Abstract
Introduction
Osteoporosis (OP) is a systemic bone disease characterized by decreases in bone density and destruction of the bone tissue microstructure, which results in increases in bone fragility and proneness to fractures. 1 The worldwide increase in ageing results in continuous increase in the incidence of OP, and OP has become a chronic disease that seriously affects the health and quality of life of middle-aged and elderly people. 2 The identification of multiple ways for maintaining healthy bone tissue homeostasis is a pressing challenge in medicine.
At present, the conventional anti-OP drugs primarily include bone resorption inhibitors, bone formation promoters and bone minerals, such as calcitonin, bisphosphonates, and fluoride. 3 However, these drugs can only improve the patient's clinical symptoms and delay the disease and cannot restore the dynamic balance of bone formation and osteoclast activity or fundamentally improve bone metabolism. 4 Traditional Chinese medicine has always played an important role in improving quality of life, and strategies with enhanced efficacy are required for the treatment of OP. Modern scientific methods have demonstrated that some Chinese medicine extracts and compounds with fewer side effects and lower costs can prevent and treat OP. 5 Therefore, an increasing number of scholars have begun to research the use of traditional Chinese medicines for the prevention and treatment of OP.
Epimedium and Anemarrhen are herbal medicines widely used in combination for the treatment of OP. Epimedium can nourish kidney homeostasis, strengthen muscles and bones, and combat rheumatism, particularly in subjects with fatigue, sexual dysfunction, rheumatic diseases, and OP. Anemarrhen reduces body heat and internal fire, generates bodily fluids, combats dryness, and is thus often used for the clinical treatment of fever, polydipsia, bone steaming, and hot flashes. Clinical studies have shown that the use of herbal formulas containing Epimedium and Anemarrhen may enhance the total effective and cure rates and alleviate pain intensity and relevant symptoms in patients with OP.6,7 However, the systematic mechanisms underlying the pharmacological activity of the Epimedium–Anemarrhen (EA) combination for OP treatment remain unexplored.
In recent years, with the continuous innovation and development of systems biology and bioinformatics, network pharmacology has integrated a large amount of information via the combination of calculation and experimental methods to discover new drug targets and molecular mechanisms. Based on the holistic and systematic interaction between the two herbal medicines, we conducted a review of potential pharmacodynamic substances, targets, and pathways in vitro, constructed a multilevel “disease-gene-target-drug” connected network, and predicted the mechanism of drug action. Exploring the relationship between drugs and disease is a novel strategy in this type of research, 8 and the systematic nature of this strategy resonates well with the overall concept of traditional Chinese medicine and the multicomponent, multipathway, and multitarget synergistic mechanisms of traditional Chinese medicines. Using network pharmacology and molecular docking methods, this study explored the effect of EA on OP at the molecular level by constructing an EA active ingredient-OP target pathway network and performing molecular docking analysis with the aim of elucidating a therapeutic mechanism. The goal of the present study was to reveal the systematic mechanisms through which the EA herbal combination can be used as a treatment for OP. The flowchart of this research study is shown in Figure 1.

Whole framework based on network pharmacology.
Materials and Methods
Compound Collection and Screening
The chemical components of the two traditional Chinese medicines were retrieved from the Traditional Chinese Medicine Database and Analysis Platform (TCMSP, https://tcmspw.com/index.php, last updated May 31, 2014) based on the pharmacokinetic characteristic parameters of the Chinese medicine system by reviewing active ingredients. 9 Their chemical components of traditional Chinese medicines were reviewed to include those with an oral availability (OB) ≥ 30% and a drug-likeness (DL) ≥ 0.18 to obtain the effective active ingredients of the drug and information on its targets.10,11 After this review, the protein targets on which the compounds act were standardized in the UniProt protein database.
Target Acquisition for Assessing Drug and Disease-Related Effects
Using OP-related diseases, such as “Osteoporosis” as keywords, the GeneCards database (https://www.genecards.org/), OMIM database (https://omim.org/), and DisGeNET database (https://www.disgenet.org/home/) were mined for potential OP treatment targets. The DrugBank database (https://go.drugbank.com/) was also used to find first-line clinical Western medicine targets for the treatment of OP.12–15 After merging the disease targets from the four databases, duplicate targets were deleted to obtain the disease targets. We then took the intersection of the effective active ingredient targets of EA and the OP-related targets, drew a Venn diagram of the drug's active ingredient targets and disease targets in the R language, and selected common genes as important potential targets of EA in the treatment of OP.
Construction of Drug Active Ingredient and Disease Target Network
The active ingredients of EA and common genes related to both the drug and the disease were imported into Cytoscape 3.7.2 (http://www.cytoscape.org/) to construct an active ingredient-target-disease network. Cytoscape software utilizes a network as its core architecture: a node in the network represents an active ingredient or target, an edge represents the interactions between nodes, and the degree of nodes represents its importance in the network. More connection points between nodes are correlated with a higher degree and thus with a higher possibility that the target will become a key target.
Protein–Protein Interaction Network Construction and Core Gene Screening
Protein–protein interaction (PPI) refers to the process in which two or more protein molecules form protein complexes through noncovalent bonds. PPIs and acquired networks are important for most biological functions and processes. The systematic analysis of protein interactions in biological systems is of great importance for understanding changes in biological signals and the functional relationships between proteins under disease conditions. 16 We inputted the selected common genes into the STRING11.0 database (https://string-db.org/) to construct a PPI network, set the biological species to “Homo sapiens,” set the minimum interaction threshold to “Highest confidence (>0.7),” and hid all disconnected nodes in the network. The other settings were maintained at their defaults. A PPI network was obtained, protein interaction network analysis was performed, and the results were downloaded in the updated TSV format. This file was then imported into Cytoscape 3.7.2; its built-in tool App CytoNCA was used to analyze the network topology parameters of active ingredients and targets, and the topological properties of nodes in the interactive network were evaluated by calculating three parameters: “degree centrality (DC),” “between centrality (BC),” and “closeness centrality (CC).” These three parameters measure the importance of nodes in the network and describe the nature of the nodes in the network. High DC, BC, and CC values for a node indicate that it plays an important role in the network.17–19 According to the results from the analysis of the topological characteristics of the abovementioned PPIs, the target above the median was selected as the core target and screened twice. Based on the above-described review, the core target in the network was obtained.
GO and KEGG Enrichment Analyses
Various bioinformatics analyses and result visualizations can be performed with R. The Entrez gene ID of the core target was determined using the “org.Hs.egdb” package and then subjected to GO functional enrichment analysis (molecular function, MF; biological process, BP; cellular component, CC) and KEGG pathway enrichment analysis with the “clusterProfiler” package. The P value cut-off = .05 and q value cut-off = .05 parameters were set, and the results for the top 20 items with the highest enrichment are presented in the form of bar graphs. Finally, using RGUI and Pathview, we downloaded the top 20 access maps of FusionExcel from the KEGG database (KEGG, https://www.kegg.jp/kegg/pathway.html, updated on July 1, 2019). We filtered the results repeatedly and manually selected the results based on relevance.
Docking Steps and Results Evaluation
Molecular docking is a convenient and effective method for exploring the interaction of small molecules with targets. Here, the GLOD module of Discovery Studio software was used to verify the prediction results of the network pharmacology analysis by molecular docking. First, “Preparation Ligand” was applied to each drug ingredient, and the parameters were set to default values and saved as molecular docking ligands. Second, by “preparing the protein,” the parameters were set to default values, the preprocessed protein structure was obtained, the original ligand was extracted from the target, and the molecule was paired as a receptor. We selected the original ligand extraction position for the receptor active site, set the docking preference to high quality in the LibDock module, set the construction method to BEST, set the parallel processing to true, and used the default values for the other parameters. The docking results are shown as scores determined by the scoring function LibDock, and higher LibDock scores are correlated with a higher binding activity of the predicted component to the target.
Results
Compound Collection and Screening
By searching the TCMSP database with the ADME screening conditions of an OB ≥ 30% and a DL ≥ 0.18, 23 types of Epimedium active ingredients and 15 types of Anemarrhen active ingredients were extracted (Supplementary Table 1), and 508 targets of the Epimedium ingredients and 188 targets of the Anemarrhen ingredients were extracted. After merging, duplicate targets were deleted, and 237 constituent targets were obtained. Finally, the UniProt database was used to identify the gene names of the targets for normalization, and 199 effective active component targets were obtained.
Target Acquisition for the Assessment of Drug and Disease-Related Effects
Major databases (GeneCards, OMIM, DisGeNET, and DrugBank) were combined to search for and supplement relevant targets. Duplicate targets were deleted after merging, and 4603 OP-related gene targets were ultimately integrated (Supplementary Table 2). We then took the intersection of the screened effective active ingredient targets of EA and the disease targets of OP and drew a Venn diagram in R, as shown in Figure 2, which resulted in the identification of 149 common genes (Supplementary Table 3).

Common genes of Epimedium–Anemarrhen and osteoporosis.
Construction of a Drug Active Ingredient and Disease Target Network
Chemical components without corresponding targets and duplicate targets were deleted, the data were then imported into Cytoscape 3.7.2, and a network diagram of 30 drug components and 149 disease-related targets was established, as shown in Figure 3. The network has a total of 179 nodes and 424 edges. The different nodes represent the active ingredients of EA and the relevant targets of OP. Figure 3 shows that one target can correspond to one or more active ingredients and that multiple targets can correspond to the same active ingredient concurrently. These results show that EA has multiple components and targets in the treatment of OP. Among the active drug ingredients, the top six with the highest degree values were quercetin, kaempferol, anhydroicaritin, luteolin, stigmasterol, and diosgenin.

The “active ingredients-common gene-disease” network. The blue quadrilateral represents the common genes, and the red quadrilateral represents the active component of Epimedium–Anemarrhen.
PPI Network Construction and Core Gene Screening
The common genes were uploaded into the STRING11.0 database to construct a PPI network, and the common genes in the network were obtained. The interactive network has 143 nodes and 1109 edges. TSV data were imported into Cytoscape 3.7.2 for visualization, and CytoNCA was used to calculate the topological parameters in the network nodes to obtain the DC, BC, and CC values (see Supplementary Table 4). The thresholds for the first screening were DC ≥ 19, BC ≥ 157.451, and CC ≥ 0.243, and a total of 54 nodes and 554 edges were obtained. Then, 54 targets were screened in more detail. The second screening threshold was DC ≥ 27, BC ≥ 315.167, and CC ≥ 0.250, and the results yielded 20 nodes and 144 edges, as shown in Figure 4. After screening, the network contained AKT1, IL6, MAPK1, MAPK8, JUN, VEGFA, CXCL8, IL1B, EGF, MYC, EGFR, MMP9, PTGS2, RELA, CASP3, ESR1, FOS, APP, AR, MMP2, and other targets; these 20 targets are likely the key targets of EA in the treatment of OP.
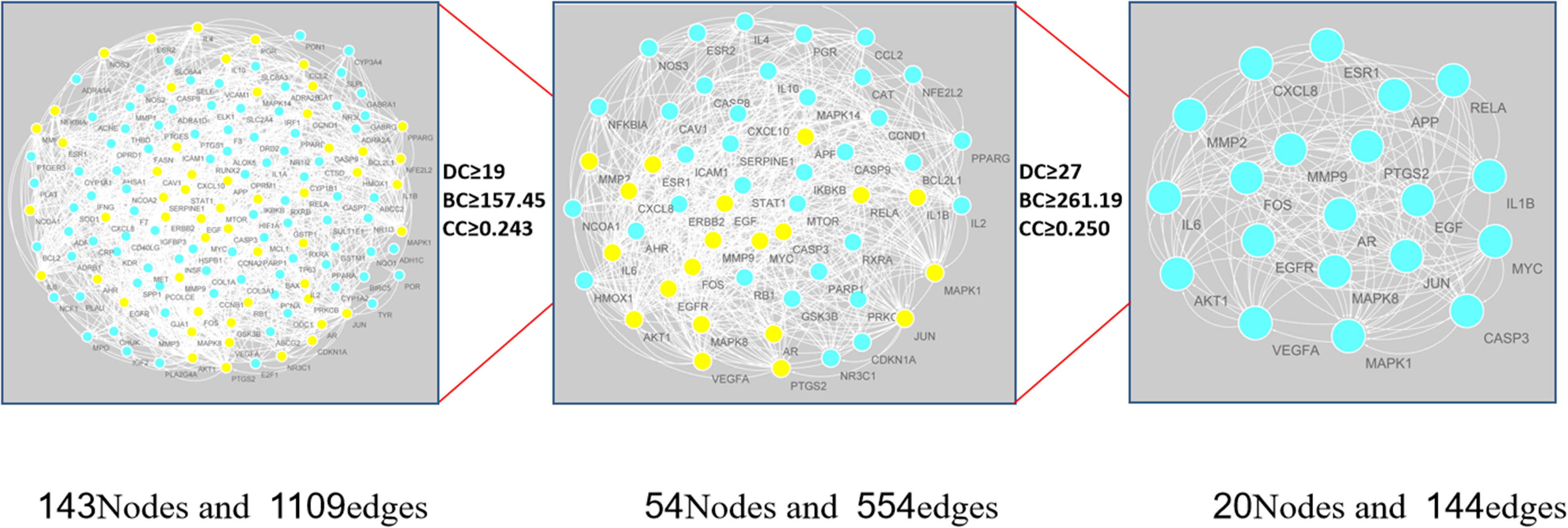
Process of topological screening for protein–protein interaction network.
GO and KEGG Pathway Enrichment Analyses
We analyzed the MFs, BPs, and CCs of 149 key targets that were primarily enriched and found that the key targets of EA for OP treatment were enriched in 156 GO terms. The biological functions and processes included nuclear receptor activity, steroid hormone receptor activity, RNA polymerase II transcription factor binding, adrenergic receptor activity, cell proliferation and apoptosis, inflammation, oxidative stress, and immunity. Other biological processes were closely related. The top 20 GO results were screened using P < .05 as the threshold, as shown in Figure 5. To clarify the roles of the therapeutic targets of EA in the signaling pathway, KEGG pathway enrichment analysis was performed using clusterProfiler, and a total of 156 pathways, including the AGE-RAGE signaling pathway in relation to diabetes complications, fluid shear stress, and atherosclerosis, were found to be enriched. The top 20 KEGG analysis results were screened using P < .05 as the threshold, as shown in Figure 6. The size and color of the bubble chart nodes indicate the number of associated genes and the P value, respectively. The size of the nodes from small to large reflect the number of associated genes from low to high, and the color range from blue to red reflects the P value from large to small. The five pathways with the smallest P values were AGE-RAGE signaling in relation to diabetes complications, fluid shear stress and atherosclerosis, hepatitis B, IL-17 signaling, and prostate cancer. In addition, the KEGG pathway enrichment analysis revealed signaling pathways closely related to OP, such as the PI3K-Akt signaling, NF-kappa B signaling, osteoclast differentiation, and Wnt signaling pathways.
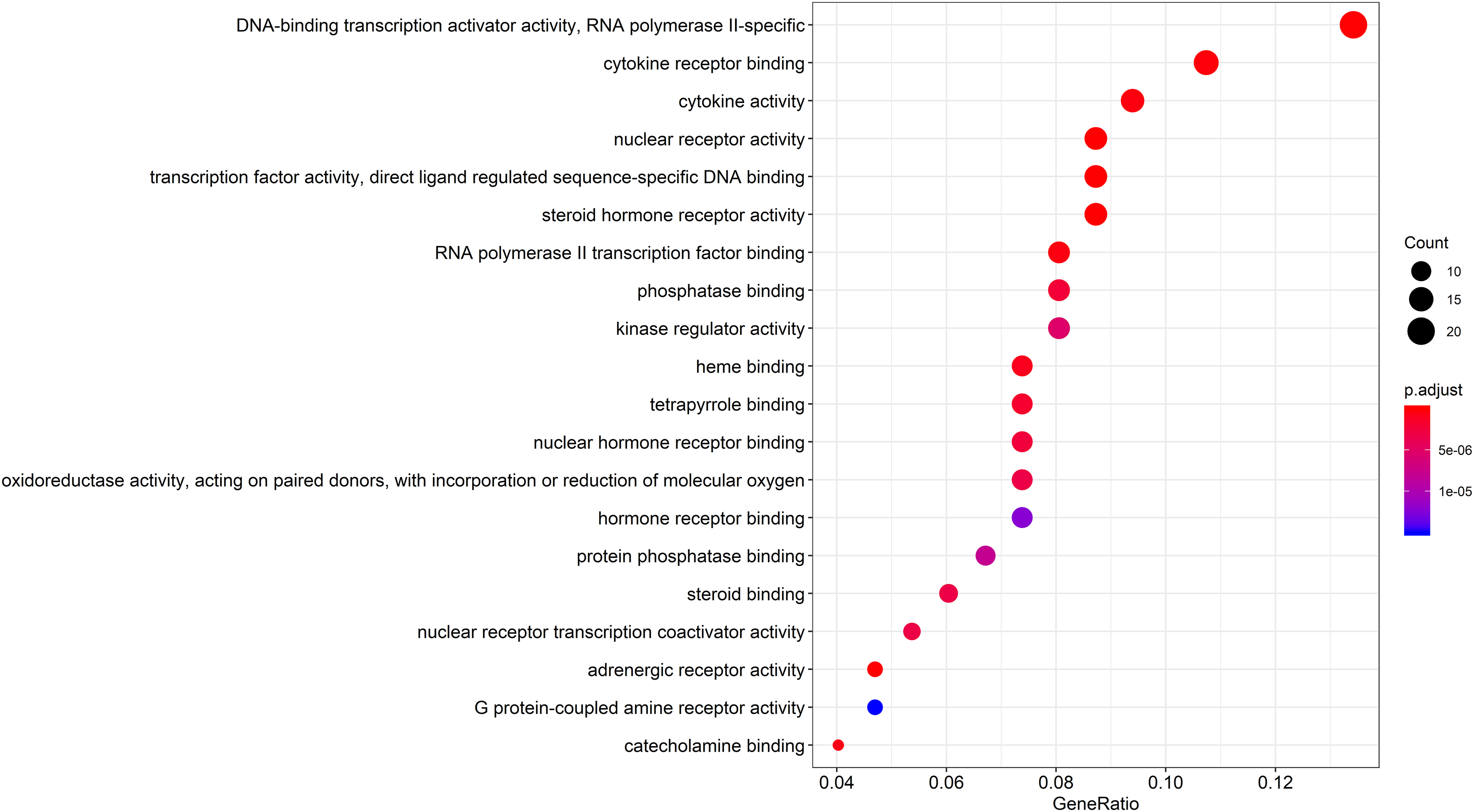
Go enrichment analysis of common genes. The larger the node and the smaller the P value, the higher the correlation.

KEGG pathway enrichment analysis of common genes. The smaller the p.adjust, the greater the correlation.
Target Pathway Analysis
The target pathway analysis tool KEGG Mapper was used to obtain the pathway map of EA in the treatment of OP, and the map included the IL-17 signaling pathway, as shown in Figure 7. The pathway targets are marked in white, and the targets used by EA in the treatment of OP are marked in red. The pathway diagram shows that EA is involved in the treatment of OP and involves the NF-κB and MAPK signaling pathways. EA has 28 effective targets in the treatment of OP. EA may be used for the treatment of OP by regulating multiple aspects, and its targets may be located in these pathways.
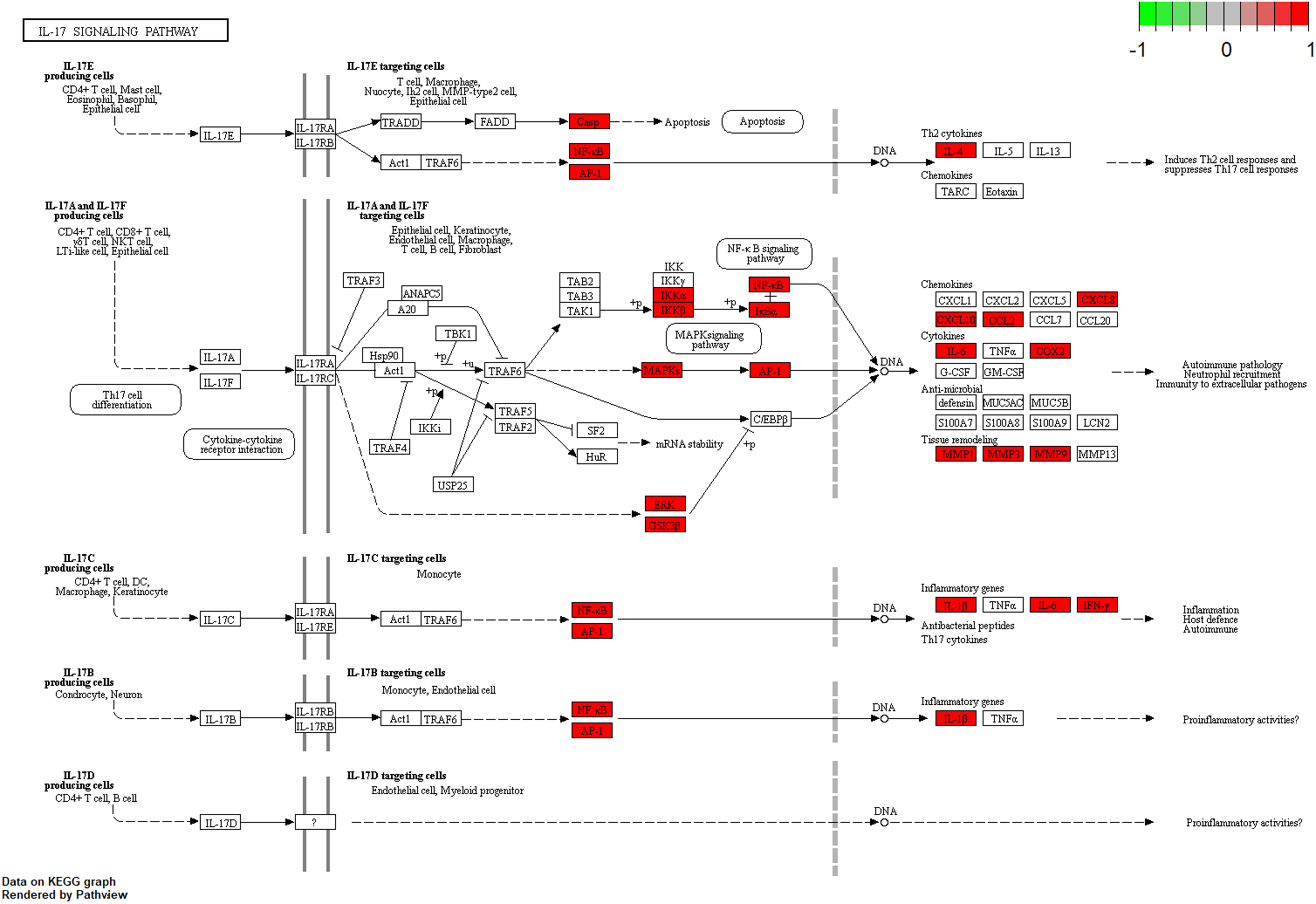
Pathway map of EA in the treatment of OP. The main targets of EA in the treatment of OP are located in the IL-17 pathway. Arrows represent the activation effect, T-arrows represent the inhibition effect, and segments show the activation effect or inhibition effect.
Molecular Docking Analysis
The LibDock score and binding mode of the docking pose were used to analyze the binding effect of the protein ligand (P-L). A lower LibDock score indicates a stronger binding ability between the molecule and its targets. As shown in Supplementary Table 5, based on the comparison of LibDock scores, we surprisingly found that the six molecules of interest combined better with MMP2. The lowest LibDock scores were obtained for the binding of luteolin, quercetin, stigmasterol, diosgenin, and anhydroicaritin with MMP2, and the LibDock score (value of 123.307) for the binding between kaempferol and MMP2 was second only to that found for the binding with JUN protein (value of 125.693). MMP2 can degrade type I collagen secreted by osteoblasts and thereby activates osteoblasts. Previous research has shown that MMP2 inhibitor 1 (MMP2-I1) plays a positive role in the osteogenesis of human bone marrow mesenchymal stem cells (hBMSCs). 20 This finding further explains that MMP2 was the main target of this series of molecules that function in the treatment of OP. From the perspective of the type of interaction, we found that electrostatic interactions (including hydrogen bond interactions and metal coordination) were the main interactions involved in the binding between luteolin and quercetin molecules, and hydrophobic interactions (in this paper, it mainly refers to Pi-alkyl) were the main interactions in the binding between stigmasterol and diosgenin molecules. Anhydroicaritin could form electrostatic and hydrophobic interactions at the same time as MMP2. As MMP2 is a potential target in the treatment of OP, the electrostatic effects of luteolin and quercetin may play a main role in treating the disease. The hydrophobic effect of the molecules stigmasterol and diosgenin may play the main role in the treatment. For anhydroicaritin, the therapeutic effect was contributed by the combination of electrostatic and hydrophobic interactions. Amino acids interact with ligands (see Supplementary Table 6). Figure 8 depicts part of the docking process.

Molecular docking of compounds with core targets. The binding mode of protein ligand predicted by docking. (a) Luteolin–MMP2 complex, (b) quercetin–MMP2 complex, (c) kaempferol–MMP2 complex, (d) stigmasterol–MMP2 complex, (e) diosgenin–MMP2 complex, and (f) anhydroicaritin–MMP2 complex. In 2D figure (left), a green dashed line represents hydrogen bond interaction, a pink or pale pink dashed line indicates pipi T-shaped and pi-Alkyl (hydrophobic) interaction, and a gray dashed line represents metal coordination.
Discussion
Modern pharmacological research shows that Epimedium mainly contains flavonoids, which have been shown to exert an anti-osteoporotic effect by numerous experiments. Moreover, the mechanism of Epimedium is closely related to immune regulation, anti-inflammatory effects, promotion of osteoblast proliferation and differentiation, and inhibition of osteoclasts.21–27 Anemarrhen has an extremely diverse structure and has been shown to exhibit a broad spectrum of biological and pharmacological activities; for example, it can improve senile dementia, exert anticoagulant, antioxidant, antitumor, anti-osteoporotic, and anti-inflammatory effects, and lower blood pressure and blood sugar.28–33 In the active ingredient-target-disease network, quercetin, kaempferol, anhydroicaritin, luteolin, stigmasterol, and diosgenin had the highest contents and may be the key ingredients in EA. Among them, quercetin, kaempferol, and luteolin are flavonoids. Quercetin is a flavonol compound with a variety of biological activities and effects, such as antioxidative stress and anti-inflammatory effects. Quercetin can promote bone synthesis by enhancing the expression of osteogenic proteins, 34 and phytoestrogens have similar effects on inhibiting bone resorption.35–37 Kaempferol can regulate estrogen receptors, promote the proliferation of mesenchymal stem cells (MSCs) and the differentiation of osteoblasts,38,39 and improve OP by inhibiting inflammation and oxidative stress and activating autophagy in osteoblasts. 40 Anhydroicariin, a cholesterol regulatory element binding protein (SREBP) inhibitor, can inhibit RANK ligand (RANKL)-induced osteoclast differentiation and improve bone loss caused by diabetes. 41 Alizarin Red staining and mineralized nodule quantification showed that anhydroicaritin has the potential to stimulate the formation of mineralization nodules and further speed up the formation of bone, which indicates that anhydroicaritin may be a potential candidate for bone regenerative medicine. 42 Luteolin exerts an antioxidant effect because it reduces oxidative stress, promotes osteoblast differentiation, and reduces glucocorticoid-induced OP by regulating the ERK/Lrp-5/GSK-3β signaling pathway in vivo and in vitro. 43 β-Sitosterol has a hormone-like effect and can inhibit inflammatory factors by regulating the hypothalamic-pituitary-adrenergic axis system to thereby exert anti-inflammatory effects. 44 β-Sitosterol can also regulate the balance of bone metabolism and antioxidation. 45 Diosgenin has anti-inflammatory and anti-immune regulatory effects, inhibits the formation of osteoclasts in vitro, stimulates the osteogenic activity of osteoblasts and exerts a certain anti-osteoporotic effect. 46 Therefore, the results obtained by network pharmacological analysis in this study are similar to those reported in the literature regarding the mechanisms of EA in the treatment of OP.
Twenty core genes were identified, analyzed in the PPI network and sorted by degree value as follows: AKT1, IL6, MAPK1, MAPK8, JUN, VEGFA, CXCL8, IL1B, EGF, MYC, EGFR, MMP9, PTGS2, RELA, CASP3, ESR1, FOS, APP, AR, and MMP2. Among these genes, AKT1 is one of three closely related serine/threonine protein kinases (AKT1, AKT2, and AKT3), which regulate many physiological processes, including metabolism, proliferation, cell survival, and angiogenesis. 47 A study published in 2012 indicated that AKT1 may regulate the differentiation and functions of osteoblasts and osteoclasts. 48 IL-6 is a proinflammatory cytokine that is importantly related to immune-mediated bone diseases and plays important roles in the immune response and bone metabolism. 49 IL-6 can strongly activate the immune system, enhance the inflammatory response, induce the expression of M-CSF and RANK, stimulate osteoclasts, and inhibit osteoblasts, which ultimately results in bone loss. 50 A study has shown that IL-6 is associated with an increase in osteoclast activity during OP after menopause. 51 In addition, animal experiments have shown that suppressing the mouse IL-6 gene can prevent bone loss after ovariectomy (OVX). 52 MAPK1 and MAPK8 belong to the MAPK signaling pathway, and BMSC-derived exosomes promote the proliferation of osteoblasts through the MAPK pathway to improve OP. 53 By stimulating the MAPK signal transduction pathway, proinflammatory factors can affect the levels of matrix metalloproteinases (MMPs), and the overexpression of MMPs is the basis of pathological processes such as OP. 54 JNK is one of the most critical pathways related to osteoclast production. 55 Lee et al treated RAW264.7 macrophages with methylglyoxal and found that JNK is the most likely factor involved in osteoclast activation. 56 VEGFA is a homologue of the VEGF family, and functional VEGF receptors exist in primary osteoblasts that allow VEGF to promote osteoblast proliferation and bone remodelling. 57 In MSCs, overexpression of VEGFA can enhance cell viability and proliferation, and the expression levels of type I and type II collagen are significantly upregulated. 58 The protein encoded by the CASP3 gene is a cysteine-aspartic protease that plays a critical role in the execution phase of apoptosis. By targeting CASP3 and activating the PI3K-Akt signaling pathway, the overexpression of miR-378 can attenuate the osteogenic differentiation inhibited by high glucose. 59 The key gene regulation of the abovementioned EA in the treatment of OP involves primary physiological processes, including inflammation, immune response, cell proliferation, differentiation, apoptosis, migration, cell cycle progression, endocrine metabolism, angiogenesis, and growth. These core genes can be used as therapeutic targets in OP.
KEGG enrichment analysis revealed that the relevant molecular mechanisms were primarily concentrated in several types of pathways, such as cell proliferation and differentiation, apoptosis, immunity, inflammation, and oxidative stress. The AGE-RAGE signaling pathway is associated with diabetes complications, including diabetic bone disease. Advanced glycation end products (AGEs) and their receptors (RAGEs) trigger oxidative stress and inflammation in osteoblasts and osteoclasts, leading to diabetic bone disease. Diabetes damages the metabolism and function of bone cells, alters bone quality, and reduces bone strength, resulting in an increased risk of fragility fractures, which may be due to AGE-RAGE signaling. However, the AGE-RAGE signaling pathway has some beneficial effects in bones, including increasing osteogenic function. Therefore, EA likely prevents and treats OP by dually regulating the AGE-RAGE signaling pathway. 60 The TNF signaling pathway is closely related to the NF-kB signaling pathway. RANK, its ligand RANKL, and the decoy receptor OPG are key regulators of osteoclast development and functional activation. RANK is a member of the TNF receptor family, and TNF-α can increase the expression of RANKL and induce osteoclast differentiation. 61 The immune system is closely related to bones. Both TNF-α and IL-6 are important pathogenic factors for immune-mediated bone diseases, including rheumatoid arthritis and postmenopausal OP, and are involved in the immune response and bone metabolism. TNF-α and IL-6 enhance the activation and antigen presentation of macrophages via different mechanisms and regulate immune function. Many research groups have reported that TNF-α inhibits the activity of osteoblasts and stimulates the proliferation and differentiation of osteoclasts at certain stages of differentiation. 62 In addition, the IL-17 signaling pathway can induce osteoclast differentiation by stimulating the expression of RANKL in osteoblasts. Concurrently, if IL-17 signal transduction is regulated accurately and effectively, inflammation can be prevented, and blocking the IL-17 signal transduction pathway can delay the inflammation process and prevent OP. 63
These studies have shown that EA can act on OP at multiple levels through a variety of mainstream signaling pathway networks mediated by direct or indirect targets. This result explains the advantages of using EA for the treatment of OP and its potential for the development of new drugs. Only a few studies have clarified the relationship between viral pathways and OP. The current research data on virus-related pathways are abundant, and the analyses of existing databases have elucidated certain discrepancies. Therefore, virus-related pathways, such as herpesvirus infection related to Kaposi's sarcoma, human cytomegalovirus infection, hepatitis C, measles virus infection, and other virus-related pathways, may also be involved in the pathogenesis of OP. All of these results must be verified in future research.
Conclusion
OP seriously threatens people's health and reduces their quality of life. This study utilized network pharmacology and molecular docking methods to elucidate the therapeutic mechanism of EA in the treatment of OP. The EA combination has 30 active ingredients that can be used to treat OP, and the important ingredients include quercetin, kaempferol, anhydroicaritin, luteolin, stigmasterol, and diosgenin. In total, 149 target genes of EA were identified as being important in OP treatment, and among these, AKT1, IL6, MAPK1, MAPK8, JUN, VEGFA, CXCL8, IL1B, EGF, MYC, EGFR, MMP9, PTGS2, RELA, CASP3, ESR1, FOS, APP, AR, and MMP2 were identified as key target genes. GO biological process and KEGG signaling pathway enrichment analyses suggested that EA can be used to treat OP based on its ability to regulate bone metabolism, reduce inflammation, reduce oxidative stress, and regulate immunity. EA may be able to treat OP by acting on the above pathways and targets, but due to the limitations of existing databases and the corresponding analysis algorithms and software functions of various platforms, experimental research is needed to confirm these results and clarify the specific mechanisms of these phenomena. The overall findings offer a comprehensive and integrative basis and evidence for the pharmacological effects of the EA combination against OP.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-docx-1-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Supplemental Material
sj-xlsx-3-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-xlsx-3-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Supplemental Material
sj-xlsx-4-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-xlsx-4-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Supplemental Material
sj-xlsx-5-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-xlsx-5-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Supplemental Material
sj-docx-6-npx-10.1177_1934578X221086904 - Supplemental material for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis
Supplemental material, sj-docx-6-npx-10.1177_1934578X221086904 for Use of Network Pharmacology and Molecular Docking Methods to Elucidate the Curative Effect of Epimedium–Anemarrhen on Osteoporosis by Bo Xu, Yu Zhou, Qing Wang, Yujiang Liu, Long Zhang, Songlin Wang, Xianda Zhang, Wenchao Li, Xiaofeng Shen and Yuwei Li in Natural Product Communications
Footnotes
Acknowledgments
We gratefully acknowledge the contributions of Xiaoqing Shi (Nanjing University of Traditional Chinese Medicine) in the study design and data analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the National Natural Science Foundation of China (No. 82074456); Jiangsu Provincial Bureau of Traditional Chinese Medicine Program (No. QN202007); Suzhou Science and Technology Bureau Project (No. SYS2020183); Suzhou Science and Technology Bureau Project (No. KJXW2020045).
Authors’ Contributions
XB and ZY contributed equally to this paper, and they designed the study. Experimental work and data collection were conducted by XB and ZL. ZY, WQ, and LYJ analyzed and interpreted the data. XB and ZY drafted the manuscript. SXF, LYW, and WQ provided critical comments and revised the manuscript. WSL, ZXD, and LWC helped perform the analysis with constructive discussions. All authors read and approved the final version of the manuscript.
Ethics Approval
Ethical Approval is not applicable for this article.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Consent for Publication
Not applicable.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Statement of Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
