Abstract
Hepatocellular carcinoma is a leading cause of cancer-related mortality worldwide. TRAIL (tumor necrosis factor–related apoptosis–inducing ligand) is a potential target for cancer therapy. However, many cancer cells are resistant to TRAIL-induced apoptosis and its mechanism is not well understood. In this study, to identify potential therapeutic targets for TRAIL-resistant cancer cells, we compared the expression levels of interferon-stimulated gene 15 in TRAIL-sensitive and TRAIL-resistant hepatocellular carcinoma cell lines. Western blot analysis showed that interferon-stimulated gene 15 expression levels were significantly higher in resistant HLCZ01and Huh7 cells than in sensitive LH86 and SMMC-7721 cells. Interferon-stimulated gene 15 knockdown in resistance cells led to TRAIL sensitivity. Conversely, interferon-stimulated gene 15 overexpression in sensitive cells resulted in TRAIL resistance. Our bioinformatics search detected a putative target sequence for microRNA miR-138 in the 3′ untranslated region of the interferon-stimulated gene 15. Real-time quantitative polymerase chain reaction analysis demonstrated that miR-138 was significantly downregulated in TRAIL-resistant cells compared to TRAIL-sensitive cells. Forced expression of miR-138 in resistant cells decreased both messenger RNA and protein levels of interferon-stimulated gene 15, and when exposed to TRAIL, activated poly(adenosine diphosphate-ribose) polymerase, indicating sensitization to TRAIL. The results suggested that miR-138 regulates the interferon-stimulated gene 15 expression by directly targeting the 3′ untranslated region of interferon-stimulated gene 15 and modulates the sensitivity to TRAIL-induced apoptosis. MiR-138 may be a target for therapeutic intervention in TRAIL-based drug treatments of resistant hepatocellular carcinoma or could be a biomarker to select patients who may benefit from the treatment.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common malignancies, comprising approximately 90% of the 850,000 estimated new liver cancer cases diagnosed annually worldwide.1–3 TRAIL (tumor necrosis factor–related apoptosis–inducing ligand) displays potent anticancer effects in a wide range of cancers resistant to conventional therapy without apparent toxic side effects on normal cells.4–7 Binding of homotrimeric TRAIL to TRAIL receptors 1 (DR4) and 2 (DR5) induces oligomerization of the receptors and promotes formation of the death-inducing signaling complex (DISC), 8 resulting in activation of the initiator caspase-8, which ultimately leads to downstream activation of caspase-3 followed by initiation of the apoptosis program.9,10 Activated caspase-3 provokes cellular destruction by cleaving several hundred cellular proteins including poly(adenosine diphosphate-ribose) polymerase (PARP). A number of TRAIL-based novel drugs targeting TRAIL receptors are in preclinical or clinical development. 9 Nevertheless, resistance to TRAIL-induced apoptosis of cancer cells is a major impediment to the use of TRAIL-based agents as antitumor drugs. Hence, it is of great importance to investigate the congenital and acquired resistance mechanisms of TRAIL-induced apoptosis, as well as to identify potential molecules that may play key roles in resistance processes in order to enhance the cytotoxic effects of TRAIL.
Interferons (IFNs) are cytokines that protect cells from viral infections, modulate the immune response, and inhibit cancer cell growth. IFNs exercise their biological functions by regulating downstream ISGs (IFN-stimulated genes) expression. Many studies have identified specific alterations in the ISG15 (IFN-stimulated gene 15) pathway in human tumors, such as bladder cancer, 11 prostate cancer, 12 breast cancer, 13 pancreatic cancer, 14 and squamous cell cancer; 15 however, the biochemical function of ISG15 remains unclear.
Sedger 16 reported that microRNAs (miRNAs) were integral to the innate response to viral infections and were important components of IFN-mediated physiological processes, which prompted us to explore the possible involvement of miRNAs in TRAIL-induced apoptosis in cancer cells.
In this study, we investigated the relationship between ISG15 and miR-138 expression levels and the sensitivity to TRAIL-induced apoptosis in HCC cell lines. We showed that the expression of ISG15 is upregulated in TRAIL-resistant cells, while downregulated in sensitive cells and inversely correlated with MiR-138 expression. ISG15 knockdown or forced expression of miR-138 in resistant cells resulted in the restoration of TRAIL sensitivity, indicating a promising therapeutic strategy for the treatment of TRAIL-resistant HCC.
Materials and methods
The study was conducted in accordance with ethical standards according to the Declaration of Helsinki and national and international guidelines, and was approved by the authors’ institutional review board.
Cell lines
The human HCC cell lines Huh7, LH86, 17 HLCZ01, 18 and SMMC-7721 were grown in Dulbecco’s Modified Eagle’s Medium (DMEM) with 10% fetal bovine serum at 37°C in a humidified atmosphere containing 5% CO2.
Reagents and antibodies
The primary antibodies used for western blot analysis were polyclonal rabbit anti-human ISG15 antibodies (Abcam, Cambridge, MA, USA) and mouse anti-β-actin antibodies (Sigma, St Louis, MO, USA). Anti-PARP antibodies were obtained from Cell Signaling Technology (Danvers, MA, USA) and TRAIL from BD Pharmingen (San Diego, CA, USA).
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay
Cell viability was analyzed by the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay. Cells were seeded in 96-well plates at a density of 10 × 103 cells per well containing 100 µL of complete medium. After overnight incubation, various concentrations of TRAIL (1, 5, 10, and 20 ng/mL) were added to each well. Separately, liver cancer cells were treated with 10 ng/mL of TRAIL for 4, 6, 8, and 12 h. Cells treated with 0.1% bovine serum albumin (BSA) for the same time periods were used as the controls. Each treatment was performed in triplicate. After TRAIL treatment, 10 µL of MTT solution (5 mg/mL) was added to each well, and the plates were incubated for another 1 h at 37°C. Then the culture medium was discarded and replaced with 150 µL of dimethyl sulfoxide (DMSO) per well to dissolve the resultant formazan crystals. Absorbance (A) was measured spectrophotometrically in a microplate reader (Bio-Rad, Herclues, CA, USA) using 570 nm as the test wavelength and 630 nm as the reference wavelength. The inhibition ratio (%) was calculated as: (1 − experimental group A570–630/control group A570–630) × 100%.
Plasmid construction and transfection
The ISG15 expression plasmid was constructed by amplifying the entire open reading frame of the human ISG15 gene with the polymerase chain reaction (PCR) using p3XFLAG-CMV-14-ISG15 plasmid (kindly provided by Dr Douglas W. Leaman of the University of Toledo) as a template and inserted into the pcDNA3.1/V5-His expression vector (Invitrogen, Carlsbad, CA, USA). The primers for amplification of ISG15 were 5′-GATCACCCAGAAGATCGGCG-3′ (forward), and 5′-GGATGCTCAGAGGTTCGTCG-3′ (reverse). The oligonucleotides, which encode a 19-mer hairpin sequence specific to the ISG15 mRNA, were incorporated into the pSilencer-neo plasmid (Ambion, Naugatuck, CT, USA). The sequence of ISG12a shRNAs targeting regions of ISG15 was 5′-TTCGTCGCATTTGTCCACCA-3′. miR-138 mimics (5′-AGCUGGUGUUGUGAAUACAGGCCG-3′) were purchased from Genepharma (Suzhou, China). All constructs were confirmed by DNA sequencing and transfected into cells using Lipofectamine 2000 reagent (Invitrogen). The transfected cells were cultured in DMEM with 10% fetal bovine serum for 24–72 h before analysis.
Real-time quantitative polymerase chain reaction
The complementary DNA (cDNA) synthesis has been previously described. 19 Briefly, miR-138cDNA was synthesized from total RNA using the stem-loop RT primer (5′-GTCGTATCCAGTGCGTGTCGTGGAGTCGGCAATTGCACTGGATACGACCGGCCT-3′), and quantified by real-time quantitative polymerase chain reaction (qPCR) using the primers 5′-ACACTCCAGCTGGGAGCTGGTGTTGTGAATC-3′ and 5′-GTGCAGGGTCCGAGGT-3′. U6 cDNA was synthesized from total RNA using a stem-loop RT primer (5′-CGCTTCACGAATTTGCGTGTCAT-3′), quantified by real-time qPCR using the primers 5′-GCTTCGGCAGCACATATACAAAAT-3′ and 5′-CGCTTCACGAATTTGCGTGTCAT-3′, and employed as the internal control. Fold variations were calculated after normalization to U6. ISG15 was quantified by real-time qPCR using the primers 5′-GATCACCCAGAAGATCGGCG-3′ and 5′-GGATGCTCAGAGGTTCGTCG-3′. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was quantified by real-time PCR using the primers 5′-TTTGTCAAGCTCATTTCCTG-3′ and 5′-TGGTCCAGGGTTTCTTACTC-3′ and employed as the internal control.
Western blot analysis
Cells were washed twice with phosphate-buffered saline (PBS) and lysed in buffer containing 20 mM Tris–HCl (pH 7.4), 137 mM NaCl, 10% (wt/vol) glycerin, 1% Triton X-100, 2 mM ethylenediaminetetraacetic acid (EDTA), and a protein inhibitor cocktail (Roche, Basel, Switzerland) for 30 min on ice. The homogenate was centrifuged at 14,000 r/min for 20 min at 4°C and the supernatants were transferred to fresh tubes. Protein samples were separated using 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto a nitrocellulose membrane. The membranes were blocked with 5% nonfat dry milk containing 0.1% Tween-20 for 1.5 h at 37°C, and then probed with primary antibodies at 4°C overnight. After washing, the membranes were incubated with horseradish peroxidase (HRP)-conjugated secondary antibody followed by ECL detection (Amersham Pharmacia Biotech, Piscataway, NJ, USA). The membranes were scanned with a LAS-4000 luminescent image analyzer (Fujifilm, Tokyo, Japan). Subsequently, to evaluate equal protein loading, the membranes were striped in 20 mM glycine (pH 2.3) and reprobed with an anti-tubulin antibody (Sigma) followed by visualization with ECL reagents (Pierce, Rockford, IL, USA), according to the manufacturer’s instructions.
Statistical analysis
Statistical analyses were performed using the two-tailed Student’s t-test, and error bars represent the standard deviation (SD). Values of p < 0.05 were considered to be statistically significant.
Results
To understand better liver tumor biology, we examined the TRAIL sensitivity of the different human HCC cell lines, LH86, Huh7, HLCZ01, and SMMC-7721, respectively. Cells were exposed to TRAIL at concentrations of 1, 5, 10, and 20 ng/mL for 6 h, and cell death was assessed using the MTT assay (Figure 1(a)).

The cytotoxic effect of TRAIL on human HCC cell lines. (a) LH86, Huh7, HLCZ01, and SMMC-7721 cells were exposed to different concentrations of TRAIL (1, 5, 10, and 20 ng/mL) for 6 h. The cell viability was determined by MTT assay and expressed as a ratio to the blank control. (b) LH86, Huh7, HLCZ01, and SMMC-7721cells were treated with 10 ng/mL TRAIL for different time lengths (4, 6, 8, and 12 h). The cell viability is shown as in (a).
Non-TRAIL-treated cells in each cell line were used as blank controls and cell viability was expressed as a ratio of the control value. Cell viabilities in all cell lines decreased as the TRAIL concentration increased, demonstrating that TRAIL-induced apoptosis increased in a dose-dependent manner.
However, the reduction in cell viability decelerated at TRAIL concentrations greater than 10 µg/mL. Next, to determine the optimum treatment time, cells were treated with 10 µg/mL TRAIL for 4, 6, 8, and 12 h (Figure 1(b)). The MTT assay revealed that cell viability decreased with the length of treatment time, showing that TRAIL-induced apoptosis increased with treatment time. After 8 h, the decrease in cell viability reached a plateau. The TRAIL treatment also demonstrated sensitivity differences among the cell lines: the TRAIL-induced apoptosis rates of HLC-Z01 and Huh7 cells were <12% in both concentration- and treatment-time experiments, whereas LH86 and SMMC-7721 cells demonstrated higher apoptosis rates, reaching over 30% with higher concentrations and longer treatment times, indicating that LH86 andSMMC-7721 cell lines are more sensitive than Huh7 and HLCZ01 cell lines to TRAIL-induced apoptosis. The results demonstrated that the optimal concentration of TRAIL and the treatment time for HCC cells was 10 ng/mL and 8 h, respectively.
Higher levels of ISG15 expression correlates with TRAIL resistance
Although some cancer cells are resistant to TRAIL, the combination of TRAIL with other reagents significantly potentiates TRAIL-induced apoptosis in resistant cell lines, indicating that the TRAIL-mediated apoptotic pathway is intact in resistant cells.20,21
Therefore, we postulated that genes that are not involved in the TRAIL-induced apoptosis pathway might play a role in TRAIL resistance as well. ISG15 is highly induced by type I IFN in many cell types and increased expression of ISG15 impacts apoptosis.14,15 Western blot analysis showed that the steady state levels of ISG15 in the TRAIL-sensitive cell lines LH86 and SMMC-7721 were lower than those of the TRAIL-resistant cell lines Huh7 and HLCZ01 (Figure 2(a)). To assess the effect of enforced expression of ISG15 on the apoptotic processes, we delivered the plasmid pcDNA3.1-ISG15 into TRAIL-sensitive LH86 cells. As expected, overexpression of ISG15 significantly decreased TRAIL-induced PARP activation as indicated by the disappearance of the cleaved fragments in the cells (Figure 2(b)). Conversely, to examine the effects of reduced ISG15 expression, we constructed the short hairpin RNA (shRNA) plasmid pSilence-ISG15 shRNA designed specifically to silence ISG15, and TRAIL-resistant Huh7 cells were transfected. ISG15 shRNA apparently decreased ISG15 expression, while induced cleavage of PARP upon TRAIL treatment indicated sensitization of resistant Huh7 cells to TRAIL-induced apoptosis (Figure 2(c)). These results suggested that the expression level of ISG15 significantly influences the sensitivity of HCC cells to TRAIL-induced apoptosis in vitro.

ISG15 expression correlates with the sensitivity of cancer cells to TRAIL treatment in vitro. (a) Western bolt analysis of the expression of ISG15 in human HCC cell lines. The ISG15 levels were visualized (left panel) and quantified by densitometry and normalized to β-actin (right panel). (b) Forced expression of ISG15 in TRAIL-sensitive cells. LH86 cells were transfected with pcDNA3.1-ISG15, followed by TRAIL treatment. Cleavage of PARP and increased expression of ISG15 was visualized by western blot (left panel). The ISG15 and the cleaved PARP levels were quantified by densitometry and normalized to β-actin (right panel). (c) ISG15 knockdown in TRAIL-resistant cells. Huh7 cells were transfected with pSilencer-ISG15 shRNA or pSilencer-control shRNA, followed by TRAIL treatment for 8 h. Cleavage of PARP and reduced expression of ISG15 were visualized (left panel). The ISG15 and the cleaved PARP levels were quantified as in (b) (right panel).
miR-138 directly targets the 3′ untranslated region of ISG15
To understand the regulatory mechanisms of ISG15 expression in HCC cell lines, we performed a bioinformatics search (TargetScanHuman 5.2) and detected a putative microRNA target sequence in the 3′ untranslated region (UTR) of the human ISG15 gene that matched the seed sequence of human miR-138 (Figure 3(a)). MicroRNAs regulate the expression of many cellular proteins and are differentially expressed in cancer cells compared to normal cells. Our findings imply that miR-138 might target and regulate ISG15 expression. We examined the steady-state expression levels of miR-138 by real-time qPCR assay in four HCC cell lines and found that miR-138 was significantly upregulated in the TRAIL-sensitive cell lines LH86 and SMMC-7721 compared to the TRAIL-resistant cell lines Huh7 and HLCZ01 (Figure 3(b)).

MiR-138 directly targets the 3′UTR of ISG15. (a) The alignment of the seed region of miR-138 with the 3′UTR of ISG15. (b) The expression levels of MiR-138 were examined by real-time qPCR and normalized to U6. The data represented the mean values of three independent experiments. *p < 0.05 and **p < 0.01 indicated comparison with HLC-Z01 cells. (c) Huh7 cells were transfected with different concentrations of miR-138 mimics (10, 25, 50, 100, and 200 ng/mL) for 36 h. The expression levels of ISG15 mRNA were examined by real-time qPCR and normalized to GAPDH. The data represented the mean values of three independent experiments. *p < 0.05 and **p < 0.01 indicated comparison with NC-treated cells. (d) Huh7 cells were transfected with 100 ng/mL of miR-138 mimics for different time periods (24, 36, 48, and 72 h). The expression levels of ISG15 mRNA were examined by real-time qPCR and normalized to GAPDH. The data represented the mean values of three independent experiments. *p < 0.05 and **p < 0.01 indicated comparison with NC-treated cells. (e) Forced expression of miR-138. LH86 and Huh7 cells were transfected with 100 ng/mL of miR-138 mimics for 48 h, and the expression of ISG15 protein was analyzed by western blot assay. The ISG15 levels were visualized (right panel) and quantified by densitometry and normalized to β-actin (right panel).
To test whether enforced expression of miR-138 affects ISG15 expression, the miR-138 mimics plasmid was introduced into Huh7 cells. The expression level of ISG15 was markedly reduced in Huh7 cells transfected with miR-138 mimics as compared to cells transfected with the control plasmid (Figure 3(c) and (d)). As the concentration of miR-138 mimics used for transfection or duration of transfection was increased, ISG15 expression decreased further. The optimum effects were observed at 100 ng/mL of miR-138 mimics and transfection duration of 48 h. Moreover, we transfected TRAIL-sensitive LH86 as well as TRAIL-resistant Huh7 cells with 100 ng/mL miR-138 mimics. Western blot assay showed that overexpression of miR-138 markedly reduced the levels of ISG15 in both Huh7 and LH86 cells (Figure 3(e)). The result suggested that miR-138 modulates ISG15 expression by directly targeting the 3′UTR of ISG15.
MiR-138 modulates the sensitivity of cancer cells to TRAIL-induced apoptosis by targeting ISG15
To examine the effect of miR-138 forced expression on TRAIL sensitivity in HCC cells, we delivered miR-138 mimics into TRAIL-sensitive SMMC-7721 as well as resistant Huh7 cells. The forced expression of miR-138 significantly reduced the endogenous level of ISG15 in both SMMC-7721 and Huh7 cells. Accordingly, the forced expression of miR-138 further enhanced the TRAIL sensitivity in SMMC-7721 cells, as indicated by an increase in PARP cleavage. In Huh7 cells, it induced the activation of PARP as shown by the appearance of a cleaved PARP fragment, implicating a conversion from TRAIL resistance to sensitivity (Figure 4(a) and (b)). To corroborate further this observation, we delivered either miR-138 mimics or pcDNA3.1-ISG15 into TRAIL-sensitive LH86 cells at the same time. Forced expression of miR-138 in LH86 cells significantly reduced the endogenous level of ISG15, while it enhanced PARP activity when cells were treated with TRAIL (Figure 4(c)).
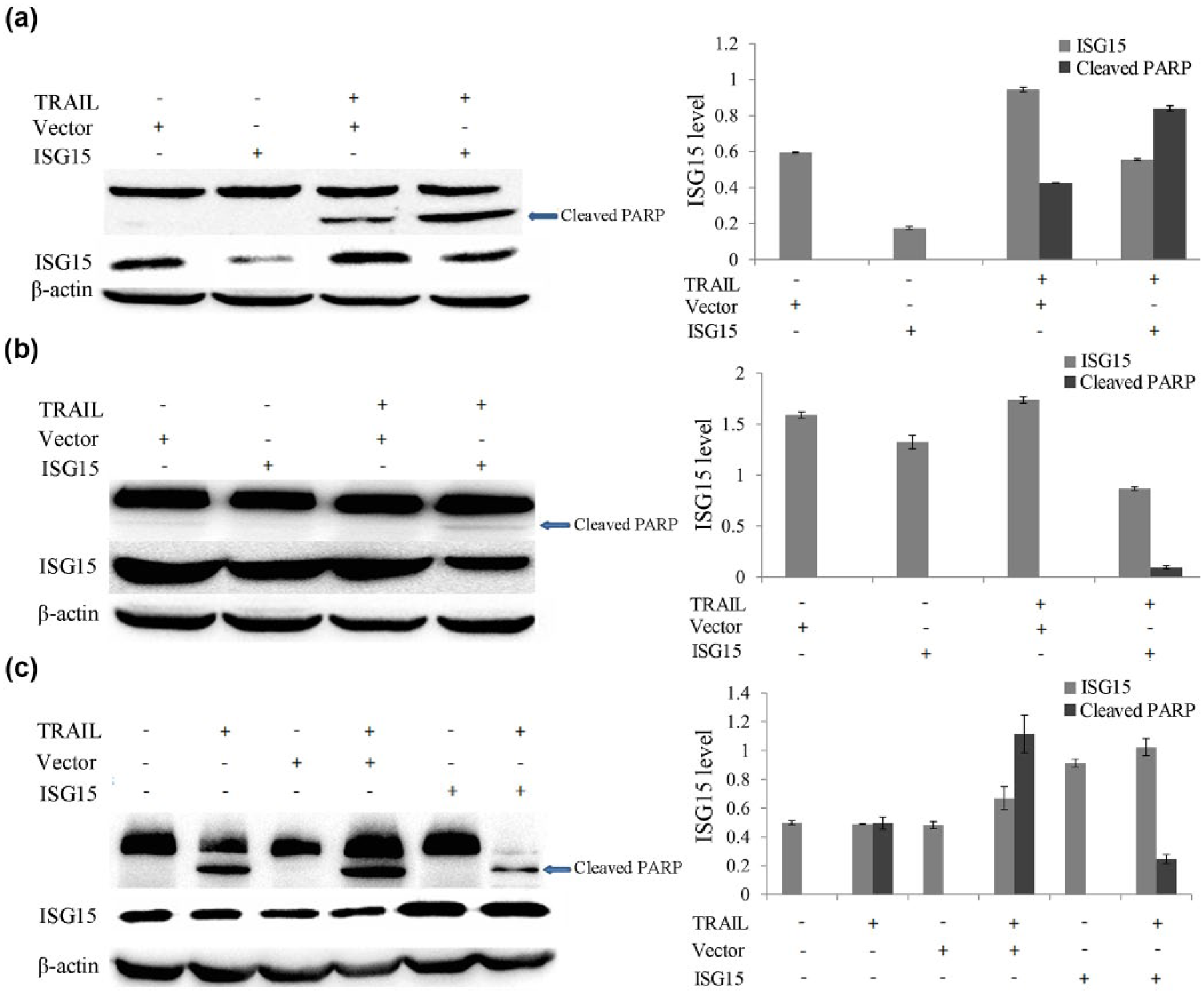
MiR-138 modulates the sensitivity of HCC to TRAIL-induced apoptosis by targeting ISG15. (a) Forced expression of miR-138 in TRAIL-sensitive cells. SMMC-7721 cells were transfected with miR-138, followed by TRAIL treatment for 8 h. Activation of PARP and expression of ISG15 protein were analyzed by western blot. A representative image is shown (left panel). The ISG15 and the cleaved PARP levels were quantified by densitometry and normalized to β-actin (right panel). (b) Forced expression of miR-138 in TRAIL-resistant cells. Huh7 cells were transfected with miR-138, followed by TRAIL treatment for 8 h. Activation of PARP and expression of ISG15 protein were analyzed by western blot. A representative image was shown (left panel). The ISG15 and the cleaved PARP levels were quantified and normalized as in (a) (right panel). (c) Forced expression of miR-138 and ISG15 in TRAIL-sensitive cells. LH86 cells were transfected either by miR-138 mimics or by pcDNA3.1-ISG15 at the same time, followed by TRAIL treatment for 8 h. Activation of PARP and expression of ISG15 protein were analyzed by western blot. A representative image is shown (left panel). The ISG15 and the cleaved PARP levels were quantified and normalized as in (a) (right panel).
In contrast, forced expression of ISG15 further increased the ISG15 level, and TRAIL-treated cells showed reduced PARP activity, indicating a conversion of the TRAIL-sensitive to the resistant phenotype. The result suggested that miR-138 modulates the sensitivity of HCC cells to TRAIL-induced apoptosis by inhibiting the expression of ISG15.
Discussion
HCC is one of the most common causes of cancer-related deaths worldwide. Hepatic resection, liver transplantation, and radiofrequency ablation are increasingly used as a curative treatment for HCC. However, only about 10%–20% of patients with HCC are currently eligible for these surgical interventions. Most patients are diagnosed at advanced stages and only receive palliative treatments; 22 however, only a few patients response well to palliative treatments. Therefore, the development of new therapeutic approaches is needed for a more effective treatment of HCC. One of the crucial factors affecting the survival rate of HCC patients is the resistance of cancer cells to therapeutic agents. Recombinant TRAIL, because of its highly specific toxicity for cancer cells, is among the most promising apoptosis-based antitumor agents.23,24 Therapy based on TRAIL is now undergoing phase 2 clinical trials in patients with different types of cancers. However, many cancer cells are resistant to TRAIL-induced apoptosis and the mechanism of TRAIL resistance remains to be established.
In an effort to identify potential therapeutic targets for TRAIL-resistant HCC cells, we found that TRAIL-resistant HCC cell lines expressed higher levels of the IFN-stimulated gene ISG15 compared to sensitive cell lines, and that a reduction of ISG15 expression in TRAIL-resistant cells restored sensitivity, while forced expression of ISG15 in sensitive cells resulted in TRAIL resistance.
miRNAs are small non-coding RNA molecules containing approximately 22 nucleotides that post-transcriptionally downregulate the expression of various target genes.25,26 Mature miRNAs are incorporated into the RNA-induced silencing complex (RISC). It subsequently serves as a complementary binding site for the 3′ UTR non-coding region of the target messenger RNA (mRNA), which leads to the degradation or translation inhibition of the mRNA, thus regulating the body’s biological functions.25–28 Many studies have shown that miRNAs are an inherent part of the innate immune system 16 and should be attractive potential drug targets since they not only regulate many cellular proteins but also are differentially expressed in malignant cells.25,26,29,30
Our bioinformatics search detected an miR-138 target sequence in the 3′UTR region of the human ISG15 gene, which implies that miR-138 regulates ISG15 expression by direct binding to the sequence, through which it may modulate the TRAIL-induced apoptosis pathway. Our data demonstrated that miR-138 is significantly upregulated in TRAIL-sensitive HCC cells compared to TRAIL-resistant cells, indicating that repression of miR-138 expression may be responsible for TRAIL-resistance. Further experiments demonstrated that forced expression of miR-138 in TRAIL-resistant cells resulted in ISG15 downregulation, suggesting that miR-138 might be a causal factor in the downregulation of ISG15 in cancer cells. The results indicated that miR-138 modulates TRAIL sensitivity in cancer cells by directly targeting ISG15. However, it is also possible that other targets of miR-138 might be involved in TRAIL resistance in cancer cells.
A limitation of our study was that first, the experiments were conducted only in vitro, which clearly could not verify the effect of ISG15 and miR-138 on the induced apoptosis of tumor cells by TRAIL in vivo. Second, the experiments only investigated the mechanisms of congenital resistance to TRAIL in HCC, but did not investigate the mechanisms of acquired resistance to TRAIL in HCC. Finally, the experiments only verified that the target gene of ISG15 was tested by miR-138 but there was no further study on the signal transduction pathways of ISG15 regulated by miR-138.
In summary, MiR-138 may be a target for therapeutic intervention in TRAIL-based drug treatment of resistant HCC. Also, it could be used as a novel indicator to select suitable cancer patients for treatment and to personalize medicine with new therapies designed for individual TRAIL-resistant tumors.
Footnotes
Acknowledgements
C.Z. and X.S. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by National Natural Science Foundation of China (grant number 81670103), The Development of Projects of Hunan Province Development and Reform Commission (grant number 2015-99), The Introduced Intelligence Project of Hunan Provincial Bureau of Foreign Experts (grant number 2016), and The Changsha City Third Science and Technology Project (grant number K1403380-31).
