Abstract
The pathogenesis of colorectal cancer remains poorly understood. Here, we show that coactivator-associated arginine methyltransferase 1 is frequently upregulated in colorectal cancer tissues and promotes cell growth in vitro and in vivo. Using bioinformatics-based prediction and luciferase reporter system, we found that coactivator-associated arginine methyltransferase 1 is post-transcriptionally targeted by microRNA-195-5p in colorectal cancer. Ectopic expression of microRNA-195-5p led to the suppression of the coactivator-associated arginine methyltransferase 1 3′-untranslated regions activity and downregulation of the endogenous coactivator-associated arginine methyltransferase 1 protein in colorectal cancer cells. Expression analysis verified that microRNA-195-5p was markedly downregulated in human colorectal cancer tissues, which was negatively correlated with the elevated levels of coactivator-associated arginine methyltransferase 1 protein. Enhanced levels of microRNA-195-5p in colorectal cancer cells resulted in a sharp reduction of cell proliferative and colony-formative capacities in vitro. Remarkably, restoration of coactivator-associated arginine methyltransferase 1 in microRNA-195-5p-transfected colorectal cancer cells partially abrogated the inhibition of cell proliferation and colony formation mediated through microRNA-195-5p. These data confirm that microRNA-195-5p might function as an anti-tumor microRNA in colorectal cancer exerting critical control over coactivator-associated arginine methyltransferase 1 expression. The newly identified microRNA-195-5p/coactivator-associated arginine methyltransferase 1 axis may act as a novel promising therapeutic target for colorectal cancer treatment.
Keywords
Introduction
Colorectal cancer is the third most frequently diagnosed cancer and the leading cause of cancer death in the world. 1 It is estimated that over 1.4 million new colorectal cancer cases and more than 693,900 deaths occurred in 2012. 2 Although much effort has been devoted to probing the pathogenesis of colorectal cancer, the exact molecular mechanisms involved in the process are still quite obscure. Hence, it is urgent to explore the potential functions of key molecules in colorectal cancer. 3
Coactivator-associated arginine methyltransferase 1 (CARM1), a member of protein arginine methyltransferase (PRMT) family, catalyzes asymmetric methylation of the arginine (Arg) residues in protein substrates. 4 Among different members of PRMT family, CARM1 was first identified to be associated with transcriptional regulation. 5 CARM1, according to the published literatures, can regulate multiple important biological processes, such as pre–messenger RNA (pre-mRNA) splicing, 6 gene regulation, 5 cell-cycle progression, 7 and the DNA damage response. 8 Given its different roles in cellular processes, it is not surprising that CARM1 is linked with cancer progression. Abnormal expression of CARM1, generally upregulated, has been reported in different types of tumors.9–11 However, the mechanism of action of CARM1 remains obscure in cancer, including colorectal cancer.
MicroRNAs (miRNAs), as post-transcriptional regulators, exert their functions by the direct binding to the mRNA 3′-untranslated regions (3′-UTR), leading to translational repression or mRNA degradation. 12 Primarily, miRNAs function as crucial factors in diverse regulation pathways, 13 including tumorigenesis, cell differentiation, proliferation, and apoptosis. Accumulating evidence indicates that miRNAs can act in human carcinogenesis as novel types of tumor suppressors or oncogenes.14–16 Although the importance of miRNAs in cancer has attracted much attention in recent years, the exact molecular functions of miRNAs and their effects on cancer are not well understood.
Here, we demonstrated that CARM1 is markedly upregulated and promotes cell growth in colorectal cancer in vitro and in vivo. Furthermore, we found that CARM1 is specifically targeted by microRNA-195-5p (miR-195-5p), which controls cell proliferation and colony formation in vitro, indicating an anti-tumor function of miR-195-5p in colorectal cancer. The newly identified miR-195-5p/CARM1 axis may function as a key pathway regulating tumor cell growth during the progression of colorectal cancer.
Materials and methods
Surgical tissue specimens
All the tumor tissues, comprising adjacent normal tissues, were derived from colorectal cancer patients. All tumor tissue sections of colorectal cancer were reviewed histologically by senior pathologists. Informed consent was obtained from all patients, and the research was approved by the Ethics Committee at Xinhua Hospital Chongming Branch. Patients’ characteristics are listed in Table 1.
Colorectal cancer patients’ characteristics.

Cell culture
Two colon carcinoma cell lines HCT15 and SW480 were obtained from Cell Bank of the Chinese Academy of Sciences (Shanghai, China). HCT15 and SW480 cells were cultured in RPMI 1640 (HCT15) and L-15 (SW480) media supplemented with 10% fetal bovine serum (FBS; GIBCO) under 5% CO2 at 37°C.
Plasmid construction and small interfering RNA transfection
In order to silence CARM1 expression stably, the short hairpin RNA (shRNA) of CARM1 and scramble shRNA were cloned into the pLL3.7 vector, respectively. The targeting oligonucleotides were 5′-GGATAGAAATCCCATTCAA-3′. The small interfering RNA (siRNA) targeting CARM1 were introduced into HCT15 and SW480 cells using Lipofectamine 3000 transfection reagent (Invitrogen). Two human CARM1 siRNA sequences were as follows:
siRNA-1: 5′-GAAGGAGAUUUGCACAGG-3′;
siRNA-2: 5′-AGACAGAGCUACGACAUCA-3′.
Real-time polymerase chain reaction
Total RNA was isolated from cells or tissues by TRIzol extraction (Invitrogen), and miRNA was extracted using commercial miRNeasy Mini kit (Qiagen). After quantification by a Nanodrop (Thermo), complementary DNA (cDNA) synthesis was carried out using a reverse transcription system (Takara). Real-time polymerase chain reaction (PCR) detection was carried out (Applied Biosystems) using the SYBR green mix (Roche) according to the manufacturer’s instructions. CARM1 expression was normalized against glyceraldehyde 3-phosphate dehydrogenase (GAPDH) or U6 for miR-195-5p. Primer sequences used for real-time PCR were as follows:
CARM1 forward primer: 5′-GAAGCCCAGCGGAAACAT-3′;
CARM1 reverse primer: 5′-GGTACCAGAAGTTGGCCTTG-3′;
GAPDH forward primer: 5′-GGTTCCCAAGTATTGTCAGCA-3′;
GAPDH reverse primer: 5′-CATGTAGTTGAGGTCAATGAAGG-3′.
Luciferase reporter assay
First, human genomic DNA was isolated from 293T cells as a PCR template. CARM1 3′-UTR fragment, containing two binding sites of miR-195-5p, was amplified by PCR and then cloned into the pMIR-REPORT™ luciferase vector (Invitrogen) downstream of the luciferase gene. Mutants were obtained using a point-mutation kit (SBS Genetech) and confirmed by sequencing. The binding sequence of miR-195-5p in the regions of CARM1 3′-UTR was mutated from UGCUGGU to GGCAGGA. For luciferase reporter assays, each well was co-transfected with 1 µg luciferase reporter plasmid, 0.5 µg β-gal plasmid (Invitrogen), together with miR-195-5p or miR-NC (100 pmol). Luciferase activity analysis was performed using a luciferase reporter assay kit (Promega) 48 h post-transfection. β-gal plasmid was used as a control to monitor transfection efficiency.
Immunoblot analysis
Total lysates extracted from cells and tissues were prepared in lysis buffer (125 mM NaCl, 1% NP-40, 50 mM Tris–HCl, 0.1% sodium dodecyl sulfate (SDS), and Protease Inhibitor Cocktail). Proteins (60 µg) were resolved in 12% SDS–polyacrylamide gel electrophoresis (PAGE) and blotted onto polyvinylidene difluoride (PVDF) membrane for 3 h at 12 V using the semidry transfer system (Bio-Rad). The membrane was then blocked in PBST (phosphate-buffered saline with Tween-20) with 5% no-fat milk for 1–2 h at room temperature. Primary antibodies, anti-CARM1 (Cell Signaling, 3379), and anti-GAPDH (MBL, M171-3) were used for incubation. After washing with PBST, the membranes were incubated with the corresponding secondary antibodies linked with horseradish peroxidase (HRP; Sigma, 1:100,000). Proteins were immune-detected by ECL detection system (Pierces).
Proliferation and colony-formation assays
Transfected HCT15 and SW480 cells were split into 96-well plate with 2000 cells per well. At the indicated time points, 10 µL of Cell Counting Kit-8 (CCK-8) reagent was added to each well and then incubated for 2 h in standard culture conditions. Proliferative activity was assessed by measurement of absorbance at 450 nm. For colony-formation assay, HCT15 and SW480 cells (1000 cells/well) were split into six-well plates and grown for 14 days. Before staining (0.1% crystal violet), cells were washed twice with ice-cold PBS and then fixed with 4% paraformaldehyde. Finally, the crystal violet solution was removed and the colonies were dried at room temperature before counting (>50 cells).
Immunohistochemistry assay
Tumor and nontumor tissues were embedded with paraffin and sectioned by a microtome. The sections were deparaffinized and rehydrated. After antigen retrieval and blocked with 3% H2O2, sections were incubated with CARM1 primary antibody (Cell Signaling, 3379) at 4°C overnight. Next, the sections were incubated with specifically secondary antibody. Visualization of CARM1 protein was performed with DAB (Invitrogen). IHC staining of CARM1 was scored independently by two pathologists who were blind to pathological data of all patients.
In vivo assay
SW480 cells (CARM1 knockdown or negative control) were digested by trypsinization and washed twice with ice-cold PBS. Before subcutaneous injection into flanks of mice, 5 × 106 cells were suspended in 100 µL PBS containing 10% Matrigel (v/v). Tumor volume measurement was performed by a caliper every 5 days, which is calculated as V = (length × width 2 )/2. After 20 days, all tumors excised from mice were analyzed immediately after euthanasia of mice.
Statistical analysis
Results are shown as mean ± standard deviation (SD), and all experiments included at least six replicates per group. Data analysis was performed using Student’s t test, and p value below 0.05 was considered statistically significant. All statistical analysis was performed with SPSS 17.0 software.
Results
CARM1 is upregulated in colorectal cancer
To assess the expression of CARM1 in colorectal cancer, we measured CARM1 expression in 45 tumor samples with immunohistochemical (IHC) staining. Remarkably, overexpression of CARM1 protein was observed in tumor tissues in comparison with their matching nontumor tissues from colorectal cancer patients (Figure 1(a) and (b), **p < 0.01), and CARM1 was located in the cytoplasm and nucleus of tumor cells. After extraction of total RNA, CARM1 mRNA analysis was carried out using quantitative reverse transcription polymerase chain reaction (qRT-PCR; Figure 1(c)). In comparison with the control group, CARM1 mRNA was significantly elevated in the colorectal cancer tissues. Together, the data from Figure 1 suggest that CARM1 is significantly increased in colorectal cancer, implying the role of CARM1 as a potential oncogene.
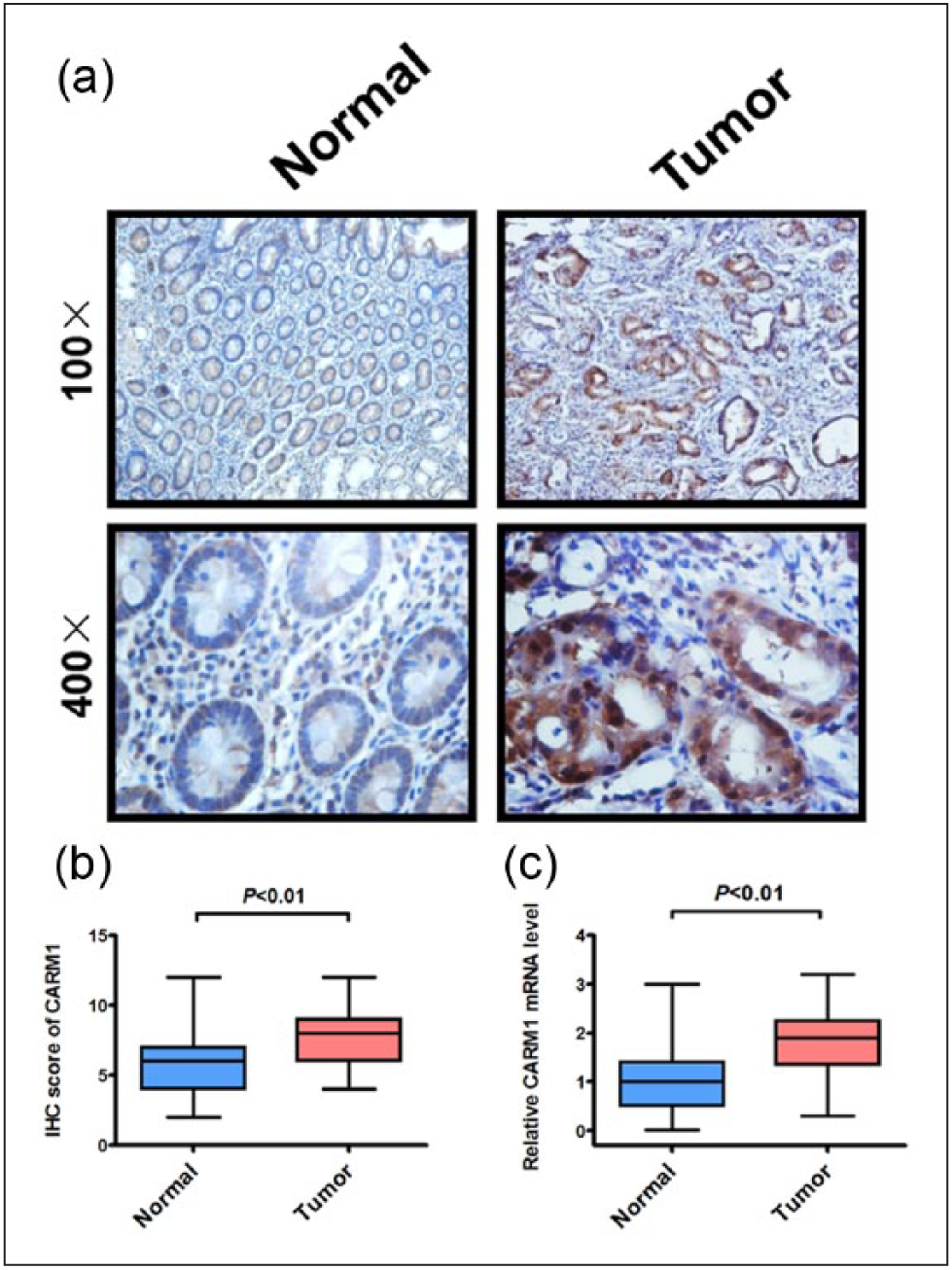
CARM1 is upregulated in colorectal cancer. (a) Immunohistochemical staining of CARM1 protein in normal and colorectal cancer tissues. Representative micrographs are shown in original magnification (100× and 400×) as indicated (left panel). (b) Histogram of mean integrated optical density (IOD) from immunohistochemical staining of cancer tissues; **p < 0.01, compared to normal control. (c) mRNA analysis of CARM1 in normal and colorectal cancer tissues (n = 45, **p < 0.01).
CARM1 depletion inhibits colorectal cancer cell growth in vitro and in vivo
We next investigated potential effect of CARM1 knockdown on colorectal cancer cells proliferation. We knocked down CARM1 expression in HCT15 and SW480 cell lines by gene-specific siRNA which reduced CARM1 protein and mRNA expression to less than 30% of the negative control (Figure 1S(a) and (b)). Remarkably, knockdown of CARM1 significantly reduced the cell proliferation rate in both cell lines (Figure 2(a)). Furthermore, knockdown of CARM1 significantly reduced the numbers of HCT15 and SW480 colonies formed in culture compared with controls (Figure 2(b)). To verify whether CARM1 has a growth-promoting effect on colorectal cancer cells, a xenograft tumor growth assay was performed in nude mice. Subcutaneous tumor growth of SW480 cells with CARM1 shRNA-mediated stable knockdown or scramble control was monitored (Figure 2(c)). Tumor volume (Figure 2(d)) and weight (Figure 2(e)) in CARM1 knockdown mice were greatly reduced in comparison with the scrambled controls (*p < 0.05). In addition, Western blot and real-time PCR analysis of xenograft tumor tissues suggested that CARM1 protein (Figure 2(f)) and mRNA levels (Figure 2(g), ***p < 0.001, n = 5) were markedly reduced compared with the controls. These data indicate that CARM1 contributes to colorectal cancer cell growth and is essential for colorectal cancer progression.

CARM1 depletion inhibits colorectal cancer cell growth in vitro and in vivo. (a) Proliferation of colorectal cancer cells transfected with siRNAs to CARM1 or negative control (NC) in both HCT15 and SW480 cells. Values at the indicated time points represent mean ± SD from six independent tests; **p < 0.01. (b) Colony-formation assay of HCT15 and SW480 cells transfected with siRNA to CARM1 or negative control; **p < 0.01. (c) Photograph of excised tumor tissues from mice. Top, tumors from scramble control; bottom, tumors from CARM1 knockdown. Scale bar, 1 cm. (d) Growth of tumors in nude mice from CARM1 knockdown or scramble control SW480 cells (*p < 0.05, n = 5). (e) Average tumor weight of xenograft tumors developed from CARM1 KD or scramble control SW480 cells (*p < 0.05). (f) Western blot assay showing CARM1 protein level in xenograft tumors following CARM1 knockdown in SW480 cells. (g) Real-time PCR assay showing CARM1 mRNA level in xenograft tumors following CARM1 knockdown in SW480 cells (***p < 0.001, n = 5).
CARM1 is a direct target of miR-195-5p
To search potential miRNAs targeting CARM1, we performed computational prediction analysis with three bioinformatics prediction programs (TargetScan, PicTar, and miRanda). Among the identified candidate miRNAs, miR-195-5p was identified by all three programs as targeting CARM1. To confirm whether miR-195-5p exerts its inhibitory effect through direct interactions with the CARM1 3′-UTR regions, the CARM1 3′-UTR regions containing two miR-195-5p binding sequence (UGCUGGU) were cloned downstream of luciferase gene in the reporter plasmid (Figure 3(a)). Afterward, luciferase activity was assessed in HCT15 and SW480 cells transfected with the reporter plasmid (β-gal as a control plasmid) and miR-195-5p or miR-NC. Notably, only transfection of miR-195-5p with the reporter plasmid resulted in about 50% reduction of luciferase activity (Figure 3(b), **p < 0.01), whereas miR-NC exerted almost no effect on the luciferase activities (#p > 0.05). Interestingly, the inhibitory function of miR-195-5p was nearly vanished when the miRNA-binding sequence was mutated from UGCUGGU to GGCAGGA. In summary, these results clearly indicate that miR-195-5p negatively regulates CARM1 expression depending on the direct interaction of miR-195-5p and CARM1 3′-UTR.

CARM1 is a direct target of miR-195-5p. (a) The diagram illustrates the construction of the luciferase reporter plasmids. Mutant: contains 3-base-mutation at the miR-195-5p-target region. (b) Luciferase reporter assays of the effect of miR-195-5p in HCT15 and SW480 cells. Results are shown as mean ± SD from three independent experiments; **p < 0.01 compared to miR-NC control. (c) Left, Western blot analysis of the effect of transfected miR-195-5p on CARM1 protein expression compared to a negative control (miR-NC) in HCT15 and SW480 cells. GAPDH served as a loading control. Right, CARM1 protein expression analysis by quantification of density of protein bands from Western blot (n = 3, **p < 0.01). (d) Quantitative real-time PCR analysis of the effect of transfected miR-195-5p on CARM1 mRNA expression compared to a negative control (miR-NC) in HCT15 and SW482 cells. GAPDH served as a loading control; #p > 0.05.
By immunoblotting analysis, the expression of CARM1 protein was evaluated in HCT15 and SW480 cells after transfecting with miR-195-5p or miR-NC. As anticipated, the expression of CARM1 protein was dramatically down-modulated in HCT15 and SW480 cells expressing miR-195-5p (Figure 3(c); n = 3, **p < 0.01). To determine whether CARM1 mRNA is affected by miR-195-5p, we performed qRT-PCR assay in HCT15 and SW480 cells after transfecting with miR-195-5p. In contrast, miR-195-5p overexpression did not affect the CARM1 mRNA levels in HCT15 and SW480 cells (Figure 3(d); #p > 0.05). Taken together, our data strongly demonstrated that miR-195-5p could negatively regulate CARM1 expression at the translation level, which is a common pattern of miRNA regulation in animal cells.
MiR-195-5p expression is downregulated in colorectal cancer tissues and inversely correlates with CARM1 protein levels
To investigate the potential roles of miR-195-5p, we checked the miR-195-5p expression in colorectal cancer specimens using qRT-PCR. As shown in Figure 4(a) and (b), miR-195-5p was drastically decreased in colorectal cancer tissues in comparison with their matching nontumor controls (n = 45, **p < 0.01). Notably, levels of miR-195-5p and CARM1 protein (Figure 4(c)) exhibited a significant inverse correlation in colorectal cancer (r = −0.3687, *p < 0.05), further demonstrating that CARM1 is a potential target of miR-195-5p in colorectal cancer.

MiR-195-5p expression is downregulated in colorectal cancer tissues and inversely correlates with CARM1 protein levels. (a and b) Relative expression of mature miR-195-5p was measured using real-time PCR in colorectal cancer samples and corresponding non-cancerous samples, and the results were normalized against an endogenous U6 RNA control. ***p < 0.001 (n = 45). (c) Pearson’s correlation scatter plot of the fold change in miR-195-5p levels and CARM1 protein levels in human colorectal cancer tissues (n = 45, p < 0.05).
MiR-195-5p inhibits colorectal cancer cell growth via targeting CARM1
Next, to determine the effects of miR-195-5p on colorectal cancer cell growth, we first confirmed the expression of miR-195-5p in HCT15 and SW480 cells by qRT-PCR (Figure S2, ***p < 0.001). The proliferation rate and colony-formation numbers of HCT15 and SW480 cells transfected with miR-195-5p were markedly decreased compared to cells transfected with scramble control (Figure 5(a) and (b)).

MiR-195-5p inhibits colorectal cancer cell growth via targeting CARM1. Restoration of CARM1 significantly reversed the suppression of (a) cell proliferation and (b) colony formation induced by miR-195-5p. Values at the indicated time points represent mean ± SD of three independent tests; **p < 0.01.
To test whether restoration of CARM1 would reverse miR-195-5p-mediated repression of cell proliferation and colony formation in colorectal cancer, we performed rescue experiments in HCT15 and SW480 cells. We found that overexpression of CARM1 by transfection of cDNA that lacked the miR-195-5p-binding sites in the 3′-UTR partially relieved miR-195-5p-mediated depression of proliferation and colony formation in HCT15 and SW480 cells (Figure 5(a) and (b)). Simultaneously, the expression of CARM1 protein was confirmed by immunoblotting assay (Figure S3). Based on these data, miR-195-5p exerted an inhibitory effect on cell proliferation and colony formation via targeting CARM1 in colorectal cancer.
Discussion
In this study, we revealed a pivotal role of CARM1 in colorectal cancer cell growth and identified the potential involvement of a miRNA-mediated mechanism in modulating CARM1 expression in colorectal cancer. Our IHC and qRT-PCR results showed that CARM1 is increased markedly in colorectal cancer tissues and promotes colorectal cancer cell growth in colorectal cancer cell lines and xenograft tumors in mice. In addition, we demonstrated that CARM1 could be post-transcriptionally targeted by miR-195-5p. MiR-195-5p is frequently downregulated and inversely correlates with CARM1 protein levels in colorectal cancer. Together, our data demonstrated that CARM1 is critical for colorectal cancer cell growth and could be directly targeted by miR-195-5p, which suggests that miR-195-5p is a potential tumor suppressor in colorectal cancer.
CARM1, as an important transcriptional coactivator, could catalyze the asymmetric di-methylation on arginine residues of histone or non-histone substrates. CARM1 is involved in the initiation and progression of various human tumors. Based on the previous studies, abnormal expression of CARM1, mostly upregulation, is associated with human prostate,10,11 breast, 7 lung, 17 and colorectal cancers. 18 Although levels of CARM1 have been reported to correlate with some cancers, the roles of CARM1 are controversial and seem to be context dependent. In breast cancer, CARM1 is of great importance for the estrogen-activated expression of E2F1, which is a critical cell-cycle transcriptional regulator. 19 However, CARM1 was found to inhibit breast cancer progression in estrogen receptor-alpha–positive tumors by activating p21cip1 and p27kip, which are negative cell-cycle regulators, instead of E2F1. 20 Ou et al. reported that CARM1 is impressively overexpressed in nine different colon cancer cell lines, and silencing of CARM1 expression could result in the reduction of cell proliferative and colony-formative capacities in colorectal cancer. Moreover, they also discovered that CARM1, as a positive modulator, is essential for the expression of Wnt/β-catenin target genes, including c-Myc, S100A4, Axin, and GRP49. 18 In this study, we found that CARM1 is highly elevated in tissues derived from colorectal cancer patients. Knockdown of CARM1 by sequence-specific siRNA led to an obvious reduction of cell proliferative and colony-formative capacities in colorectal cancer cells. Based on our evidence, CARM1 appears to be a promising therapeutic target for developing novel treatments in colorectal cancer.
As is already apparent, cancer is driven by progressive events associated with complicated networks of abnormal gene expression and tumor microenvironment alterations, in which miRNAs play important roles. 21 Not only can miRNAs serve as potential biomarkers, but they also have promise as therapeutics. A specific miRNA targeting CARM1 has not been previously identified in colorectal cancer. In a bioinformatics search for potential miRNAs targeting CARM1, we identified miR-195-5p as one of the most promising miRNAs. With overexpression of miR-195-5p, CARM1 protein expression was slightly downregulated in HCT15 and SW480 cells. However, we found no obvious changes in levels of the mRNA of CARM1 when miR-195-5p was upregulated in HCT15 and SW480 cells. Nevertheless, we could not exclude the possibility that other miRNAs may regulate CARM1 expression, such as miR-15a 22 and miR-181, 23 which was not examined in our study.
MiR-195-5p belongs to the miR-15–497 family. Members of this miRNA family share the same 3′-UTR binding seed sequence and have been reported to cooperatively regulate a number of targets. 24 Moreover, the sequence of mature miR-195-5p is conserved across mammalian species. 25 Previous studies have shown aberrant miR-195-5p expression in multiple cancers, including breast cancer, 26 hepatocellular carcinoma, 27 colorectal cancer,28,29 gastric cancer, 30 and non–small cell lung cancer (NSCLC). 31 However, the biological function of miR-195-5p in colorectal cancer, to our knowledge, remains unclear. In addition to CARM1, miR-195-5p also regulates other mRNAs. Recently, miR-195-5p is reported to suppress cell cycle and tumorigenesis through the control of molecules involved in the G1/S phase transition,32,33 including Cyclin D1, 34 CDK4, 32 and E2F3. 35 Earlier experiments also suggest that miR-195-5p has additional targets, such as MYB, 31 BCL-2, 36 IKKa, and TAB3. 37
Our experiments show that increased miR-195-5p expression could reduce cell proliferative and colony-formative abilities in colorectal cancer. Similar observations have been reported in a number of cancer cells.37–39 The expression of CARM1 can be regulated by miR-15a and miR-181 in acute coronary syndrome and human embryonic stem cells reported by two groups, respectively.22,23 However, no studies have yet examined that CARM1 is a direct target of miR-195-5p in colorectal cancer. Our investigation not only demonstrated that miR-195-5p interacted with CARM1 mRNA and suppressed its protein expression in cancer cells but also showed that miR-195-5p inhibited cell growth and was downregulated in colorectal cancer tissues. Based on the negative regulation effect of miRNA on target mRNA, we speculated that decreased miR-195-5p may lead to the upregulation of CARM1 in colorectal cancer. In this study, although we have demonstrated that miR-195-5p is obviously downregulated in our series of colorectal cancer specimen, further studies based on a large group of cases and controls will be needed.
Conclusion
Our study demonstrates that CARM1 promotes cell growth and is directly targeted by miR-195-5p. To our best knowledge, this is the first report showing that miR-195-5p could suppress colorectal cancer through, at least partially, downregulating the expression of CARM1, a newly discovered target in colorectal cancer. Furthermore, it is novel to know that miR-195-5p and CARM1 may be independent prognostic factors in colorectal cancer. Thus, of particular interest, the miR-195-5p/CARM1 regulatory axis might represent a new molecular target for colorectal cancer treatment.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Shanghai Committee of Science and Technology, China (Grant No. 134119b3100).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
