mechanism of miR-136 in human gastric cancer, especially in gastric cancer peritoneal metastasis. In this report, we have, for the first time, established the role of miR-136 in gastric cancer–specific peritoneal metastasis.
Bioinformatics analysis showed that miR-136 could post-transcriptionally downregulates its target HOXC10 expression by directly targeting its 3′-UTR. Homeobox genes (i.e. HOXC10, HOXC6, and HOXD3) serve as important regulators of morphogenesis and differentiation during normal embryonic development.31–33 Several recent studies have suggested that dysregulation of specific homeobox genes is involved in cancer development, invasion, and metastasis, and altered expression patterns of homeobox genes have been observed in several different tumor types, including cancers of the lung, prostate, ovary, breast, colorectum, and cervix, 34 which indicates that HOXC10 may play an important role in the development of cancer and the epigenetical regulation involved. Our results showed that HOXC10 could promote gastric cancer peritoneal metastasis. Moreover, we reported that miR-136 can bind to one site of the 3′-UTR of HOXC10 and dramatically decrease the level of HOXC10 protein expression, which provides the first line of evidence in support of a mechanism for HOXC10 regulation at the post-transcriptional level.
We know that only a fraction of the cells in the particular gastric cancer population studied here have the ability to form highly aggressive, peritoneal metastases. Thus, metastatic ability is not a uniform trait in a gastric cancer cell population. In this study, we showed that miR-136 can significantly inhibit gastric cancer–specific peritoneal metastasis through downregulating its functional target of HOXC10. The newly identified miR-136/HOXC10 axis provides a new insight into the pathogenesis of gastric cancer peritoneal metastasis and represents a new, prognosis marker and potential therapeutic target for gastric cancer peritoneal metastasis.
Footnotes
Acknowledgements
J.Z., P.G., X.L., and J.W. have contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (nos 81000890, 3097134, and 81272344).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
