Abstract
Oral squamous cell carcinoma is the most aggressive cancer that is associated with high recurrence, metastasis, and poor treatment outcome. Dysregulation of long non-coding RNAs has been shown to promote tumor growth and metastasis in several cancers. In this study, we investigated the expression of 11 selected long non-coding RNAs that are associated with cell proliferation, metastasis, and tumor suppression in oral squamous cell carcinomas and normal tissues by quantitative real-time polymerase chain reaction. Out of the 11 long non-coding RNAs profiled, 9 were significantly overexpressed in tumors with tobacco chewing history. Moreover, the long non-coding RNA profile was similar to the head and neck cancer datasets of The Cancer Genome Atlas database. Linc-RoR, a regulator of reprogramming, implicated in tumorigenesis was found to be overexpressed in undifferentiated tumors and showed strong association with tumor recurrence and poor therapeutic response. In oral squamous cell carcinomas, for the first time, we observed linc-RoR overexpression, downregulation of miR-145-5p, and overexpression of c-Myc, Klf4, Oct4, and Sox2, suggesting the existence of linc-RoR-mediated competing endogenous RNA network in undifferentiated tumors. Taken together, this study demonstrated the association of linc-RoR overexpression in undifferentiated oral tumors and its prognostic value to predict the therapeutic response.
Keywords
Introduction
Oral cancer is the most common cancer in head–neck region affecting predominantly male with 75% of diagnosed cases around 60-year-olds, of which 90% are oral squamous cell carcinoma (SCC). Every year more than 11 million people are diagnosed with cancer and 8.2 million people died from cancer worldwide. 1 In India, oral cancer ranks number one among all cancers in male patients and number three among females. Oral cancer has been associated with tobacco chewing with betel quid, slaked lime, and areca nut in India and other Asian countries, whereas in western countries, cigarette smoking and alcohol consumption are the major risk factors. 2 Human papillomavirus (HPV), the major causative factor of SCC of uterine cervix, has also been linked with a subset of oral cancer development. 3 Even with the advances in the treatment, oral cancer remains a major health threat and overall survival rate has not improved from 5 years, and the most effective radiotherapy (RT) regimens attain only 45%–55% of tumor regression rates. 4 Although remarkable progresses have been achieved in drug discovery to treat cancer, drug resistance remains as a foremost clinical barrier in managing the disease and results in poor clinical outcomes for the patients. 5
Oral carcinogenesis is a multi-step process leading to the transformation of a normal cell into a cancer cell through a wide range of genetic alterations that include gene rearrangements, point mutations, and gene amplifications, leading to deregulation in molecular pathways modifying cell growth, survival, and metastasis. Recent advances in high-throughput technologies identified dysregulation of non-coding RNA in several type of cancers. Long non-coding RNAs (lncRNA), one of the largest and diverse classes of cellular transcripts >200 bp in length, plays an important role in embryonic development and cancer. Differential expression of non-coding RNAs is increasingly documented as a new hallmark feature in cancer. LncRNAs located in the nucleus modulate the transcription by recruiting or sequestering transcription factors (TFs) or chromatin-modifying enzymes and regulates cellular function in
Efforts are on to explore the lncRNA profile of various cancers with advanced high-throughput RNA sequencing technologies. Recent reports established the versatile function of lncRNAs and their role in tumorigenesis. However, there is no report on the lncRNA expression profile in Indian oral tumors. In this study, we profiled the expression pattern of 11 selected lncRNAs using real-time quantitative polymerase chain reaction (PCR) and analyzed the profile to associate with patient’s clinicopathological characters and therapeutic response. We found that majority of the oral SCCs with tobacco chewing/smoking history showed dysregulation of lncRNA, and for the first time, we report the linc-RoR ceRNA network in undifferentiated tumors and its association with therapeutic response.
Methods and materials
Clinical specimens
This study was approved by the Institutional Ethics Committee (IEC), Madras Medical College, Chennai (No. 04092010) and was conducted within the ethical framework of Dr ALM PG Institute of Basic Medical Sciences, Chennai. In total, 60 oral SCC tissue samples were collected from Royapettah Government Hospital, Chennai. The patient’s contextual and clinicopathological characteristics were documented with standard questionnaire following the IEC guidelines, and written informed consent was obtained from each patient after explaining about the research study. The tumor specimens (
RNA isolation and quality control
Tissues percolated with RNAlater solution were washed twice with ice cold 1× phosphate-buffered saline (PBS) to make it free from residual RNAlater solution. Homogenization of tissue samples was carried out in a Micro Smash MS-100 automated homogenizer (Tomy, Japan) with zirconium beads. RNA was isolated using RNeasy Mini Kit (Qiagen, Germany) with supplier protocol and instructions. The RNA was eluted in 30 µL of nuclease-free water and quantified using NanoDrop 2000 UV-Vis spectrophotometer (Thermo Fisher Scientific, USA). The integrity of RNA was verified by resolving in 1% agarose gel containing 0.5 µg/mL ethidium bromide in Mupid gel electrophoresis (TaKaRa, Japan).
Complementary DNA synthesis
Complementary DNA (cDNA) was synthesized from total RNA (2 µg) using custom-designed universal oligo (dT) reverse primer for 11 lncRNAs, random hexamer primer for TFs, and miRNA seed-specific stem loop primer for miR-145-5p.12,13 RNA samples were added with respective primers and preincubated at 65°C for 20 min to denature the RNA secondary structures and placed on ice. cDNA conversion reactions were done using reagents from the Invitrogen Reverse Transcription Kit (Invitrogen, USA) following the manufacturer’s protocol.
Quantitative real-time PCR
Relative quantification (RQ) was performed using the TaqMan® (ABI, USA) custom assays. LncRNA forward primers specific to the last exon/3′ end of the lncRNAs and a universal reverse primer were used for the lncRNAs (Table S2 and S3). Similarly, miRNA145-5p-specific forward primer and a universal reverse primer were used to profile the mature miRNAs. After reverse transcription, the cDNAs were diluted 25 times and RT-qPCR was carried out in 384-well optical plates (in triplicates) in 10 µL reaction with TaqMan® 2× Universal Master Mix (no AmpErase uracil N-glycosylase (UNG)), specific forward primer, universal reverse primer, and universal fluorescein amidite (FAM)-labeled minor groove-binding (MGB) probe (5′-CAGAGCCACCTGGGCAATTTT-3′) for lncRNAs and miRNA. Predesigned assay for glyceraldehyde 3-phosphate dehydrogenase (GAPDH), p53, Klf4, and miR-143-3p gene expression analysis and custom-designed assay for c-Myc were procured from ABI (Table S4). All the assays were performed following vendor’s protocol. Expression of Oct4 and Sox2 was quantified using SYBR® Green dye following vendor’s protocol (Table S5). The experiments were carried out in a 7900HT Real-Time PCR System. A negative control without cDNA was also included in parallel for all assays. GAPDH and RNU44 were used as endogenous references for lncRNAs/coding genes and for miRNAs, respectively. The expression level was calculated using 2−ΔΔCt calculation.
14
Each experiment was done in triplicates, and the mean was used for the analysis. Numerical data were presented as mean and standard error of mean (±SEM) in graphs. Differences between means were analyzed using Student’s
Analysis of the 11 selected lncRNA expression in The Cancer Genome Atlas RNA sequencing datasets
To compare the expression levels of the 11 selected lncRNAs of this study population with other populations, biomining work was carried out. RNA sequencing data of 523 oral cancer samples from The Cancer Genome Atlas (TCGA) database (updated January 2016) were obtained using cBioPortal interactive online cancer genomics database (http://www.cbioportal.org/). The real-time PCR data were log2 transformed and were compared with the TCGA RNA sequencing data. All statistical analyses were performed using GraphPad Prism 6 (GraphPad Software Inc, USA). Hierarchical clustering heat map was generated using Expander V7.0, developed by Tel Aviv University, Israel (©2003). All tests were two-tailed and a
Clinical evaluation of the therapeutic response
All the 60 patients were treated with RT (total radiation dose 50–60 Gy) first and with concurrent combination chemotherapy for three courses with cisplatin and 5-fluorouracil/paclitaxel. Tumor response to chemotherapy/RT was evaluated and the clinical response was demarcated 4 weeks after the initial treatment as follows: complete response (CR)—tumor regressed totally; partial response (PR)—50% tumor reduced compared to primary tumor size; stable disease (SD)—tumor size not reduced and no disease progression; and progressive disease (PD)—increase in primary tumor volume or appearance of new lesions. Patients who were evaluated as CR and PR were categorized as treatment responder group, and the remaining patients were designated as poor responders or resistant group. The patients were followed post-treatment for 13 months to study the tumor recurrence and therapeutic outcome.
Results
Clinical characteristics of oral SCCs
In total, 60 oral SCC samples were used for this study, and the clinicopathological features of the tumor samples were listed in Table S1. The mean age of cancer patients was 52.75 years (±11.12 years), and 45% of them were below the mean age and 55% are above the mean age. Males were overrepresented (71.67%) than the females (28.33%). Tumor of buccal mucosa (
Consolidated demographic and clinicopathological details of 60 oral squamous cell carcinoma patients.
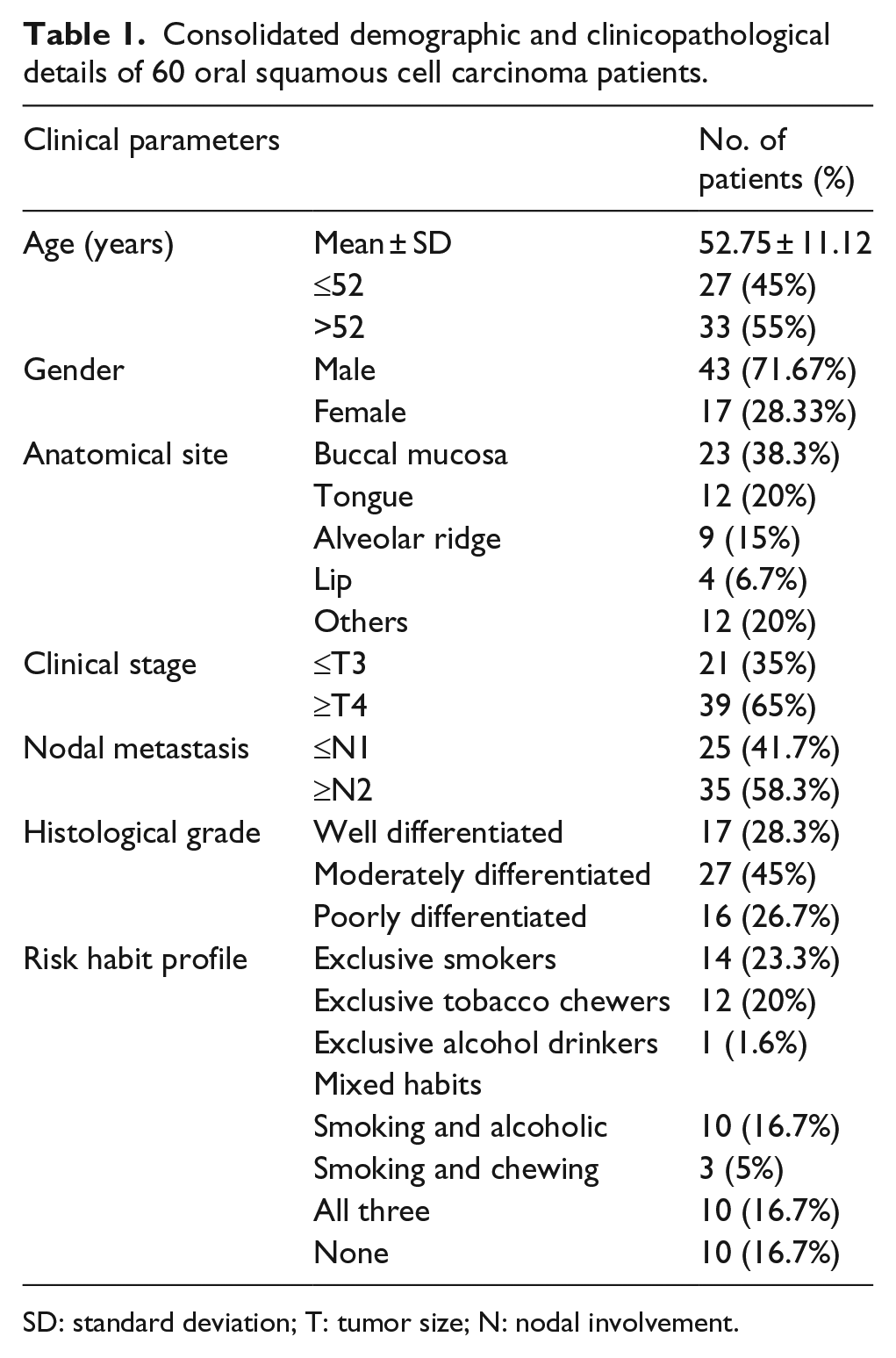
SD: standard deviation; T: tumor size; N: nodal involvement.
LncRNAs were differentially expressed in oral SCCs
We profiled the expression of 11 lncRNAs differentially expressed in various tumors. Six lncRNAs (CDKN2B-AS1, H19, HOTAIR, AP5M1, linc-RoR, and FALEC) associated with cell proliferation, two metastasis-associated lncRNAs (LINC00312 and MALAT1), and three tumor-suppressor lncRNAs (MEG3, POU3F3, and PANDAR) were selected for the expression profiling. The expression of these lncRNAs was analyzed in oral SCCs using real-time PCR with custom-designed primer/probes. Using log2-transformed 2−ΔΔCt values of lncRNA expression, unsupervised hierarchical clustering was performed (Figure 1). The clustering showed a differential expression of lncRNAs across the tumor samples, but no significant clustering pattern was observed either with the expression levels or clinical features, but overall dysregulation of lncRNA expression was common in tumors.

The unsupervised clustering of 60 OSCC primary tumor samples. The color scale shown at the bottom left illustrates the relative expression of lncRNAs across all samples: red color represents an expression level above mean and green color represents expression level lower than mean. Atop the cluster map, the boxes represent the clinicopathological features of each oral cancer sample. The blue shaded box represents the positive/presence of the clinicopathological features.
When comparing the expression of lncRNAs with normal samples, nine were found to be upregulated, and linc-RoR expression was statistically significant (
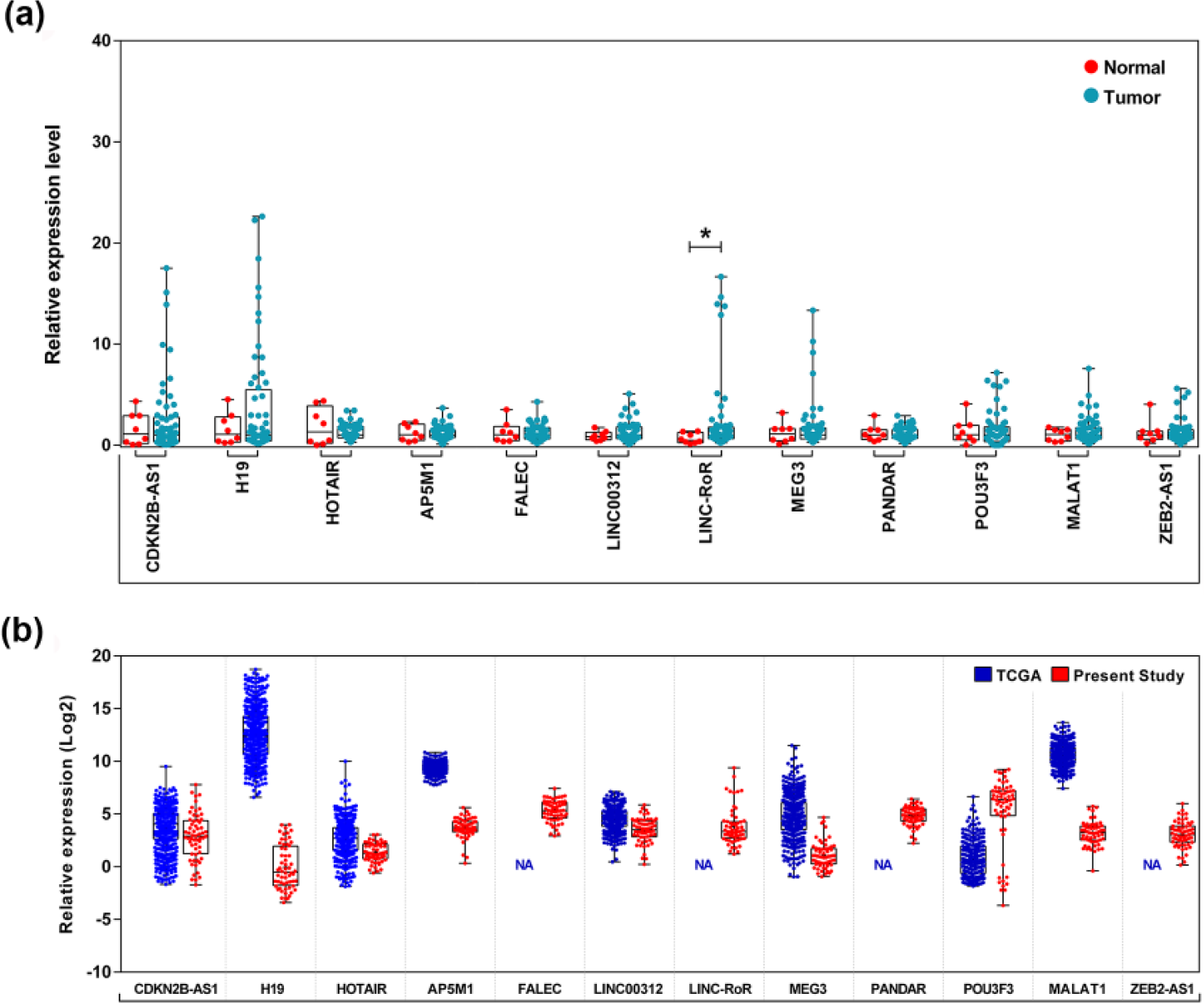
The expression profile of 11 selected lncRNAs in oral cancer. (a) Expression of lncRNAs in oral normal tissues and tumor tissues. Linc-RoR was significantly overexpressed in tumors (
LncRNAs were overexpressed in patients with the history of tobacco chewing/smoking
In order to understand the role of differentially expressed lncRNA in oral SCCs, the expression level of each lncRNA was tested for their association with the clinicopathological parameters by univariate analysis (Table 2). Interestingly, in tobacco chewers, nine lncRNAs CDKN2B-AS1, HOTAIR, AP5M1, FALEC, LINC00312, linc-RoR, PANDAR, POU3F3, and MALAT1 were expressed at significantly high level, whereas H19 alone was found to be downregulated in tobacco chewers (
Association of lncRNA levels with clinicopathological characteristics of oral tumors.

LncRNAs with significant
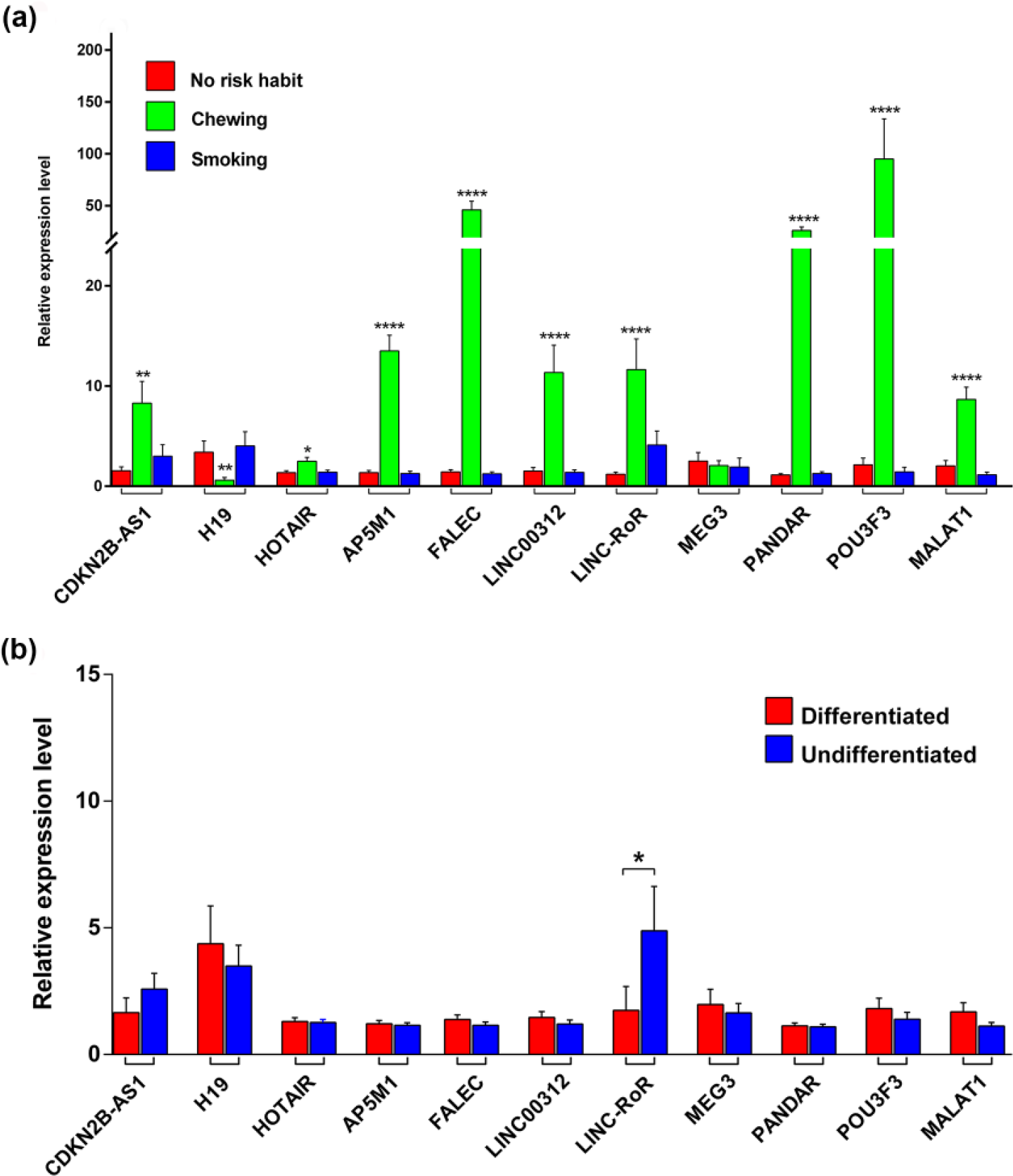
The differential expression of candidate lncRNAs in oral tumors with clinicopathological features. (a) Expression of lncRNAs in oral tumor patients with tobacco chewing and smoking habits. The student’s
linc-RoR ceRNA network is associated with undifferentiated oral SCCs
To understand functional significance of linc-RoR overexpression in undifferentiated tumors, we checked the expression levels of linc-RoR ceRNA network components, miR-145 and its downstream targets, and the core pluripotent TFs Oct4, Nanog, Sox4, Klf4, and c-Myc. The expression of miR-145-5p was significantly downregulated in undifferentiated tumor samples that overexpressed the linc-RoR (
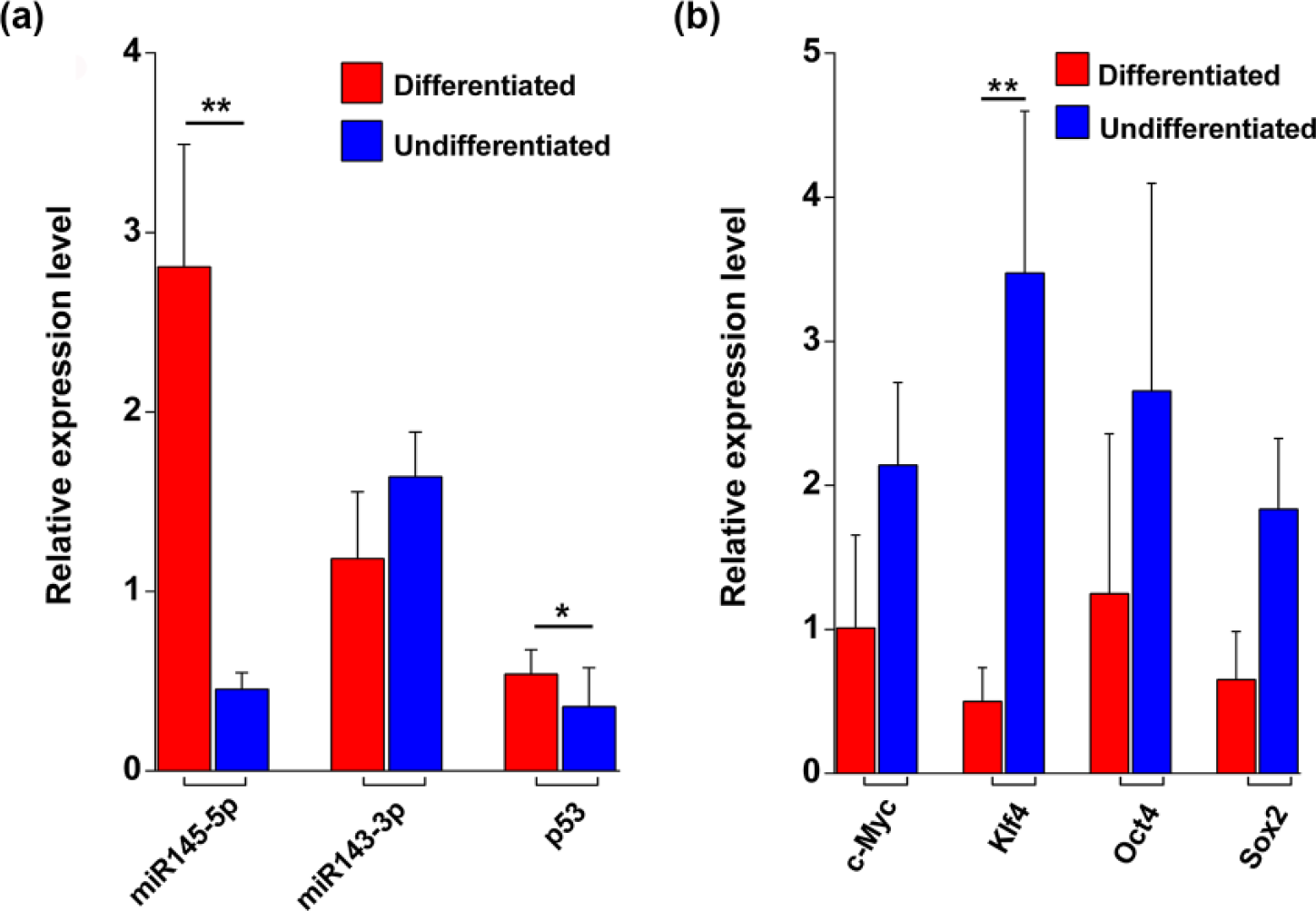
Expression of linc-RoR ceRNA network–associated genes. (a) Relative expression levels of miR-145-5p, miR-143-3p, and p53 mRNA between differentiated and undifferentiated tumor types. MiR-145-5p (
Linc-RoR is induced by p53, and linc-RoR in turn affects the levels of p53 through a negative feedback regulation.9,11 We tested the expression of p53 in the available oral SCCs and observed that the p53 expression was downregulated in undifferentiated tumor samples (
LncRNA overexpression determines the therapeutic response and tumor recurrence
The lncRNAs were analyzed with reference to the therapeutic response and tumor recurrence. We are able to collect treatment follow-up data only for 23 (38%) patients due to poor patient follow-up. We observed that 68% of the patients showed tumor recurrence and 50% of them had tumor relapse within 6 months post surgery. In the treatment outcome, CR was achieved in 11.8% of the patients, PR in 23.6%, SD in 35.3%, and PD in 29.3% patients. Patients with linc-RoR overexpression showed poor response to therapeutic regimen (

LncRNAs expression and its association with disease recurrence and therapeutic response. (a) Linc-RoR is significantly overexpressed in patients with poor treatment response (
Discussion
Oral cancer is the 11th most common cancer worldwide, and in India, it ranks number one among all cancers in male patients and number three among females. The major obstacles of oral cancer treatment are the development of resistance, disease recurrence, and metastasis. Epithelial-to-mesenchymal transition (EMT) is suggested to play an important role in oral cancer invasion and metastasis and also chemo–radio resistance. 16 The cancer stem cells (CSCs) are also suggested to play a vital role in cancer relapse and resistance to therapy. 17 LncRNAs, a diverse classes of cellular non-coding transcripts, play an important role in development and cancer and function by acting as signals, decoys, guides, and scaffolds and as a repressor or activator of gene transcription and translation.18–21
In this study, we profiled the expression of 11 selected lncRNAs that are associated with cell proliferation, metastasis, and tumor suppression and reported to be dysregulated in various cancers using real-time PCR in 60 oral SCCs. Notably, six lncRNAs (CDKN2B-AS1, H19, HOTAIR, AP5M1, linc-RoR, and FALEC) were associated with cell proliferation, two lncRNAs (LINC00312 and MALAT1) were associated with metastasis, and three lncRNAs (MEG3, POU3F3, and PANDAR) reported as tumor suppressor were selected for the expression profiling.6,7 Expression of lncRNAs was spatiotemporally regulated, and the dysregulation of lncRNAs was also reported as context specific. Majority of the profiled lncRNAs were overexpressed in tumor samples. In contrast, the expression of H19, AP5M1, and MALAT1 was downregulated in this study compared with TCGA-HNSC datasets. Therefore, we suggest that the underexpression of these three lncRNAs in our study samples may be due to the influence of tobacco or the low basal expression level in the study population. Analyzing the expression level individually with the clinical and pathological features revealed a significant association of lncRNA expression with the tobacco chewing and smoking habit. Out of 11 lncRNAs analyzed, 9 were overexpressed in the tumors from tobacco chewers. The lncRNA H19 was found to be overexpressed in tumor samples with smoking history, but was downregulated in tobacco chewers. The lncRNA H19 has been reported to express at higher levels in metastatic bladder cancer and hepatocellular carcinoma (HCC).20,22 Zhang et al. 23 reported that H19-enforced expression altered the miR-200 pathway, induced the mesenchymal-to-epithelial transition (MET), and suppressed tumor metastasis in HCC. Therefore, H19 downregulation observed in tobacco chewers of this study could result in maintenance/activation of EMT in the oral SCCs. FALEC and LINC00312 were co-expressed in oral SCCs. FALEC is associated with BIM1 and represses p21.24,25 LINC00312 (NAG7) associated with lymph node metastasis and H-ras pathway associated tumor progression in nasopharyngeal carcinoma.26,27 Moreover, H-ras has been reported to have relatively high incidence of mutational activation in Indian oral cancer.28,29 The overexpression of FALEC and LINC00312 in this study and their association with the BIM1 and H-ras pathway suggest their important role in oral SCCs.
Another interesting association was found between linc-RoR and cellular differentiation. Linc-RoR was expressed at higher levels in tumors with undifferentiated pathology. Linc-RoR is transactivated by p53, and linc-RoR in turn can suppress p53 during DNA damage through direct interaction with the hnRNP I and inhibits p53-mediated cell cycle arrest and apoptosis, 11 thus allowing the cells to escape from chemotherapy/RT. Endogenous linc-RoR functions as a ceRNA for core pluripotent TFs Oct4, Nanog, and Sox2 by sequestrating tumor-suppressive miR-145 which targets the core TFs.8,9 MiR-145 has been shown to be downregulated in variety of tumors including HNSC.30,31 In this study, we observed very low level expression of miR-145-5p in undifferentiated oral SCCs. In addition, the upstream regulator of linc-ROR, the p53, was found to be expressed at low level in undifferentiated tumor samples suggesting that upregulated linc-RoR could sequester miR-145-5p as well as transcriptionally repress the p53 in negative feedback manner.
Over the past decade, the understanding of stem cell biology and the role of stem cells in carcinogenesis have been remarkable. A minor population of cancer cells undergoes self-renewal and has an ability to differentiate into invasive cells inside the tumor environment. We tested the expression of four core TFs (Klf4, c-Myc, Oct4, and Sox2) associated with pluripotency in the oral tumors and observed their upregulation in the undifferentiated tumors. To understand the association of lncRNA expression with drug resistance and tumor relapse, we compared the expression levels with treatment outcome. Remarkably, we found that linc-RoR was significantly overexpressed in oral cancer patients with poor treatment response and recurrence. Enrichment of linc-RoR in nasopharyngeal carcinoma has been reported to result in chemoresistance through suppression of p53 signal pathway. 32 Takahashi et al. 33 reported that transforming growth factor beta (TGFβ) signaling during stress selectively enriched linc-RoR and directed extracellular vesicle-mediated transfer of linc-RoR to create chemoresistance in HCC.
In conclusion, this study shows that lncRNAs were dysregulated in oral SCCs, and majority of them were significantly associated with tobacco chewing/smoking. For the first time in oral SCCs, we identified the linc-RoR ceRNA network in oral tumors and its strong association with undifferentiated cellular phenotypes and treatment outcome. The sponging of miR-145-5p by overexpressed linc-RoR may relieve the post-transcriptional control of target genes c-Myc, Klf4, Oct4, and Sox2, which facilitates the maintenance of undifferentiated state. Taken together, this study demonstrated the association of linc-RoR overexpression with undifferentiated oral tumors and its prognostic value to predict the therapeutic response. Survival analysis from TCGA-HNSC samples showed no significant change in overall survival rate with linc-RoR expression. However, our treatment follow-up data showed a significant importance of linc-RoR with therapeutic response and recurrence suggesting that linc-RoR may function in context-dependent manner in Indian ethnicity. Therefore, this study has limitation as most of the patient survival data are not available because of poor patient follow-up and requires further investigation with larger sample size and experimental validation to understand the mechanism of linc-RoR in oral tumorigenesis. To the best of our knowledge, this is the first study from India to profile the lncRNA expression and to show their association with tobacco chewing/smoking and also to report the linc-RoR ceRNA network and its association with therapeutic response.
Footnotes
Acknowledgements
The authors thank all the patients for providing valuable clinical samples and their participation in this study. G.A., S.R., V.V., K.A., M.M., and A.K.D.M.R. gratefully acknowledge the Government of India’s University Grant Commission (UGC) and Council of Scientific and Industrial Research (CSIR) for providing research fellowships. We also thank DST-FIST, UGC-SAP, and DHR-MRU for providing infrastructure facilities.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The study was supported by grants received by corresponding author from Board of Research in Nuclear Sciences of Department of Atomic Energy, Government of India (No. 35/14/10/2014-BRNS/0210) and Department of Biotechnology, Ministry of Science and Technology, Government of India (BT/PR4820/MED/12/622/2013).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
