Abstract
Dysregulation of long non-coding RNAs has been found in many human cancers, including colorectal cancer that is still the third most prevalent cancer worldwide and related to poor prognosis; along with this, robust testimony has identified that long non-coding RNAs can take charge of tumor suppressor genes or oncogenes. This review summarizes nowadays research achievements on the character of long non-coding RNAs in the prognosis and diagnosis of colorectal cancer. On the basis of the results acquired in the last decade, some long non-coding RNAs are rising as biomarkers of colorectal cancer for prognosis, diagnosis, even prediction of therapeutic result, and have crucial effects in the regulation of colorectal cancer cell functions such as proliferation, invasion, apoptosis, metastasis, and drug resistant ability. Also, long non-coding RNAs, circulating in body fluids, could act as novel, affordable, lightly accessible, non-invasive detection tools for the personal health management of patients with colorectal cancer. Especially, circulating long non-coding RNA profiles may be demonstrating preferable prognostic and diagnostic capability and better accuracy than respective long non-coding RNAs in colorectal cancer.
Background
Colorectal cancer (CRC) is the third most diagnosed cancer in this blue planet, whose incidence is increasing in young adults, especially in the developing countries. 1 As the fourth main cause of cancer-related deaths in the world, CRC contributes to a serious threat to human health. To descend mortality of CRC, in past decades, researchers strive to study the mechanisms of this disease. Unfortunately, the prognosis of this malignancy remains pessimistic in numerous patients who even received surgery, and the cancer reappear or metastasis come up after several months. One of the leading challenges of this disease is the clinical diagnosis may be late. 2
Therefore, biological biomarkers for the identification of detection and prediction may have clinical significance. Especially, in past decade, extensive molecular research has demonstrated that long non-coding RNAs (lncRNAs) may be novel and significant biological biomarkers for early diagnosis and prognosis of colon cancer. 3
As its name indicates, LncRNAs are a group of non-coding RNA molecules whose lengths are larger than 200 nucleotides and lack protein-coding capability. LncRNAs are conducive to chromosome dosage compensation, epigenetic regulation, imprinting, nuclear and cytoplasmic trafficking, splicing, transcription, translation, cell cycle control, and cell differentiation among other functions.4,5 Studies also reveal that lncRNA aberrant expression is closely related to numerous diseases including cancer.6,7 At present, scores of lncRNAs are involved in the progression and development of cancer; hence, they may serve as novel biological biomarkers for pre-testing and predicting colon cancer.
Previous molecular mechanism research in CRC except lncRNA
People endeavored to study the occurrence and progression of CRC in the past. First and foremost, one of the greatest risks of rising CRC is the mutations of the genes. NRAS, BRAF, and KRAS genes are well known to be crucial in the advancement and development of CRC.8,9 Second, protein has also been involved in studies of CRC. Carcinoembryonic antigen (CEA) is a kind of glycoprotein from the CRC tissue and is used as prognosis and diagnosis tools in CRC. 10 Moreover, RNAs, including messenger RNA (mRNA) and microRNA, simultaneously become the research objection. MicroRNA-19a can regulate inflammation to drive CRC progression. 11
Biological functions of lncRNAs
The major role of lncRNAs has been described as regulators in the expression of protein-coding genes by influencing its neighboring genes (in cis) or influencing distant genes on other chromosomes (in trans). 12 At multiple levels such as transcription, translation, and protein function, lncRNAs could control individual genes and gene expression procedure through changing the fundamental transcriptional mechanism or by epigenetic regulation (Figure 1). Explicitly, (1) lncRNAs, at the upstream promoter area, can affect downstream gene expression while transcribing or affect gene expression by mediating histone modification and chromatin remodeling; (2) as the precursors of microRNA or small interfering RNA (siRNA), lncRNA could control gene expression through these small RNAs; (3) by directly binding specious proteins, lncRNA could regulate protein activity, serve as structural component of RNA–protein complexes, and change specific protein localization; and (4) lncRNA can product different splice variants via multiple alternative splicing modes (Figure 1). 13 Having functional diversity, in fact, unusual expression of lncRNAs could constitute the pathogenesis of sickness, involving cancer. So, it is unquestionable that lncRNAs have clinical importance in colon cancer biology and prognosis. 14
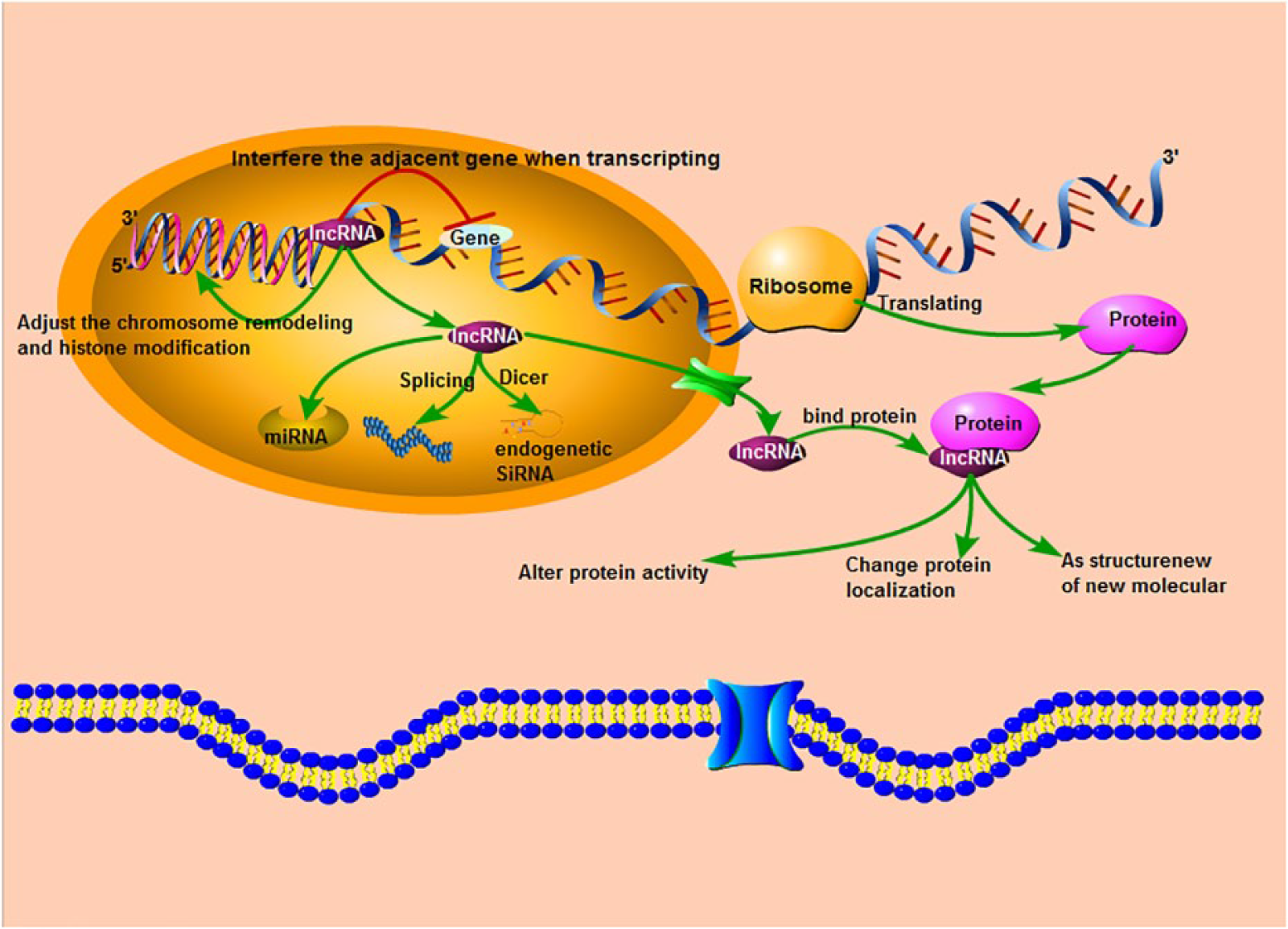
An overview of the lncRNA biogenesis and its functional mechanism. RNA polymerase II produces two nucleotide transcripts, called miRNA of approximately 20 nucleotides in length and siRNA by Dicer. LncRNA could move from the nucleus into cytoplasm, bind to protein to shift the character of this protein, and make interference on adjacent gene expression when transcribing from DNA to RNA.
LncRNAs in biology of colon cancer
Colon cancer with poor prognosis not just constantly grows and diffuses regionally but spread through the circulatory or lymphatic systems to other organs. Proliferation, invasion, and metastasis are the main biological characteristics of colon cancer-related deaths, which can contribute to poor prognosis. 15 Recent studies have implied that lncRNAs are widely involved in proliferation, invasion, and metastasis, which directly augment the likelihood of lncRNA-based diagnostics and prediction of prognosis in colon cancer. 16
LncRNAs contribute to poor prognosis in colon cancer
Including colon cancer, a substantial number of malignant diseases show certain characters that define the malignant cell phenotype. Increasing trials indicate that these features of colon cancer, such as proliferation, invasion, and metastasis, are regulated by lncRNAs. In the following sections, we describe several lncRNAs that promote these features, particularly those that facilitate poor prognosis in colon cancer (summarized in Table 1).
LncRNAs with experimental data supporting the functions of contributing to poor prognosis in CRC.

LncRNA: long non-coding RNA; CRC: colorectal cancer.
Actin filament–associated protein 1 antisense RNA 1
Actin filament–associated protein 1 antisense RNA 1 (AFAP1-AS1) also has been demonstrated to be associated with a risk of lymph node metastasis, distant metastasis, overall survival (OS), relapse-free survival, progression-free survival, and poor prognosis in various cancers.37,38 Particularly in CRC, a research conducted by Wang et al. manifested that AFAP1-AS1 was abnormally overexpressed in CRC tissues and positively associated with tumor–node–metastasis (TNM) stage, tumor size, and distant metastasis. In addition, in patients with overexpressed AFAP1-AS1, poorer OS and disease-free survival are more likely to occur. 17
Bladder cancer–associated transcript 1
Bladder cancer–associated transcript 1 (BLACAT1), also known as linc-UBC1, is a lncRNA with a length of 2616 bp, which was first characterized in bladder cancers. 39 It was sequentially studied as a tumor pro-oncogenic factor in gastric 40 and colon cancers. 20 Jun Su et al. clarified, both in vitro and in vivo, that overexpression of BLACAT1 is in CRC tissues than the controls, and higher BLACAT1 expression level is correlated with large tumor size, greater tumor depth, more lymph node metastasis, advanced TNM stages, and poor prognosis of CRC. 21
CRC-associated lncRNA
CRC-associated lncRNA (CCAL), also named as CCAT, is located at 8q24.21; just as its name, showed a remarkably increasing trend of expression levels from the normal colorectal tissues to adenoma and CRC. 41 The study conducted by Ma et al. 22 suggests that estimating the prognosis effect of high CCAL expression levels in CRC patients was associated with poor survival prognosis detected by quantitative reverse transcription polymerase chain reaction (qRT-PCR); high CCAL expression predicted poor therapeutic outcomes in 252 CRC patients. 22
Zinc finger antisense 1
Zinc finger antisense 1 (ZFAS1) is a non-coding RNA gene located at chromosome 20 (chr20) in the antisense direction, and 11 kb in scale. In 2011, ZFAS1 first appeared, to play a cancer-suppressive role, in a research report of breast cancer conducted by Askarian-Amiri et al. 42 However, in subsequent studies, ZFAS1 is revealed as an oncogene with overexpression in hepatocellular and gastric cancer, 43 involving CRC. In vivo, a recent study showed that higher ZFAS1 expression group in CRC patients was notably related with higher TNM stage, lymphatic invasion, and poorer relapse-free survival, compared to the lower ZFAS1 expression group. According to this relationship between ZFAS1 expression and CRC clinicopathological features, ZFAS1 could be regarded as a potential prophetic factor for CRC relapse, lymphatic invasion, and cancerous metastasis, even involving TNM stage. 36
Urothelial carcinoma associated 1
Human urothelial carcinoma associated 1 (UCA1) was first regarded as a biomarker for the prognosis of bladder cancer and was overexpressed in bladder cancer. 44 UCA1 might play an important cancer-promoting role in human cancers, such as bladder cancer, breast cancer, ovarian cancer, gastric cancer, lung cancer, tongue squamous cell carcinomas, and CRC.45,46 Especially in CRC, recent preliminary studies suggested that UCA1 can promote growth of CRC cells and the drug resistance of CRC cells to 5-fluorouracil (5-FU) by decreasing apoptosis, in vitro and in vivo; UCA1 can bundle endogenous miR-204-5p and restrain its activity to increase the protein levels of cyclic adenosine monophosphate responsive element–binding protein 1 (CREB1), negatively related to survival time and clinical outcome in CRC patients. 35
FEZ family zinc finger 1 antisense RNA 1
FEZ family zinc finger 1 antisense RNA 1 (FEZF1-AS1), a recently discovered lncRNA, is located on the counterpart of gene FEZF1. 23 Either in vivo or in vitro, Chen et al. 23 found that the upregulation of FEZF1-AS1 visibly participates in colorectal tumorigenesis and associates with CRC metastasis and poor prognosis.
Nuclear-enriched abundant transcript 1
Nuclear-enriched abundant transcript 1 (NEAT1), located on 11q13.1, is a recently detected oncogenic lncRNA whose abnormal elevation has been identified in different types of cancer, including CRC, nasopharyngeal carcinoma, gastric cancer, and osteosarcoma. 47 Relative to matching normal counterparts, NEAT1 expression level is elevated in CRC tissues and is positively associated with poor differentiation, invasion, distant metastasis, TNM staging, and shorter disease-free survival in CRC patients. 30
Promoter of CDKN 1A antisense DNA damage activated RNA
Promoter of CDKN 1A antisense DNA damage activated RNA (PANDAR), located on 6p21.2, contributes to chromosomal instability and metastatic progression in multiple cancers containing bladder cancer, gastric cancer, hepatocellular carcinoma, and non–small cell lung cancer (NSCLC).48,49 Latterly, Lu et al. 31 clarified that PANDAR expression can support migration and invasion in CRC, compared with normal tissue, and predict poor prognosis in patients with CRC.
PVT-1
LncRNA PVT-1 is encoded by oncogene PVT-1 and maps to chromosome 8q24. Somel’s studies have suggested that PVT-1 participates in cancer pathophysiology to affect diagnosis and prognosis. 50 By transfecting with PVT-1-specific siRNA in vitro, Takahashi et al. suggested that PVT-1 knockdown contributes to significant loss of proliferation and invasion capabilities of CRC cell. And then, in vivo, the low PVT-1 expression group showed less venous invasion, lymph node metastasis, and survival time compared to the high PVT-1 expression group. 32
H19
H19, located at chromosome 11p15.5, is a paternally imprinted gene involved in carcinogenesis, frequently exhibits inordinately high expression either in tumor cells or cancer tissues, including endometrium, ovary, stomach, and bladder cancer. 51 Likewise, overexpression of H19 was detected in CRC to promote cell viability and proliferation of CRC cells through the upregulation of miR-138, which targets vimentin genes. 24
HOX antisense intergenic RNA
The lncRNA HOX antisense intergenic RNA (HOTAIR) was identified as a non-coding RNA gene, in opposite direction, of the gene encoding homeobox C12 (HOXC12) in 2007. In the last few years, in cancer cells, overexpression of HOTAIR has been gradually discovered in multiple cancer types either primary or metastatic lesions. 52 In these studies, HOTAIR expression was associated with poor prognosis and metastasis, especially in CRC. 25 Svoboda et al. 26 clarified that HOTAIR is a negative prognostic factor in primary CRC. But in the blood of CRC patients, HOTAIR also plays a crucial role as a malignant predictor.
Metastasis-associated lung adenocarcinoma transcript 1
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) is a lncRNA derived from chromosome 11q13 and is more than 8000 nucleotides in length, which is thought to be crucial in the processing of pre-mRNAs and is revealed to be upregulated in liver, cervical, breast, and colon cancers. 53 One of these recent studies, conducted by Zheng et al., demonstrated that MALAT1 was upregulated in CRC than normal tissues. Moreover, for CRC, patients with high level of MALAT1 expression showed significantly poor prognosis than patients with low MALAT1 expression. 28
ATB
LncRNA-ATB is a novel molecular marker that was first demonstrated in hepatocellular cancer. 54 However, in CRC, Yue et al. suggested that, compared to cases with decreased ATB expression, elevated ATB expression was a significant independent indicator for more CRC recurrence and reduced survival. 18 Iguchi et al. recently clarified that lncRNA-ATB overexpression was markedly related with big tumor size, depth of lymphatic invasion or tumor invasion, and lymph node metastasis. In addition, the level of lncRNA-ATB expression had been involved in poorer outcomes of patients with CRC. 19
Highly upregulated in liver cancer
Highly upregulated in liver cancer (HULC) is a long non-coding RNA that was initially demonstrated at the most overexpressed level in hepatocellular carcinoma. Subsequently, HULC has been identified as an oncogenic long non-coding RNA in other cancer types, including osteosarcoma pancreatic cancer, gastric cancer, and hepatic metastasis of CRC.55,56 Recently, studies have also shed new light on the oncogenous role of HULC in colorectal carcinoma progression. 27
Small nucleolar RNA host gene 12
Small nucleolar RNA host gene 12 (SNHG12), located on the chromosome region of 1p35.3 and includes six exons, is a novel upregulated lncRNA that emerged in certain types cancer including breast cancer, osteosarcoma, and hepatocellular carcinoma.57,58 Similarly, a recent study illuminated that the level of SNHG12 in CRC tissues is higher than the adjacent normal mucosa as detected by qRT-PCR. Furthermore, upregulation of SNHG12 was distinctly associated with larger tumor size, advanced tumor stage, and worse 5-year survival rate in CRC patients. 33
Taurine-upregulated gene 1
Taurine-upregulated gene 1 (TUG1) was first reported in mouse retinal cells. 59 It is reported that high lncRNA-TUG1 expression was associated with cell proliferation of various cancer. One recent research conducted by Liang Wang suggested that TUG1 also indicates a poor prognosis of CRC patients by affecting cell proliferation and migration. Similarly, another research conducted by Junfeng Sun established that TUG1 is a negative prognostic predictor in CRC patients. 34
Pathways of partial lncRNA in human CRC

A simple model of pathways of partial lncRNAs affecting CRC prognosis and contributing to CRC diagnosis. Green arrows represent upregulation or promotion, red lines represent downregulation or demotion. Violet nodes represent lncRNAs that regulate target gene, RNA, and protein via either the same or different ways, which facilitates the function of diagnosis and predicts prognosis. LncRNAs can be detected in the plasma, because of entering the lymph and blood plasma through cytomembrane.
Pathways of lncRNAs affecting CRC diagnosis and prognosis.

LncRNA: long non-coding RNA; CRC: colorectal cancer; EMT: epithelial–mesenchymal transition.
Decrease in the level of lncRNAs contributes to poor prognosis in colon cancer
Every coin has two sides. Comparing lncRNAs with the oncogenic function, some lncRNAs have been established as negative process regulators of poor prognosis of CRC. In the following sections, these anticancer lncRNAs along with their correlated expression in CRC are enumerated (summarized in Table 3).
LncRNAs with experimental data repressing the functions contributing to poor prognosis in CRC.
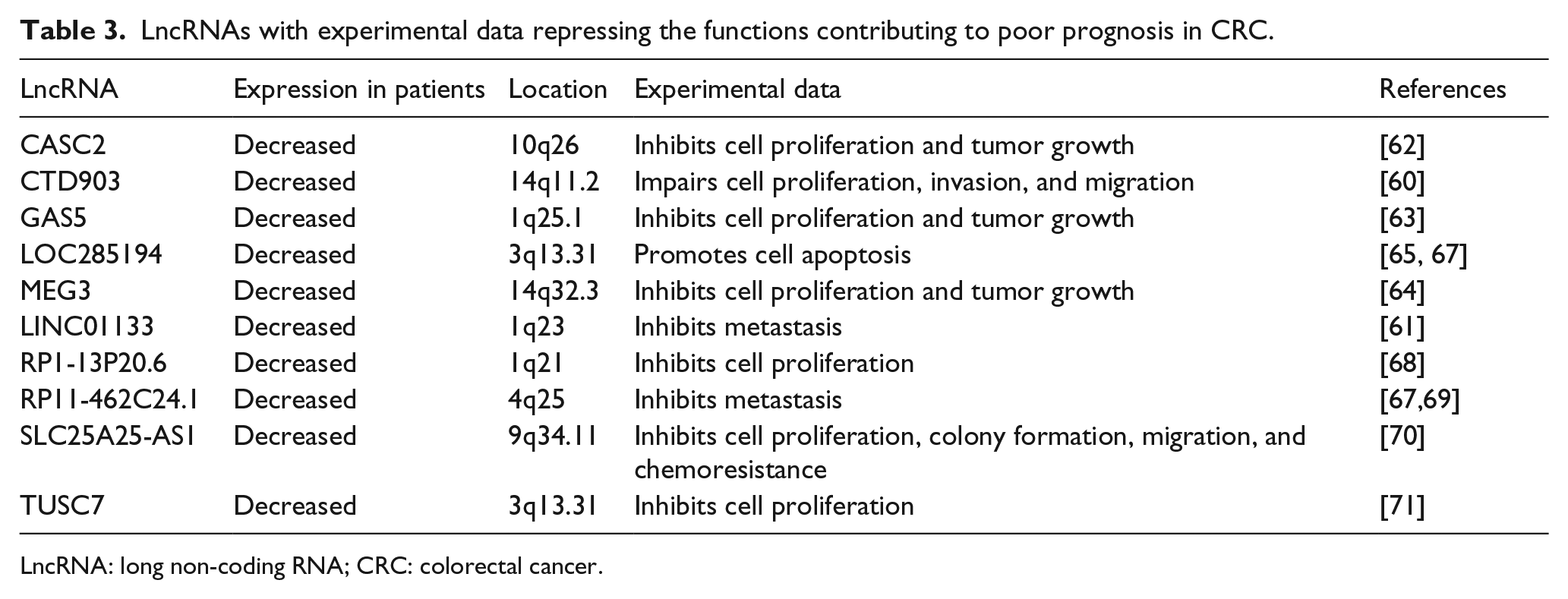
LncRNA: long non-coding RNA; CRC: colorectal cancer.
Cancer susceptibility candidate 2
Cancer susceptibility candidate 2 (CASC2), located at chromosome 10q26, is a tumor suppressor in gastric cancer, endometrial cancer, renal cancer, NSCLC, as well as CRC, either in vitro or in vivo. 72 Huang et al. 62 recently conducted a study that suggests that CASC2 expression was not only visibly reduced in CRC cell lines and CRC tissues but also more obviously decreased in patients with advanced CRC.
CTD903
LncRNA-CTD903 located on chromosome 14q11.2 got its name as this transcript is 903 bp in length. The studies conducted by Yuan et al. 60 reveal that CRC cells with CTD903 low expression have more aggressive biological behavior than high expression cells.
The growth arrest specific transcript 5
The growth arrest specific transcript 5 (GAS5) is transcribed at 1q25 and the length of GAS5 is approximately 630 nucleotides. Many studies have identified that GAS5, as a tumor suppressor, is downexpressed in prostate cancer cells, breast cancer cells, and renal cell carcinoma cells. 73 In comparison with the control group, Yin et al. observed that the cell cycle progression of CRC was significantly arrested at the G1–G0 phase by GAS5 through flow cytometric analysis. Similar results were also found in patients with high levels of GAS5 expression who had significant longer survival time than those with low levels. 63
LOC285194
LOC285194 is also known as “LSAMP antisense RNA 3,” consists of four exons, and located on 3q13.31 with >2 kb length. 67 LOC285194 was remarkably lower in CRC tissues, compared with adjacent normal mucosa. Lower expression level of LOC285194 was related to greater CRC size, more malignant grade, more distant metastasis, and poorer disease-free survival. 65 As a tumor suppressor, lncRNA LOC285194 can inhibit CRC cell growth by directly interacting with p53 gene that has been involved in a broad range of cell-biological processes. 66
Maternally expressed gene 3
The maternally expressed gene 3 (MEG3) is one of the most studied lncRNAs in time domain as a cancer suppressor molecule. 74 Because MEG3 is expressed in normal cells, the downregulation of MEG3 was investigated in various tumor types, including gastric cancer, NSCLC, and hepatocellular cancer. 75 In CRC tissues, MEG3 expression level is lower than that in the matched normal tissues and is negatively associated with tumor invasion depth, histological grade, and TNM stage. 64
LINC01133
LINC01133, located on 1q23, is underexpressed in lung adenocarcinoma but not in lung squamous cell cancer. 76 But in CRC, LINC01133 is an antimetastasis lncRNA, whose expression is memorably reduced in CRC tissues compared with the counter normal mucosa. Moreover, Kong et al. 61 certified that lower LINC01133 expression appears in 219 CRC patients with poor survival.
RP1-13P20.6
LncRNA-RP1-13P20.6, on the basis of the University of California, Santa Cruz (UCSC), Genome Biological information (http://genome.ucsc.edu/index.html), is located on the reverse strand of 1q21 region, and its length is 1997 nucleotides with one intron and two exons. By real-time polymerase chain reaction, Liu suggested that RP1-13P20.6 was downregulated obviously in the CRC tissues compared to their matched normal adjacent tissues. Furthermore, it was observed that lncRNA RP1-13P20.6 was decreased dramaticlly in CRC patients than the control. 68
RP11-462C24.1
RP11-462C24.1, located on chr4q25, is a lncRNA that is composed of four exons with the length of 1136 bp. The research conducted by Shi et al. shed light on that RP11-462C24.1 is downregulated in CRC tissues compared to adjacent normal mucosa, is lower in patients with CRC metastasis than these without metastasis. RP11-462C24.1 expression level is negatively correlated with the malignant degree of CRC and more distant metastasis. 69
SLC25A25-AS1
SLC25A25-AS1 consists of five exons and is located at 9q34.11. SLC25A25-AS1 is transcribed in the opposite direction of the gene SLC25A25 that can encode a calcium ion–sensitive mitochondrial carrier (CaMC) protein and is classified as 53 members of the mitochondrial inner membrane family. 77 From carcinoma tissues and counter adjacent normal tissues of 30 CRC patients, from the serum of 109 CRC patients and 99 normal individuals, Li et al. found that SLC25A25-AS1 was obviously downregulated in both serum of CRC patients and the tumor tissues. Decrease of SLC25A25-AS1 in CRC cells can enhance the resistant ability to common chemotherapeutics. 70
Tumor suppressor candidate 7
Tumor suppressor candidate 7 (TUSC7), located on 3q13.31, is a newfangled lncRNA that is identified to be downregulated in several cancer cells such as hepatocellular carcinoma, gastric cancer, osteosarcoma, and NSCLC. 78 Likewise, as a tumor suppressor, TUSC7 was downregulated in CRC compared with non-tumor tissues of the same patient. Meanwhile, low expression of TUSC7 indicates worse survival in CRC patients. 71
Conclusion and perspectives
CRC currently ranks third in tumor incidence rates, with annual new cases of more than 1,000,000. This situation of CRC brings huge economic and sprint burden to the people’s life. Although great advancements for CRC treatment have been developed, the low survival rates of CRC patients still exist, which may be promoted by the lack of a specific and efficient screening method. On account of difficulty to recognize symptoms, patients with CRC at an advanced stage are often treated. Therefore, it is momentous to identify molecular biomarkers for early diagnosis and predicting disease outcome. This condition starves for exploring corresponding indexes. Currently, some molecular biomarkers for CRC, as a convenient and economical screening method for CRC, are widely used in clinical medicine, for example, CEA, CA199, and CA125, which are also used for other tumor types, such as pancreatic, gastric, and lung cancers. Nonetheless, the specificity and sensitivity of these conventional molecular biomarkers for CRC are dissatisfactory to clinical diagnosis and prognosis. Thus, searching for effective biomarkers is indispensable because such biomarkers would be a beneficial supplement to the detection and assessment of CRC.
Recently, lncRNAs have attracted extensive attention and have been studied broadly to ascertain their complex mechanisms and functions. Here, we have highlighted on the late advancements that lncRNAs are involved in CRC and on the ability of diagnosis and prognosis in patients with CRC, which may satisfy actual clinical requirements. Accumulating evidence has shown that lncRNAs regulating cancer cell proliferation, migration, and apoptosis might also serve as biomarkers for CRC diagnosis and prognosis in clinical setting either in tissue samples or in serum. First, in tissue samples, for instance, the lncRNA-AFAP1-AS1 expression was upregulated in CRC tissues and cells, which can promote cancer cell proliferation and metastasis. 17 Second, in serum samples, for example, LOC285194 and RP11-462C24.1 can provide a new method that could differentiate patients with CRC from healthy controls. 67 Furthermore, several dysregulated lncRNAs are present in both cancer tissues and patient serum samples, such as BANCR, NR_034119, NR_029373, and NR_026817. 79 In conclusion, lncRNA is involved in the development and progression of CRC and shows that lncRNA may be a potential diagnostic and new prognostic target in patients with CRC.
But a wide range of clinical application of lncRNA in CRC may be still on the road. First, up to now, there did not appear a diagnostic panel based on the result of lncRNA research, which can directly, early, accurately diagnose and prognose patients with CRC. This diagnostic panel may have a consistent pattern that is composed of dozens of dysregulated lncRNAs in CRC cells. Based on the aberrant expression data of lncRNAs, researcher designs score-marking rules that could directly calculate the malignant degree, TNM stage, transfer, as well as the risk of recurrence of CRC, the disease-specific survival rate of CRC patients, and guide the further treatment of choice including operative programme, chemotherapy regimen, and palliative treatment. Second, lncRNAs in serum samples of patients are less researched than lncRNAs in CRC tissues and CRC cells, although the detection of lncRNAs in serum also has been studied. Because auxiliary tools in diagnosing and predicting cancers require non-invasive sampling methods for detection and a low economic burden, circulating lncRNAs, as diagnosing biomarkers, serve the purpose due to its easier accessibility. Studies have demonstrated that lncRNAs are stably present in serum for an extended time, even after exposing to multiple freeze-thaw cycles, room temperature, and high or low pH. 80 These added new requirements and theoretical guarantee to future research for circulating lncRNAs. More research on circulating lncRNAs, as powerful CRC indicators, must be conducted in the future.
According to the above fact, in order to optimize the prognostic and diagnostic use of lncRNAs in CRC, increase its effect and make up for its deficiency, further research in the field of lncRNAs is required.
Footnotes
Acknowledgements
H.D., M.L., and J.M.W contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
