Abstract
Background
The prevalence of Hemagglutinin 9 Neuraminidase 2 (H9N2) avian influenza virus (AIV) in poultry in Asia has raised serious concerns about its zoonotic transmission and the potential for a pandemic, underscoring the need for effective therapeutics. Chikusetsusaponin IVa (CHS), a bioactive triterpenoid saponin from Radix saposhnikoviae, has exhibited anti-inflammatory activity in preclinical studies. We hypothesized that CHS could mitigate immunopathology during tumor necrosis factor (TNF)-infection by suppressing excessive inflammation.
Objectives
CHS is a natural saponin with anti-inflammatory activity. This research aimed to explore the in vitro and in vivo protective effects and mechanisms of CHS against H9N2 AIV infection.
Materials and Methods
We employed Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis and Gene Ontology (GO) analysis for the prediction of the underlying mechanisms, followed by a network analysis approach. In animal experiments, the toll-like receptor 4 (TLR4)/nuclear factor kappa B (NF-κB) signaling pathways and associated proteins were assessed using immunohistochemistry and western blot analysis.
Results
In A549 cells, CHS treatment reduced H9N2 AIV-induced apoptosis in a dose-dependent manner. CHS was found to have a suppressive effect on the mRNA expression of various inflammatory cytokines (including TNF-α, interleukin (IL)-6, IL-1β, and IL-8) as well as mediators such as inducible nitric oxide synthase (iNOS) and prostaglandin-endoperoxide synthase 2 (PTGS2). It can be seen that CHS does have anti-inflammatory effects. Bioinformatics analysis indicated that CHS might exhibit anti-inflammatory effects by modulating cytokine–cytokine receptor interaction and the NF-κB signaling pathways. Building upon this, it was found that CHS blocked the activation of TLR4 and NF-κB signaling pathways in A549 cells infected with H9N2 AIV. In vivo, CHS prolonged the survival time and increased the survival rate of H9N2 AIV-infected mice. CHS was found to have a protective effect on the lungs of infected mice, resulting in reduced damage and lower levels of viral load and inflammation. CHS inhibited the production of proinflammatory cytokines (IL-1β, TNF-α, IL-6, and IL-8) and inhibited the activation of TLR4 and NF-κB signaling pathways in the pulmonary tissues of mice that were infected.
Conclusion
CHS exerts protective effects against H9N2 AIV infection in vitro and in vivo. The anti-inflammatory mechanisms may involve the inhibition of cytokine production and the blockade of TLR4 and NF-κB signaling pathways.
Introduction
The Hemagglutinin 9 Neuraminidase 2 (H9N2) avian influenza virus (AIV) subtype has become widespread in poultry across various Asian countries and poses a significant risk to public health as it has the potential to spread from animals to humans (Liu et al., 2020; Zhou et al., 2022). H9N2 AIV can infect humans and cause mild to severe flu-like symptoms. Although human infections are typically non-fatal, the virus may become more adapted to humans and cause a pandemic (Basavaraju et al., 2022; Carnaccini & Perez, 2020; Ren et al., 2022). There is currently no effective treatment for the H9N2 AIV infection. Finding safe and affordable treatments is therefore a major global public health priority.
CHS is a naturally occurring triterpene saponin derived from Radix Saposhnikoviae. It has been traditionally used in Chinese medicine to address inflammation conditions (Guo et al., 2021). CHS has shown anti-inflammatory and immunomodulatory effects in various cell types and animal models. Anti-inflammatory activities of CHS probably include inhibition of proinflammatory mediator synthesis, including cytokines, chemokines, and prostanoids; blockade of activation of transcription factors like Nuclear Factor Kappa B (NF-κB) and receptor-typoikine signaling pathways (Liu et al., 2021; Ma et al., 2021; Shao et al., 2020; Wang et al., 2015). Given its ability to modulate inflammation and boost immunity, CHS has the potential to be used for the treatment of viral infections.
Influenza virus infections trigger excessive inflammatory responses that contribute to tissue damage and severe disease (Klomp et al., 2021). Controlling inflammation is therefore crucial in the management of influenza (Hulme et al., 2021; Park et al., 2020). We hypothesized that the anti-inflammatory effects of CHS may help control inflammation and reduce tissue injury in H9N2 AIV infections.
This paper demonstrates the protective effects of CHS against H9N2 AIV infection and elucidates the anti-inflammatory mechanisms of action, which may lead to new therapeutic options for this disease. CHS could be a promising agent for controlling inflammation and tissue injury in H9N2 AIV infection.
Materials and Methods
Materials
The human lung carcinoma A549 cells were acquired from Procell Life Science & Technology Co. (Wuhan, China, Cat CL-0016. The H9N2 AIV was obtained from National Center for Veterinary Culture Collection (CVCC) (Beijing, China). Chikusetsusaponin IVa (CHS) (purity ≥ 98%, Cat CFN92516) and oseltamivir phosphate (Lot 6002207009) were purchased from ChemFaces (Wuhan, China) and Yangtze River HEC Changjiang Pharmaceutical Co., Ltd (Yichang, China), respectively. Antibodies against glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Cat bs-0755R), toll-like receptor-4 (TLR4) (Cat bs-20594R), myeloid differentiation primary response 88 (MyD88) (Cat bs-1047R), NF-κBp65 (Cat bs-20160R), p-NF-κBp65 (Cat bs-5661R), interleukin (IL)-1β (Cat bs-0812R), IL-6 (Cat bs-0782R), IL-8 (Cat bs-0780R), and tumor necrosis factor-alpha (TNF-α) (Cat 60291-1-Ig) were acquired from Bioss Antibodies (Beijing, China), Dako (Glostrup, Denmark), and Proteintech Group (Wuhan, China). Major instruments used were quantitative PCR (Bio-Rad CFX Connect, Model 1855485), flow cytometry (Beckman Coulter Cytoflex, Model B710256), western blotting (Bio-Rad Mini-PROTEAN Tetra Cell, Model 1658005), chemiluminescence imaging (OI VisionBlue 600, Model A17162), fluorescence microscopy (Olympus TH4-200, Model TH4-200-LA-H), and confocal microscopy (Olympus FV3000, Model FV31S-H).
Flow Cytometry Analysis
The Annexin V-fluorescein isothiocyanate (V-FITC) propidium iodide (PI) Apoptosis Kit was employed to label A549 cells, and the level of apoptosis was assessed using flow cytometry.
The A549 cells (1×106 cells/well) were seeded onto sterilized slides and cultured for 12 h. After being infected with 100TCID50 H9N2 AIV, the cells were incubated with or without CHS in a fresh medium for 48 h. The cells were divided into five groups: control group, model group, or three concentrations of CHS (15, 30, and 60 µg/mL). The digestion was terminated by ethylenediaminetetraacetic acid (EDTA)-free trypsin. The liquid above was removed following a 5-min centrifugation (800 rpm, 4℃), and precooled; 200 µL 1× phosphate-buffered saline (PBS) was used to suspend the cell pellets. This process was repeated twice. the samples were resuspended in 100 µL 1× Binding Buffer, 5 µL V-FITC, and 5 µL PI at room temperature for 15 min; 300 µL of 1× Bing Buffer resuspended cells were transferred into a flow tube, which was protected from light. The Flow Jo software (Version 10) was used to interpret the results.
RT-qPCR Analysis of Gene Expression in A549 Cells Treated with CHS
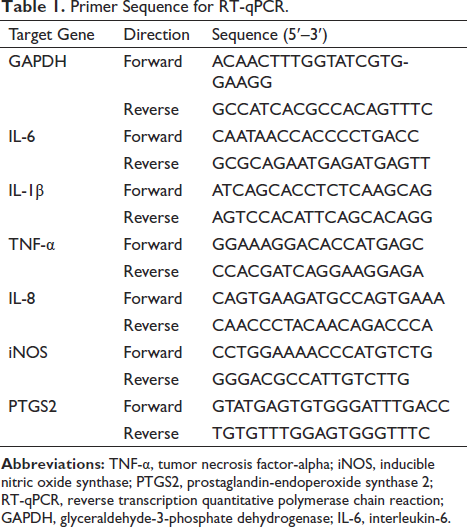
Primer sequences for TNF-α, IL-6, IL-1β, IL-8, inducible nitric oxide synthase (iNOS), prostaglandin-endoperoxide synthase 2 (PTGS2), and GAPDH genes (Table 1) were designed using Primer 5.0. A549 cells were subjected to the protocol outlined in the “Flow cytometry analysis” section and collected for quantitative reverse transcription polymerase chain reaction (RT-qPCR) analysis. The extraction of total RNA from A549 cells was performed using Trizol reagent, followed by the synthesis of cDNA utilizing the iScript cDNA Synthesis Kit. The iTaqTM Universal SYBR Green Supermix was used for RT-qPCR. The conditions were: cycles at 95℃ for 10 min, then 40 cycles at 95℃ for 10 s, at 60℃ for 30 s. The results were computed using the 2−∇∇ct method with GAPDH as the endogenous control for normalization.
Primer Sequence for RT-qPCR.

Identification and Enrichment Analysis of CHS Targets Against H9N2-induced Inflammatory Response
In order to identify the potential targets of CHS against H9N2-induced inflammatory response, we employed the STITCH database (
Network Analysis of CHS Targets Using Cytoscape
The combined score and gene names were imported into Cytoscape to obtain an initial network graph. Pathway information, including NF-κB signaling pathways and Toll-like receptor signaling pathways, was marked by matching with the KEGG database, with these pathways highlighted in dark yellow while others were marked in light yellow. The network layout was adjusted using a force-directed layout. Nodes directly connected to CHS were enlarged to highlight first-level interactions, and core targets with higher degree values were selected for further analysis.
Detection of TLR4/NF-κB Pathway-related Protein Expression by Western Blotting
Cells were treated according to the grouping detailed as before and lysed with a buffer containing protease and phosphatase inhibitors. The cells that were broken down were gently scraped using a specialized tool, and the resulting mixture was subjected to centrifugation at a speed of 12,000 revolutions per minute for a duration of 10 min at a temperature of 4°C. Protein concentrations in the supernatants were determined by bicinchoninic acid assays, and the remaining supernatant was boiled in water for 10 min to obtain protein samples. The total protein was separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto a polyvinylidene difluoride membrane. The membrane was obstructed using 5% skim milk for 1 h at room temperature, followed by overnight incubation at 4℃ with primary antibodies (diluted to a ratio of 1:10,000 for GAPDH and 1:1000 for other primary antibodies). After rinsing the membrane with 1× Tris-buffered saline with 0.1% Tween 20 detergent (TBST) for three cycles, it was subjected to incubation with secondary antibodies (diluted at a ratio of 1:3000) at ambient temperature for 1 h. It was subjected to wash again with 1× TBST three times, and protein bands were detected using an enhanced chemiluminescence (ECL) kit. The protein bands were quantitatively analyzed using ImageJ software to determine their grayscale values.
Immunofluorescence Analysis
A549 cells were treated as before and harvested for IF analysis. After the slides were slightly dried, 4% precooled paraformaldehyde was fixed for 15 min, a tissue pen was used to draw a circle in the middle of the cover slide where cells were evenly distributed. One hundred microlitres of film breaking working solution was added and incubated for 10 min at room temperature. The cells were washed with PBS for 3 times, and 3% hydrogen peroxide solution was dropped in the circle. The cells were coated with primary antibody NF-κBp65 (1:200) diluted with 5% bovine serum albumin (BSA) in a certain proportion, and incubated at 4℃ overnight. Then, FITC-conjugated Affinipure Goat Anti-Rabbit IgG (1:200) was added to the ring to cover the cells, and incubated at room temperature for 50 min. After 100 µL of 4′,6-diamidino-2-phenylindole staining, the plates were sealed by adding an anti-fluorescence quench sealing agent and imaged with a fluorescence microscope.
Experimental Design and Animal Preparation
Animal and Virus Preparation
SPF-level male BALB/c mice (6–8 weeks old, weighing 20 ± 2 g) were purchased from Wuhan Biological Product Research Institute Co., Ltd. (Production License No. SCXK (E) 2021-0026, Experimental Use License No. SYXK (E) 2017-0067). Mice underwent 1 week of familiarization in their cages held within the same basic environment inside the animal testing facility consisting of constant temperature at 22 ± 2℃, relative air humidity ranging from 40% to 70%, and alternating light/dark cycles lasting 12 h each. All animal experiments were approved by the Animal Ethics Committee of Hubei University of Traditional Chinese Medicine and conducted according to the ethical guidelines for the care and use of laboratory animals provided by the National Research Council. After 1 week of acclimatization, mice were lightly anaesthetized with ether and intranasally infected with 50 µL of H9N2 AIV urine (3LD50) for the model and treatment groups, while the normal group received 50 µL of sterile urine. The 60 mice were randomly divided into six groups, 10 mice in each group, including the normal group, model group, CHS (15 mg/kg) group, CHS (30 mg/kg) group, CHS (60 mg/kg) group, and positive drug group. The mortality of each mouse was recorded daily from 24 h after randomization to 14 days after virus infection. The survival curve of each group was plotted using GraphPad Prism 8 software. On day 5, after infection and 2 h after drug administration, the mice were euthanized by inhalation of ether. The lungs, thymus, and spleen were quickly removed and washed with precooled PBS, and the surface moisture was absorbed with absorbent paper before being weighed. The lung index, thymus index, spleen index, and lung index inhibition rate were calculated. The upper lobe of the right lung was fixed with 4% paraformaldehyde solution for pathological sectioning, while the left lung supper lobe were utilized to assess the viral load in pulmonary tissue.
IHC Detection of Protein Expression
Lung tissue paraffin blocks were deparaffinized with xylene and rehydrated with water. To retrieve the antigen, the tissue was subjected to microwaving in EDTA buffer (pH = 9.0) for 8 min at a moderate temperature. The tissue was then washed with PBS (pH = 7.4) three times for 5 min each, followed by incubation with 3% hydrogen peroxide at room temperature in the dark for 25 min, and then washed again with PBS (pH = 7.4) three times for 5 min each. The tissue was incubated with 3% BSA for 30 min at room temperature, and then incubated with primary antibodies against TNF-α (1:500), IL-1β (1:200), IL-6 (1:200), and IL-8 (1:200) overnight at 4°C. After washing the tissue three times with PBS (pH = 7.4) for 5 min each, it was incubated with a species-specific secondary antibody conjugated with horseradish peroxidase (HRP) at room temperature for 50 min. The tissue was then washed with PBS and incubated with freshly prepared diaminobenzidine (DAB) chromogenic solution for 10 min. Positive expression was indicated by brown–yellow staining. The slides underwent a 3-min hematoxylin counterstaining process, followed by dehydration and mounting using neutral resin. Images were captured using a laser-scanning confocal microscope.
Detection of TLR4 and NF-κB Signaling Pathways by Western Blotting and Immunohistochemistry (IHC)
Western blotting was employed to measure the expression levels of TLR4, MyD88, TRAF6, NF-κB p65, and p-NF-κB p65 in lung tissues. The lung tissue samples from each group were gathered and blended in RIPA buffer. The BCA protein assay kit was utilized to determine the concentration of proteins, followed by the separation of equal protein quantities through SDS-PAGE and subsequently transferred onto PVDF membranes. The membrane was dealed with 5% non-fat milk for 1 h at room temperature and then incubated with primary antibodies against TLR4, MyD88, TRAF6, NF-κB p65, p-NF-κB p65 (1:1000), and GAPDH (1:10 000) overnight at 4℃. The membranes were subsequently rinsed with TBST and exposed to HRP-conjugated secondary antibodies (diluted 1:3000) for a duration of 1 h at ambient temperature. ECL detection reagents were used for protein band visualization and quantification was performed using ImageJ software.
The immunohistochemical analysis was performed to assess the TLR4 and NF-κB p65 expression in lung tissues. Lung tissue paraffin sections were prepared as before. The sections were subsequently subjected to overnight incubation at 4℃ with primary antibodies targeting TLR4 (1:200) and NF-κB p65 (1:200), followed by a 1-h incubation at room temperature with secondary antibodies conjugated to HRP. The sections were then stained with DAB chromogenic solution and counterstained with hematoxylin. The presence and analysis of TLR4 and NF-κB p65 in lung tissues were observed under a light microscope, revealing a brown-yellow positive expression.
Statistical Analysis
All experiments were three replicates, and the data were expressed as the mean ± standard deviation of the three replicates (n = 3). SPSS 26.0 software was used for statistical analysis, and single-factor analysis of variance test analysis of variance was used to compare the statistical significance among the groups. A p-value <0.05 was considered statistically significant (p < 0.05 was significant; p < 0.01 was extremely significant).
Results
Effects of CHS on Apoptosis of H9N2 AIV-infected
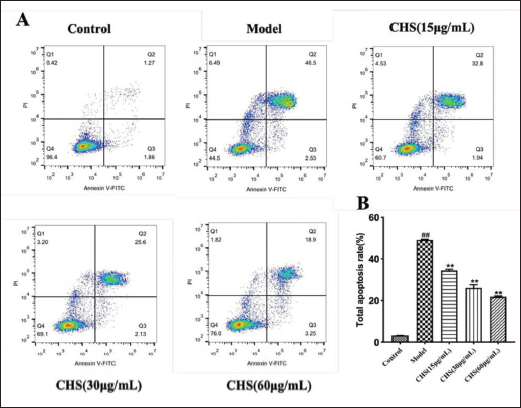
In order to assess the protective effect of CHS on A549 cells infected with H9N2 AIV, we examined the alteration in apoptosis rate following treatment with different concentrations (15, 30, and 60 µg/mL) of CHS for 48 h in H9N2 AIV-infected A549 cells. The model group exhibited a significant increase in apoptosis rate following H9N2 AIV infection compared to the control group (p < 0.01). However, the apoptosis of A549 cells showed a dose-dependent reduction, which was statistically significant (p < 0.01) (Figure 1).
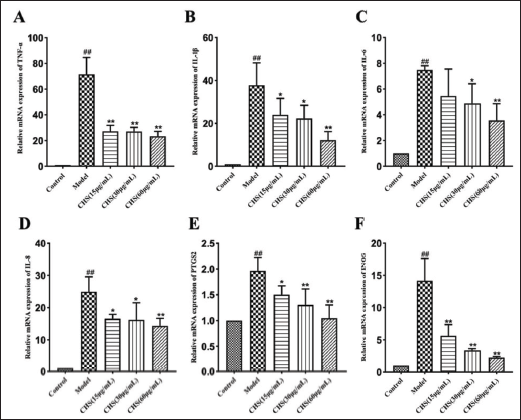
CHS Inhibited H9N2 AIV-induced Inflammatory Response
The H9N2 AIV infection can induce a strong inflammatory response by increasing cytokine production (Zhang et al., 2019). CHS has anti-inflammatory properties against various stimuli (Xu G. et al., 2020). In the model group, the mRNA expression levels of proinflammatory factors, TNF-α, IL-6, IL-1β, IL-8, iNOS, and PTGS2 were significantly elevated after H9N2 AIV infection (p < 0.01). However, in each treatment group, the mRNA expression of these inflammatory factors decreased. Notably, the CHS (60 µg/mL) group significantly inhibited the transcription level of inflammatory factors induced by H9N2 AIV in A549 cells and inhibited the development of inflammation (Figure 2).
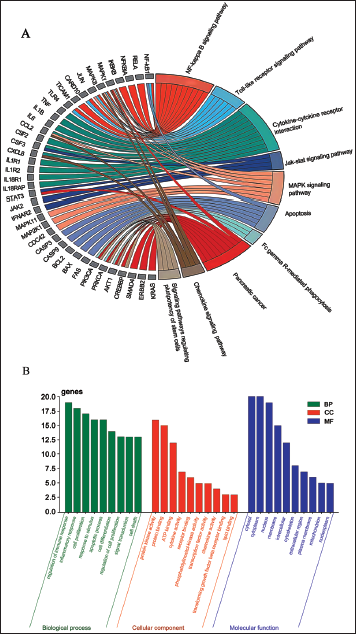
Bioinformatics Analysis Revealed the Anti-inflammatory Mechanisms of Chikusetsusaponin IVa
Our bioinformatics analysis using the STITCH database identified 50 target genes of CHS. Enrichment analysis performed using KEGG and GO demonstrated that the cytokine–cytokine receptor interaction and NF-κB signaling pathways were significantly enriched, suggesting their involvement in the regulation of inflammation mediated by CHS (Figure 3A). The mitogen-activated protein kinase signaling pathway and TLR signaling pathway were also enriched with target genes related to inflammation. The target genes were mainly involved in biological processes related to inflammatory response, cytokine regulation, and cell proliferation/apoptosis. The MFs of target genes are mainly related to protein kinase activity, ATP binding, and cytokine activity, all of which are involved in inflammation. The target genes were mainly located in organelles such as cytoplasm and nucleus, suggesting that CHS may primarily regulate inflammation by affecting events in these locations. Overall, our GO analysis effectively revealed the major biological functions and subcellular localizations involved in the target genes of CHS related to the regulation of inflammation (Figure 3B).
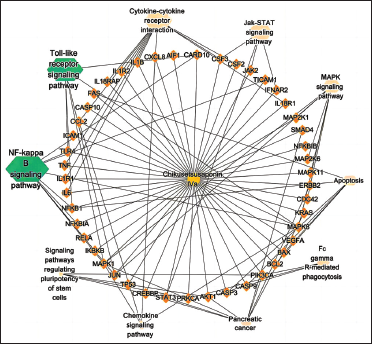
Analysis of the “Compound-Targets-Pathways” Network for CHS Anti-H9N2 Action
A “compound-target-pathway” network was constructed by screening active ingredients, cross targets, and top pathways. This network demonstrated the synergistic effects of the compound on numerous targets when CHS was used as an anti-H9N2 agent.
To analyze the network, Cytoscape 3.9.2 was utilized, and pathways with higher degree values were found to be associated with more genes. It was suggested that pathways with high degree values played a crucial role in CHSs central correlation during its anti-H9N2 action. The results showed that the NF-κB signaling pathways and TLR signaling pathway exhibited the highest degree values in the network, suggesting that they could be the primary pathways for CHSs antiviral effect against H9N2 (Figure 4).
CHS Inhibited H9N2 AIV-induced TLR4 and NF-κB Expression
We detected TLR4 and NF-κB signaling pathway-related proteins. In this study, H9N2 AIV infection in A549 cells significantly upregulated the expressions of TLR4, MyD88, TRAF6, and p-NF-κB p65 (p < 0.01). Following intervention with CHS, their expression levels were reduced compared to the model group, indicating significant differences (p < 0.05) (Figure 5A).
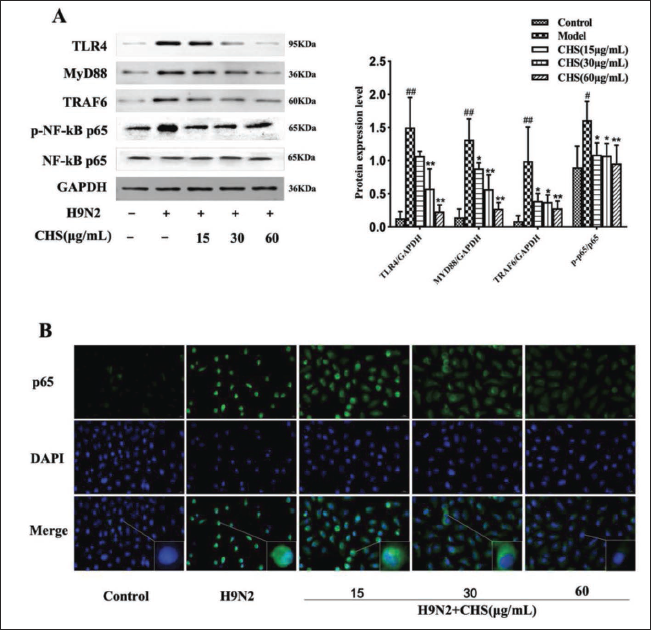
Moreover, immunofluorescence staining demonstrated that the model group exhibited rapid nuclear translocation of p65 due to NF-κB pathway activation, while CHS treatment effectively inhibited this translocation (Figure 5B).
CHS Treatment Reduced Lung Damage and Viral Load in H9N2 AIV Infected Mice
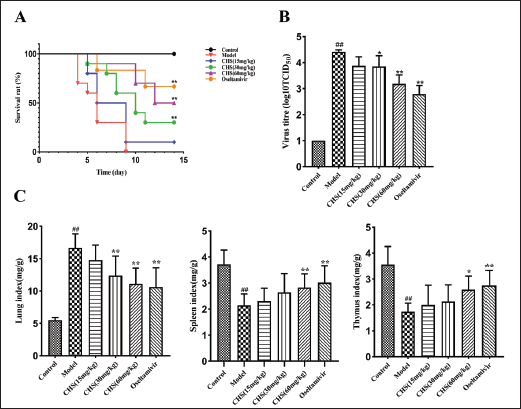
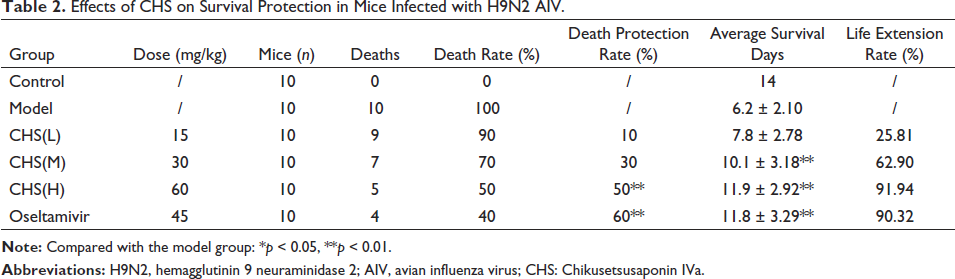
Mice in the model group exhibited mortality starting from the fourth day post-H9N2 AIV infection and all died on the ninth day. Compared with the model group, CHS (60 mg/kg) could increase the survival rate of mice by 50% (p < 0.01) and prolong the survival time of mice (11.9 ± 2.92 days), with a 91.94% increase in life span; CHS (30 mg/kg) and CHS (15 mg/kg) increased the survival rate of mice by 30% and 10%, respectively. Oseltamivir significantly increased the survival rate of mice by 60% (p < 0.01), and significantly prolonged the survival time of mice (11.8 ± 3.29 days). Its protective effect was not significantly different from that of CHS (60 mg/kg), indicating that CHS (60 mg/kg) had an anti-H9N2 AIV effect similar to oseltamivir in prolonging the survival rate of mice (Figure 6A and Table 2).
Effects of CHS on Survival Protection in Mice Infected with H9N2 AIV.
The results of lung, thymus, and spleen indices and lung tissue virus titers of H9N2 AIV-infected mice with different concentrations of CHS (15, 30, and 60 mg/kg) are shown in Figure 6B–C. In Figure 6B, the lung tissue virus titer of mice in the model group exhibited a significant increase compared to the normal group (p < 0.01). However, both the CHS (60 mg/kg) group and the oseltamivir group showed a significant decrease in the lung tissue virus titers compared to the model group (p < 0.01).
In Figure 6C, the lung index of mice in the model group showed a significant increase compared to the normal group (p < 0.01). However, treatment with CHS (60 mg/kg), CHS (30 mg/kg), and oseltamivir resulted in a significant decrease in lung indices compared to the model group (p < 0.01). It is noteworthy that despite the decrease, the lung indices of all treatment groups remained higher than those of the normal group.
Inhibition of H9N2 AIV-induced Inflammatory Cytokines in Lung Tissues by CHS
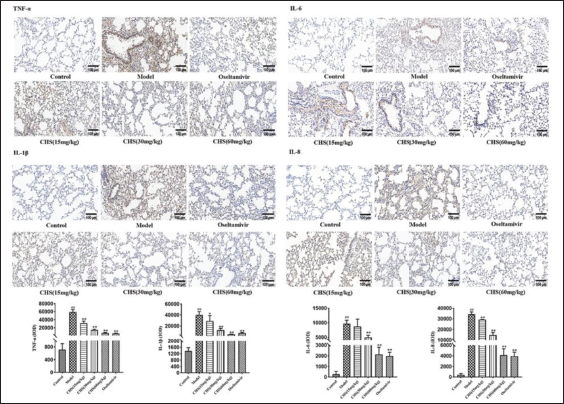
IHC was employed to measure the levels of inflammatory cytokines IL-1β, TNF-α, IL-6, and IL-8 in lung tissues (Figure 7). In comparison to the control group, the model group displayed a significant increase in the brown–yellow positive expression of them in lung tissues, while these cytokines were expressed at low levels in the normal group. All administered doses could suppress the expression of these four cytokines to varying degrees compared to the model group. As the CHS dose concentration increased, the brown–yellow positive expression of the four cytokines gradually decreased. These results suggest that CHS (60 mg/kg) significantly inhibits the massive secretion of pulmonary inflammatory cytokines induced by H9N2 AIV.
Suppression of TLR4 and NF-κB Signaling Pathways by CHS in H9N2 AIV-induced Lung Tissues
The results of western blotting showed that the model group exhibited a significant increase in the expression levels of TLR4, MyD88, TRAF6, and p-NF-κB p65 (p < 0.01), compared with the control group (Figure 8A). Following CHS intervention, the expression level of NF-κB p65 remained unaltered. The expression levels of TLR4, MyD88, TRAF6, and p-NF-κB p65 decreased in the CHS (60 mg/kg) group, displaying significant differences (p < 0.01) when compared to the model group.
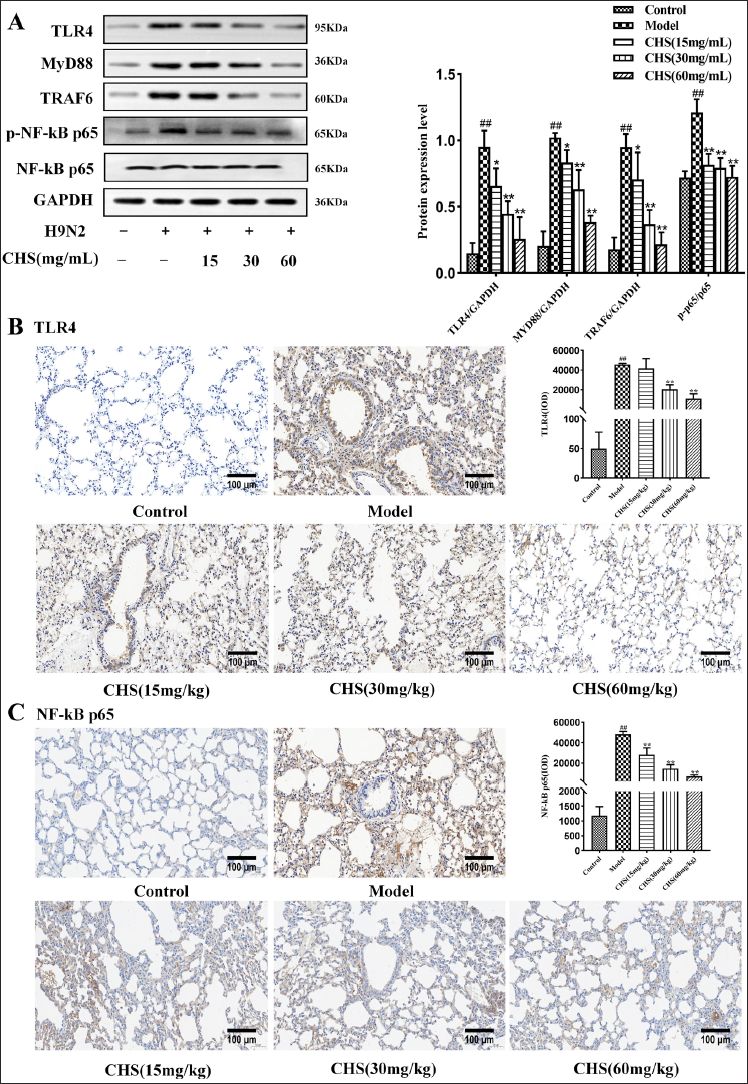
The results of the immunohistochemistry analysis indicated that the model group displayed a significant increase in brown-yellow positive expression of TLR4 and NF-κB p65 in lung tissues,compared to the control group (Figure 8B and C). Conversely, these proteins were expressed at relatively low levels in the normal group. All administered doses of CHS were found to exhibit varying degrees of suppression of their expression of them compared to the model group. As the concentration of CHS increased, there was a gradual decrease in their brown–yellow positive expression of them.
Discussion
We found that CHS treatment decreased apoptosis and inhibited inflammatory responses in H9N2 AIV-infected A549 cells. CHS also prolonged survival, decreased lung damage, and suppressed inflammation in H9N2 AIV-infected mice. The anti-inflammatory mechanisms of CHS may involve suppression of cytokine production and blockade of TLR4 and NF-κB signaling pathways.
Overproduction of proinflammatory cytokines contributes to immunopathology in cases of influenza virus infections (Carty et al., 2021; McElvaney et al., 2021; Pan et al., 2022; Tian et al., 2021). CHS effectively reduced the expression of TNF-α, IL-6, IL-1β, and IL-8 in both H9N2 AIV-infected A549 cells and mice, which may help prevent inflammation-induced tissue damage (Guo et al., 2020; Wang et al., 2020; Wang et al., 2021). By suppressing these key inflammatory cytokines, CHS could restrain excessive inflammation in the lungs during H9N2 AIV infection. In line with prior research, we discovered that CHS suppressed the activation of TLR4 and NF-κB signaling pathways, which are crucial for initiating inflammatory responses (Barnabei et al., 2021; Capece et al., 2022; Ciesielska et al., 2021; Li Y. et al., 2021; Wu et al., 2021; Xu X. et al., 2020; Zhang et al., 2022). By inhibiting these signaling pathways, CHS may suppress cytokine expression and mitigate inflammatory responses (Li S. et al., 2021).
The in vivo results demonstrated the ability of CHS to protect mice from lethal H9N2 AIV infection. CHS prolonged survival time, reduced lung viral load, and decreased lung damage in infected mice. The protective effects are likely due to the ability of CHS to control inflammation and support normal lung structure and function (Yi, 2019). Although the effects of CHS did not surpass those of the antiviral drug oseltamivir, CHS may facilitate lung recovery after infection by modulating inflammatory responses. Oseltamivir only targets viral replication but does not have anti-inflammatory effects, while CHS may regulate uncontrolled inflammation and reduce tissue damage (Świerczyńska et al., 2022; Si et al., 2021; Walsh et al., 2022). The combination of CHS and oseltamivir could be synergistic and more effective in treating H9N2 AIV infection.
In conclusion, this study reveals that CHS possesses protective effects against H9N2 AIV infection via anti-inflammatory mechanisms. CHS warrants further research for development as an immunomodulatory drug. The effects and mechanisms of CHS against other influenza A virus subtypes should also be investigated. Future work should focus on in-depth mechanisms, optimal dosage and administration regimens, as well as possible synergistic effects with anti-viral drugs.
Overall, this research provides evidence that CHS could be useful for controlling inflammation and protecting against H9N2 AIV infection. The anti-inflammatory and immunomodulatory effects of CHS could also apply to other respiratory virus infections associated with cytokine storms and severe immunopathology (Gu et al., 2021; Tay et al., 2020).
This study evaluated the anti-H9N2 effects of CHS primarily in cell and mouse models, which has limitations in translating the results to humans. First, differences exist between mice and humans in anatomy, physiology, and metabolism, which may impact the pharmacokinetics of CHS. Clinical pharmacokinetic studies should be conducted to establish suitable dosing regimens in humans. Second, safety cannot be fully assessed from mouse models alone; more comprehensive toxicology studies are required. Third, we only explored the monotherapy efficacy of CHS instead of combination therapy with existing antivirals. Finally, there may be individual differences in responses to CHS among clinical populations that need to be identified. In summary, this study provides preliminary evidence for the anti-influenza potential of CHS, but systematic assessments of its efficacy and safety profiles in preclinical pharmacology studies and clinical trials are warranted before clinical use in humans.
Conclusion
The preliminary experimental data in this study indicate that CHS may alleviate H9N2-induced lung injury by inhibiting TLR4/NF-κB activation and cytokine production. However, this study did not comprehensively assess the long-term therapeutic effects of CHS or its safety on other organs. Therefore, CHS may have the potential to treat H9N2 infection by controlling immunopathology, but further research is warranted to systematically evaluate its long-term efficacy and safety.
Summary
CHS reduced apoptosis and suppressed the expression of inflammatory cytokines and mediators in H9N2-infected A549 cells. Bioinformatics analysis predicted that the anti-inflammatory mechanisms of CHS may involve cytokine–cytokine receptor interactions and inhibition of TLR4/NF-κB pathways. Experiments confirmed CHS inhibited TLR4/NF-κB activation in infected cells. In mice, CHS prolonged survival, decreased lung damage, and suppressed inflammation in H9N2-infected mice. CHS inhibited pulmonary expression of TLR4/NF-κB and inflammatory cytokines. Molecular docking predicted potential binding between CHS and TLR4/NF-κB. In summary, CHS protected against H9N2 infection via anti-inflammatory effects by blocking TLR4/NF-κB signaling and cytokine production. CHS could be a promising agent for controlling immunopathology and tissue damage in H9N2 AIV infection.
Abbreviations
AIV: Avian influenza virus; CHS: Chikusetsusaponin IVa; TNF-α: Tumor necrosis factor-alpha; H9N2: Hemagglutinin 9 neuraminidase 2; iNOS: Inducible nitric oxide synthase; PTGS2: Prostaglandin-endoperoxide synthase 2; TLR4: Toll-like receptor 4; PI: Propidium iodide; EDTA: Ethylenediaminetetraacetic acid; PBS: Phosphate buffered saline; V-FITC: Annexin V-fluorescein isothiocyanate; RT-qPCR: Reverse transcription quantitative polymerase chain reaction; GAPDH: Glyceraldehyde-3-phosphate dehydrogenase; MyD88: Myeloid differentiation primary response 88; TRAF6: TNF receptor-associated factor 6; SDS-PAGE: Sodium dodecyl sulfate-polyacrylamide gel electrophoresis; PVDF: Polyvinylidene difluoride; ECL: Enhanced chemiluminescence; BSA: Bovine serum albumin; HRP: Horseradish peroxidase; DAB: Diaminobenzidine; RIPA: Radio immune precipitation assay; MAPK: Mitogen-activated protein kinase; ATP: Adenosine triphosphate; IL-6: Interleukin-6; IL-8: Interleukin-8; IL-1β: Interleukin-1 beta; NF-κB: Nuclear factor-kappa B; IHC: Immunohistochemistry; IF: Immunofluorescence.
Footnotes
Acknowledgments
The authors are immensely thankful to the Key Laboratory of Traditional Chinese Medicine Resources and Chemistry of Hubei Provinc for offering all the necessary facilities and infrastructure for uninterrupted conduct of this work.
Authors’ Contributions
DQ: Resources, Methodology, Funding acquisition, Writing-Original Draft. ZX: Investigation, Conceptualization, Validation, Writing-Original Draft. WST: Validation, Formal analysis. HX: Investigation, Project administration, Methodology. LYW: Supervision, Formal analysis. WY: Investigation, Data Curation. ZXQ: Project administration, Methodology, Visualization. YPT: Software, Data Curation. All the authors read and approved the final version of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Statement of Ethical Approval and Informed Consent
Compliance with Ethical Standards: All applicable international, national, and/or institutional guidelines for the care and use of experimental animals were followed. This study involved no human participants or samples obtained from patients.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Youth Project of Hubei University of Chinese Medicine (No.2021ZZX020). This research was supported by National Natural Science Foundation of China (Grant No.30470194).
