Abstract
Long non-coding RNAs have been revealed to play important roles in the progression of hepatocellular carcinoma. However, the detailed mechanisms underlying their activities are not fully understood. Using microarray technology, a number of long non-coding RNAs were previously identified to be aberrantly expressed in hepatocellular carcinoma. In this study, one of these long non-coding RNAs, designated lncRNA-PE (lncRNA promotes epithelial–mesenchymal transition), was further explored to study its expression profile and function. A cohort of human hepatocellular carcinoma tissue samples combined with benign controls and established human hepatocellular carcinoma cell lines were examined for the expression of lncRNA-PE. The biological functions of lncRNA-PE were examined by wound-healing and Transwell assays, which revealed that lncRNA-PE promotes cell invasion and migration. By detecting the level of epithelial–mesenchymal transition markers, lncRNA-PE was revealed to promote epithelial–mesenchymal transition in hepatocellular carcinoma cells. Further study suggested that lncRNA-PE downregulated miR-200a/b by repressing the primary transcript expression, enhanced ZEB1 expression, and promoted epithelial–mesenchymal transition of hepatocellular carcinoma cells. All these data imply that lncRNA-PE might play an important role in hepatocellular carcinoma development via the miR-200a/b-ZEB1 pathway.
Introduction
One of the most common and aggressive human malignancies, 1 hepatocellular carcinoma (HCC), has a poor prognosis and a high recurrence rate, largely due to its high rate of metastases. 2 Improvements in early detection are urgently needed, as is the identification of new therapeutic targets. 3 Metastasis of HCC cells is known to be influenced by many cell-specific and microenvironmental factors, 4 although the underlying molecular mechanisms of metastasis have not been completely elucidated. The epithelial–mesenchymal transition (EMT) has been proved to be an important early event of tumor cell metastatic dissemination, in which cells are endowed with a more motile, invasive potential. 5 The microRNA 200 (miR-200) family regulates the epithelial phenotype by targeting ZEB1 messenger RNA (mRNA), and changes in miR-200 expression may be an important step in tumor progression. 6
Long non-coding RNAs (lncRNAs) are RNAs longer than 200 nucleotides with limited protein coding potential and have been shown to regulate a variety of cellular processes,7,8 such as cell proliferation, apoptosis, cell migration,9,10 and tumor metastasis. 11 Recently, the specific roles of lncRNAs in regulating EMT are being studied by increasing numbers of researchers; 12 however, further studies on EMT-related lncRNAs need to be carried out to improve our understanding of their roles in the invasion-metastasis cascade of HCC. Thus, in this study, we aimed to explore the expression level and the functional roles of a novel EMT-related lncRNA in HCC and to evaluate its potential as a new diagnostic marker for HCC.
Materials and methods
GEO data set analysis
Gene expression data (GSE45267, GSE55092, and GSE40873 profiling data) were downloaded as raw signals from Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo), interpreted, normalized, and log2 scaled using software R.
Tissue samples and cell lines
Human HCC cell lines BEL-7402 and SK-Hep-1, as well as the normal human liver cell line LO2, were obtained from the American Type Culture Collection (ATCC, USA) and were cultured at 37°C in an atmosphere containing 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM) plus 10% fetal bovine serum (FBS; Gibco, USA).
HCCs and surrounding control tissue specimens were obtained from 24 patients at General Hospital of PLA (Beijing, P.R. China) after surgical resection with informed consent. The tumor tissues and adjacent normal tissues were frozen in liquid nitrogen after resection. No patient in this study received chemotherapy or radiation therapy before the surgery. Liver samples were fully clinically characterized (Table 1). This study was performed with the approval of the Medical Ethical Committee of General Hospital of PLA, which were previously used in the other study. 13
Patient clinical features.

HCV-ab: hepatitis C virus antibodies; HBsAg: hepatitis B surface antigen.
Clinicopathological characteristics of the patients with HCC are presented. All samples were obtained during surgery.
RNA extraction and real-time polymerase chain reaction
Total RNA of cells and tissues was isolated using TRI Reagent (Sigma, USA). First-strand complementary DNA (cDNA) was generated using the ImProm-II Reverse Transcription System (A3800; Promega, USA) and random primers or oligo-dT primers. We used SYBR Premix Ex Taq (RR420A; TaKaRa, Japan) and gene-specific primers for reverse transcription polymerase chain reaction (RT-PCR). The gene-specific primers are shown in the following. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was employed as an endogenous control for mRNA and lncRNA. For microRNA (miRNA) analysis, quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed as described before 14 and U6 small nuclear RNA (snRNA) was used for normalization. The relative expression levels of RNAs were calculated using the comparative Ct method.
Primers used for real-time PCR are as follows: lncRNA-PE-F: 5’AGGATAGGATGGGCTTTTTCTGT-3’; lncRNA-PE-R: 5’-GCAATTGCCCTAGACCCAGT-3’; pri-miR-200b-F: 5’-CAGCCGTGGCCATCTTACT-3’; pri-miR-200b-R: 5’-CCGCCGTCATCATTTACCAGG-3’; GAPDH-F: 5’-TCAGTGGTGGACCTGACCTG-3’; GAPDH-R: 5’-TGCTGTAGCCAAATTCGTTG-3’; U6-F: 5’-CGCTTCGGCAGCACATATACTA-3’; U6-R: 5’-CGCTTCACGAATTTGCGTGTCA-3’; miR-200b: 5’-TAATACTGCCTGGTAATGATGA-3’; miR-200a: 5’-CATCTTACCGGACAGTGCTGGA-3’; and miR-451a: 5’-AAACCGTTACCATTACTGAGTT-3’.
Small interfering RNAs and miRNAs
Small interfering RNAs (siRNAs) for lncRNA promoting EMT (lncRNA-PE) were purchased from GenePharma (China), the sequences of siRNAs are as follows: silncRNA-PE-1: 5’-CCGGCCCAUAUGUUGAAAUTT-3’ and 5’-AUUUCAACAUAUGGGCCGGTT-3’; silncRNA-PE-2: 5’-CCAUGCGAACACACAGCAATT-3’ and 5’-UUGCUGUGUGUUCAUGGTT-3’. For miR-200a/b overexpression studies, 20 nM miR-200a/b oligonucleotide duplex (miRBase Release 21) was purchased from GenePharma.
RNA immunoprecipitation
Cells were harvested using phosphate-buffered saline (PBS) and lysed in immunoprecipitation (IP) lysis buffer (Life Technologies, USA) containing an RNase inhibitor (RRI, 0.2 U/µL; TaKaRa, Japan) and protease inhibitors (Roche, Switzerland) for at least 10 min at 4°C. To perform IP experiments, the supernatants of lysates were first incubated with antibodies for 4 h at 4°C, and then, beads were added into the mixture and rotated overnight at 4°C. All precipitates were washed thrice with a wash buffer (20 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), 150 mM NaCl, 5 mM MgCl2, and 1% NP40) and then boiled with sodium dodecyl sulfate (SDS)-loading buffer for 10 min. For RNA IP, RNAs were extracted by mixing the TRI Reagent with beads and then subjected to reverse transcription and real-time PCR analysis.
Luciferase assay
Dual-luciferase assays were performed according to the Dual-Luciferase Reporter Assay System guidelines (Promega, USA). Cells cultured in 24-well plates were incubated with passive lysis buffer (Promega) for 15 min at room temperature. Cell lysates were transferred into EP tubes together with Luciferase assay reagent II (LAR II), and an equal volume of the Stop&Glo Reagent was added into the EP tubes. The fluorescence signals of cell lysates were analyzed using a GloMax Luminometer (Promega). The data are presented as means ± SD of three independent experiments.
Cell migration and invasion assay
In the wound-healing assay, cells were scratched in the monolayer and cultured under normal conditions. The migrated distances were measured at 0, 24, and 48 h after scratching. For the invasion assay, BEL-7402 and SK-Hep-1 cells were serum starved for 24 h in DMEM containing 0.1% FBS. Serum-starved cells were resuspended in DMEM containing 0.1% FBS, and 2 × 105 cells were added to the upper chamber of each well (6.5 mm in diameter, 8-µm pore size; Corning, USA) coated with 30 mg/cm2 Matrigel (ECM Gel; Sigma). Medium containing 0.1% FBS and supplemented with hepatocyte growth factor (HGF; 20 ng/mL) was placed in the lower compartment of the chamber. After 24 h at 37°C, cells on the upper membrane surface were removed by careful wiping with a cotton swab and the filters were fixed by treatment with 95% ethanol for 30 min and stained with 0.2% crystal violet solution for 30 min. Invasive cells adhering to the lower surface of the filter were then counted (five high-power fields/chamber) using an inverted microscope. The migration assay is the same as the invasion assay, except that no Matrigel was used and the permeating time for cells was 12 h.
Western-blot analysis
Cells were lysed with radioimmunoprecipitation assay (RIPA) buffer containing protease inhibitors. After bicinchoninic acid (BCA) quantification, equal amounts of protein were separated by 10% SDS–polyacrylamide gel electrophoresis (PAGE), electrophoretically transferred to nitrocellulose membrane (Millipore, USA), and incubated with primary antibodies against the target proteins followed by appropriate secondary antibodies (Pierce, USA). Signals were visualized with SuperSignal® West Pico Chemiluminescent Substrate (Pierce) and film exposure.
Antibodies for western blotting
For western blot, antibodies against ZEB1 (sc-10570; Santa Cruz Biotechnology, USA), E-cadherin (sc-8426; Santa Cruz Biotechnology), vimentin (sc-6260; Santa Cruz Biotechnology), N-cadherin (sc-7939; Santa Cruz Biotechnology), ZO-1 (sc-10804; Santa Cruz Bio-technology), and GAPDH (10494; Proteintech, USA) were used.
Vector construction
The cDNA encoding lncRNA lncRNA-PE was PCR-amplified by KOD Plus Neo DNA Polymerase (Toyobo, Japan) and subcloned into the pcDNA3 vector (Invitrogen, USA). The promoter region of miR-200a/b was PCR-amplified and subcloned into BglII and NheI sites of pGL3-enhancer (Promega) vector. Primers used for the construction of vectors are as follows: lncRNA-PE-KpnI: 5’-AGCTGGTACCGACGGAGTCTCGCTCTGTCGCCCA-3’; lncRNA-PE-XhoI: 5’-CGCACTCGAGATCATTTAATTTTCCACAATTGAT-3’; miR-200a/b-promoter-NheI: 5’-ACCTGCTAGCCATTCCCGGACAGTCACAG-3’; and miR-200a/b-promoter-BglII: 5’-CAGGAGATCTCCTGGCACAGGAAGTCAGTT-3’.
Isolation of cytoplasmic and nuclear RNA
Cytoplasmic and nuclear RNA were isolated and purified using the RiboTrap Kit (RN1011; MBL, Japan) according to the manufacturer’s instructions.
Statistical analysis
Microsoft Excel software and GraphPad Prism 5 were used to assess statistical differences between experimental groups. The MO-free survival (MFS) was evaluated by the survival pack of R. The median expression level for each gene was used as a cutoff value calculated by Cutoff Finder. 15 Statistical significance was analyzed using t tests, and p < 0.05 was considered statistically significant (*p < 0.05, **p < 0.01, and ***p < 0.001).
Results
LncRNA-PE is upregulated in HCC tissues
We sought to screen for lncRNAs that can promote the progression of HCC and EMT. By combining microarray data from our previous study with data in the article that presented the discovery of lncRNA-atb, 12 we serendipitously uncovered a lncRNA, BC013423 (1454-bp long). It is upregulated both in HCC 16 and upon transforming growth factor-beta (TGF-β; treatment. 12 We inferred that this lncRNA may have a role in promoting HCC progression and EMT and sought further validation. For convenience, we call this lncRNA lncRNA-PE in the later parts of this article. First, we used the online database Coding Potential Calculator 17 to calculate its coding potential, which was 0.663685, compared with the score of lncRNA SNHG1, 0.951527, and that of MALAT1, 0.348632, which indicated that it had very weak coding potential. To determine the role of this lncRNA in HCC progression, we first verified the expression of lncRNA-PE in HCC cells, which was increased in two HCC cell lines relative to the expression in the normal human liver cell line LO2 (Figure 1(a)). We next analyzed its expression level in HCC tissues compared with their normal counterparts in GSE45267 and GSE55092 and discovered that lncRNA-PE was upregulated in HCC tissues (Figure S1(A) and (B)). Then, we validated its expression levels in 24 pairs of HCC tissues and adjacent liver tissues using quantitative real-time PCR. The lncRNA-PE level was significantly upregulated (>2 fold) in 50% of HCC tissues (12 of 24 patients; Figure 1(b)) compared with adjacent normal tissues. Moreover, Figure S1(C) indicates that the cumulative recurrence-free survivals were significantly associated with lncRNA-PE expression in GSE40873 (log-rank test, p = 0.014). These data indicate that ectopic lncRNA-PE expression may be related to HCC progression and EMT, and lncRNA-PE could be a potential HCC prognosis maker.
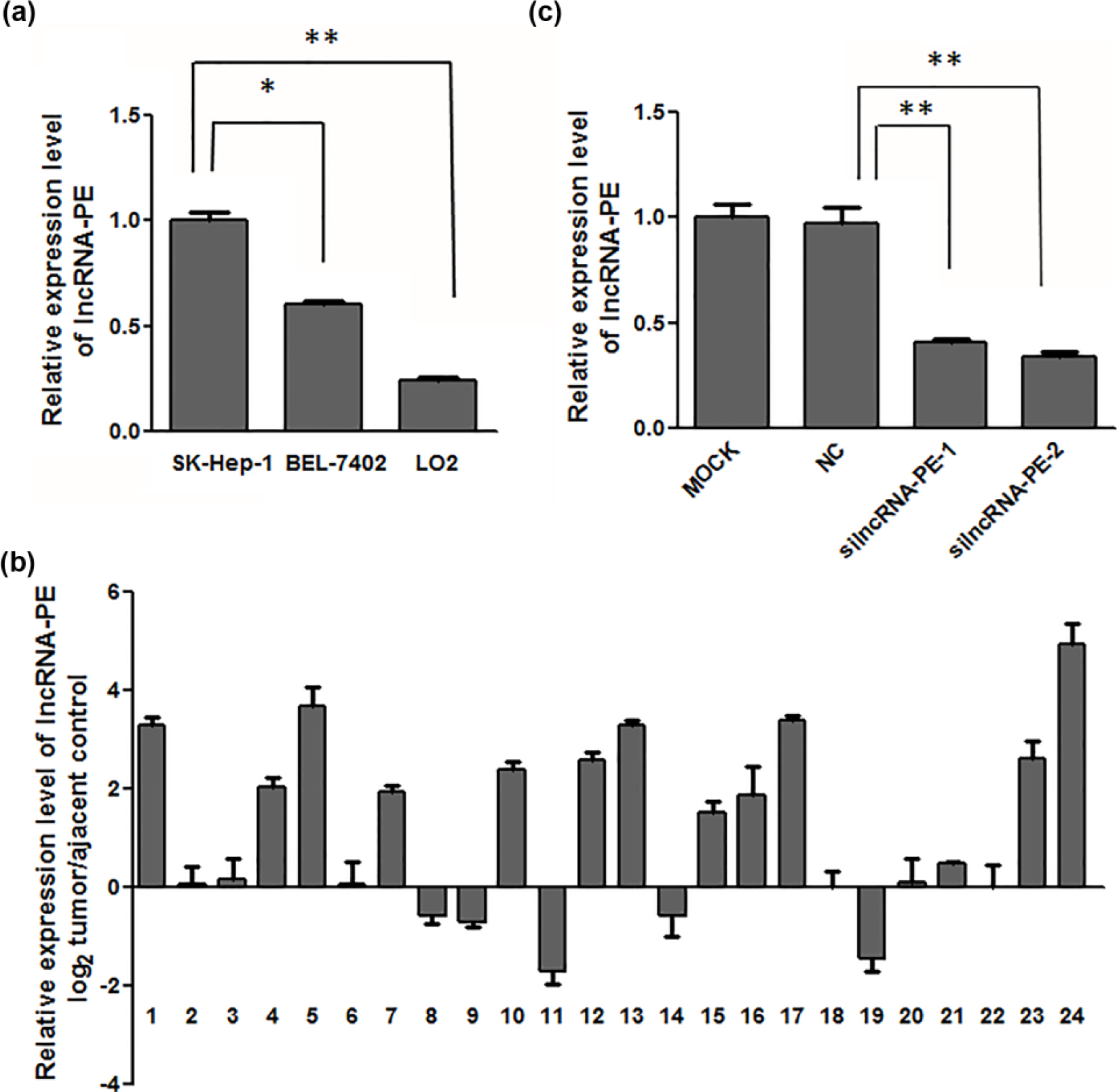
LncRNA-PE levels were upregulated in hepatocellular carcinoma cells and tissues. (a) LncRNA-PE expression was assessed by real-time PCR in hepatocellular carcinoma cell lines and a normal liver cell line. lncRNA-PE levels were normalized to GAPDH levels (*p < 0.05). (b) LncRNA-PE expression was assessed in hepatocellular carcinoma tissues and adjacent normal tissues (n = 24). LncRNA-PE expression was assessed by real-time PCR, and values were normalized to those for GAPDH. Bars represent the ratio between expression in adjacent normal and hepatocellular carcinoma tissues (log scale). (c) The mRNA levels of lncRNA-PE after lncRNA-PE siRNAs were transfected in SK-Hep-1 cells.
LncRNA-PE promotes HCC cell migration and invasion
To explore the role of lncRNA-PE in cell migration and invasion, we hypothesized that there would be a loss-of-function effect if this upregulated lncRNA was inhibited by silencing RNAs (siRNAs). As shown in Figure 1(c), lncRNA-PE levels were significantly decreased in SK-Hep-1 cells transfected with siRNA against lncRNA-PE (Figure 1(c)). Moreover, downregulation of lncRNA-PE significantly inhibited SK-Hep-1 cell migration and invasion (Figure 2(a) and (b)), whereas overexpression of lncRNA-PE in BEL-7402 cells was found to have the opposite effect (Figure S2). These data suggest that lncRNA-PE can promote cell motility and invasion in HCC.

LncRNA-PE knockdown decreased hepatocellular carcinoma cell invasion and migration. (a) Wound-healing assay used to detect cell migration changes in hepatocellular carcinoma cells. Decreased cell migration was observed in SK-Hep-1 cells after knocking down lncRNA-PE with siRNAs for 2 days, and columns in each groups were normalized with the column of 0 h. (b) Cell migration and invasion assays. For the cell migration assay, SK-Hep-1 cells were transfected with lncRNA-PE siRNAs and normal control RNA and then subjected to Transwell migration assays as described. After 12 h, migrating cells were counted after staining with crystal violet. Migrating cell numbers are the average count of three random microscopic fields. Each bar represents the mean ± SD of the counts from a single representative experiment (*p < 0.05). For the cell invasion assay, cells transfected with lncRNA-PE siRNAs and normal control RNA were subjected to Transwell invasion assays as described. After incubation for 24 h, cells that invaded through the pores to the bottom surface of the membrane were fixed, stained, and counted. Invading cell numbers are the average count of three random microscopic fields. Each bar represents the mean ± SD of the counts from a single representative experiment (*p < 0.05).
LncRNA-PE induces EMT of HCC cells
EMT has been proved to be important in the early events of tumor cell metastatic dissemination by endowing cells with more motility and potential for invasiveness. We therefore investigated whether lncRNA-PE regulates EMT. The expression levels of the epithelial marker (E-cadherin and ZO-1) and the mesenchymal marker (N-cadherin and vimentin) were analyzed by western blot. The results revealed that knockdown of lncRNA-PE reduced N-cadherin and vimentin, whereas induced E-cadherin and ZO-1 at the protein levels (Figure 3(a)). Consistent with this, overexpression of lncRNA-PE had the opposite effects (Figure 3(b)).
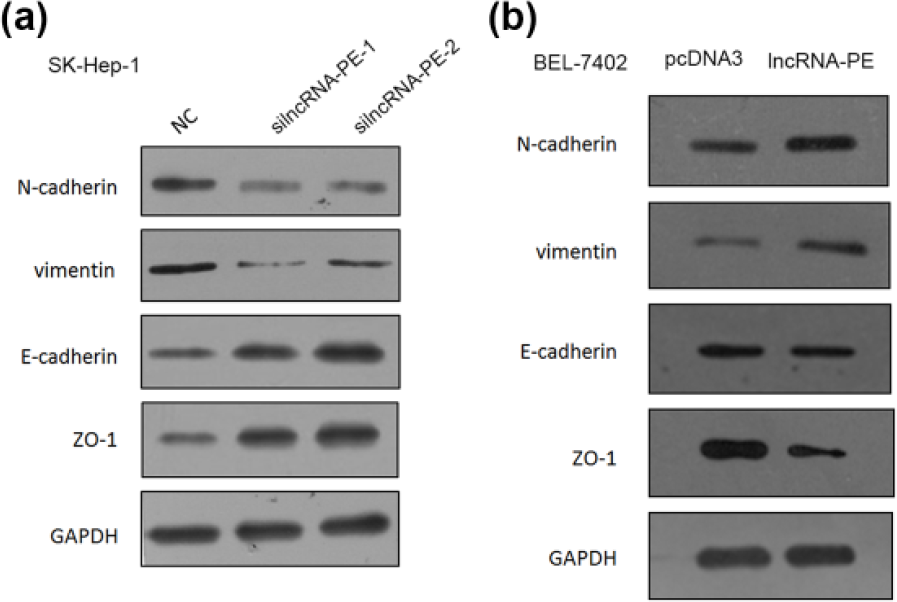
Effects of lncRNA-PE knockdown and overexpression on EMT markers in vitro. (a) The protein levels of four EMT markers after lncRNA-PE siRNAs were transfected into SK-Hep-1 cells. (b) The protein levels of four EMT markers after lncRNA-PE were overexpressed into BEL-7402 cells.
LncRNA-PE induces EMT via the miR-200a/b-ZEB1 pathway
We hope to seek the mechanistic insight into the role of lncRNA-PE in regulating HCC cell motility and EMT. As miR-200a/b can downregulate ZEB1, 6 an important EMT trigger, we wonder whether this lncRNA took part in miR-200-ZEB1 pathway. Therefore, we applied siRNA knockdown of lncRNA-PE in SK-Hep-1 cells and found that miR-200a and miR-200b were upregulated, with no influence on miR-451a (Figure 4(a)), with the opposite effects being caused by overexpression of lncRNA-PE (Figure S4(A)). Continuing our investigation of the molecular mechanism by which lncRNA-PE regulated miR-200, we tested whether lncRNA-PE regulated miR-200 via a competing endogenous RNA (ceRNA) mechanism. We analyzed the distribution of lncRNA-PE in the cell, and it was localized both in the nucleus and the cytoplasm (Figure S3(A)), which indicated that it may have the ability to act as an miRNA sponge. The ability to bind with protein AGO2, a key component of the miRNA-containing RNA-induced silencing complex (RISC), is a distinctive feature of ceRNA. 18 Therefore, we conducted an RNA IP experiment with AGO2 and proved that lncRNA-PE has poor affinity with AGO2 (Figure S3(B)), ruling out the possibility that lncRNA-PE utilized a ceRNA mechanism, and must regulate miR-200 via other mechanisms. We hypothesized that knockdown of lncRNA-PE could either induce the transcription of miR-200 or enhance its stability. We performed a dual-luciferase-reporter assay to ascertain whether the miR-200a/b upregulation observed after knockdown of lncRNA-PE depends upon regulation of the miR-200a/b promoter to enhance its transcription. pGL3-enhancer luciferase reporters containing the miR-200a/b promoter (pGL3-enhancer-miR-200a/b promoter) were constructed, and luciferase plasmids (pGL3-enhancer-miR-200b promoter or the control reporter pGL3-enhancer) were transfected into the different BEL-7402 and SK-Hep-1 cells with either lncRNA-PE or its siRNAs. Knockdown of lncRNA-PE increased the luciferase activity of the pGL3-miR-200b promoter (Figure 4(b)). Reciprocally, overexpression of lncRNA-PE had the opposite effect (Figure S4(B)). Therefore, we draw a preliminary conclusion that lncRNA-PE can inhibit the transcription of miR-200a/b. Next, we tested whether knockdown of lncRNA-PE can alter the expression of primary miR-200b and miR-200a which share the same promoter with miR-200b, and the data showed that the primary miRNA transcript of miR-200b and miR-200a was upregulated when lncRNA-PE was knocked down (Figure 4(a) and (c)), with the opposite effects being caused by overexpression of lncRNA-PE (Figure S4(A) and (C)). Additionally, both the RNA and protein level of ZEB1 were downregulated by siRNA against lncRNA-PE (Figure 4(d) and (e)), and overexpression of lncRNA-PE had the opposite effect (Figure S4(D)–(E)). To test whether lncRNA-PE regulated ZEB1 through miR-200a/b, we overexpressed lncRNA-PE in hepatoma cells; in the meantime, miR-200a/b mimics were transfected to compensate its effects. Results shown that both miR-200a and miR-200b mimics successfully downregulated ZEB1 (Figure S4(F)) and could compensate lncRNA-PE’s effects on ZEB1 (Figure S4(G)), which verified that lncRNA-PE regulated ZEB1 through miR-200a/b.

LncRNA-PE knockdown can alter EMT via the miR-200b-ZEB1 pathway. (a) The miRNA levels of miR-200b, miR-200a, and miR-451a after lncRNA-PE siRNAs were transfected into SK-Hep-1 cells. (b) Luciferase assays indicated that lncRNA-PE regulated the expression of miR-200b via its promoter. (c) The mRNA level of pri-miR-200b after lncRNA-PE siRNAs was transfected into SK-Hep-1 cells. (d) The mRNA level of ZEB1 after lncRNA-PE siRNAs was transfected into SK-Hep-1 cells. (e) The protein levels of ZEB1 after lncRNA-PE siRNAs were transfected into SK-Hep-1 cells. Each bar represents the mean ± SD of the counts from a single representative experiment (*p < 0.05).
Discussion
HCC is the third leading cause of cancer-related death worldwide. Since clinical symptoms are not easily observed during the early stage, the prognosis is poor at the time of diagnosis, which, in most cases, is during the advanced stage. 19 Thus, it is of much significance to explore new diagnostic and therapeutic molecular targets for HCC.
LncRNAs are transcripts of more than 200 nucleotides with limited protein encoding functions 20 and have been implicated in the molecular mechanisms of carcinogenesis. 21 As regulators of genetic information flow that interact with the epigenetic, transcriptional, and posttranscriptional pathways, lncRNAs promote tumor formation, progression, and metastasis in many human malignancies.22,23 However, HCC-related lncRNAs are still an emerging field, with only a few lncRNAs known to be involved in HCC tumorigenesis. Maternally expressed gene 3 (MEG3) expression was markedly reduced in HCC cells and contributed to HCC growth inhibition through upregulating p53; 3 highly upregulated in liver cancer (HULC) plays an important role in tumorigenesis and is upregulated by miR-372 in HCC. 24
In this study, we identify a novel lncRNA, lncRNA-PE, which is upregulated in HCC cell lines and tissues, and its high expression is correlated with poor prognosis in GEO datasets, which indicates that it could be a potential prognostic biomarker in HCC patients. Recurrence determines the postoperative prognosis with HCC, and it was reported that miR-200 family suppression is also reported to be related to poor prognosis of some colorectal, which may result from ZEB1-mediated cancer metastasis. 25 Moreover, there are much to investigate how the transcription of lncRNA-PE is controlled in HCC. Whether the expression of lncRNA-PE is regulated by transcriptional or epigenetic mechanisms? Therefore, future investigation of the upstream regulatory machinery that acts on lncRNA-PE in HCC could provide additional insight into HCC progression.
LncRNA-PE promotes HCC cell invasion by upregulating ZEB1, the master inducer of EMT via inhibiting transcription of miR-200a/b (Figure 5). It has been reported that ZEB1 is a target of miR-200a/b, 26 which indicates that ZEB1 triggers an miRNA-mediated feedforward loop promoting EMT and invasion of cancer cells. 27 Also, loss of expression of miR-200 family may repress E-cadherin by ZEB1 during EMT, which enhances migration and invasion during cancer progression. 28 All these data support our conclusion that lncRNA-PE has effects on HCC cell metastasis and EMT.
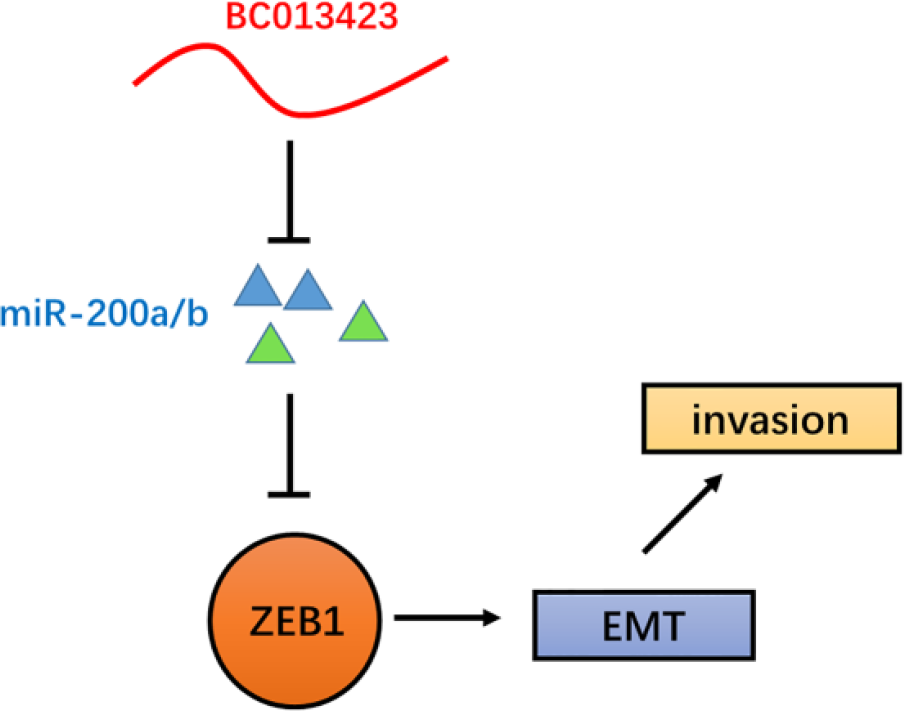
Schematic model of lncRNA-PE functions. LncRNA-PE promotes HCC cell invasion by activating the miR-200 family, upregulating ZEB1, and then inducing EMT.
Yuan et al. 12 reported that lncRNA-ATB acts as a ceRNA 29 and shares miR-200 response elements with ZEB1, which induces EMT in HCC. We proposed that lncRNA-PE could play a role as a ceRNA; however, our data show that it does not bind with AGO2, which precluded this possibility. Further exploration of the mechanism by which lncRNA-PE regulates miR-200b revealed that lncRNA-PE is distributed both in the nucleus and the cytoplasm (Figure S3(A)). Therefore, it could either influence the stability of miR-200b in the cytoplasm or induce its transcription in nucleus. We found that lncRNA-PE enhanced the promoter of miR-200b, but we still have not fully elucidated the exact mechanism. We speculated that lncRNA-PE could inhibit the binding of transcription factors with the promoter of miR-200b, but this idea will need further validation, such as with chromatin immunoprecipitation (ChIP) assay technology. As metastasis is a complex multistep process and EMT is just an early stage, 12 it remains possible that lncRNA-PE may act via additional pathways to alter tumor invasion or other aspects of cancer pathogenesis.
Taken together, our research demonstrated that lncRNA-PE is upregulated in HCC cell lines and tissues and acts as a key regulator of the miR-200-EMT pathway, thus improving our understanding of HCC progression. As direct targets of lncRNA-PE, miR-200 and ZEB1 mediated the role of lncRNA lncRNA-PE in local invasion. Furthermore, the effects of lncRNA-PE on the early stages of metastasis suggest that lncRNA-PE could be a target for anti-metastasis therapies and a prognostic biomarker in HCC patients.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported partially by Chinese State Key Projects for Basic Research (2013CB910801) and Chinese National Natural Science Foundation projects (91540202, 31370760, 31270836, and 31470782).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
