Abstract
In recent years, abnormal RNA editing has been shown to play an important role in the development of esophageal squamous cell carcinoma, as such abnormal editing is catalyzed by ADAR (adenosine deaminases acting on RNA). However, the regulatory mechanism of ADAR1 in esophageal squamous cell carcinomas remains largely unknown. In this study, we investigated
Introduction
The incidence of esophageal cancer ranks eighth among cancers worldwide. 1 A total of 80%–85% of esophageal cancer cases occur in developing countries, including China. The majority of esophageal cancer cases in China are esophageal squamous cell carcinoma (ESCC), 2 and it has a remarkable geographic distribution. 3 Previous studies indicated that ethnic and genetic factors as well as lifestyle play roles in ESCC development. The incidence of esophageal cancer is threefold higher in men than in women. 4 From a molecular oncology perspective, more than 83% of ESCCs contained a somatic mutation in TP53. 5 Over-expression of epidermal growth factor receptor (EGFR) occurs in 59.6%–76% of ESCCs and is associated with a poor prognosis.6,7 In addition, the purported risk factors for ESCC include a low consumption of fruits and vegetables; deficiency of selenium, zinc, and vitamin E; high level of exposure to areca nuts or polycystic aromatic hydrocarbons; and poor oral hygiene.8,9 The prognosis and 5-year overall survival of patients with ESCC remain very poor. 10 The above reports suggest that the progression of ESCC is complex. Therefore, identifying additional molecular mechanisms involved in ESCC is of utmost importance.
The family of adenosine deaminases acting on RNA (ADAR), including ADAR1, ADAR2, and ADAR3, is characterized by the high conservation of the C-terminal region, which functions in deaminase catalysis, and an N-terminal region that can combine with dsRNA.
11
ADAR1 and ADAR2 have the capacity to catalyze RNA editing, while previous reports suggest that ADAR3 cannot.12–14 RNA editing is discovered by a common post-transcriptional modification to change the sequence of nucleotides that generated various proteins.15,16 In recent years, several papers revealed that RNA editing appears to be specific to cell types or cell environments. Both hyper- and hypo-editing of coding regions can contribute to human diseases, including a variety of cancers.17,18 For example, hyper-editing of antizyme inhibitor 1 (
Here, we performed integrated experiments to detect the relationships between
Materials and methods
Cell lines and clinical samples
Seven Japanese ESCC cell lines (KYSE30, KYSE70, KYSE150, KYSE180, KYSE410, KYSE450, and KYSE510) were cultivated in 50% RPMI 1640 medium (HyClone, Logan, UT, USA) and 50% F12 medium (HyClone), mixed with 10% fetal bovine serum (Gibco, New York, NY, USA), 100 IU/mL penicillin and 100 IU/mL streptomycin (HyClone). HET-1a, a cell line derived from normal esophageal squamous epithelium cells, was maintained in BEGM (Bronchial Epithelial Cell Growth Medium) with several growth factors (Lonza, Basel, Switzerland) added. All cell lines were confirmed by short tandem repeat (STR), and mycoplasma contamination was monitored.
A total of 15 paired primary ESCC tumor tissues and their matched non-tumor tissues were obtained from the Cancer Institute and Hospital of the Chinese Academy of Medical Science between 2007 and 2008 for DNA and RNA extraction (Table S1). All clinical samples used in this study were approved by the medical ethics committee of the National Cancer Center/Cancer Hospital.
IFN and inhibitor treatment
Human ESCC cell lines seeded in six-well plates were treated with the following doses of IFN: 1000 U/mL of Recombinant Human IFN-alpha A/D (
Cells seeded in six-well plates were treated with S-ruxolitinib, inhibiting JAK1 expression (Selleckchem, Houston USA), as well as s31-201, inhibiting STAT3 expression (Selleckchem, Houston, USA). After being treated with inhibitors for 24 h, cells were mixed with IFN for culture.
DNA and RNA preparation
DNA from tumor and matched normal fresh-frozen tissues as well as from cell lines was isolated using the Qiagen QIAamp DNA Mini Kit (Qiagen, Venlo, Netherlands). The total RNA was isolated from cultured cell lines or clinical samples using TRIzol reagent (Invitrogen, Carlsbad, CA, USA), and the complementary DNA (cDNA) was prepared with a Fermentas RevertAid™ Premium First Strand cDNA Synthesis Kit (Thermo, Rockford, IL, USA) according to the manufacturer’s protocol.
Quantitative real-time polymerase chain reaction
Quantification of
Western blot analysis
The cultured cell lines with and without IFN treatment were lysed by radioimmunoprecipitation assay (RIPA) buffer mixed with protease and phosphatase inhibitor mixtures (Thermo). Equal amounts of proteins were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) electrophoresis, transferred to nitrocellulose filter membranes (Whatman GmbH, Maidstone, Kent, UK) and blocked with Tris-buffered saline (TBS) buffer containing 5% nonfat milk for 1 h at room temperature. The membranes were incubated overnight at 4°C with primary antibodies and then washed with Tris-buffered saline with Tween 20 (TBST) buffer with three times. After washing, the membranes were incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature and then washed in TBST buffer three times. The protein bands were detected using enhanced chemical luminescence (Millipore, Billerica, MA, USA), and the immunoblots were quantified using ImageJ software. The following antibodies were used: mouse anti-ADAR1 antibody (1:480, ab88574; Abcam), mouse anti-β-actin antibody (1:2000, ab8227; Abcam), rabbit anti-STAT1 antibody (1:1000, CST, #14994), anti-P-STAT1 antibody (1:1000, CST, #7649), anti-STAT2 antibody (1:1000, CST, #72604), and anti-P-STAT2 antibody (1:1000, CST, #88410).
High-throughput sequencing
DNA exome sequencing of clinical samples was performed by BGI (Shenzhen, China), while DNA exome sequencing of cultured cell lines was performed by WuXi AppTec (Wuxi, China). RNA sequencing (RNA-seq) of clinical samples was completed by BGI, and RNA-seq of cultured cell lines with and without IFN treatment was completed by Novogene (Beijing, China).
RNA editing analysis
First, duplicate reads were removed using Picard MarkDuplicates based on BAM files generated from RNA-seq data alignment, as was done for the exome data. The preliminary list of SNPs in RNA was generated using SAMtools (version 0.1.19), and the DNA genotypes for those SNPs were determined. The putative RNA editing sites were obtained by comparing the DNA and RNA sequences. To reduce the false-positive sites, we selected RNA editing candidate sites using the following criteria: (1) the site was supported by more than 10 mapped reads in RNA-seq data, and those reads were not duplicative; (2) the RNA mutant genotype was supported by more than two unique sense and antisense strand mapped reads, separately; (3) at least 10% of the RNA mapped reads supported the mutant allele; and (4) the site was homozygous in the corresponding DNA data and supported by more than 10 uniquely mapped reads. In addition, we excluded all RNA variants detected at HLA genes empirically. If the fold change of editing degree between tumor and normal tissue was more than two, a given RNA editing site was defined as RNA overediting in ESCC tumors. In view of previous papers showing reduced accuracy of the identification of non-A→G and T→C RNA editing using transcriptome data because of mapping errors, only A→G and T→C editing sites were retained for the RNA editing analysis.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 6.0 software. Data are presented as the mean ± standard error of the mean (SEM), which was obtained from triplicate experiments. Student’s t-test was used to estimate the significance of difference; p < 0.05 was considered significant.
Results
ADAR1 was significantly over-expressed and abnormal RNA editing sites were detected in primary ESCC tumors
First, we found

Then, we analyzed A-to-I RNA editing by comparing DNA and RNA sequences from the ESCC tumor and normal samples. We observed 815 RNA editing events in 15 ESCC tumors, with a total of 1353 such events in all samples, including both tumor and normal RNAs. Of these, only a fraction of RNA editing sites occurred both in tumor and in normal tissue for a given patient (Figure 1(e)). Furthermore, 609 events could be classified as RNA hyper-editing in tumors when compared with RNA editing loci in matched normal tissues (Table S2; Figure 1(f)). Specific RNA editing sites, such as
ADAR1 expression was correlated with the JAK/STAT pathway in ESCC
We aimed to determine how

The expression of
Type I IFN regulated ADAR1 expression through the JAK/STAT pathway
Seven ESCC cell lines and one normal esophageal squamous epithelium cell line, named HET-1a, were treated with type I IFN-α or IFN-β to confirm the causality of ADAR1 expression induced through such molecule. The results of quantitative real-time PCR showed that
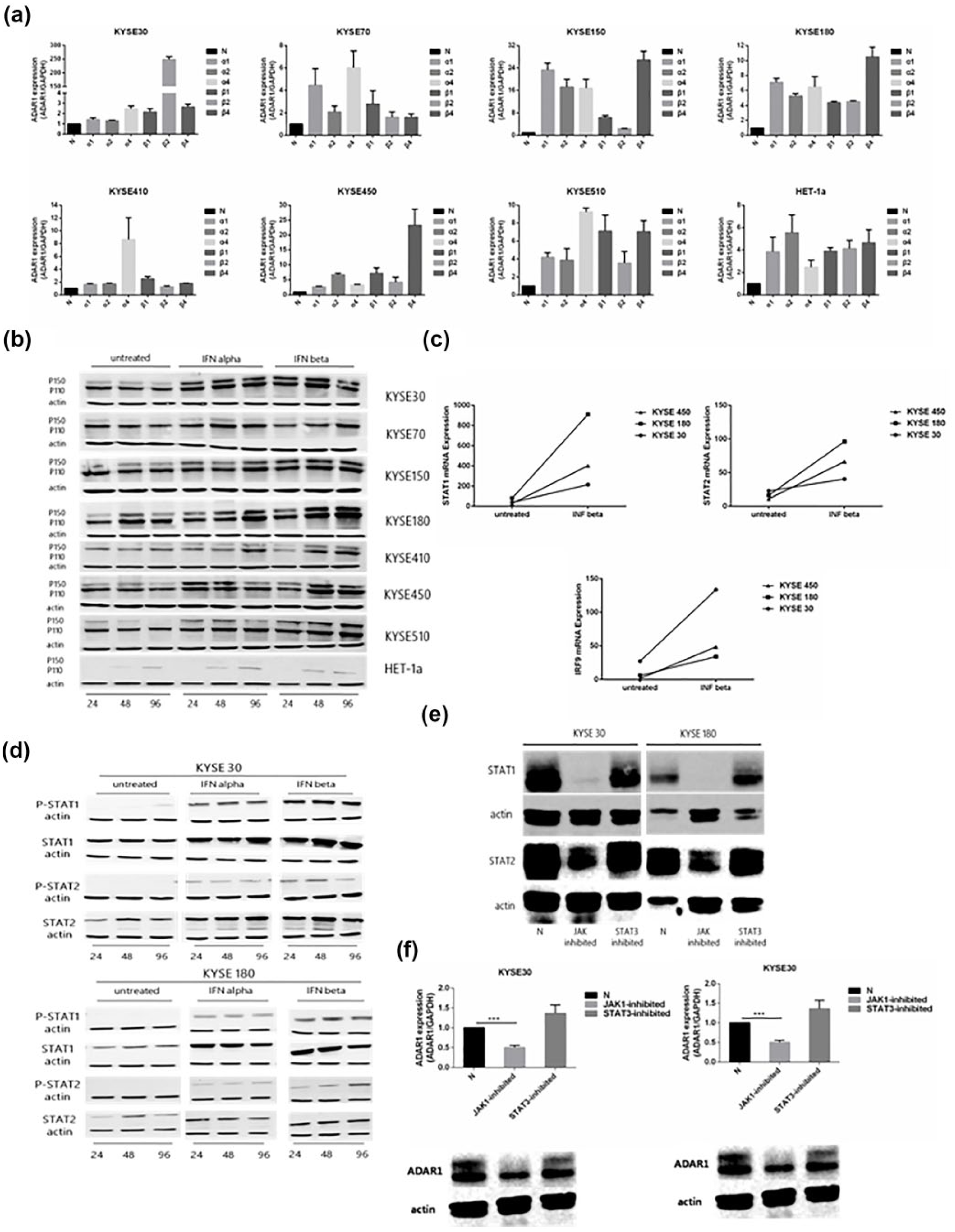
Type I interferon regulated ADAR1 expression through the JAK/STAT pathway. (a)
However, how type I IFN regulates ADAR1 expression remains unclear. Therefore, we cultured ESCC cell lines with type I IFN for 72 h and sequenced their transcriptome. The RNA-seq analysis showed the mRNA expression levels of
The JAK/STAT pathway induced A-to-I mRNA editing in ESCC cell lines through the upregulation of ADAR1 expression
The above evidence illustrated that type I IFN can affect ADAR1 expression through the JAK/STAT pathway. Next, we determined RNA editome using RNA-seq compared with IFN-β-treated and normal ESCC cells of KYSE30, KYSE180, and KYSE450. First,

Type I interferon response affected A-to-I mRNA editing in ESCC through ADAR1 induction. (a)
Finally, we analyzed the edited degree of specific A-to-I mRNA editing sites and found that type I IFN conferring an increasing level of those A-to-I editing sites in three ESCC cell lines:
Discussion
The ADAR family includes three members: ADAR1, ADAR2, and ADAR3. ADAR1 and ADAR2 express in most human tissues, while ADAR3 was detected only in the brain. 26 ADAR1 and ADAR2 can catalyze RNA editing, but ADAR3 does not have such a function.12–14 Previous studies reported that all three ADAR enzymes were downregulated in glioblastoma, conferring a proliferation rate increase through the reduction of A-to-I RNA editing. 27 Nevertheless, Chan et al. 21 found that ADAR1 upregulation and ADAR2 downregulation resulted in a disturbance of A-to-I RNA editing, which participated in the progression of HCC. Moreover, the over-expression of ADAR1 was detected in non–small cell lung cancer 28 and breast cancer, 29 while the upregulation both of ADAR1 and ADAR2 was confirmed in prostate cancer. 30 In this study, we observed over-expression of ADAR1 in ESCC tissues compared to their matched non-tumor tissues, but no differences were observed in ADAR2 between ESCC tissues and their matched non-tumor tissues.
Previous reports revealed the relationship between ADAR1 expression and cancer, and the mechanisms for the regulation of ADAR1 also emerged. Qin et al.
20
reported that the increase in
Not only is the mechanism of ADAR1 regulation unclear, but the target genes of ADAR1 can also be highly complex. A-to-I RNA editing mediated by ADAR is the change of nucleotide sequence of RNA transcripts relative to that of the encoding DNA and can enhance the RNA and protein diversity.
33
A-to-I RNA editing has two types: special site editing and hyper-editing.
34
A type of A-to-I RNA editing was found to occur in the coding region, leading to a change in nucleotide sequence,
35
and affecting the progression of cancer. For example, the chloride channel GABRA3 is necessary and sufficient for invasion and metastasis of breast cancer. A-to-I editing on the
However, there are several questions that remain unanswered. We observed a correlation between ADAR1 expression and JAK/STAT pathway-related genes, but how STAT1 and STAT2 regulate ADAR1 activation is unclear. Modulation of editing at an individual site is entangled with many processes, and the mechanism of ADAR1 regulation could be very important. The widespread editing induced by ADAR1 in combination with the conservation of editing sites across tissues and patients suggests that there may be clinical and therapeutic implications for a wide range of cancer patients. Some RNA editing events may be served as “driver” mutations, and further efforts are required to characterize the function of clinical relevant RNA editing in ESCC.
Footnotes
Acknowledgements
We thank Fang Zhou for the careful work in collecting information on ESCC patients. We are also grateful to Meihua Xiong and all the staff in the Lab and Tumor Bank of the Department of Thoracic Surgery for their support during the study.
Compliance with ethical standards
Samples were obtained with informed consent, and the study was approved by the medical ethics committee of the National Cancer Center/Cancer Hospital.
Data deposition
The sequencing data supporting findings in this manuscript can be found in the European Genome-Phenome Archive (EGA) under accession EGAS00001001731.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by National Natural Science Foundation of China (81502514, 81402300), The National Key Basic Research Development Plan973 Plan (2015CB553901), and CAMS Innovation Fund for Medical Sciences (CIFMS) (2016-I2M-1-001)
