Abstract
Recent studies have shown that some members of the tripartite motif-containing protein family function as important regulators for carcinogenesis. In this study, we investigated whether tripartite motif-containing protein 44 acts as a cancer-promoting gene through its overexpression in esophageal squamous cell carcinoma. We analyzed esophageal squamous cell carcinoma cell lines to evaluate malignant potential and also analyzed 68 primary tumors to evaluate clinical relevance of tripartite motif-containing protein 44 protein in esophageal squamous cell carcinoma patients. Expression of the tripartite motif-containing protein 44 protein was detected in esophageal squamous cell carcinoma cell lines (8/14 cell lines; 57%) and primary tumor samples of esophageal squamous cell carcinoma (39/68 cases; 57%). Knockdown of tripartite motif-containing protein 44 expression in esophageal squamous cell carcinoma cells using several specific small interfering RNAs inhibited cell migration and invasion, but not cell proliferation. Immunohistochemical analysis demonstrated that the overexpression of the tripartite motif-containing protein 44 protein in the tumor infiltrated region was associated with the status of lymph node metastasis (
Introduction
Esophageal cancer is the fifth leading cause of cancer-related death worldwide, 1 and esophageal squamous cell carcinoma (ESCC) accounts for approximately 90% of esophageal carcinomas diagnosed in Asian countries. Despite recent improvements in diagnostic techniques and peri-operative management, survival rate of ESCC patients remains extremely poor. Since molecular targets as a treatment for ESCC were suggested to improve the survival of patients with this lethal disease, studies have attempted to identify biological factors involved in the malignant potential of ESCC. However, few molecular targeted agents have been specifically developed for its treatment, and thus, novel genes associated with the progression of ESCC need to be identified in order to conquer this aggressive malignancy.
Ubiquitination (the ubiquitin–proteasome system) is a post-translational protein modification. It plays important roles in the regulation of intracellular biological phenomena, such as the cell-cycle regulation, signal transduction, and transcriptional control.2–5 Recently, Really Interesting New Gene (RING) finger ubiquitin E3 ligases, one of the E3 ubiquitin ligases, have been reported to play important roles in cell-cycle regulation and carcinogenesis in malignancy.6–11 Several recent studies have demonstrated that several members of the tripartite motif-containing protein (TRIM) family, which are characterized by a conserved RING finger, B-box, and coiled-coil domains, function as important regulators in carcinogenesis.12–16
In this study, we focused on tripartite motif-containing protein 44 (TRIM44: 11p13). Fitzgerald et al.15,17 conducted two studies on TRIM44 in esophageal adenocarcinoma including junctional and gastric cardial adenocarcinoma. Jarvinen et al. 18 demonstrated that TRIM44 plays an important role in the malignant behavior of head and neck squamous cell carcinoma. Our group also previously demonstrated that TRIM44 is crucially involved in tumor cell proliferation through its overexpression in gastric cancer. 16 However, studies have not yet examined its clinical significance or functions that contribute to ESCC. We herein demonstrated that TRIM44 was overexpressed more frequently in primary ESCC tumors than in normal esophageal tissues, and the prognoses of patients overexpressing the TRIM44 were significantly poorer than those of patients with non-expressing tumors. We also showed that the down-regulated expression of TRIM44 suppressed cell migration and invasion, but not cell proliferation in ESCC cell lines. Our results provide novel evidence for the potential of TRIM44 as an important molecular biomarker to predict malignant outcomes and also as a target for molecular therapy in ESCC.
Materials and methods
Cell lines and primary tissue samples
Cell culturing procedure was performed as described in our previous reports.16,19–22 A total of 14 ESCC cell lines, TE1, TE2, TE4, TE5, TE6, TE8, TE9, TE10, TE11, TE13, TE14, TE15, KYSE70, and KYSE170, were used in this study. All of TE and KYSE 23 series cell lines authenticated by RIKEN were purchased from RIKEN Cell Bank (Tsukuba, Japan). All cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium (Sigma, St. Louis, MO, USA) with 100 mL/L fetal bovine serum (FBS; Trace Scientific, Melbourne, VIC, Australia). Primary tumor tissue samples of ESCC had been obtained from 68 consecutive ESCC patients, who underwent curative esophagectomy (R0 or R1), without any preoperative treatment such as chemotherapy or radiotherapy, at our institution between 1998 and 2009. All the samples were embedded in paraffin after 24-h formalin fixation. Written consent was always obtained in the formal style and after approval by the local Ethics Committee. Relevant clinical data were available for all patients, and disease stage was defined in accordance with the International Union against cancer tumor–lymph node–metastases (TNM) classification. 24 The median follow-up period in the current cohort was 45.2 months (ranging from 3.9 to 157.0 months). In the current cohort, total number of the death cases was 23, and 22 cases of them died of ESCC. All of the 22 cases had recurrences. The “prognostic marker” study is conducted according to the REMARK guidelines. 25
Western blotting and quantitative real-time reverse transcription polymerase chain reaction
Western blotting (WB) and quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) were performed as described in our previous reports.16,19–22 For WB analysis, a primary anti-TRIM44 antibody (BC024031) was purchased from Proteintech Group, Inc. (Chicago, IL, USA) and an anti–glyceraldehyde 3-phosphate dehydrogenase (GAPDH) antibody was purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). TaqMan Gene Expression Assays (Hs00214040_m1 for TRIM44; Hs99999903_m1 for beta-actin; Applied Biosystems, CA, USA) were used for qRT-PCR according to the manufacturer’s instructions. Each assay was performed in duplicate for each sample.
Loss-of-function by small interfering RNA
Loss-of-function screening was performed using small interfering RNAs (siRNAs) targeting TRIM44 (Stealth Select RNAi™ siRNA; Invitrogen, St. Louis, MO, USA) and negative control siRNA (Stealth RNAi™ siRNA Negative Control Med GC; Invitrogen). Each siRNA (20 µM) was transfected into ESCC cells using Lipofectamine RNAiMAX (Invitrogen) according to the manufacturer’s instructions. The knockdown of the targeting gene was confirmed by both of WB and qRT-PCR.
Cell growth assay, Transwell migration and invasion assays, and wound-healing assay
Cell growth assay was performed using colorimetric water-soluble tetrazolium salt (WST) as described in our previous reports.16,19,21,22 Transwell migration and invasion assay was performed using 24-well modified Boyden chambers (uncoated membrane for migration, Matrigel-coated membrane for invasion assay; Transwell chamber; BD Transduction Laboratories, Franklin Lakes, NJ, USA) as described in our previous reports.16,19,21,22 Each of the siRNA transfectants (siRNA-TRIM44 and siRNA-Control) was transferred into the upper chamber (4 × 104 cells per well). Following 48-h incubation after siRNA transfection, the migrated or invasion cells were counted directly in triplicate. Wound-healing assays were performed using Culture-Insert in µ-Dish 35 mm (ib80206; Ibidi, Martinsried, Germany). Each of the siRNA transfectants (5 × 105 cells/mL, 70 µL/well) was transferred into each insert and incubated for 24 h. After cultivation, the inserts were gently removed and culture medium with 10% FBS was slowly injected over the cell patches. We evaluated wound healing as horizontal migration potential after subsequent 10-h incubation.
Immunohistochemistry
The precise methodology for immunohistochemistry is described in our previous report. 20 Immunohistochemistry was performed using the avidin–biotin–peroxidase complex system (VECTASTAIN Elite ABC Universal Kit; Vector Laboratories Inc., Burlingame, CA, USA) with anti-TRIM44 (1:1000) primary antibody. Formalin-fixed ESCC cell line overexpressing TRIM44 (TE13, data not shown), in which >50% of cells showed staining for each protein, was used as a positive control, whereas formalin-fixed gastric cancer cell line with the weak expression of TRIM44 (HGC27, data not shown) and TE13 staining without the TRIM44 antibody was used as a negative control. 16 In order to evaluate relationship between TRIM44 expression and tumor invasive potential, we evaluated the staining intensity of the TRIM44 protein in the infiltrated region of the each case. In the scoring of TRIM44 expression, three-step scoring of all areas in each slide including tumor and non-tumor regions was blindly provided using high-power (200×) microscopy by an independent pathologist.16,19–22 In this study, we evaluated TRIM44 expression levels especially for the infiltrated region of invasive front in each tumor. TRIM44 protein expression was identified as negative in most of the non-tumorous esophageal squamous cells, and staining intensity was divided into three steps (scores 0, 1, and 2) in the infiltrated region of invasive front in each case as described before.16,19–22
Statistical analysis
Clinicopathological factors of the corresponding patients were evaluated for significance by the chi-squared test or Fisher’s exact test. For survival analysis, Kaplan–Meier survival curves were constructed for groups based on univariate predictors, and differences between the groups were evaluated by the Log-rank test. Univariate and multivariate survival analyses were performed using Cox proportional hazards model. Each statistical difference was considered significant at the p < 0.05 level using a two-sided test.
Results
Protein expression of TRIM44 in ESCC cell lines
A WB analysis was performed using a TRIM44-specific antibody (Figure 1(a)) in order to determine TRIM44 protein levels in 14 ESCC cell lines. Of the 14 cell lines, the overexpression of TRIM44 was observed in 8 (TE2, TE8, TE9, TE10, TE11, TE13, KYSE70, and KYSE170) cell lines (8/14 lines, 57%), suggesting that this gene is a target for activation in these cell lines.

(a) Western blotting analysis in 14 ESCC cell lines showed the overexpression of TRIM44 in 8 cell lines (8/14 lines, 57%). (b) The expression level of both protein and mRNA of TRIM44 was more efficiently knocked down after the transient introduction of a TRIM44-specific siRNA.
Suppression of cell migration and invasion by the knockdown of TRIM44
In order to assess the oncogenic role of TRIM44, we first conducted a cell proliferation assay using siRNA specific to TRIM44. In KYSE170 cells which overexpress TRIM44 protein, the expression level of the TRIM44 protein and messenger RNA (mRNA) was more efficiently suppressed after the transient transfection with the siRNA-TRIM44 than with the siRNA-Control (Figure 1(b)). A migration and invasion assay using KYSE170 cells showed that the knockdown of TRIM44 inhibited both of migration and invasion potential in ESCC (Figure 2(a)). Moreover, horizontal cell migration potential was inhibited after the knockdown of endogenous TRIM44 in the wound-healing assay (Figure 2(b)). However, the cell proliferation potential of KYSE170 cells transfected with siRNA-TRIM44 was the same as that of cells transfected with the siRNA-Control (Figure 2(c)).
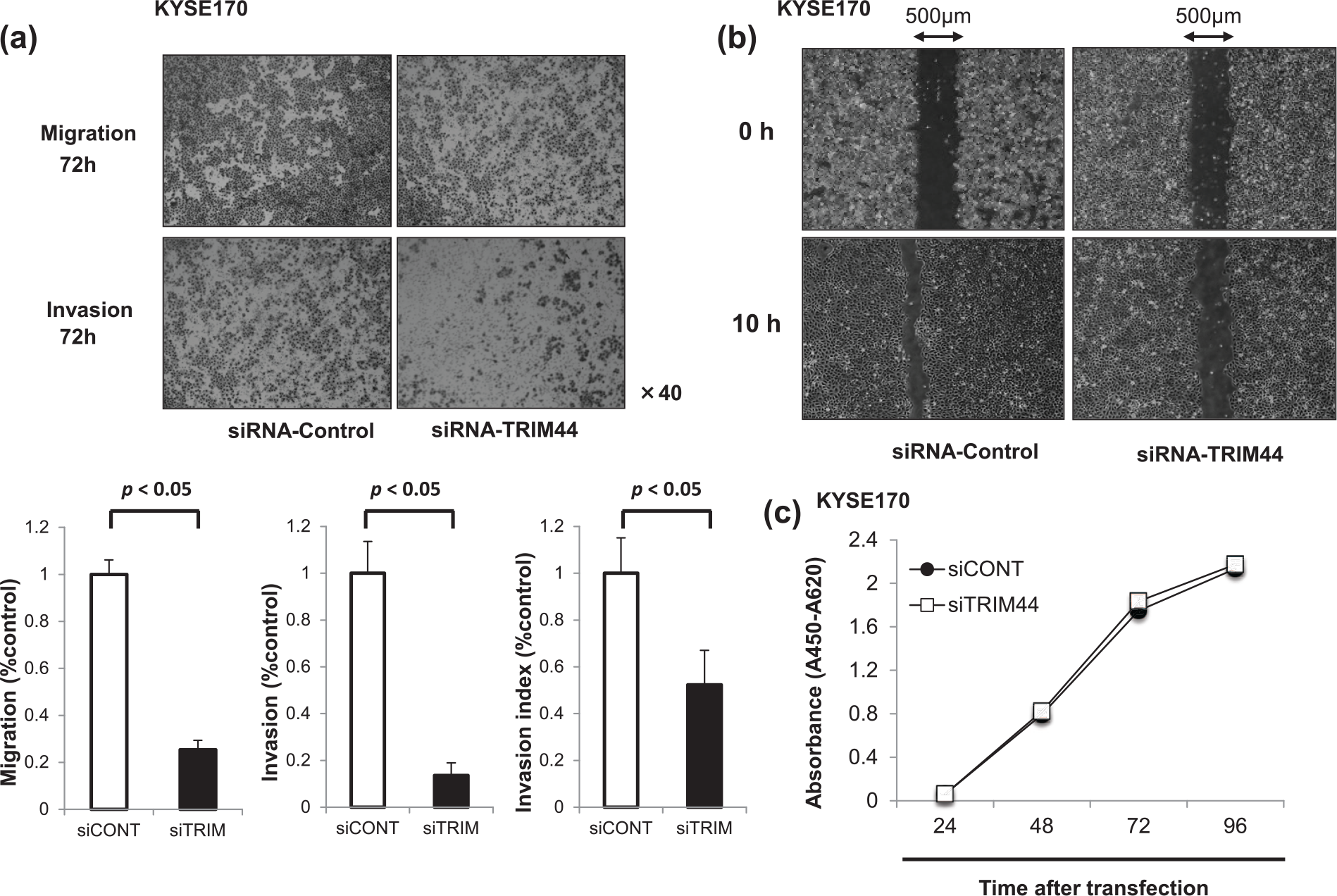
In vitro analysis of loss-of-function by small interfering RNA (siRNA): (a) invasion and migration assay, (b) wound-healing assay, and (c) cell proliferation assay.
Immunohistochemical analysis of TRIM44 expression in primary tumors samples of ESCC
We examined the clinicopathological significance of TRIM44 expression in primary tumor samples of ESCC based on the immunohistochemical staining pattern of TRIM44 protein. We classified 68 ESCC tumors into two groups, expression level of positive and negative groups, according to the staining intensity of the TRIM44 protein in the infiltrated region in the each tumor as described in section “Materials and methods.” TRIM44 protein expression was identified as negative in most of the non-tumorous esophageal squamous cells, and staining intensity was divided into three steps (scores 0, 1, and 2) in the infiltrated region of each case (Figure 3(a)). When score 0 (29 cases) was defined as negative and score 1 (21 cases) and score 2 (18 cases) were positive, TRIM44 positive cases showed significantly worse prognoses by the Kaplan–Meier survival curve and Log-rank test (
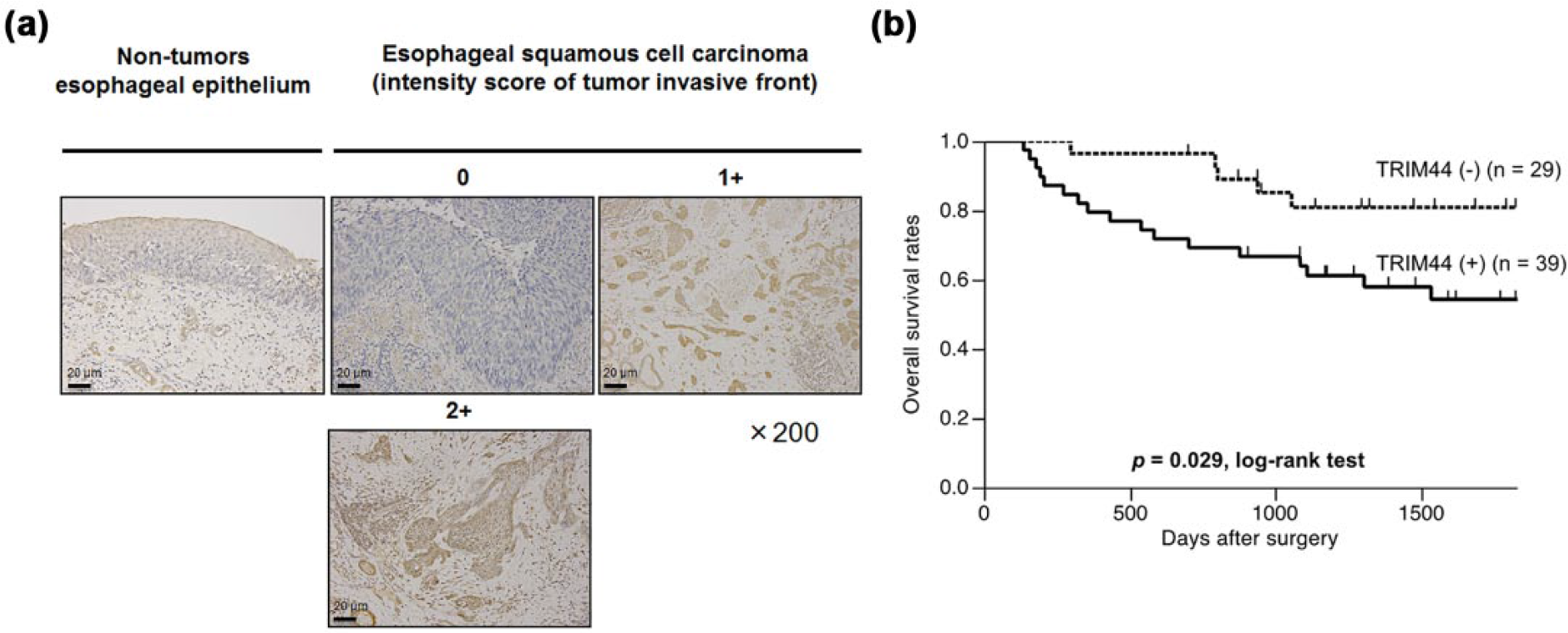
Staining intensity of TRIM44 in non-tumor esophageal epithelium and ESCC tumor was analyzed. (a) Immunohistochemical analysis of TRIM44 expression in primary tumors of ESCC may be divided into several steps (scores 0, 1, and 2) in the infiltrated region of each case. (b) Patients with positive immunoreactivity (score 1 and 2) for TRIM44 showed significantly worse prognoses than negative immunoreactivity (score 0) by the Kaplan–Meier survival curve and Log-rank test (
Relationship between TRIM44 protein expression levels and clinicopathological characteristics in primary cases of ESCC
The relationship between the TRIM44 protein expression levels and clinicopathological features is summarized in Table 1. The expression of the TRIM44 protein was associated with lymph node metastasis (
Correlation between the clinicopathological features and the expression level of TRIM44 in ESCC patients.

ESCC: esophageal squamous cell carcinoma; TNM: Tumor–node–metastasis.
The values given in boldface represent statistical significance.
The
Pathological tumor depth by the seventh TNM staging.
Pathological nodal metastasis.
Survival analysis of the clinicopathological features and TRIM44 expression in ESCC patients.
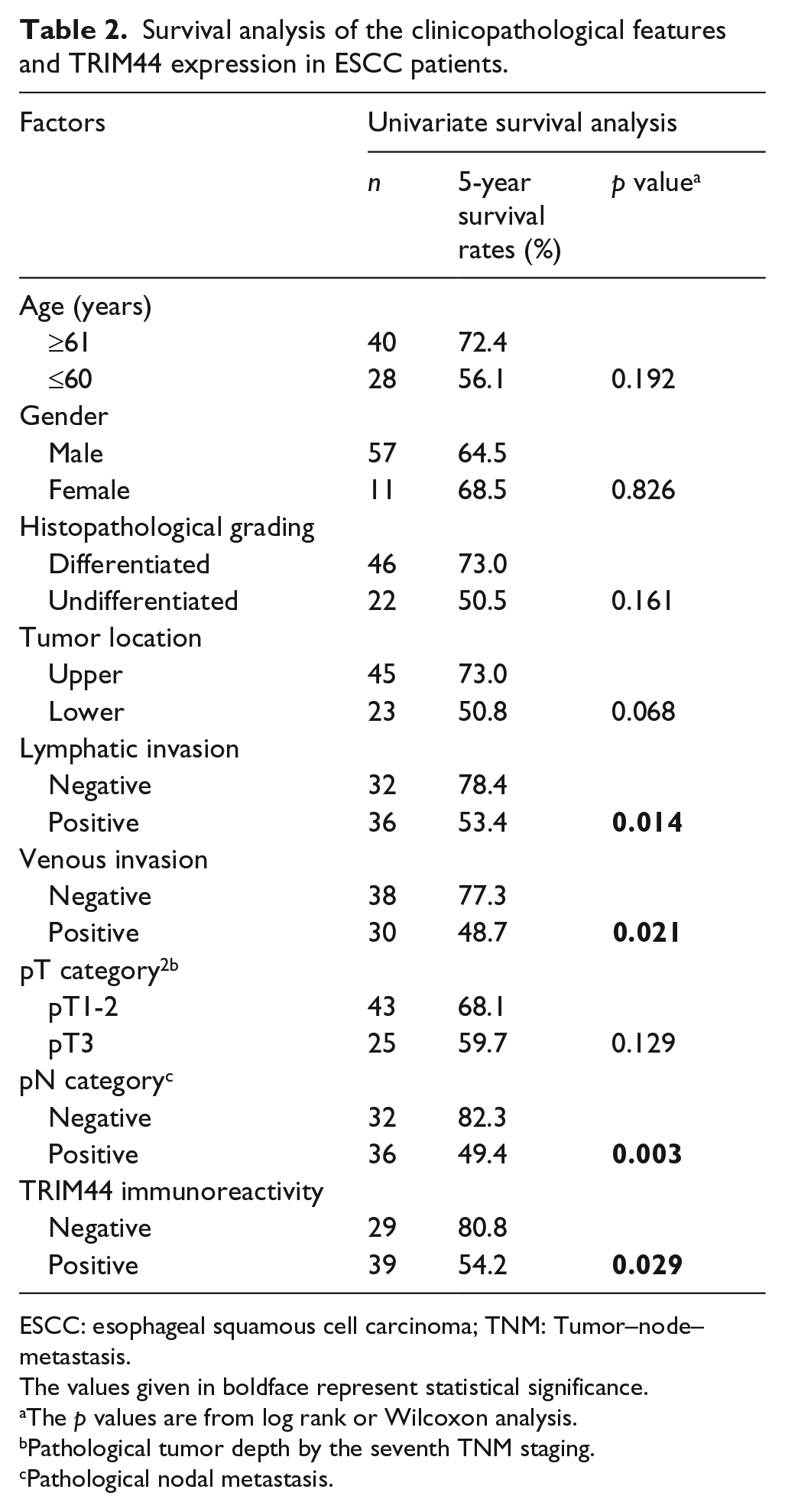
ESCC: esophageal squamous cell carcinoma; TNM: Tumor–node–metastasis.
The values given in boldface represent statistical significance.
The
Pathological tumor depth by the seventh TNM staging.
Pathological nodal metastasis.
Moreover, a multivariate prognostic analysis using Cox proportional hazards model identified the positive expression of TRIM44 in primary tumors as an independent worse prognostic factor (hazard ratio = 2.815;
Multivariate prognostic analysis using the Cox proportional hazard model of the clinicopathological features and TRIM44 expression in ESCC patients.

HR: hazard ratio; CI: confidence interval; TNM: tumor–node–metastasis; ESCC: esophageal squamous cell carcinoma.
The values given in boldface indicate statistical significance.
The
Pathological tumor depth by the seventh TNM staging.
Pathological nodal metastasis.
Discussion
Ubiquitination (the ubiquitin–proteasome system) is a post-translational protein modification and plays important roles in the regulation of intracellular biological phenomena, such as the cell-cycle, signal transduction, and transcriptional control.2–5 A large number of specific enzymes associated with ubiquitination have recently been reported including ubiquitin-activating enzyme E1, ubiquitin-conjugating enzyme E2, and ubiquitin-protein ligase E3. E3 ubiquitin ligases are generally defined to exhibit ubiquitin ligase activity and possess a substrate targeting function and comprise a potentially large number of diverged multi-subunit protein complexes.26,27 RING finger ubiquitin E3 ligases, E3 ubiquitin ligases, were previously reported to play important roles in cell-cycle regulation and carcinogenesis in malignancy.6–11
Several recent studies have demonstrated that some TRIM proteins, which are characterized by a conserved RING finger, B-box, and coiled-coil domains, function as important regulators for carcinogenesis.12–16 Our group previously reported that TRIM44, one of the TRIM family members, played a crucial role in tumor cell proliferation through its overexpression in gastric cancer. 16 Fitzgerald et al.15,17 conducted two studies on TRIM44 in esophageal adenocarcinoma including junctional and gastric cardial adenocarcinoma. Järvinen et al. 18 also detected a high-level amplified region of 11p13, including TRIM44 and CD44 overexpression, in head and neck squamous cell carcinoma. Ong et al. 28 showed that genomic amplification drives TRIM44 overexpression in esophago-gastric cancers and breast cancers, and targeting the mechanistic target of rapamycin (mTOR) pathway provides a potential therapeutic option for TRIM44-amplified tumors. Recently, Zhu et al. 29 reported that high expression of TRIM44 is associated with enhanced cell proliferation, migration, invasion, and resistance to doxorubicin in hepatocellular carcinoma. Taken together, these previous findings suggest that TRIM44 plays an important role in carcinogenesis and cancer development. Therefore, we hypothesized that the expression of TRIM44 may also be associated with tumor malignancy in clinical samples, and an appreciable change in a functional analysis may be observed in an in vitro study on ESCC.
In this study, the expression of the TRIM44 protein was detected in ESCC cell lines (8/14 cell lines; 57%) and primary tumor samples of ESCC (39/68 cases; 57%). The knockdown of TRIM44 expression in ESCC cells using several specific siRNAs inhibited cell migration and invasion, but not cell proliferation. In an immunohistochemical analysis, the overexpression of the TRIM44 protein in the infiltrated region of the tumor correlated with the status of lymph node metastasis, and overall survival rates were significantly worse in patients with TRIM44-overexpressing tumors than in those with non-expressing tumors (
However, the limitation of this study includes relatively small number of the patients. While knockdown of TRIM44 gene in KYSE170, which overexpressed TRIM44 protein, significantly induced inhibition of cell migration and invasion without any inhibition of cell proliferation using in vitro assay, the association between TRIM44 expression levels and microscopically lymphatic or venous invasion did not show any significance in our cohort. There is a discrepancy in these results; therefore, we should recognize the limitation of small patient number of the current cohort.
Although there were several limitations to this study, the results obtained suggest that TRIM44 contributes to the malignancy of ESCC.
Di et al. demonstrated that TRIM11 may be an indicator of glioma malignancy and possesses an oncogenic function mediated through the epidermal growth factor receptor (EGFR) signaling pathway. A strong correlation has been reported between the expression of TRIM11 and that of CD133 (an epithelial-to-mesenchymal transition (EMT) marker30,31), and the overexpression of TRIM11 has been shown to promote a stem-like phenotype in glioma. 14 They also found that the overexpression of TRIM11 potentially leads to a more aggressive glioma phenotype, along with increased malignant tumor growth and poor survival in glioma. 14 Luo et al. 32 recently demonstrated that TRIM44 promoted the migration and invasion of human lung cancer cells through the activation of nuclear factor-κB (NF-κB) signaling by upregulating the expression of C-X-C motif chemokine ligand 16 (CXCL16) and matrix metalloproteinase (MMP)-9. The immunohistochemical results obtained in this study in addition to the in vitro data of the cell migration/invasion assay or wound-healing assay indicate that the expression of TRIM44 in ESCC is related to tumor malignancy and aggressiveness. However, the locus of TRIM44 is 11p13, at the same locus of the cancer stem marker CD44. CD44 is a cell surface marker for cancer stem cells, and CD44-expressing cancer cells have the ability to initiate tumors in some types of cancers. 33 Our preliminary data suggest that the expression pattern of TRIM44 is similar to that of CD44 in ESCC cell lines, and changes in the expression of an EMT-related molecule 34 were observed after the knockdown of TRIM44 (data not shown). Therefore, TRIM44 may play an important role in the tumor malignancy of ESCC through some of the molecular mechanisms involved in cell migration and invasion processes. However, our results are still preliminary, and thus, a more precise analysis of the process of malignant potential is needed in order to clarify the role of TRIM44 in ESCC.
A study by Li et al. revealed that TRIM44 regulates ubiquitination and stabilizes the protein. A zinc-finger ubiquitin binding protein (ZF UBP) domain at the N-terminal region of the TRIM44 module has also been found in deubiquitinating enzymes containing ubiquitin-specific peptidase 33 (USP33)/the von Hippel–Lindau tumor suppressor protein-interacting deubiquitinating enzyme 1 (VDU1), and USP20/VDU2, members of the ubiquitin-specific protease family.35,36 Another study by Li et al. 37 demonstrated that USPs deubiquitinate their substrates. USP20/VDU2 was reported to deubiquitinate and stabilizes its associated protein, HIF1a. These findings suggest that TRIM44 functions as an “USP-like TRIM” and acts as a cancer-promoting gene that regulates the deubiquitination and stabilization of oncogenes. However, the precise molecular role of TRIM44 in cancer progression is not clear.
In conclusion, although our results are preliminary, they suggest that TRIM44 plays a crucial role in tumor invasion through its overexpression and highlights its usefulness as a predictor and potential therapeutic target in ESCC.
Footnotes
Acknowledgements
The authors cordially appreciate Daniel Mrozek (Medical English Service, Kyoto, Japan) for language editing. T.K. and S.K. have contributed equally to this work.
Author contributions
T.K. and S.K. helped in conception and design of the study; T.K. and S.K. participated in the development of methodology; T.K., Y.N., S.H., H.K., A.S., H.F., K.O., and H.T. helped in acquisition of data (provided animals, acquired and managed patients, provided facilities, and so on); T.K., S.K., and H.T. analyzed and interpreted data (e.g. statistical analysis, biostatistics, and computational analysis); T.K. and S.K. wrote, reviewed, and/or revised the manuscript; T.K., Y.N., S.H., H.K., A.S., H.F., and K.O. provided administrative, technical, or material support (i.e. reporting or organizing data, constructing databases); and S.K., D.I., and E.O. supervised the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
