Abstract
Background
Kurarinone, a compound derived from Sophora flavescens, has been widely studied for its anti-inflammatory properties, as well as its potential in treating autoimmune encephalomyelitis and inflammation-related musculoskeletal diseases. However, the specific molecular mechanisms by which Kurarinone affects osteoporosis are still not fully understood.
Objectives
To explore the potential pharmacological targets and molecular mechanisms underlying the therapeutic effects of Kurarinone on osteoporosis.
Materials and Methods
To confirm the impact of Kurarinone on osteoporosis, network pharmacology techniques were used to analyze the mechanisms of Kurarinone on osteoporosis. Molecular docking was performed to examine the binding of Kurarinone to key targets involved in osteoporosis. Then, we conducted TRAcP staining, quantitative polymerase chain reaction (q-PCR), and Western blot experiments to validate the results.
Results
Utilizing network pharmacology approaches coupled with docking analysis, our investigation revealed that Kurarinone was involved in the osteoclast differentiation and MAPK signaling pathways. According to
Conclusion
This study suggests that Kurarinone may treat osteoporosis by inhibiting MAPK phosphorylation and inhibiting osteoclast formation.
Introduction
The skeletal system is an indispensable element within the human musculoskeletal framework, fulfilling a critical function in the mechanics of corporeal motion. Osteoclast-mediated bone resorption is pivotal in the regulation of skeletal homeostasis, and it also serves as the root cause for the onset of metabolic bone diseases, such as bone loss, degradation of microarchitectural integrity within bone tissue, and increased bone fragility (Black & Rosen, 2016; Ma et al., 2023a, 2023b). This condition presents a significant global burden on women and has substantial economic implications. Moreover, studies have shown that approximately 20% of men face an increased risk of osteoporotic fractures (LeBoff et al., 2022). Despite advancements in osteoporosis treatment, the drugs available often come with numerous reported adverse effects, which have led to a decrease in public willingness to engage in treatment. Of particular concern is that exogenous estrogen replacement therapy may portend potential adverse events such as fever, joint muscle pain, and hypocalcemia, with a possible increase in the risk of cardiovascular events (Muhammad et al., 2018; Mo et al., 2022). Consequently, there is an escalating demand for innovative and more benign options, particularly those derived from traditional herbal medicine, within the field of osteoporosis research (Ma et al., 2023a, 2023b).
The emergence of bioinformatics has revolutionized network pharmacology, moving away from the traditional “single-mode (target + drug)” approach to a more comprehensive “multi-component (target + drug)” model, which leverages progressive pharmacological research methods to understand potential mechanisms, target predictions, and toxicology of nature products (Zhang et al., 2022). In line with this advancement, our research utilizes network pharmacology, molecular docking, and subsequent experimental validation to predict and validate pathways and targets of Kurarinon in treating osteoporosis.
Given the significant role of osteoclasts in bone metabolism disorders such as osteoporosis, as well as the pharmacological properties of Kurarinone, this study aims to explore the potential mechanisms of Kurarinone in the prevention and treatment of osteoporosis. To achieve this goal, the study plans to explore potential target proteins and related mechanisms involved in Kurarinone’s intervention in osteoporosis through network pharmacology methods and validate them through molecular docking and
Materials and Methods
Materials and Reagents
Kurarinone, obtained commercially from ChemFaces, was provided with a purity of 98% or higher. The Kurarinone powder was stored at –80° after dissolving in dimethyl sulfoxide (DMSO) as 100 mM stock solution. Alpha minimum essential medium (α-MEM), along with 100 U/ml penicillin, 100 U/ml/streptomycin (P/S), and fetal bovine serum (FBS), was obtained from Gibco. The Cell Counting Kit-8 (CCK-8) was procured from Glpbi. Receptor activator of nuclear factor-κ B ligand (RANKL, Cat #315-11C) was purchased from PeproTech (Rocky Hill, NJ). The RAW264.7 cell lineage was acquired from Guangzhou Jennio Biotech Co., Ltd.
Structural Formula and Related Targets of Kurarinone
To obtain relevant target information for Kurarinone with its chemical name and CAS number (34981-26-5), we examined the PubChem Data Bank (
Obtain Osteoporosis-related Targets
To identify specific targets related to osteoporosis, we conducted comprehensive searches using the keyword “osteoporosis” in prominent disease databases such as DISEASES, OMIM, CTD, and Genecards. The objective was to identify genes associated with osteoporosis. After carefully removing duplicates and organizing the obtained results, we compiled a set of osteoporosis-related genes that were shared among these four public databases, providing valuable resources for future research.
Subsequently, we utilized the online platform draw Venn diagram to compare the potential targets of Kurarinone with the disease-related targets. This platform generated two intersected graphs, allowing us to identify the common targets shared by the drug and the disease. To visually represent this overlap, we generated informative Venn diagrams illustrating the convergence of drug and disease targets.
Protein–protein Interaction (PPI) and Enrichment Analysis
To perform PPI network analysis, the acquired target proteins were imported into the STRING database, generating relevant data. Subsequently, visualization of the PPI network was achieved using Cytoscape software. The common targets were subjected to gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis using the Metascape database. A
Molecular Docking between the Core Targets and Kurarinone
Kurarinone, a ligand, was selected for analysis, and its two-dimensional mol2 structure was obtained from the TCMSP website. The receptor, corresponding to the core gene mentioned in PPI network, was designated, and its respective protein structure was retrieved in PDB format from the RCSB PDB database. For molecular docking, AutoDock software was utilized. The binding energy served as the evaluation parameter to assess the docking effectiveness between Kurarinone and the target protein.
In vitro Osteoclastogenesis Assay
The RAW264.7 cells at the fifth passage were cultured in a complete medium comprising 1% penicillin/streptomycin (P/S) and 10% FBS, at a temperature of 37°C with 5% CO2. When the cells reached a confluence of 90–95%, they were subcultured.
Cytotoxicity Assays
RAW264.7 cells were seeded in a 96-well plate at a density of 1 × 104 cells per well. On the second day, the cells were treated with different concentrations of Kurarinone (0, 5, 10, and 15 µM) and cultured for 2 days. After 48 hours, 10 µl of CCK-8 solution was added to each well as instructed by the CCK-8. The plates were then incubated in the dark for 1 hour. The absorbance values at 450 nm were measured using an enzyme-linked immunosorbent assay reader after a 2-hour incubation, and the recorded data were analyzed statistically.
TRAcP Staining
RAW264.7 cells were seeded at a density of 3 × 103 cells/well in a 96-well plate and cultured overnight. On the second day, the cells were induced to differentiate into osteoclasts with the addition of 50 ng/mL of RANKL, and they were concurrently treated with different concentrations of Kurarinone (0, 5, 10, and 15 µM) as interventions. PBS was used as a positive control with an equal volume, and the cells were cultured until osteoclast differentiation was mature. The TRAcP staining solution was prepared according to the manufacturer’s instructions. The culture medium in the 96-well plate was aspirated, and the plates were washed with 1 × PBS three times. Then, the cells were fixed with 4% paraformaldehyde (Macklin, Shanghai, China) for 20 minutes and washed with 1 × PBS three times. One hundred microliters of TRAcP staining solution were added to each well, and the cells were incubated at 37°C for 30 minutes. Subsequently, the plates were washed with 1 × PBS three times. After staining, the positive cell numbers were observed and counted under a light microscope (Olympus, Japan), followed by analysis.
Ribonucleic Acid (RNA) Extraction and Real-time Polymerase Chain Reaction (PCR)
The 6-well plates were used to culture RAW264.7 cells (8 × 104 cells/well), which were stimulated with RANKL in the presence of Kurarinone (0, 5, 10, and 15 µM) until the osteoclasts reached maturity. Total RNA was isolated from the cells using Trizol reagent, and complementary deoxyribonucleic acid (cDNA) was prepared from 1 µg of total RNA template through reverse transcription. Below are the primer sequences for real-time PCR (Table 1):
Relevant Primer Sequences.
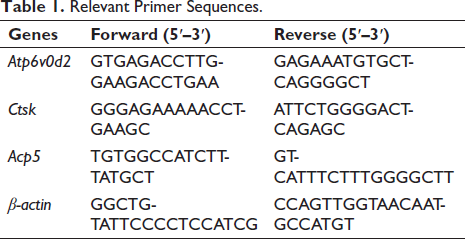
Western Blot Assay
RAW264.7 cells were seeded into 6-well plates at a density of 2 × 105 cells per well. RANKL at a concentration of 50 ng/mL was added to each well on days 1, 3, and 5 to induce osteoclast differentiation, concurrently with 15 µM of Kurarinone and an equivalent volume of PBS as a control intervention. The culture medium was replaced every 2 days, continuing until osteoclast maturation, at which point proteins were extracted to assess the expression of osteoclast function-related proteins.
To investigate signaling pathway proteins, RAW264.7 cells were seeded into 6-well plates at a density of 2.5 × 105 cells per well and cultured overnight. The cells were then subjected to a 2-hour starvation period, followed by pretreatment with Kurarinone (15 µM) for 1 hour before stimulation with RANKL (50 ng/mL) for a designated duration. Following stimulation, the cells were lysed with RIPA buffer to extract proteins. Subsequently, the proteins were separated via SDS-PAGE and transferred onto a nitrocellulose membrane. The membrane was then blocked with a 5% BSA solution to prevent nonspecific binding. The membrane was incubated overnight at 4°C with primary antibodies and subsequently with the corresponding secondary antibodies for 1 hour. Quantitative analysis of the images was performed using Gel-pro and ImageJ software.
Results
The Common Targets between Kurarinone and Osteoporosis
The chemical structure of Kurarinone is C26H30O6, and its molecular formula is depicted in Figure 1A. Through comprehensive searching and meticulous sorting, a total of 359 disease targets related to osteoporosis were identified. By intersecting the target mappings, we obtained 53 common targets, as demonstrated in Figure 1B.
Analysis of Common Targets between Kurarinone and Osteoporosis. (A) Molecular Formula of Kurarinone; (B) Venn Diagram of the Kurarinone–Osteoporosis Intersection Targets; (C–D) Protein–protein Interaction (PPI) Network of the Potential Targets.
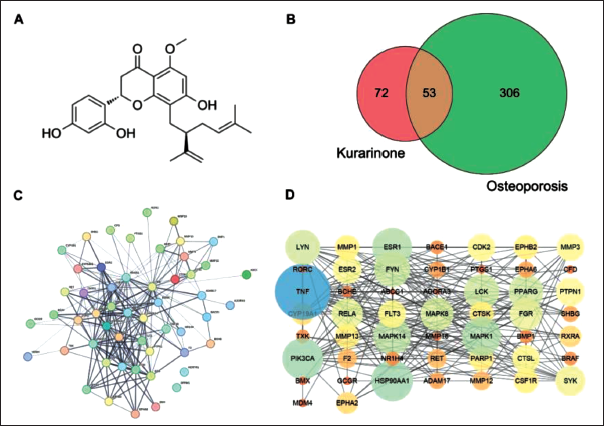
PPI Network Construction
Upload the 53 intersecting targets to the STRING online platform for protein interaction analysis (Figure 1C), and utilize Cytoscape to construct the Kurarinone-targeted PPI network for treating osteoporosis (Figure 1D). In Figure 1, there are a total of 51 nodes (protein genes) involved, with 243 edges (interactions between protein genes). The average node degree value is 6.47, indicating that the mechanism of Kurarinone in treating osteoporosis is based on the synergistic effects of multiple genes and multiple targets. Nodes that are larger and darker in color have higher degree values.
Biological Function
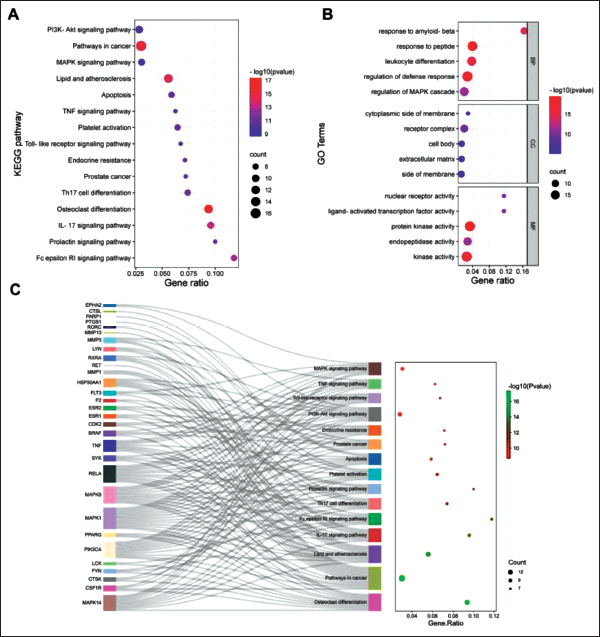
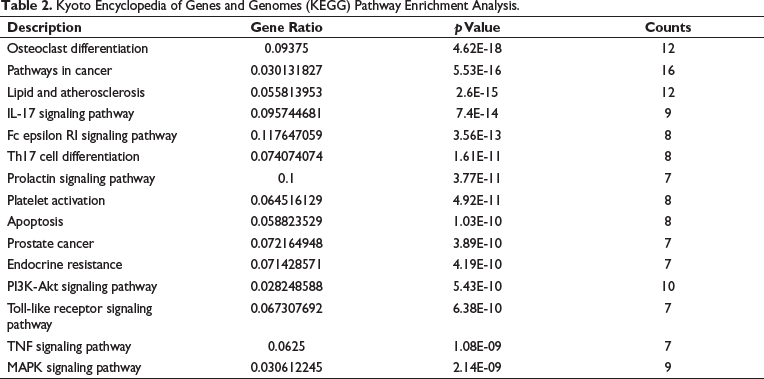
The KEGG analysis revealed a total of 111 enriched signaling pathways, with osteoclast differentiation, PI3K-Akt, pathways in cancer, and MAPK signaling pathways emerging as the most enriched (Figure 2A, Table 2). In addition, the GO analysis identified 412 enrichments, of which 352 were related to biological processes, including response to peptide, regulation of defense response, and regulation of MAPK cascade. Sixteen enrichments pertained to cellular components, such as receptor complex, membrane surface, and extracellular matrix. Meanwhile, 44 enrichments were associated with molecular functions, including protein kinase activity, nuclear receptor activity, and endopeptidase activity (Figure 2B). Figure 2C illustrates the distribution of key targets in the KEGG pathways. Overall, the results of the GO and KEGG analyses suggest that Kurarinone may modulate osteoclast differentiation by targeting specific proteins through the PI3K-Akt and MAPK pathways.
Enrichment Analysis of Potential Targets. (A) Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analysis; (B) Gene Ontology (GO) Enrichment Analysis; (C) Sankey Diagram for KEGG Enrichment Terms.
Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis.
Molecular Docking
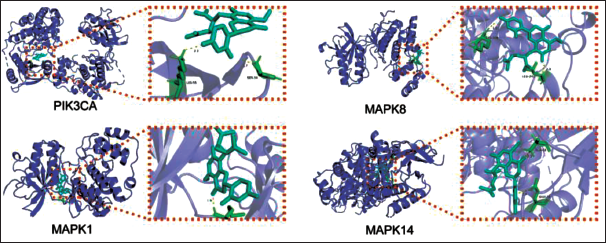
Among 53 common targets in the process of Kurarinone treating osteoporosis, the core target proteins in the MAPK signaling pathway were chosen for molecular docking, including PIK3CA, MAPK1, MAPK8, and MAPK14. AutoDock Vina software was used to predict the docking interactions between Kurarinone and the key targets, namely, PIK3CA (−7.9 kcal/mol), MAPK1 (−8.1 kcal/mol), MAPK8 (−7.1 kcal/mol), and MAPK14 (−7.6 kcal/mol). The molecular docking results demonstrated the stability of these targets and Kurarinone after docking, as depicted in Figure 3.
Kurarinone Docking with the Core Target Molecule.
Kurarinone inhibits RANKL-induced Osteoclastogenesis
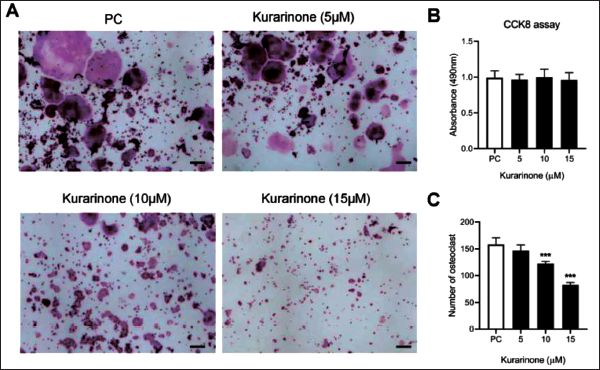
In order to evaluate the impact of Kurarinone on osteoclast differentiation, cellular experiments with Kurarinone were also conducted in the present study. The cellular cytotoxicity of Kurarinone was assessed via the CCK-8 assay to discern whether its potential involvement in the formation of osteoclastogenesis included detrimental impacts on cellular proliferation. The results corroborated that within the concentration spectrum employed in the current investigation (5–15 µM), Kurarinone failed to manifest any deleterious effects on the proliferation of RAW264.7 cells. Subsequently, Kurarinone was added in a concentration-dependent manner during RAW264.7 cells cultured with RANKL. It was observed that higher doses of Kurarinone impacted osteoclast formation, as evidenced by a reduction in TRAcP-positive osteoclast count. The lowest number of osteoclasts was observed at a concentration of 15 µM (Figure 4A–C). Furthermore, the expression of pivotal osteoclast-specific genes such as
Kurarinone Suppresses Receptor Activator of Nuclear Factor-κ B Ligand (RANKL)-induced Osteoclastogenesis In Vitro . (A) Representative Images of RAW264.7 Cells Treated with Varying Concentrations of Kurarinone and TRAcP Staining were Captured using Light Microscopy. (B) RAW264.7 Cells Treated with Different Concentrations of Kurarinone were Assessed using the Cell Counting Kit-8 (CCK-8) Method. (C) Osteoclast Counts Revealed a Reduction in the Number of Osteoclasts with Increasing Kurarinone Concentrations. All Bar Graphs are Presented as Mean ± Standard Deviation (SD). n = 3, ***p < 0.001 Relative to the Control Group. Scale Bar = 200 µm.
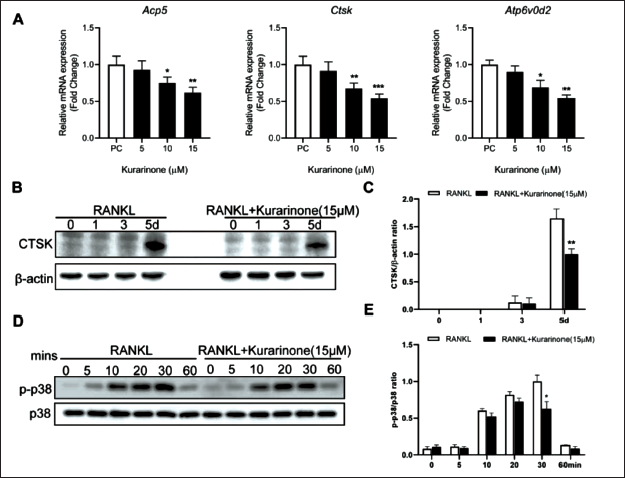
Kurarinone Significantly Inhibited the Amplification of Osteoclast-specific Genes and Proteins. (A) Kurarinone (15 µM) Significantly Inhibited the Expression of Osteoclast-specific Genes. (B–C) Kurarinone Significantly Reduced the Expression of CTSK Protein. (D–E) Kurarinone Significantly Reduced the p38 Phosphorylation on 30 mins. All Bar Graphs are Presented as Mean ± Standard Deviation (SD). n = 3, *p < 0.05, **p < 0.01, ***p < 0.001 Relative to the Control Group.
Discussion
Excessive increases in osteoclast-mediated bone resorption influence the balance of bone metabolism and cause various bone diseases, including Paget’s disease, osteoporosis, rheumatoid arthritis, and metastatic cancers (Ono & Nakashima, 2018). Osteoclasts, which are large specialized cells, originate from the fusion of multiple monocytes and macrophage progenitors, either in the vicinity or on the surface of bones. Once mature, osteoclasts secrete proteolytic and acidic enzymes such as cathepsin K (CTSK), acid phosphatase 5 (Acp5), and matrix metalloproteinases (MMPs), which are involved in the degradation of bone matrix proteins produced by osteoblasts during the process of bone resorption (Wang et al., 2022). In pathological conditions, excessive bone resorption often leads to osteoporosis, disrupting the normal bone remodeling process and resulting in significant loss of bone mass (Mo et al., 2022).
Recently, there has been a burgeoning enthusiasm for delving into alternative therapeutic strategies aimed at treating osteoporosis. This focus has been primarily directed toward the identification and assessment of novel, safe, and natural compounds. The quest for these ingredients has emerged as a vital priority in the field. Notably, certain traditional Chinese medicine (TCM), which shares similarities between medicine and food, has become a focus of research in the prevention and treatment of osteoporosis due to its low cost and minimal adverse reactions (He et al., 2023; Liu et al., 2023). Of particular importance is Kurarinone, a natural bioactive compound extracted from the rhizome of
Beyond the strides made in contemporary information technology and the continuous inquiry in bioinformatics and pharmacology, there has been a fruitful merge of network pharmacology with TCM. This amalgamation includes a detailed examination of the active elements within TCM and the formation of a “drug-target-pathway-disease” network. This approach empowers researchers to discern the intricate biological roles and operational mechanisms tying medications to their associated maladies (Jiao et al., 2021). This novel comprehension of TCM’s pharmacological impacts and the underlying mechanisms offers a fresh perspective for delving into TCM’s pharmacological specifics (Ye et al., 2023). Such insights pave the way for innovative directions in the advancement of TCM research.
In our current study, we have demonstrated the inhibitory effect of Kurarinone on osteoclast differentiation; however, the underlying mechanism remains unclear. To address this, we identified 53 common targets shared between Kurarinone and osteoporosis. Through the analysis of PPI network topology, we observed that these targets are associated with the PI3K-Akt, MAPK pathways, and osteoclast differentiation. Notably, the key targets PIK3CA, MAPK1, MAPK8, and MAPK14 showed a strong binding affinity with Kurarinone in molecular docking experiments, suggesting their potential significance in the treatment of osteoporosis using Kurarinone.
The PI3K/Akt signaling pathway is an enzymatically linked receptor-mediated signaling cascade that regulates various cellular activities. This pathway is not only involved in modulating the progression of osteoporosis by suppressing inflammatory responses and osteoclast formation but also plays a role in inhibiting osteoporosis by promoting osteoblast proliferation, differentiation, and bone formation (Xi et al., 2015). Consequently, the PI3K-AKT signaling pathway is vital for maintaining bone homeostasis.
Upon binding of RANKL to RANK, a series of signaling cascades is triggered, initiating the regulation of mature osteoclast development. The MAPK pathways, including three major members (p38, JNKs, and ERKs), are implicated in osteoclast differentiation and formation stimulated by RANKL (Zhang et al., 2022). P38 plays a significant role in differentiating osteoclast precursors into mature osteoclasts. Studies have demonstrated that mice with specific deletion or inhibition of p38 show reduced expression of NFATC1 and CTSK proteins, resulting in decreased bone resorption by osteoclasts and increased bone mass (Lee et al., 2016). These findings indicate the significant role of p38 in regulating osteoclast function and bone metabolism. The Western blot analysis unveiled Kurarinone’s capability to inhibit p38 phosphorylation. Therefore, Kurarinone may hinder osteoclast formation by obstructing the p38/MAPK pathway.
Conclusion
In this study, we utilized bioinformatics methods to identify target genes of Kurarinone for the treatment of osteoporosis. Through enrichment analysis, we identified the MAPK signaling pathway as highly significant.
Abbreviations
TCM: Traditional Chinese medicine; PPI: Protein–protein interaction; KEGG: Kyoto Encyclopedia of Genes and Genomes; BP: Biological processes; MF: Molecular functions; CC: Cellular components; MMPs: matrix metalloproteinases.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Ethical Approval and Informed Consent
Not applicable.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
