Abstract
Previous meta-analysis has not shown different effects of miR-608 rs4919510 polymorphism in specific cancer types and reported no significant association between rs4919510 and cancer risk among Chinese. However, more recent findings have been inconsistent. Therefore, we performed an updated meta-analysis to examine whether this polymorphism is associated with cancer risk based on ethnicity and type. A total of 18 case-control studies, comprising 12,517 cases and 15,624 controls, were included in our study. Surprisingly, in contrast with previous meta-analysis, a significant association between the rs4919510 polymorphism and cancer risk was observed in Chinese (CG vs CC: odds ratio = 1.11; 95% confidence interval = 1.04–1.19). In further stratified analyses based on cancer type, rs4919510 was significantly associated with an increased risk of papillary thyroid cancer (CG vs CC: odds ratio = 1.25; 95% confidence interval = 1.01–1.54) and exhibited borderline significant associations with increased risk of gastric cancer (GG vs CC: odds ratio = 1.27; 95% confidence interval = 1.00–1.62) and lung cancer (CG vs CC: odds ratio = 1.14; 95% confidence interval = 0.99–1.32), but a decreased risk of colorectal cancer (GG vs CC: odds ratio = 0.74; 95% confidence interval = 0.60–0.91). Moreover, the RegulomeDB database indicated that rs4919510 may affect the expression of two nearby genes (SEMA4G and MRPL43), and the Cancer Genome Atlas database revealed that the expression level of SEMA4G was significantly lower in colorectal cancer and lung cancer tissues than that in adjacent non-tumour tissues, while the expression level of SEMA4G was significantly higher in gastric cancer tissues than that in adjacent non-tumour tissues. These findings provide evidence that the miR-608 rs4919510 polymorphism may modify cancer susceptibility in a type-specific manner. Furthermore, SEMA4G may function as an oncogene or tumour suppressor to regulate tumour development in a type-specific manner. Further studies with experimental evaluations are warranted.
Introduction
Cancer is the leading cause of death worldwide; in 2005, 14% of all deaths were due to cancer, while this value increased to 16% in 2015. 1 Despite significant reductions in cancer mortality in some countries, cancer cases increased by 33% from 2005 to 2015, and the expected increase will continue for years and possibly decades, which has sparked public concern. 2 Admittedly, there are various factors contributing to the occurrence of tumour development, but it is undeniable that the interaction of environment/lifestyle and host genetic factors (primarily single nucleotide polymorphisms (SNPs)) is categorised as one of the major elements responsible. 3
MicroRNAs (miRNAs) comprise a large and growing family of short noncoding RNAs that are frequently dysregulated in malignancies. Because miRNAs play essential roles as oncogenes or tumour suppressors, they likely contribute to cancer initiation and progression.4,5 According to the latest version of the miRNA database (miRBase), over 2500 mature miRNAs have been identified in humans. Genetic variants in miRNAs may modify cancer risk, as was demonstrated by various miRNAs including miR-608 rs4919510. However, the results of previous studies that explored the association between the miR-608 rs4919510 polymorphism and cancer risk were inconclusive. For instance, one previous study reported that the rs4919510 variant G allele was significantly associated with an increased risk of breast cancer in Chinese populations. 6 However, among Iranian people, the same variant showed no significant association with the development of breast cancer, and the effect value was in the opposite direction relative to the previous study in a Chinese population. 7 Even so, an increasing number of studies have focused on the association between miR-608 rs4919510 and several common types of cancer, including lung cancer,8–10 colorectal cancer,11–13 gastric cancer,14,15 head and neck cancer, 16 breast cancer,6,7,17,18 papillary thyroid cancer,19,20 hepatocellular carcinoma, 21 oesophageal squamous cell carcinoma, 22 and nasopharyngeal carcinoma. 23 Moreover, a recent study summarised the associations of miR-608 rs4919510 with cancer risk by meta-analysis. 24 However, there were only eight included studies, and the sample size in most of these studies was relatively small, which may have limited the statistical power. Clearly, the association between the miR-608 rs4919510 polymorphism and cancer susceptibility requires further analysis. Therefore, we performed an updated meta-analysis, including all published data to date, to more precisely characterise the association between miR-608 rs4919510 polymorphism and cancer susceptibility.
Materials and methods
Identification and eligibility of relevant studies
Relevant literature was collected by searching the PubMed and Web of Science databases (the last search update was 20 January 2017) using the following keywords: (‘microRNA 608’, ‘microRNA-608’, ‘miR-608’ or ‘rs4919510’) and (‘cancer’, ‘carcinoma’, ‘tumor’, ‘tumour’ or ‘neoplasm’) and (‘polymorphism’, ‘variation’, ‘variant’ or ‘mutation’). This meta-analysis included only publications written in English with available full-text articles. In this meta-analysis, the studies were required to meet the following standards: (1) they involved the miR-608 rs4919510 polymorphism and cancer risk; (2) they were designed as case-control studies and (3) they contained available genotype frequency data. The primary reasons for study exclusion were as follows: (1) the studies did not involve miR-608; (2) they did not involve rs4919510 polymorphism research; (3) they were not related to cancer research and (4) no relevant genotype data were reported. Ultimately, the data for this meta-analysis included 18 case-control studies, comprising 12,517 cases and 15,624 controls (Table 1).
Characteristics of literature included in the meta-analysis.

HWE: Hardy–Weinberg equilibrium; HNSCC: head and neck squamous cell carcinoma; ESCC: oesophageal squamous cell carcinoma.
Including Lithuania and Latvia.
Including Lithuania, Latvia and Germany.
Data extraction
Two investigators (S.W. and W.Y.) independently extracted the data and reached a consensus on all of the items. The following information, including the first author’s name, year of publication, country of origin, ethnicity, type of cancer, numbers of cases and controls and genotyping platform, was obtained from each article. Subject ethnicity was categorised as either Chinese or non-Chinese.
Functional annotation based on the RegulomeDB and the Cancer Genome Atlas databases
RegulomeDB (http://regulome.stanford.edu) is a new database that can be used to predict whether a variant affects transcription factor binding and gene expression, with categories ranging from 1 to 6 based on the confidence that a variant is functional. Category 1 is further divided into six subcategories (1a–f), and lower scores indicate increasing evidence that a variant is located in a functional region; for example, a variant scored as 1a has the highest likelihood of affecting binding and being linked to the expression of a target gene (Table 2). The gene expression data of tumour and adjacent non-tumour tissues and corresponding personal clinical information for each patient were obtained from the Cancer Genome Atlas (TCGA; http://cancergenome.nih.gov) database portal.
The RegulomeDB scoring scheme for a single SNP.

SNP: single nucleotide polymorphism.
Statistical analysis
The risk of cancer associated with the miR-608 rs4919510 polymorphism was estimated for each study using the odds ratio (OR) and its 95% confidence interval (95% CI). The intrastudy heterogeneity was examined with a chi-square-based Q statistic test, and p < 0.05 was considered to be statistically significant. When the heterogeneity between studies was absent, we pooled the results using fixed-effect models; otherwise, a random-effects model was chosen. Subsequently, we evaluated the risks of the heterozygous and variant homozygous genotypes relative to the wild-type homozygous genotype and then assessed the risks of the combined heterozygous as well as variant homozygous genotypes relative to the wild-type homozygous genotype while assuming the dominant effects of the variant allele, together with assessing the risks of the variant homozygous genotype relative to the combined wild-type homozygous and heterozygous genotypes while assuming the recessive effects of the variant allele; furthermore, we also assessed the allele model (G vs C). Additionally, we performed stratification analyses based on ethnicity (divided into Chinese and non-Chinese) and cancer type. Begg’s and Egger’s tests were utilised to evaluate publication bias. Differences in expression between tumour and adjacent non-tumour tissues were assessed by Student’s t-test. All analyses were performed using Stata version 12.0 software (Stata Corporation, College Station, TX, and USA).
Results
Characteristics of the published studies
Following the application of the strict screening criteria, 18 case-control studies that contained a total of 12,517 cases and 15,624 controls were ultimately included in the current quantitative analysis. The general characteristics of the included studies are listed in Table 1. Among the included studies, 13 studies were conducted among Chinese populations and 5 studies were conducted among non-Chinese populations. The distribution of genotypes in the controls was consistent with Hardy–Weinberg equilibrium (HWE) for 17 studies, except the study performed by Li et al. Altogether, 4 studies reported the effects of miR-608 rs4919510 on breast cancer, 3 studies each on colorectal cancer and lung cancer, 2 studies each on gastric cancer and papillary thyroid cancer, 1 on head and neck squamous cell carcinoma, 1 on oesophageal squamous cell carcinoma, 1 on nasopharyngeal carcinoma and 1 on hepatocellular carcinoma. Genotyping was performed using TaqMan in 8 studies, Sequenom in 6 studies, SNaPshot in 1 study, PCR-RFLP in 1 study, SNPstream in 1 study and Illumina Infinium Human Exome BeadChip in 1 study. The distributions of miR-608 rs4919510 polymorphism genotypes in each individual study are listed in Supplementary Table 1.
Quantitative synthesis
The evaluations of the associations of miR-608 rs4919510 with cancer risk are presented in Table 3. In total, the variant G allele showed a significantly increased cancer risk (CG vs CC: OR = 1.10; 95% CI = 1.03–1.16; p = 0.16 for the heterogeneity test; I2 = 25.1%; Figure 1). The results of other tested models are also listed in Table 3.
Summary ORs of the miR-608 rs4919510 polymorphism and cancer risk.

OR: odds ratio; CI: confidence interval.
Random-effects model was used when p value for heterogeneity test <0.05; otherwise, fixed-effect model was used.
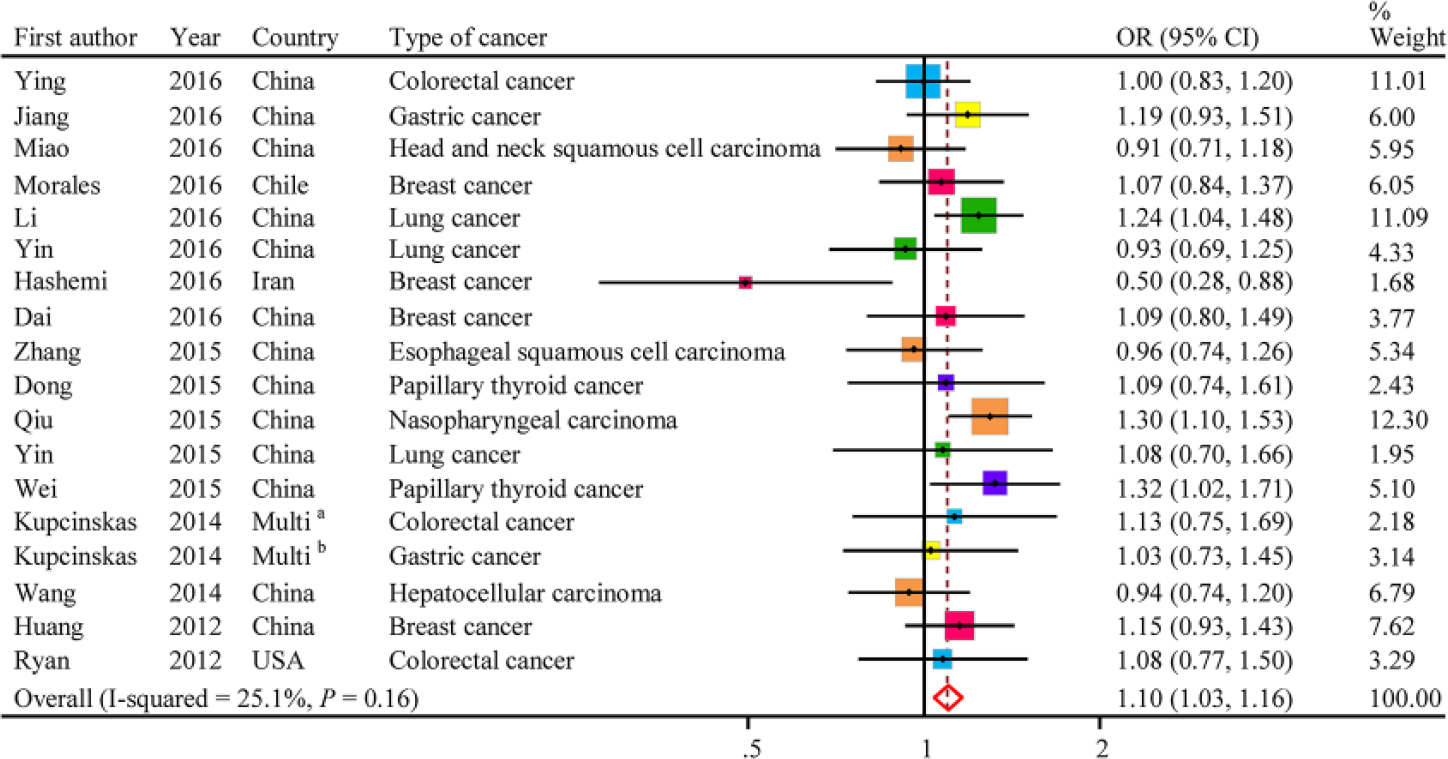
Forest plot of the miR-608 rs4919510 polymorphism and cancer risk (CG vs CC).
We next evaluated the effect of the miR-608 rs4919510 polymorphism on cancer risk among subgroups (Table 3). In our stratified analyses, the miR-608 rs4919510 SNP showed a significant association with increased cancer risk among Chinese (CG vs CC: OR = 1.11; 95% CI = 1.04–1.19; p = 0.24 for the heterogeneity test; I2 = 20.1%). Beyond that, the rs4919510 was significantly associated with an increased risk of papillary thyroid cancer (CG vs CC: OR = 1.25; 95% CI = 1.01–1.54; p = 0.42 for the heterogeneity test; I2 = 0.0%) and exhibited borderline significant associations with increased risk of gastric cancer (GG vs CC: OR = 1.27; 95% CI = 1.00–1.62; p = 0.20 for the heterogeneity test; I2 = 37.9%) and lung cancer (CG vs CC: OR = 1.14; 95% CI = 0.99–1.32; p = 0.25 for the heterogeneity test; I2 = 27.3%). However, for colorectal cancer, the result was not concordant with gastric cancer, papillary thyroid cancer and lung cancer, as the variant G allele was significantly associated with a decreased colorectal cancer risk (GG vs CC: OR = 0.74; 95% CI = 0.60–0.91; p = 0.25 for the heterogeneity test; I2 = 28.0%).
Sensitivity analysis and publication bias
To test the stability of the rs4919510 results, we conducted sensitivity analysis by sequentially removing each eligible study. The study by Hashemi et al. that focused on breast cancer was identified as the major contributor of heterogeneity (CG vs CC: I2 = 25.1%; p = 0.16 for heterogeneity). After the removal of this study, the heterogeneity was significantly reduced (I2 = 0.0%; p = 0.50 for heterogeneity), indicating that the study by Hashemi et al. significantly changed the pooled OR (Supplementary Table 2). We further utilised funnel plot, Begg’s test and Egger’s test to evaluate potential publication biases of the studied literatures. The shape of the funnel plot was symmetrical (Supplementary Figure 1). Moreover, Begg’s and Egger’s tests provided no statistical evidence for the existence of publication bias (G vs C: p = 0.47 and p = 0.21, respectively).
Functional annotation
To investigate the functional effect of miR-608 rs4919510 on corresponding genes, we first searched the RegulomeDB scoring system. Interestingly, the miR-608 rs4919510 SNP was found to have a score of 1f, suggesting its link with the expression of two genes (SEMA4G and MRPL43). Next, we quantified the expression levels of SEMA4G and MRPL43 in four cancer types with a previous significant association with rs4919510 using the TCGA database. As shown in Figure 2, the expression level of SEMA4G was significantly higher in gastric cancer tissues compared to adjacent non-tumour tissues (p = 1.05 × 10−3). However, for colorectal cancer and lung cancer, the direction of expression was not concordant with gastric cancer, as the expression level of SEMA4G was significantly lower in colorectal cancer tissues and lung cancer tissues compared to corresponding adjacent non-tumour tissues (p = 4.84 × 10−20 and p = 6.23 × 10−17, respectively; Figures 3 and 4). In addition, the level of MRPL43 expression was significantly lower in gastric cancer tissues compared to adjacent non-tumour tissues (p = 1.76 × 10−5). However, no significant difference in expression was observed for MRPL43 in colorectal cancer tissues and lung cancer tissues compared with corresponding adjacent non-tumour tissues. In addition, we did not observe any significant association for SEMA4G and MRPL43 expression in papillary thyroid cancer.

The expression level of SEMA4G in gastric cancer tissues and adjacent non-tumour tissues.

The expression level of SEMA4G in colorectal cancer tissues and adjacent non-tumour tissues.

The expression level of SEMA4G in lung cancer tissues and adjacent non-tumour tissues.
Stratified analyses of SEMA4G expression
Stratified analyses were also conducted for SEMA4G expression levels based on ethnicity, gender, age and tumour–node–metastasis (TNM) stage. As shown in Table 4, significant associations for gastric cancer were observed in Asians, males, older individuals (age ⩾60 years) and early tumour stage (TNM I/II). For colorectal cancer, significant associations were observed for each stratification. Similarly, for lung cancer, significant associations were observed for each stratification, with the exception of late-stage lung cancer (TNM III/IV).
Stratified analyses of the expression levels of SEMA4G in tumour tissues and adjacent non-tumour tissues.
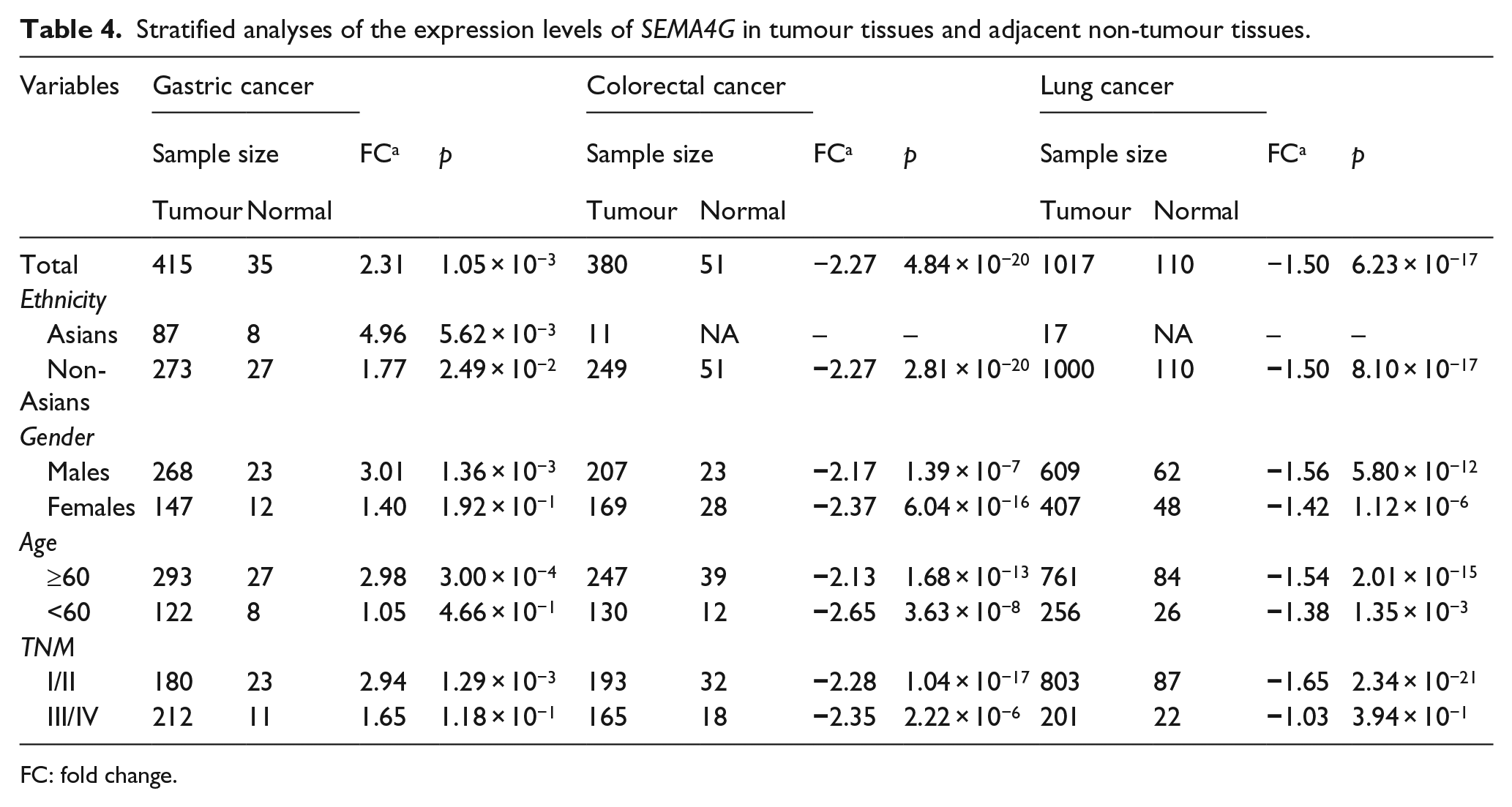
FC: fold change.
Discussion
Previous meta-analysis has not revealed any significant relationship between the miR-608 rs4919510 SNP and cancer risk in Chinese. However, more recent findings have been inconsistent. Therefore, we conducted an updated meta-analysis to examine whether this polymorphism is associated with cancer risk, especially in Chinese. In this study, we performed a meta-analysis by pooling 18 case-control studies with a total of 12,517 cases and 15,624 controls, and our findings demonstrated that the variant G allele was significantly associated with an increased cancer risk. More importantly, in stratified analyses, in contrast with previous meta-analyses, we reported for the first time that rs4919510 was clearly associated with an increased cancer risk in Chinese.
The rs4919510 SNP at 10q24.31 is located in the intron of the SEMA4G gene. Furthermore, the RegulomeDB scoring system suggested that the miR-608 rs4919510 SNP may modify the expression of SEMA4G. Additionally, the TCGA database suggested that the expression level of SEMA4G was significantly lower in colorectal cancer tissues compared to adjacent non-tumour tissues, indicating that SEMA4G may play as a tumour suppressor gene of colorectal cancer, which was in perfect agreement with a study performed by Wang et al. 25 that demonstrated SEMA4G may be a tumour suppressor gene related to colorectal cancer. The rs4919510 variant G allele exhibited a significant association with a decreased risk of colorectal cancer in our meta-analysis, leading us to hypothesise that the rs4919510 variant G allele in the SEMA4G gene might increase the expression of SEMA4G in the colon and rectum. Therefore, the elevated expression of SEMA4G in the colon and rectum may decrease the colorectal cancer risk. Similarly, SEMA4G expression was detected at low levels in lung cancer tissues compared with adjacent non-tumour tissues. Moreover, as the rs4919510 variant G allele exhibited a significant association with an increased risk of lung cancer in meta-analysis, it is plausible that the variant G allele of rs4919510 may decrease the expression of SEMA4G in the lung. Therefore, the decreased expression of SEMA4G in the lung may increase the lung cancer risk. However, for gastric cancer, the expression level of SEMA4G was significantly higher in gastric cancer tissues compared to adjacent non-tumour tissues. Because the rs4919510 variant G allele exhibited a significant association with an increased risk of gastric cancer in meta-analysis, it is plausible that the variant G allele of rs4919510 may increase the expression of SEMA4G in the stomach. Therefore, the increased expression of SEMA4G in the stomach may increase the gastric cancer risk, indicating that SEMA4G may play the role of an oncogene in gastric cancer development. The most striking implication of the above evidence is that SEMA4G may act as either a tumour suppressor gene (in colorectal cancer and lung cancer) or an oncogene (in gastric cancer) in a type-specific manner and that the abnormal expression of SEMA4G might be associated with tumourigenesis or be important for tumour progression and maintenance. Further experimental studies are needed to verify this hypothesis, especially whether the expression level of the SEMA4G gene in tumour tissues was associated with different genotypes of rs4919510.
SEMA4G (semaphorin 4G), which is known to be a DNA damage-binding and repair gene, 26 belongs to the semaphorin family that contains more than 20 genes divided into 7 subfamilies. Some semaphorins have been found to function as tumour suppressors and may inhibit tumour progression by various mechanisms, whereas other semaphorins may function as inducers and promoters of tumour progression. 27 Specifically, previous studies have demonstrated that SEMA4G may act as a tumour suppressor gene, as the expression of SEMA4G was at low levels in colorectal cancer tissues compared with normal tissues. 25 Further pathway analysis based on the KEGG database suggested that SEMA4G may function in the axon guidance pathway. Indeed, recent evidence has indicated a role for the dysregulation of some axon guidance genes in tumour initiation and progression,28,29 such as the axon guidance factors SEMA6B and SEMA5A.30,31 In addition, alignment of SEMA4G with its most closely related family member, 32 SEMA4C, was reported to be a potential serum marker for the diagnosis as well as the risk of breast cancer and cervical cancer metastasis. Wei et al. 33 suggested that by releasing high levels of soluble SEMA4C, tumour-associated lymphatic endothelial cells not only promote lymphangiogenesis but also promote the proliferation and migration of tumour cells, thus facilitating lymphatic metastasis.
A previous meta-analysis suggested that miR-608 rs4919510 was significantly associated with cancer risk in Caucasians but showed no significant relationship in Chinese. However, in contrast to these findings, our present meta-analysis suggested that miR-608 rs4919510 was significantly associated with cancer risk in Chinese but not in non-Chinese. There are several reasons for these inconsistent results. First, the difference may be due to genetic heterogeneity between different ethnicities, as 13 of the total 18 included studies were performed in China, and most of the subjects were of Han Chinese decent. Second, the relatively small sample size for non-Chinese (a total of 5 studies including 1401 cases and 2,224 controls) may have limited the statistical power. Therefore, further studies with larger sample sizes are needed to evaluate our conclusions regarding non-Chinese populations.
The merit of our meta-analysis is that we systemically reviewed the relationship between the miR-608 rs4919510 polymorphism and cancer susceptibility using all data published to the present day, and in contrast with previous meta-analysis, we identified for the first time that the variant G allele of rs4919510 may increase lung cancer, gastric cancer and papillary thyroid cancer risk but decrease colorectal cancer risk. Additionally, the well-designed functional annotation together with the result of our meta-analysis further provides evidence that SEMA4G may act as either a tumour suppressor gene or oncogene to regulate tumour development in a type-specific manner. However, there are some study limitations that need to be addressed. First, no expression data for miR-608 in cancer tissues and adjacent non-tumour tissues were obtained from the TCGA database, which further limits the functional annotation of miR-608 in cancer development. Second, although the RegulomeDB scoring system indicated that the miR-608 rs4919510 SNP was significantly associated with SEMA4G expression, we could not obtain detailed data on the relative expression of SEMA4G according to rs4919510 genotypes in corresponding tissues. Therefore, we cannot confirm whether the variant G allele of rs4919510 increases or decreases the expression of SEMA4G in corresponding tissues. Further experimental confirmation is therefore required. Third, since Chinese and Non-Chinese subgroups differ considerably regarding on cancer types, the significant association of miR-608 rs4919510 with cancer risk in Chinese may be primarily the cancer type, instead of ethnicity effect. Finally, since the 95% CI of OR for gastric cancer and lung cancer includes the value of 1.00, the point of no effect, we cannot exclude the possibility that the reported OR for gastric cancer and lung cancer may due to a random chance and not to some real cancer risk effect.
Conclusion
This study provides evidence that miR-608 rs4919510 may modify cancer susceptibility in Chinese. More importantly, the results of our present meta-analysis together with the TCGA expression data provide evidence that SEMA4G may act as either a tumour suppressor gene or oncogene to regulate tumour development in a type-specific manner. However, further studies with experimental evaluations are warranted.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the National Natural Science Foundation of China (81502876, 81572259, 81272602, 81302011 and 81602021), the Natural Science Research of Jiangsu Higher Education Institutions (15KJB330006), the International Science and Technology Cooperation Program of China (No. 2014DFA31940), the Science and Technology Program of Nantong City (MS22015088), the Jiangsu Leadership Health Management Research Project (BJ15018) and the Science and Technology Program of Changshu Health and Family Planning Commission (csws201612). The funding sources had no role to play in the study design, the collection and interpretation of the data, writing of the report or decision to submit this paper for publication.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
