Abstract
UNC5C is a member of the UNC5H family of transmembrane receptors and functions as a dependence receptor. The expression of UNC5C is lost or markedly reduced in a large proportion of cancers at the messenger RNA level. However, there is little information available regarding the protein expression of UNC5C, the relationship between UNC5C protein expression and UNC5C methylation, and the correlation between patient clinical features and UNC5C protein expression in colorectal cancer. In this study, the methylation and protein expression of UNC5C were examined in 36 adenomatous polyps, 73 colorectal cancers, and 28 corresponding normal mucosa, and the correlation between the methylation, as well as protein expression status, and the clinicopathologic features was evaluated. Furthermore, the relationship between the methylation and protein expression of UNC5C, and correlation between UNC5C protein expression and overall survival were analyzed. The results showed that aberrant methylation of UNC5C was observed in colorectal cancers (78%) and adenomatous polyps (64%). The methylation-specific polymerase chain reaction results were confirmed by bisulfite sequencing of UNC5C promoter region. UNC5C methylation was significantly higher in early tumor, node, metastasis stage (I + II) of colorectal cancers. Compared with the corresponding normal tissues, protein expression of UNC5C was significantly lower in colorectal cancers (42%) and adenomatous polyps (81%). Protein expression of UNC5C was significantly higher in early tumor, node, metastasis stage (I + II) of colorectal cancers compared with advanced tumor, node, metastasis stage. Furthermore, patients with UNC5C-negative expression had a poorer prognosis than those with UNC5C-positive expression through Kaplan–Meier survival analysis (p = 0.038), univariate (p = 0.044) and multivariate analysis (p = 0.045). According to Spearman rank correlation analysis, UNC5C methylation and protein expression were negatively correlated (r = −0.461, p < 0.001). Together, these results suggest that UNC5C methylation may be an earlier event in the development of colorectal cancer, which was negatively correlated with protein expression. UNC5C may have a critical role in the pathogenesis of colorectal cancers and be a valuable prognostic factor of colorectal cancers patients. UNC5C may be identified as an attractive therapeutic target for the treatment of colorectal cancers in the further studies.
Introduction
Colorectal cancer (CRC) is one of the most common malignancies and a leading cause of cancer deaths worldwide. Recent statistics show that there are more than 1 million CRC cases with about nearly 700 thousand deaths every year.1,2 The prognosis of CRC is related to the stage at diagnosis, with a 5-year survival rate of 90% at early diagnosis and less than 10% when distant metastases develop. 3
UNC5C is a member of the UNC5H family of transmembrane receptors and functions as a dependence receptor. When the ligand netrin-1 is present, UNC5C transmits positive signals of proliferation and migration. On the other contrary, when the ligand is absent, UNC5C sends proapoptotic signals to trigger cell death. 4 The expression of UNC5C is lost or markedly reduced in a large proportion of cancers,5–7 including CRC. Inactivation of UNC5C would contribute not only to tumor progression but also to the initiation of tumor formation. So, recent several studies suggest that UNC5C is likely to be a tumor suppressor.8,9
CRC develops as a consequence of genetic and epigenetic alterations. Gene mutations have long been known to be important in cancer formation. However, epigenetic alterations have only recently been recognized as significant contributors to cancer development. Accumulation of epigenetic changes including aberrant DNA methylation, abnormal histone modification, and noncoding RNAs was found in CRC and other tumors.10,11 Epigenetic changes may serve as a diagnostic, prognostic, and chemo-sensitive marker. Because epigenetic changes are reversible under certain circumstances, UNC5C becomes a possible cancer preventive or therapeutic target by epigenetic-related reagents.12,13
In this study, the methylation and protein expression status of the UNC5C were examined in 36 adenomatous polyps (APs), 73 primary CRCs, and 28 corresponding normal mucosa, and the correlation between the methylation, as well as protein expression status, and the clinicopathologic features was evaluated. Furthermore, the relationship between the methylation and protein expression of UNC5C, and correlation between UNC5C protein expression and overall survival were analyzed.
Materials and methods
Sample collection and DNA isolation
A total of 73 CRC tissue specimens, 36 AP tissue specimens, and 28 corresponding normal tissue specimens were collected consecutively at the First Affiliated Hospital of Jinzhou Medical University from patients during colorectal surgery. No patient underwent radiotherapy or chemotherapy before surgery. All the tissue specimens were confirmed histologically. All the clinical information and follow-up data were based on studies from our tumor registry service. Postoperative follow-up has lasted at least 5 years for 73 CRCs patients. This study was approved by the Ethics Committee of Jinzhou Medical University, and written informed consent was acquired from each patient. The clinicopathologic profiles of the patients enrolled in the study are shown in Table 1.
Characterization of the tissue samples enrolled in the study.

Methylation-specific polymerase chain reaction
DNA was prepared and modified according to the procedure described by Lee et al. 14 Briefly, genomic DNA was denatured with NaOH. Then, hydroquinone (Sigma, St. Louis, MO) and sodium bisulfite (pH 5.0; Sigma) were added and mixed well, and samples were incubated at 50°C for 16 h. Modified DNA was purified using Wizard DNA clean-up kit (Promega, Madison, WI).
After bisulfite modification, methylation-specific polymerase chain reaction (MSP) was performed. The primer sequences 8 of UNC5C gene for unmethylated products were 5′-AAATTCAACTAAAACTTTACC AACACAACA-3′ (upstream) and 5′-TAATTGGGGGTGGGAGTATGTTTGTTT-3′ (downstream). The primer sequences of UNC5C gene for methylated products were 5′-CGACTAAAACTTTACCGACGCGACG-3′ (upstream) and 5′-CGGGGGCGGGAGTACGTTCGTTC-3′ (downstream). The 25-µL total reaction volume contained 2.5 µL TaKaRa Taq HS, 2.5 µL MgCl2 (25 mM), 4 µL deoxynucleotide triphosphate (dNTP) mix (2.5 mM each), 1 µL upstream primer (10 µM), 1 µL downstream primer (10 µM), and 50–100 ng bisulfite-modified DNA. Polymerase chain reaction (PCR) amplification was performed as follows: 95°C for 15 min, 35 cycles of 95°C for 30 s, annealing at 68°C (methylated primers) or 62°C (unmethylated primers) for 30 s, and finally a 30-s extension at 72°C. A final 10-min extension at 72°C completed each PCR. A total of 10 µL of each PCR product was loaded into 2% agarose gel and was visualized after staining with ethidium bromide.
Cloning and sodium bisulfite sequencing
Bisulfite-modified DNA from five methylated tumor specimens and five unmethylated normal samples selected randomly were amplified using primers 8 specific to the bisulfite-modified sequence in the CpG island (upstream: 5′-CCTCCCTAAACTTCCCAAATAAAA-3′, downstream: 5′-GTAGGGTTAGGGTA GGTAGTA-3′). Each of the PCR products subsequently was cloned using the pMD18-T cloning system (Takara, Kyoto, Japan). Positive clones identified by double enzyme digestion with PCR products were sequenced to confirm methylation status at the CpG sites in the analyzed region.
Immunohistochemistry for UNC5C protein
Immunostaining for UNC5C was performed using a monoclonal antibody to UNC5C protein (MAB1005, dilution 1:100; R&D Systems Inc., Minneapolis, MN). Tissue sections (4 µm thick) were deparaffinized with xylene and rehydrated with an ethanol gradient. The sections were retrieved and treated with hydrogen peroxidase to block endogenous peroxidase activity. The primary antibodies were incubated at 4°C overnight. Thereafter, sections were incubated with a secondary anti-mouse antibody conjugated with horseradish peroxidase and treated with 3,3-diaminobenzidine Kit (Zhongshan Biotechnology Inc., Beijing, China). Last, sections were counterstained with hematoxylin. All slides were independently examined and scored by two pathologists who were blind to the clinical and pathological information of the subjects. For UNC5C immunohistochemical staining assessment, the ratio of positive cells per specimen and staining intensity was analyzed. The UNC5C immunoreactivity level was classified by the proportion of positive cells: 0, <5% positive cells; 1+, 5%–30% positive cells; 2+, 30%–50% positive cells; and 3+, >50% positive cells. The intensity of UNC5C expression was also scored: 0, negative to weak; 1, moderate; and 2, strong. The score was the sum of the intensity and the percentage of positive cells. A score of ≤1 was applied as a cut-off point for loss of UNC5C expression.
Statistical analysis
Methylation or protein expression of UNC5C in APs, CRCs, and corresponding normal mucosa was analyzed using chi-square tests. The associations between UNC5C methylation or expression and clinicopathologic parameters were analyzed also using chi-square tests. Relationships between methylation and protein expression of UNC5C gene were analyzed using Spearman rank correlation analysis. Cox regression was adopted to analyze the effect of several risk factors on survival. Differences were considered significant when p value was <0.05.
Results
Methylation status of UNC5C gene
The isolated genomic DNA of samples and amplification examples detected by MSP of corresponding normal tissue and tumor were analyzed by agarose gel electrophoresis. Samples showing only unmethylated signals were scored as unmethylated, and samples showing only methylated signals were scored as methylated. If samples show both unmethylated and methylated signals, they were scored as methylated. Each sample was measured three times, and only samples with consistent methylated signals were regarded as methylated (Figure 1).

Methylation-specific polymerase chain reaction. (a) Gel electrophoresis analysis of extracted genomic DNA. (b) One example of normal tissue showed only unmethylated UNC5C band. (c) One example of tumor tissue showed only methylated band. (d) One example of tumor tissue showed both unmethylated and methylated bands. (1: methylated allele; 2: unmethylated allele; M: 100 bp DNA ladder).
Aberrant methylation of UNC5C gene was present in 57 out of 73 CRC cases (78%) and 23 out of 36 APs cases (64%). However, in 28 corresponding normal mucosa of CRC, methylation of the UNC5C gene was not detected. The UNC5C gene was significantly more often methylated in CRC and AP than in corresponding normal mucosa. But there was no significant difference of UNC5C methylation between CRC and APs (Table 2).
Methylation of UNC5C in APs, CRCs, and corresponding normal mucosa.

Cloning and sodium bisulfite sequencing
Five methylated tumors and five unmethylated normal samples were randomly selected for bisulfite sequencing PCR analysis. Gel electrophoresis analysis of the amplification products revealed a single band with expected size (368 base pairs (bp)). The PCR clones identified by double enzyme digestion were sequenced (Figure 2). Comparing the results with the original sequence showed that CpG island methylation was not found in unmethylated normal samples, but the percentage of methylation at the CpG sites in the analyzed region was high (>90%) in methylated tumors (Figure 3).
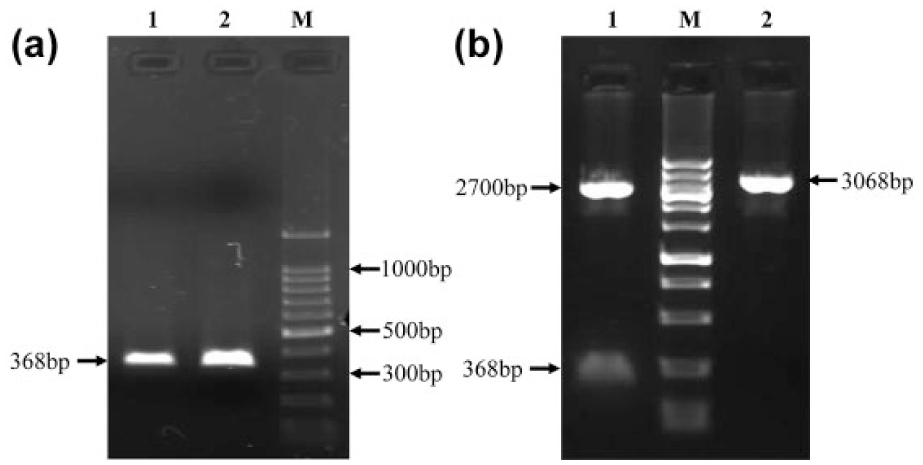
Gel electrophoresis analysis of bisulfite sequencing PCR. (a) Products of BSP (lanes 1 and 2). (b) Recombinant plasmid pMD18-T was digested by EcoRI and HindIII and target fragment were found (lane 1). (M: DNA ladder).

Change in MeCpG content in the region of the UNC5C promoter. PCR clones were sequenced from five different tumors (methylated) and from five corresponding normal tissues (unmethylated) to confirm methylation at the CpG sites in the analyzed region.
Correlation between UNC5C gene methylation and clinicopathologic parameters in CRCs
We examined associations between UNC5C gene methylation status with clinicopathologic parameters in CRCs. We found significantly more UNC5C methylation in elderly (≥60 years) compared with younger (<60 years) patients, whereas there were no significant associations were found when patient gender, tumor location, or tumor differentiation were compared in methylated versus unmethylated CRCs. In addition, we found that UNC5C methylation was significantly greater in tumor, node, metastasis (TNM) stage I and stage II CRCs compared with TNM stage III + IV CRCs (Table 3).
Correlation between UNC5C methylation and clinicopathologic parameters of CRC (n = 73).

CRC: colorectal cancer; TNM: tumor, node, metastasis.
Protein expression of UNC5C
Immunohistochemical staining showed that UNC5C protein was predominantly localized in the cytoplasm (Figure 4). Among these CRC and AP patients, the protein expression of UNC5C was positive in 31 out of 73 CRC cases (42%) and 29 out of 36 APs cases (81%). However, in 28 corresponding normal mucosa of CRC, the protein expression of UNC5C was all positive (100%). Compared with the corresponding normal tissues, protein expression of UNC5C was significantly lower in CRC and AP. There was also significant difference of UNC5C expression between CRCs and APs (Table 4).

Immunostaining of UNC5C expression. (a) Adenomatous polyps, (b) highly differentiated CRC, (c) moderately differentiated CRC, and (d) poorly differentiated CRC.
Protein expression of UNC5C in APs, CRCs, and corresponding normal mucosa.

Correlation between UNC5C protein expression and clinicopathologic parameters in CRCs
The associations between UNC5C protein expression and clinicopathologic parameters were analyzed in CRCs. Results showed that UNC5C immunostaining was positive in 6 (60%) out of 10 TNM stage I cases, positive in 14 (58%) out of 24 TNM stage II cases, and positive in 11 (28%) out of 39 TNM stage III + IV cases. Protein expression of UNC5C was significantly higher in early TNM stage (stage I and stage II) compared with advanced TNM stage (stage III + IV) patients. However, there were no significant associations found between UNC5C expression and age, gender, tumor location, or tumor differentiation. Detailed results are shown in Table 5.
Correlation between UNC5C expression and clinicopathologic parameters of CRCs (n = 73).
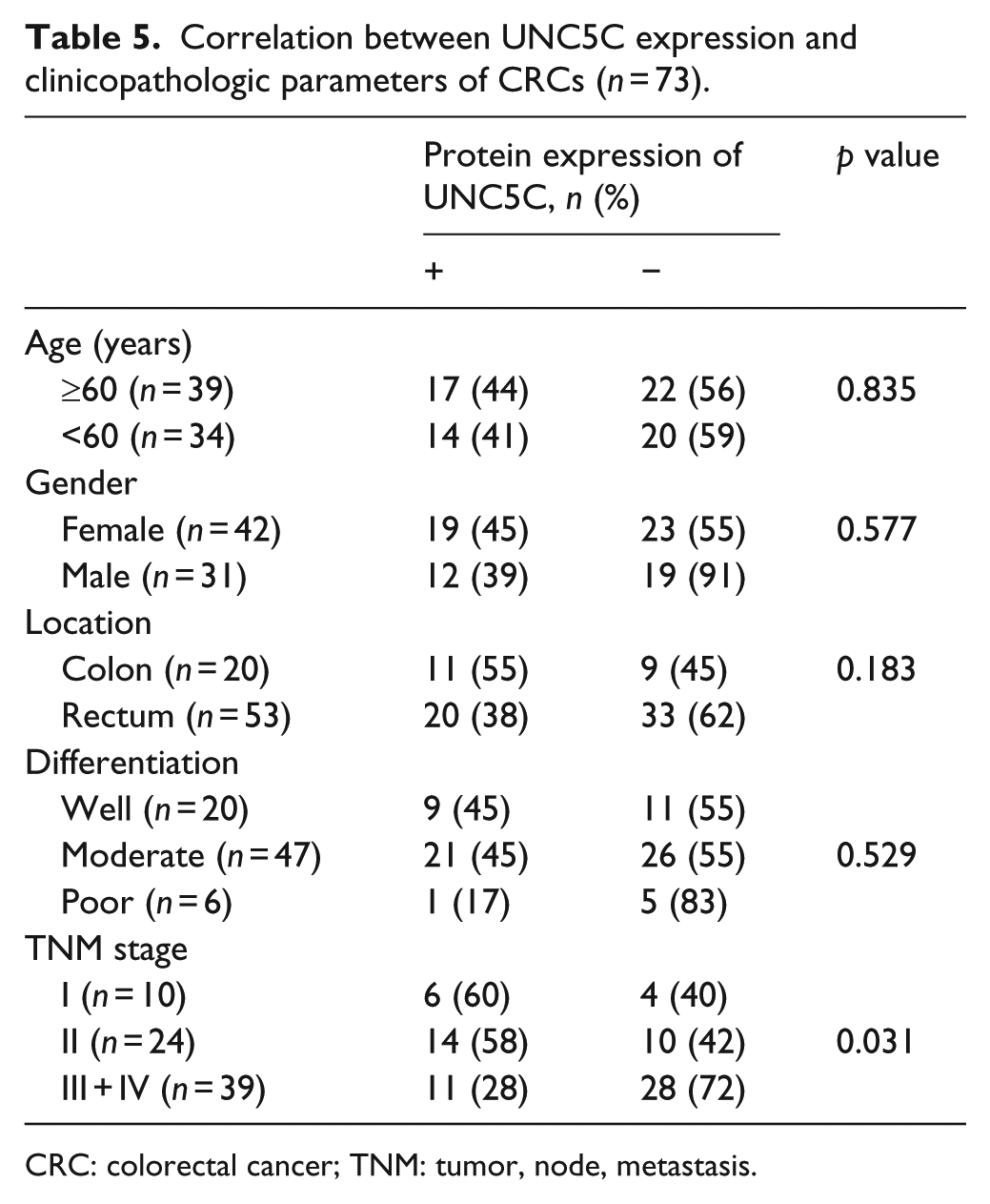
CRC: colorectal cancer; TNM: tumor, node, metastasis.
Relationship between the methylation and protein expression of UNC5C in CRC and AP
In 80 UNC5C methylated cases of APs and CRC, 33 cases of UNC5C protein expression were positive and 47 cases were negative. While, in 29 UNC5C unmethylated cases of APs and CRC, 27 cases of UNC5C protein expression were positive and 2 cases were negative. According to Spearman rank correlation analysis, UNC5C methylation and protein expression were negatively correlated (r = −0.461, p < 0.001; Table 6). In addition, the relationship of UNC5C methylation and protein expression in different TNM stages was analyzed. Results showed that UNC5C protein expression was positively correlated with methylation, although this association did not achieve statistical significance (r = 0.500, p = 0.667).
Positive expression and methylation of UNC5C in CRCs and APs (n = 109).
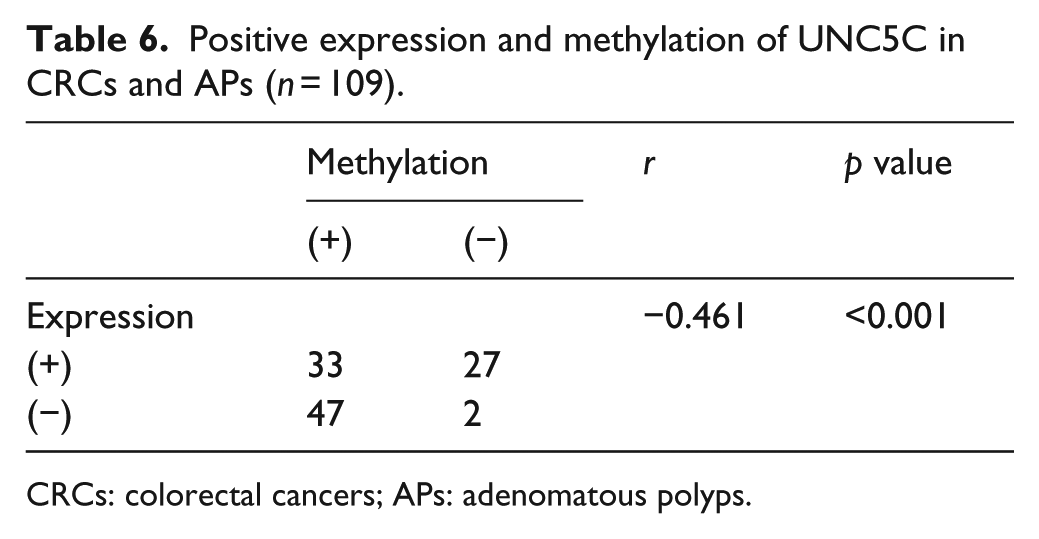
CRCs: colorectal cancers; APs: adenomatous polyps.
Correlation between expression of UNC5C and overall survival
In the 73 followed up patients, the 5-year survival rate for patients with positive expression of UNC5C was 61.29% compared to 40.48% in patients with negative expression. According to the Kaplan–Meier survival analysis, patients with UNC5C-negative expression had a poorer prognosis (p = 0.038; Figure 5(a)) than those with UNC5C-positive expression. Cox proportional hazards model was adopted to do univariate analysis and multivariate analysis of survival. Through the univariate analysis, we found TNM stage and UNC5C expression (p = 0.044) were associated with overall survival. In multivariate analysis, the expression of UNC5C emerged as an independent and significant factor associated with better 5-year survival (hazard ratio (HR) = 2.242; 95% confidence interval (CI) = 1.017–4.944; p = .045). (Figure 5(b), Table 7).
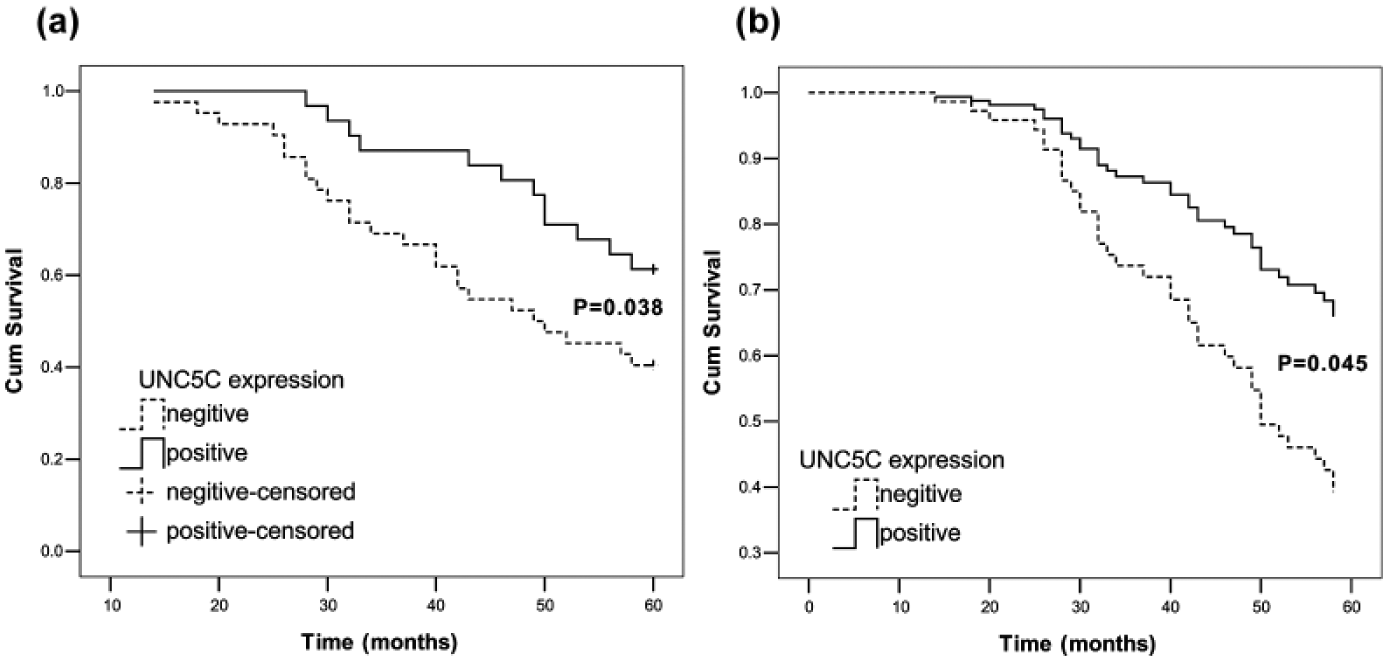
Survival curves by patients’ UNC5C expression status. (a) Kaplan–Meier curves showing 5-year overall survival of CRC patients in relation to UNC5C expression. Log-rank test, p = 0.038. (b) Multivariate Cox proportional hazard models for overall survival.
Univariate and multivariate Cox proportional hazard models for overall survival.

HR: hazard ratio; CI: confidence interval; TNM: tumor, node, metastasis.
Discussion
The tumorigenesis of CRC is a complicated process with multi-step events of the molecular pathway involving the activation of oncogenes and inactivation of tumor suppressor genes. Four different genomic and epigenomic instabilities including chromosome instability, microsatellite instability (MSI), CpG island methylator phenotype (CIMP), and DNA global hypomethylation have been described for CRC. 15 These epigenetic changes affect every phase of tumor development from initiation to metastasis. DNA methylation is known to be abnormal in all forms of cancer, which is a way of inactivation of tumor suppressor genes. DNA methylation can also lead to abnormal expression of genes related to cell proliferation and differentiation, resulting in loss of control of the normal growth of cells and the occurrence of malignant tumor formation.
UNC5C is a proapoptotic molecule that governs axon migration in cooperation with its ligand, netrin-1. 16 UNC5C expression is not restricted to the nervous system but can be detected in thyroid, kidney, ovary, uterus, stomach, colon, lung, spleen, bladder, and breast tissues, suggesting a role for these receptors out of the neuronal guidance system. 17 Research shows that the expression of UNC5C is lost or markedly reduced in RNA level in a large proportion of cancers,5–7 including CRC. But there is little information available regarding the protein expression status of UNC5C, the relationship between UNC5C protein expression and UNC5C methylation, and the correlation between patient clinical features and UNC5C protein expression in CRC.
Prior studies reported that aberrant methylation of the netrin-1 receptor genes including UNC5C were frequently observed in primary colon cancers.9,18 In this study, aberrant methylation of UNC5C gene was observed in CRCs (78%) and APs (64%) than in corresponding normal mucosa (p < 0.001). To confirm MSP results, we also performed bisulfite sequencing of UNC5C promoter region in five pairs of tumor/corresponding normal tissues. Indeed, we observed that the UNC5C is highly methylated in tumor samples, while no methylation occurs in normal tissues (Figures 2 and 3). These results are consistent with previous studies. The correlation between the methylation status of UNC5C and the clinicopathologic parameters was further evaluated in CRCs. A significant difference in TNM stage (p = 0.041) was found. UNC5C methylation was significantly higher in early TNM stage (I + II) of CRCs compared with advanced stage (III + IV). This result suggests that UNC5C methylation may be an earlier event in the development of CRC. However, Hibi et al. 18 reported that UNC5C was more frequently methylated in advanced colorectal carcinomas, while the result of Shin et al. 9 showed no significance between UNC5C methylation and tumor stage (Dukes’). But, Shin et al. 9 found that methylation-induced silencing of UNC5C by itself was significantly more frequent in Dukes’ stage A neoplasms. These different results may be perhaps because of the samples from different regions. Whether UNC5C methylation is an earlier event or late event needs to be further studied in the future.
Some studies show that the expression of UNC5C is lost or markedly reduced in a large proportion of cancers5–7 at the messenger RNA (mRNA) level, including CRC. Previous studies reported that the inactivation of Unc5h3 in mice, the human UNC5C ortholog, was shown to enhance progression of intestinal tumors. 8 More recently, the function of the UNC5 family of receptors has been implicated in the regulation of cell death processes in bladder cancers.19,20 And, UNC5C has been confirmed as a new risk factor for Alzheimer’s disease (AD) recently. 21 In our study, the protein expression of UNC5C was detected using immunohistochemistry (IHC). Compared with the corresponding normal tissues (100%), protein expression of UNC5C was significantly lower in CRCs (42%) and APs (81%). There was also significant difference of UNC5C expression between CRCs and APs. Being consistent with previous reports,9,18 these results also suggested tumor suppressive effects of the UNC5C gene due to its downregulation in colorectal malignancies. The associations between UNC5C protein expression and clinicopathologic parameters were analyzed in CRCs. Result showed that the protein expression of UNC5C was significantly higher in early TNM stage (I + II) compared with advanced TNM stage (III + IV). This result also suggested tumor suppressive effects of UNC5C. Furthermore, the relationship between UNC5C expression and the 5-year overall survival was evaluated. Kaplan–Meier survival analysis (log-rank, p = 0.038), univariate (HR = 2.030; 95% CI = 1.019–4.047; p = 0.044) and multivariate analysis (HR = 2.242; 95% CI = 1.017-4.944; p = 0.045) showed that patients with UNC5C-negative expression had a poorer prognosis than those with UNC5C-positive expression. These results suggest that UNC5C may have prognostic value and may be identified as an attractive therapeutic target for the treatment of CRCs.
Finally, the relationship between the methylation and protein expression of UNC5C in CRC and AP was analyzed. According to Spearman rank correlation analysis, UNC5C methylation and protein expression were negatively correlated (r = −0.461, p < 0.001). In addition, the relationship of UNC5C methylation and protein expression in different TNM stages was analyzed. However, we did not find significant association. These results suggest that gene methylation may cause the downregulation of UNC5C. Other mechanism such as loss of heterozygosity (LOH), leading to decreased or absent protein expression of UNC5C in colorectal tumors has been reported. 8 The specific mechanism of decreased expression of UNC5C is still need to be further studied.
In summary, this study found that aberrant methylation of UNC5C gene was observed in CRCs and APs. UNC5C methylation was significantly higher in early TNM stage. The protein expression of UNC5C was significantly lower in CRCs and APs and which was also significantly higher in early TNM stage (I + II) compared with advanced stage (III + IV). UNC5C protein expression was negatively correlated with methylation status (r = −0.461, p < 0.001). UNC5C methylation may cause decreased protein expression. Furthermore, patients with UNC5C-negative expression had a significantly poorer prognosis than those with UNC5C-positive expression.
Footnotes
Acknowledgements
The authors thank all researchers who contributed to this work, and the authors apologize to the researchers whose works could not be discussed here due to space limitations.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Written informed consent was obtained from all patients. The study was approved by the Ethics Committee of Jinzhou Medical University.
Funding
This work was supported financially by Science research project from Department of Education of Liaoning Province, No. L2015322 (to Jie Wu), and President Foundation of Jinzhou Medical University (No. XZJJ20140119).
