Abstract
TIG3 (tazarotene-induced gene 3) has been reported to suppress the progression of several malignancies, where this gene is universally downregulated. However, the expression of TIG3 in primary glioblastoma and its relevance to patient’s prognosis have not been elaborated. Thus, this study was aimed to evaluate TIG3 expression level in primary glioblastoma and investigate the prognostic value of TIG3 for patients. The Cancer Genome Atlas database was first utilized to analyze the expression and prognostic potential of TIG3 in 528 glioblastoma cases. Compared with control group, glioblastoma showed significantly elevated TIG3 expression (p < 0.001). Log-rank analysis revealed that higher expression of TIG3 was associated with shorter overall survival (358vs 383 days, p = 0.039). Furthermore, TIG3 protein expression detected by immunohistochemistry confirmed positive correlation of TIG3 expression and glioma grade and upregulation of TIG3 in our cohort of 101 primary glioblastoma patients compared to 16 normal brains. Finally, Kaplan–Meier analysis and Cox regression analysis identified high TIG3 expression as an independent risk factor for overall survival of primary glioblastoma patients (overall survival, 10 vs 13 months, p = 0.033; hazard ratio = 1.542, p = 0.046). Together, this study indicated that increased expression of TIG3 in primary glioblastoma is a novel biomarker for predicting poor outcome of patients. We then hypothesize that TIG3 may function in a different pattern in glioblastoma.
Introduction
Malignant gliomas account for approximately 80% of primary malignant brain tumors 1 and exhibit rapid and infiltrative growth and easy recurrence. 2 Primary glioblastoma (GBM) is the most common subtype of glioma, which has limited effective therapies and poor diagnosis, along with a median survival time of 14 months post diagnosis. 3 Studies have shown that primary GBM displays numerous genetic alterations and lots of signaling pathways are implicated in GBM oncogenesis. 4 Remarkably, three core signaling pathways, receptor tyrosine kinase (RTK)/RAS/phosphatidylinositol 3-kinase (PI3K), p53, and retinoblastoma (RB) protein pathways, were identified by The Cancer Genome Atlas (TCGA) research network recently. 5 However, a plenty of single or combined therapeutic approaches targeting these effectors still failed to improve overall survival (OS).4,6 Given these difficulties, identification of molecular biomarkers will enable prognostic stratification and facilitate personalized medicine. Currently, commonly used markers, such as 1p/19q loss and IDH1 mutation, are restricted in low-grade gliomas or secondary GBM to determine prognosis, due to the low frequency of these molecular events in primary GBM.7–9 Thus, there is a great need for new molecular targets and biomarkers of primary GBM to enable prognostic evaluation and novel therapeutic strategies.
Tazarotene-induced gene 3 (TIG3), also known as retinoic-acid receptor responder 3 (RARRES3) or retinoid-inducible gene 1 (RIG1), belongs to the class II tumor-suppressor gene HREV107 family.10,11 TIG3 ubiquitously expresses in most normal tissues and at the highest levels in several terminals, differentiated tissues including suprabasal epidermis, hair follicles, sebaceous gland, and terminal, differentiated colorectal epithelial cells.12–14 Previous studies have shown that TIG3 played a major role in cell proliferation, differentiation, and apoptosis and loss of TIG3 function would contribute to malignant transformation.15–18 Consistently, decreased expression of TIG3 was detected in many types of cancers, such as breast cancer, epidermal squamous cell carcinoma, colorectal cancer, B-cell lymphocytic leukemia, hepatocellular carcinoma, and cholangiocarcinoma.19–23 Furthermore, restoration of TIG3 expression could significantly suppress hepatocellular carcinoma growth via ERK1/2 inhibition by promoting apoptosis and inhibiting proliferation and migration, antagonize breast cancer cell growth and stemness properties through regulation of WNT proteins acylation and signaling activities, and inhibit metastasis of colorectal cancer via suppression of MTDH to regulate epithelial–mesenchymal transition.21,24,25 However, TIG3 expression and its underlying mechanism in primary GBM have not been elucidated, as well as its association with clinical outcome.
Here, we first searched a public tumor database TCGA to learn TIG3 in GBM. Consequently, a significant elevation of TIG3 expression was found in GBMs compared to normal brains, and its higher expression was correlated to poorer OS. Given the verified condition of TIG3 as a tumor-suppressor gene and downregulation in several malignancies,19,21–23 these results seemed intriguing.
We further applied immunohistochemistry and tissue microarray analysis to examine the expression of TIG3 in 101 tumor tissues of patients with primary GBM and 16 normal brain tissues. Then, we investigated the correlation between TIG3 expression and the clinicopathological characteristics of primary GBM patients. Our results showed that there was an increased expression of TIG3 in primary GBM tissues compared to the normal ones and a poorer prognosis of primary GBM patients was associated with higher TIG3 expression.
Materials and methods
Acquisition of human tissue specimens
The acquisition of tissue samples was approved by the Specialty Committee on Ethics of Biomedical Research, Second Military Medical University. Glioma tissue samples were obtained from patients with glioma who underwent surgery at Changzheng Hospital between 1999 and 2010. The diagnosis of glioma was made by two experienced pathologists independently based on the 2007 World Health Organization (WHO) Classification of Tumors of the Central Nervous System. 26 Normal brain tissue specimens were obtained from trauma patients with severe head injuries, on whom partial resection of the brain was performed to decompress the intracranial pressure to survive. All the patients and legal guardians signed the informed consent forms of human tissue acquisition used in this study by National Regulations on the Use of Clinical Samples in China. The subject selection was based on the following criteria: (1) the patient was diagnosed with primary GBM and without a history of other tumors; (2) complete clinical data including age, gender, clinical manifestations, magnetic resonance imaging (MRI) imaging data, extent of resection, and adjuvant therapy were collected for each participant; and (3) the patient underwent evaluation by enhanced head MRI scans for tumor relapse or progression after surgery at least once every 6 months. Patients receiving chemotherapy or radiotherapy before surgery were excluded.
Tissue microarray and immunohistochemistry
Tissue microarray analysis was performed on 268 glioma specimens and 16 normal brains with rabbit antibody against TIG3 (1:50, ab134823; Abcam Co., Ltd, Cambridge, MA, USA) as previously described.27,28 The immunohistochemistry staining was scored by two pathologists independently by intensity and extent of TIG3 expression. The intensity scored 0 to 3 (0: negative; 1: light brown; 2: medium brown; 3: dark brown). The percentage of positive cells was scored as follows: 0, no staining; 1, <20%; 2, 20%–70%; and 3, ≥70%. The scores of intensity and positive percentage were multiplied to estimate the TIG3 expression level, which was classified as high (total score ≥2) and low (total score <2).
Statistical analysis
All statistical analyses were performed with the SPSS 18.0 software (SPSS Inc, Chicago, IL, USA), and results were presented as mean ± standard deviation (SD). TIG3 gene expression values of 528 GBM cases and 10 controls based on the Affymetrix microarrays (Human gene U133A), as well as the follow-up information, were collected from TCGA database (www. cancaergenome.nih.gov). The association between the TIG3 expression and the GBM prognosis was studied in the gene expression profile dataset and our tissue microarray profiling, respectively. Student t-test and Mann–Whitney U-test were used to compare TIG3 messenger RNA (mRNA) and protein expression levels in GBM and control groups, respectively. Spearman analysis was used to examine the relationship between TIG3 expression and glioma grade. Progression-free survival (PFS) and OS curves were plotted by the Kaplan–Meier method and compared by log-rank test. The median TIG3 expression value (7.90) and its vicinal values in TCGA cohort were tried as the cutoff to find out the potential of TIG3 for predicting GBM patient prognosis. Expression value of 7.50 was selected to define patient as high (≥7.50) or low (<7.50) TIG3 group. The identification of relevant prognostic factors in our cohort was performed by univariate analysis, multivariate analysis, and stepwise backward Cox regression model. Factors with a result of p < 0.2 on univariate analysis were added into multivariate analysis for potential association with OS and PFS. OS was measured from the time of diagnosis to time of death. PFS was defined as the period from the date of diagnosis to time of tumor progression on MRI or death of the patient from glioma. All statistical tests were two-tailed, and the results were considered statistically significant when the p values were less than 0.05.
Results
Demographic characteristics and clinicopathological outcomes of participants
A total of 268 glioma patients (including 12 Grade I, 108 Grade II, 47 Grade III, and 101 Grade IV glioma cases) were eligible for further evaluation in our study, and the demographic and clinicopathological features of the primary GBM patients were listed in Table 1. The period of follow-up was from 0.5 to 64 months. The recurrence was found in 95 (94.1%) cases, and only 8 (7.9%) patients were still alive. The median OS and median PFS were 11 months (95% confidence interval (CI): 9.406–12.594) and 10 months (95% CI: 7.725–12.275), respectively.
Demographic and clinicopathological characteristics of the patients with primary GBM.
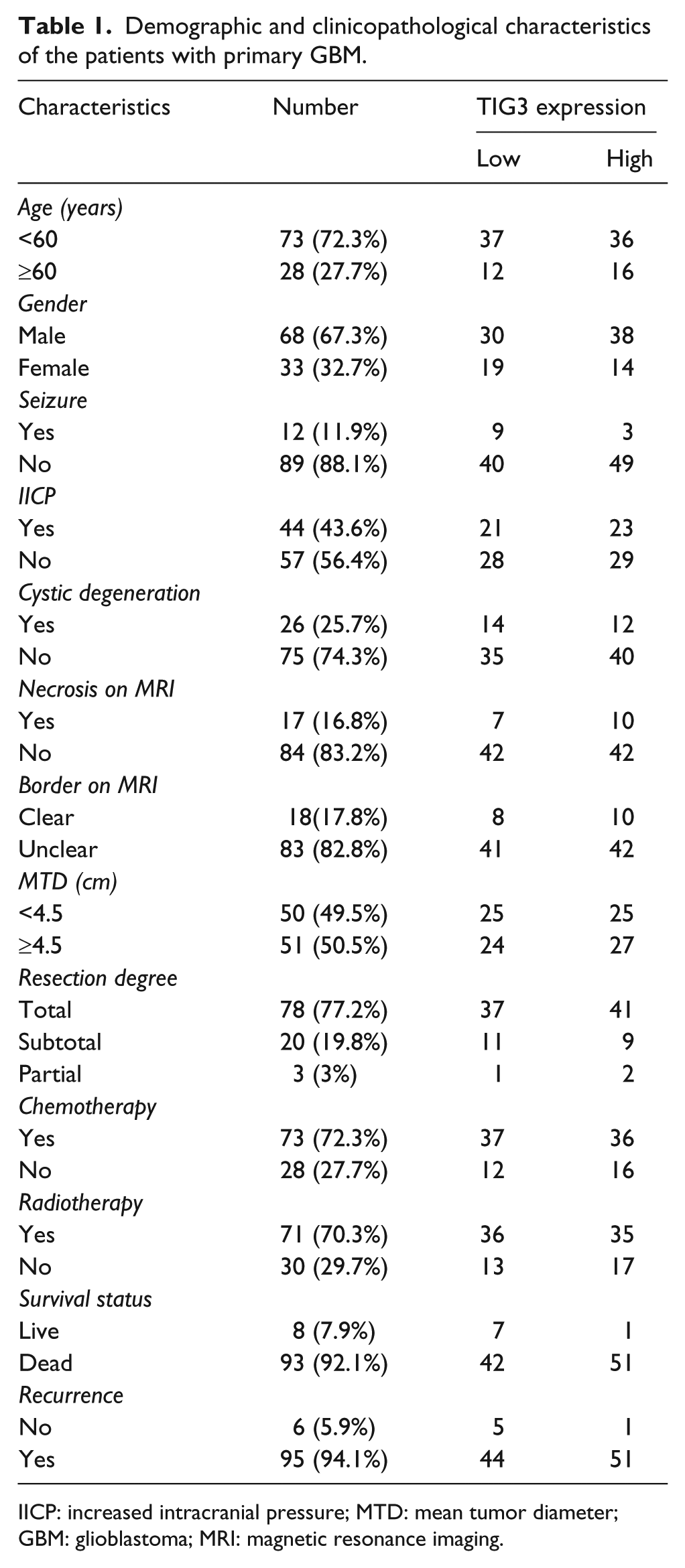
IICP: increased intracranial pressure; MTD: mean tumor diameter; GBM: glioblastoma; MRI: magnetic resonance imaging.
TCGA dataset analysis identifies TIG3 highly expressed in GBM and associated with patient survival
The association between TIG3 expression level and GBM was studied in 528 GBM cases deposited in TCGA database. TCGA performed genome-wide profiling of 528 GBMs and 10 normal brain tissues. It was found that TIG3 showed a moderately higher transcriptional level in GBMs compared with normal brains (7.87 ± 1.20 vs 5.83 ± 0.55, p < 0.001; Figure 1(a)). Furthermore, the TIG3 expression was found to be related to the OS of GBM patients (p = 0.039; Figure 1(b)). Patients with high TIG3 expression had a significantly short OS (358 vs 383 days). Although no statistically significant result was obtained in the PFS analysis, there was still a moderate difference between high and low TIG3 subgroups (median PFS: 186 vs 228 days, p = 0.126; Figure 1(c)).

Upregulation of TIG3 is in GBM of TCGA database and associated with poor patients’ survival. (a) TIG3 mRNA expression level detected in 528 GBM cases is higher than that of 10 normal brains (7.87 ± 1.20 vs 5.83 ± 0.55, p < 0.001). (b and c) Kaplan–Meier survival analysis in 528 GBMs stratified by TIG3 expression shows that high expression of TIG3 predicts a poor prognosis, and the difference in OS between the two groups is statistically significant (358 vs 383 days, p = 0.039), but not for PFS (median PFS: 186 vs 228 days, p = 0.126).
Large TMA confirms TIG3 as an independent risk factor for primary GBM patients
The expression level of TIG3 was further examined by tissue microarray in our cases and controls. It is found that TIG3 almost exclusively appeared in the cytoplasm of GBM cells with sparse expression in the nucleus (Figure 2(a)–(c)). Compared with normal brains, TIG3 expression was noticeably increased in primary GBM tissues (1.653 ± 1.228 vs 0.375 ± 0.619, p < 0.001; Figure 2(e)). Noticeably, TIG3 expression in all glioma positively correlated with pathological grade (p < 0.001; Figure 2(e)), which indicated that TIG3 might be involved in the progression of malignant glioma. The relationship between TIG3 expression and clinical outcome was further investigated. During the follow-up, 51 out of 52 primary GBM patients died in the high TIG3 group, while 42 out of 49 primary GBM patients with low TIG3 had the same ends. As determined by the log-rank test, high TIG3 expression was identified as a risk factor for OS (10 vs 13 months, p = 0.033) and PFS (8 vs 11 months, p = 0.030) of primary GBM patients (Figure 2(f)–(g) and Table 2). Meanwhile, among other clinical features, radiotherapy and resection degree were found to be protective factors for OS, as well as age less than 60 years and radiotherapy for PFS (Table 2).

TIG3 expression is elevated in primary GBM based on TMA data and correlates with survival time of patients. Representative staining images in primary GBM samples and normal brain are shown in (a–c) and (d), respectively. Most TIG3 protein expresses in the cytoplasm of GBM cells. (e) Immunohistochemical staining score of TIG3 expression in 101 primary GBMs is significantly higher than that in 16 normal brain tissues (1.653 ± 1.228 vs 0.375 ± 0.619, p < 0.001). TIG3 expression is positively related to pathological grade of glioma (p < 0.001). (f and g) Kaplan–Meier survival analysis in 101 primary GBM cases stratified by TIG3 expression indicates both shorter OS (10 vs 13 months, p = 0.033) and PFS (8 vs 11 months, p = 0.030) in patients with higher TIG3 expression.
Univariate analysis of factors associated with survival and progression of primary GBM patients.

IICP: increased intracranial pressure; MTD: mean tumor diameter; HR: hazard ratio; OS: overall survival; PFS: progression-free survival.
Furthermore, variables with p < 0.2 in the above univariate analysis were added into Cox proportional hazards regression model to estimate the ability of TIG3 to predict survival. As a result, high expression of TIG3 (HR = 1.542, p = 0.046), small extent of resection, and none post-surgery radiotherapy were demonstrated as independent risk factors for shorter OS in primary GBM patients (Table 3). For PFS, post-surgery radiotherapy was also an independent protective factor. However, low expression of TIG3 (HR = 0.657, p = 0.053) did not reach statistical significance (Table 3).
Multivariate analysis of factors associated with survival and progression of primary GBM patients.

HR: hazard ratio; OS: overall survival; PFS: progression-free survival; CI: confidence interval.
Variables were adopted for their prognostic significance by univariate analysis (p < 0.2).
Discussion
Recent studies have demonstrated decreased TIG3 expression in various human malignancies, indicating a function as a tumor suppressor.19–23 However, our study found that TIG3 expression was positively related to glioma grade and upregulated in GBM, where its high expression was associated with an unfavorable outcome of patients. To our knowledge, this is the first time that TIG3 expression is identified in an opposite way in malignant glioma and can be used as a prognostic factor.
Previous studies have shown that some signaling pathways such as Notch can exert both oncogenic and tumor-suppressive functions depending on the cellular context. 29 A key issue regarding TIG3 is how this protein interplays with signaling pathways involved in the development of GBM. A study in breast cancer has found that TIG3 expression is negatively regulated by 17β-estradiol (E2) through estrogen receptor. 30 However, there exist no typical estrogen response elements in the genomic sequence of TIG3 regulatory regions, which indicates an indirect regulatory relationship between E2 and TIG3. Though GBM expresses high E2, estrogen receptors are present at a low level, which is different from that in breast cancer.30–33 Accordingly, the indirect regulatory mechanism between TIG3 and E2 may be affected by other effectors or may not occur in GBM. Recently, p53 was reported to directly regulate TIG3 transcription via a p53 response element (p53RE) in the TIG3 promoter. 34 It is important to note that TIG3 promoter activity is greatly enhanced by wild-type but not mutated p53 protein, which can be abolished by sequence-specific mutation of p53RE as well. 34 Given the high frequency of p53 mutation or function loss in GBM, 5 the direct regulatory relationship between p53 and TIG3 seems to be in a subordinate status in the tumor cells. Moreover, IDH1 mutation was found to be significantly correlated with underexpression of TIG3 in GBM. 35 It is demonstrated that glioma patients with the IDH1 mutation have a better outcome than those with wild-type IDH1 gene. 9 Though IDH1 mutation occurs at a low frequency in primary GBM, 9 low expression of TIG3 predicting longer survival in this study is consistent with these previous findings. The TIG3 expression is also influenced by retinoid acid (RA) signaling.11,36 Unlike other tumors, GBM exhibits a unique aberrant expression profile of RA signaling genes, such as upregulation of the retinol-binding protein CRBP1, RA synthesizing protein ALDH1A1, and RA-binding protein FABP5, with augmented intratumoral RA level and alternative, prosurvival RA usage, which then can promote TIG3 expression.37,38 Taken together, our results suggest that increased TIG3 may be involved in the altered RA signaling pathway and promote gliomagenesis.
TIG3 exerts its suppressing function such as inhibition of RAS activation, which is dependent on the membrane localization of the TIG3 protein mediated by the hydrophobic C-terminal region. 15 The N-terminal hydrophilic region was demonstrated to target TIG3 to the centrosome to inhibit daughter centrosome separation during cell division and lead to cessation of cell proliferation. 20 The intracellular locations of TIG3 compete in a mutually exclusive fashion possibly due to the protein conformation change. 20 Recently, the conserved N-terminal NlpC/P60 domain of TIG3 was also found to exhibit calcium-independent phospholipase A1/2 activity, which indicated that TIG3 might be involved in the process of phospholipid metabolism.12,39 Interestingly, the inhibitory effect on cell survival induced by the C-terminal domain must be independent of the phospholipase A1/2 activity. 39 Based on these findings, we suspect that TIG3 may not provide the tumor-suppressing activity through the mechanism of subcellular localization, but acts as a phospholipid-metabolizing enzyme on the cellular context of GBM. Moreover, calcium-independent phospholipase A2 enzymes have been implicated in tumorigenesis with in vitro studies, which supports our hypothesis that TIG3 plays an oncogenic role in GBM, though the loss of TIG3 phospholipase A1/A2 activity was showed to promote breast cancer lung metastasis.19,40
In addition, a previous study reported surprising data that mutation of TIG3 amino acids 124–164 would result in p53 level reduction possibly mediated by endoplasmic reticulum stress response. 41 A primary function of p53 is to regulate several cellular programs including the cell-cycle arrest, response of cells to DNA damage, senescence, apoptosis, and differentiation, and dysregulation of p53 will contribute to the progression of GBM. 4 Thus, the possibility cannot be ruled out that some unknown regulation or mutation of TIG3 could induce endoplasmic reticulum stress response to reduce p53 protein level in GBM.
In summary, we demonstrated that high expression of TIG3 was associated with poor prognosis of primary GBM patients, and TIG3 could work as an independent risk factor for OS. Different from previous studies that TIG3 could suppress tumor progression in types of cancers, our results indicated that TIG3 might function as an oncogenic effector and be involved in the progression of GBM, which needs more investigations in the future.
Footnotes
Acknowledgements
H.W. and H.X. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Natural Science Foundation (81572501 and 81272781), Program for Academic Leaders of Shanghai (No. 043), and “Pu Jiang Talent” Project of Shanghai (PJ[2014]0002617).
