Abstract
Background
Codonopsis Radix (CR), a renowned traditional Chinese medicine (TCM) formula, has been widely applied for its immunomodulatory, antitumor, antioxidant, neuroprotective, and antiviral effects. However, the multitarget mechanism of CR in ovarian cancer (OC) remains to be elucidated.
Objectives
We applied bioinformatics and molecular docking techniques to explore possible pharmacological targets, bioactivities, and molecular mechanisms of CR for OC treatment.
Materials and Methods
We identified 40 common genes associated with CR and OC and obtained core genes through a protein–protein interaction network.
Results
Enrichment analysis revealed that mitochondrial electron transport was the key biological process involved. Based on the analysis, we selected estrogen receptor 1 (ESR1) and phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) as the key target genes for molecular docking. In the final verification analysis, we evaluated the effect of the PIK3CA mutation on the survival rate of patients with OC and determined that the phosphoinositide 3-kinase/protein kinase B (PI3K/AKT) pathway was the key pathway in the OC treatment.
Conclusion
These results suggest that CR inhibits the activity of mitochondrial complex II, reduces adenosine triphosphate (ATP) production by mitochondrial electron transport, inhibits PI3K/AKT phosphorylation, and promotes apoptosis in OC cells.
Introduction
Ovarian cancer (OC) is one of the most common and deadly gynecological malignancies worldwide (Azribi et al., 2021). Around the world, 230,000 women are annually diagnosed with OC, and 150,000 die from this disease. The main contributor to the high death-to-incidence ratio in OC is the advanced disease stage at the time of diagnosis (Fillon, 2022). According to the World Health Organization’s classification, OC is divided into five main subtypes: high-grade serous carcinoma (HGSC), low-grade serous carcinoma, clear cell carcinoma, endometrioid carcinoma, and mucinous carcinoma (Redondo et al., 2021). HGSC is the subtype with the highest mortality (Schwickert et al., 2021). Since the early symptoms of OC are non-specific, this disease is susceptible to misdiagnosis and often overlooked during the early stages (Kemppainen et al., 2019). Currently, OC is treated with surgery and chemotherapy (Morand et al., 2021). However, patients with advanced disease are highly susceptible to recurrence years after treatment due to chemotherapy resistance, and the 5-year survival rate is 47% after diagnosis (Yang et al., 2020).
Chinese medicine and natural drugs have unique advantages regarding limiting tumor growth and reducing side effects after radiotherapy owing to their low toxicity, multi-targeting, and high efficacy (Liu et al., 2020; Zhang et al., 2020). Studies have revealed that many traditional Chinese medicine (TCM) extracts and monomer compounds, such as lignans, triterpenes, and flavonoids, significantly inhibit tumor growth (Buck et al., 2011; Lee et al., 2022; Vitelli Storelli et al., 2019). These TCMs have been widely accepted as mainstream supplements and alternative therapies (Xiang et al., 2019). Codonopsis Radix (CR) is a TCM that invigorates the spleen and nourishes the lungs and blood (Gao et al., 2019). Modern medicine has shown that polysaccharides from CR play an important role in immunomodulation, antitumor, antioxidation, neuroprotection, and antiviral effects (Luan et al., 2021). In addition, the active compounds in CR can inhibit the expression of matrix metalloproteinase, activate the reactive oxygen species (ROS)-mediated p38 pathway, induce caspase-dependent apoptosis, and inactivate the Ras/phosphoinositide 3-kinase/protein kinase B (PI3K/AKT) pathway, thereby exerting cytotoxic effects on cancer cells (Ahn et al., 2020; Lee et al., 2005; Wang et al., 2022). Although CR has rich experimental data and clinical practice in the treatment of cancer, the active components, corresponding target genes, and potential mechanisms of action of CR in OC treatment have not been fully elucidated.
Therefore, this study aimed to investigate the mechanisms and targets of CR in OC treatment using bioinformatics and molecular docking methods and provided new insights regarding the clinical intervention and treatment of OC.
Materials and Methods
Screening of Active Ingredients and Targets for CR
The active ingredients and targets of CR were screened using the Bioinformatics Analysis Tool for Molecular Mechanism of Traditional Chinese Medicine (BATMAN-TCM Database,
Acquisition of Target Genes for OC
OC-related target genes were acquired from the GeneCards database (
Construction of Targets Interacting with the Active Ingredients of CR
OC-related target genes and targets for CR were imported into Venny (Venny 2.1.0,
Establishment of the Protein–Protein Interaction (PPI) Network
In the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING 11.5,
Construction of the Interaction Network and Analysis
The targets of CR among the OC-related targets identified using STRING were further analyzed using Cytoscape software v3.8.0 to visualize and analyze the interaction network. We ranked by the DEGREE method using the CytoHubba plug-in.
Enrichment Analysis
First, the targets were imported into the DAVID Bioinformatics Resources (DAVID database,
Molecular Docking
By screening target genes with higher degree values in the PPI core network and enriched in the KEGG pathway, we identified the active ingredients associated with these target genes and used the active ingredients as ligands and the target genes as receptors for molecular docking. The docking process combined AutoDock Tools 1.5.7 with AutoDock Vina. The parameters of the mating box were obtained from AutoDock Tools 1.5.7, and docking was performed using AutoDock Vina. Finally, docking models with the lowest binding affinity were visualized. Hydrogen bonds and intermolecular force connection lines were displayed using PyMOL.
Validation of Key Genes
The cBioPortal (
Results
Identification of OC-associated Genes and Intersection with CR
Based on the set conditions, we screened 53 drug components and 295 drug targets from the BATMAN-TCM database and 1,473 disease targets from the GeneCards database and identified 40 target genes where drugs intersected with diseases (Figure 1A).
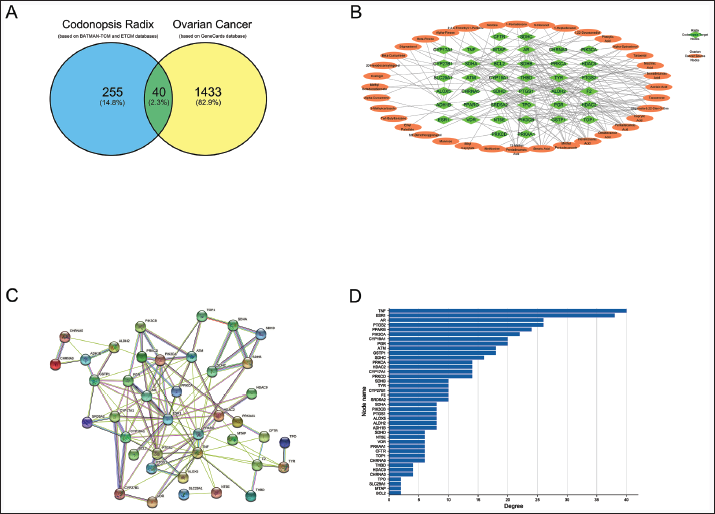
Interaction Network Analysis of Target Genes
Forty target genes as target nodes and 36 active ingredients corresponding to the target genes as attribute nodes were imported into Cytoscape 3.8.0 to construct the “CR active ingredient-target genes” interaction network (Figure 1B and Supplementary Table 1).
Visualization of PPI Networks and Further Screening of Target Genes
Analysis of PPI networks constructed from the 40 target genes in the STRING database revealed 40 nodes, 121 edges, an average node count of 6.05, and a PPI enrichment p-value < 1.0e−16 (Figure 1C). The results of the PPI network analysis were imported into the Cytoscape software, and the 40 target genes were ranked by node degree using the CytoHubba plugin (Figure 1D).
Enrichment Analysis, Pathway Annotation, and Mechanism Prediction of Target Genes
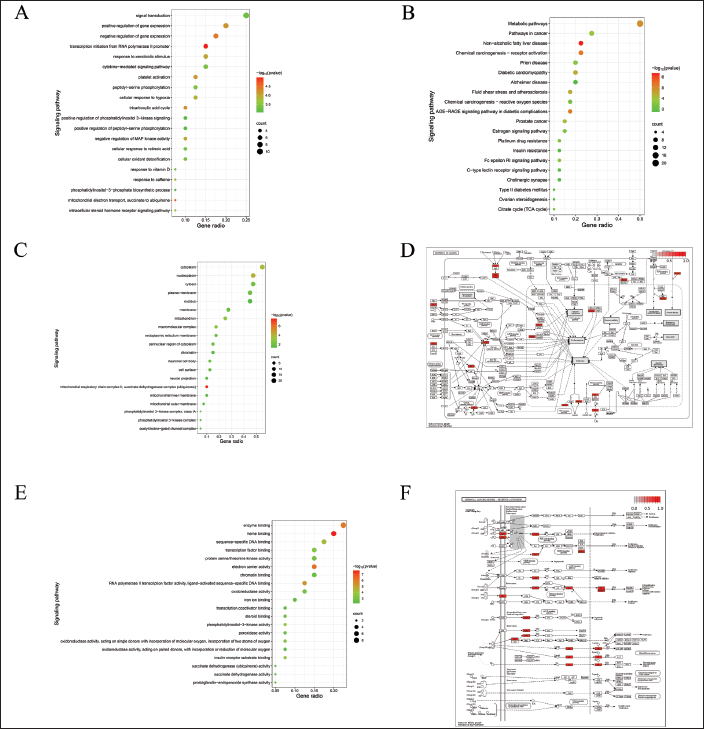
GO enrichment analysis was performed using the DAVID and Metascape databases. The results of GO analysis based on biological processes, cellular composition, and molecular function were visualized by taking the 20 entries with the smallest p-values. The results revealed that the biological processes enriched by the target genes primarily included transcription initiation from the RNA polymerase II promoter, mitochondrial electron transport from succinate to ubiquinone, and negative regulation of gene expression (Figure 2A). The cellular composition of the target genes mainly included the mitochondrial respiratory chain complex II and the succinate dehydrogenase complex (ubiquinone) (Figure 2C). The molecular functions of the target genes included heme binding, electron carrier activity, and enzyme binding (Figure 2E). Twenty entries with the smallest p-values were used for visualization of the KEGG pathway analysis (Figure 2B), and the results showed that the pathway in cancer (Figure 2D) and the chemical carcinogenesis-receptor activation pathway (Figure 2F) were significantly associated with OC.
Enrichment Analysis and Path Annotation. (A) Biological Processes Terms. (B) Kyoto Encyclopedia of Genes and Genomes Pathway Enrichment Analysis. Node Color was Displayed as a Gradient From Red to Green in Descending Order of p-values. The Size of the Nodes was Arranged in Ascending Order of the Number of Genes. (C) Cellular Component Terms. (D) Pathway in Cancer. (E) Molecular Function Terms. (F) The Chemical Carcinogenesis-receptor Activation Pathway. The Red Node Represented the Target Genes Enriched in the Pathway.

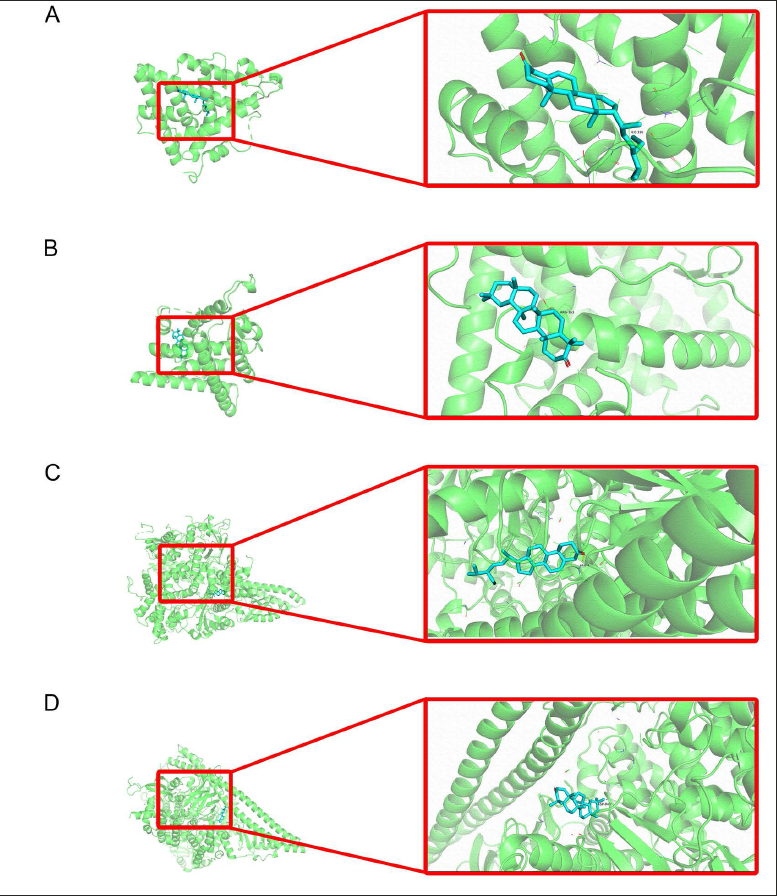
Molecular Docking
Combining the degree values and the results of KEGG analysis, we found that estrogen receptor 1 (ESR1) and phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) genes could be significantly expressed in OC treatment and that these two genes corresponded to two of the same active components of CR, stigmasta-5,22-dien-3-one and taraxerone. We then identified the corresponding target proteins of ESR1 and PIK3CA as 1SJ0 and 4JPS, which were docked to the small-molecule structures of stigmasta-5,22-dien-3-one and taraxerone, respectively (Figure 3). Molecular docking results indicated that all docking complexes had binding energies ≤ −7 kcal/mol, suggesting stable binding capacity in humans (Supplementary Table 2).
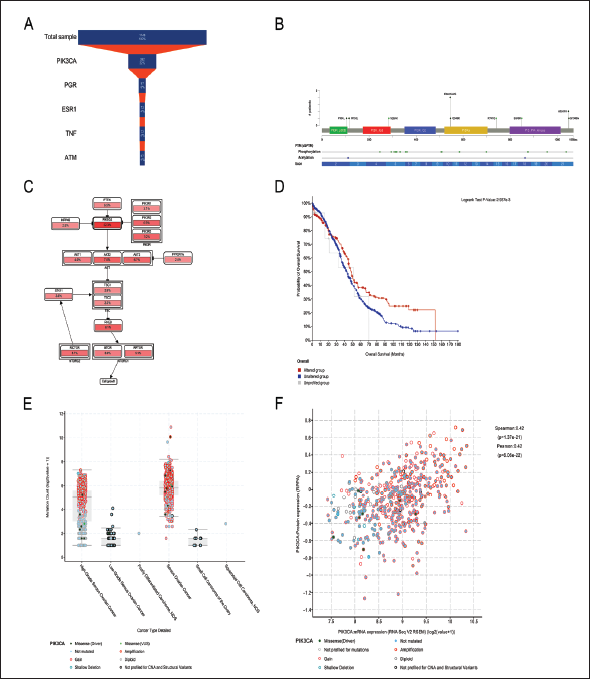
Genetic Alterations and Survival Analysis of Key Targets
The top 10 target genes with degrees were selected from the cBioPortal database for validation analysis of the 1,748 OC samples (Supplementary Table 3). We observed that PIK3CA mutations were more frequent than the other mutations (progesterone receptor [PGR], ESR1, tumor necrosis factor [TNF], ATM Serine/Threonine Kinase [ATM]) (Figure 4A). Therefore, we selected PIK3CA as a target for our study. We used posttranslational modification (PTM) to activate the molecular mechanism of PIK3CA and identified 18 missense mutations (including four duplication mutations in patients with multiple samples). Two point mutations, P104L and R115L, have been reported in PIK3CA exon 2; N284K mutation was observed in exon 5; E545K, E545A, E545G, and Q546K mutations were detected in exon 10; R741Q and E849K mutations were found in exons 15 and 18, respectively; and H1047R and G1049A were mutated in exon 21 (Figure 4B). Compared to the other samples, the mutation frequency of E545A in PIK3CA was the highest. Kaplan–Meier survival curves revealed that upregulation of PIK3CA expression significantly increased patient survival (Figure 4D and Supplementary Table 4), with a mean increase in the survival time of 4.96 months (95% confidence interval). Clinical evaluation showed a large number of missense mutations and copy number amplification mutations of PIK3CA in patients with high-grade serous OC and serous OC (HGSOC), as well as diploid mutations in patients with HGSOC, low-grade serous OC, poorly differentiated carcinoma, serous OC, and squamous cell carcinoma (Figure 4E). We also identified an association between mRNA and protein expression of PIK3CA mutation types (Figure 4F). Finally, we identified the genes associated with PIK3CA in the PI3K pathway (Figure 4C), from which we concluded that CR may inhibit PIK3CA mutations in the PI3K pathway in OC treatment.
Discussion
OC is the most lethal gynecological cancer. Chemotherapy resistance and the selection of effective therapies for patients remain major challenges (Schwickert et al., 2021). Although the survival rate for early-stage OC exceeds 90%, there is no evidence indicating that OC screening can save lives (Menon et al., 2021). Therefore, there is an urgent need to develop new treatments and therapeutic drugs to impede disease progression (Liu et al., 2020). In this study, data were mined and analyzed using bioinformatics and molecular docking techniques to reveal the biological activity, core therapeutic target genes, and potential mechanisms of CR in OC.
We screened the top 10 genes with the highest degrees in all OC studies in the cBioPortal database for evaluation and identified five genes with the largest sample sizes (TNF, ESR1, PIK3CA, PGR, and ATM), which could be potential targets for CR therapy in OC. TNF-α activates the transcription of relaxin, and targeting the relaxin/RXFP1 pathway can overcome chemoresistance in platinum-resistant ovarian tumors (Burston et al., 2021). ESR1 can inhibit the PIK3CA mutation in the PI3K pathway, upregulate the pro-apoptotic gene BIM, retain normal cells, and induce cancer cell toxicity (Stratikopoulos et al., 2019). PGR is a ligand-activated transcription factor that has been identified as a pivotal mediator of many processes associated with ovarian and uterine functions. Studies have shown that luteinizing hormone mediated by the PI3K/AKT signaling pathway can induce PGR expression in preovulatory follicles (Ogiwara et al., 2021). ATM is a PI3K kinase that plays an important role in DNA damage repair (Arnould et al., 2021). It has been proven that the PI3K signaling pathway is critical in this study.
Enrichment analysis revealed that CR inhibited OC by affecting mitochondrial electron transport and respiration. Adenosine triphosphate (ATP) is produced during oxidative phosphorylation in the mitochondrial electron transport chain (ETC) (Zhao et al., 2019). Ubiquinol produced by mitochondrial complexes I and II, which supply electrons to ubiquinone, reportedly promotes the regeneration of nicotinamide adenine dinucleotide (NAD+) and flavin adenine dinucleotide (FAD). Mitochondrial complex III is responsible for oxidizing ubiquinol back to ubiquinone, thereby promoting tumor growth and allowing complexes I and II to continue to function. When complex II-mediated respiration is inhibited, ubiquinol production is also inhibited. Consequently, the functions of mitochondrial complexes I and III are weakened, adversely affecting mitochondrial oxidative phosphorylation and ATP synthesis and impairing the growth of cancer cells (Bandara et al., 2021; Martínez-Reyes et al., 2020).
By combining the degree values and the results of enrichment analysis, we selected ESR1 and PIK3CA as the key target genes for molecular docking and were surprised to find both of them in the active components of stigmasta-5,22-dien-3-one and taraxerone. Stigmasta-5,22-dien-3-one is a derivative of stigmasterol, which has been shown to activate endoplasmic reticulum stress sensor protein and endoplasmic reticulum–mitochondrial axis protein in OC cells and thus treat OC through signaling cascade inhibition of the PI3K/MAPK pathway (Bae et al., 2020). Taraxerone can inhibit the proliferation and colony formation of cancer cells, demonstrating antitumor potential through apoptosis and cytotoxicity in a dose-dependent manner (Barboza et al., 2020). The results of docking studies showed that stigmasta-5,22-dien-3-one and taraxerone could bind effectively to ESR1 and PIK3CA, and the docking binding energy of taraxerone and PIK3CA was the lowest with the most stable structure.
PIK3CA, the p110α catalytic subunit of PI3K, is an upstream component of the PI3K pathway (Srinivas et al., 2019). Studies have shown that the most common molecular alterations in the PI3K pathway are mutations in the PIK3CA gene that induce hyperactivation of p110α (Verret et al., 2019). In advanced HR +/HER2-breast cancer, PIK3CA mutations have been shown to be chemoresistant and have a poor prognosis (Mosele et al., 2020). Therefore, the inhibition of PIK3CA mutations is key to the treatment of OC by CR. When PIK3CA displays abnormalities, AKT is activated, and phosphatidylinositol 3,4,5-trisphosphate is elevated, causing the production of cancer cells (Li et al., 2019). PIK3CA mutations can occur in any of the four domains of the gene: the adaptor-binding, C2, helical, and catalytic (kinase) domains. Most mutations focus on the E542 and E545 sites in the helical domain and the H1047 site in the kinase domain. E542 and E545 are commonly replaced with lysine, whereas H1047 is frequently substituted with arginine (Pearson et al., 2018). Previous studies reported that activation of p53 by cisplatin via PTM leads to attenuation of PIK3CA mutations and subsequently reduced AKT phosphorylation, resulting in reduced protein synthesis, survival, proliferation, and upregulation of the apoptotic machinery (Thakur & Ray, 2016). Interestingly, our study found that CR not only inhibits PIK3CA mutations but also exhibits lower toxicity than chemotherapy drugs. As such, it could be a potential candidate for OC treatment.
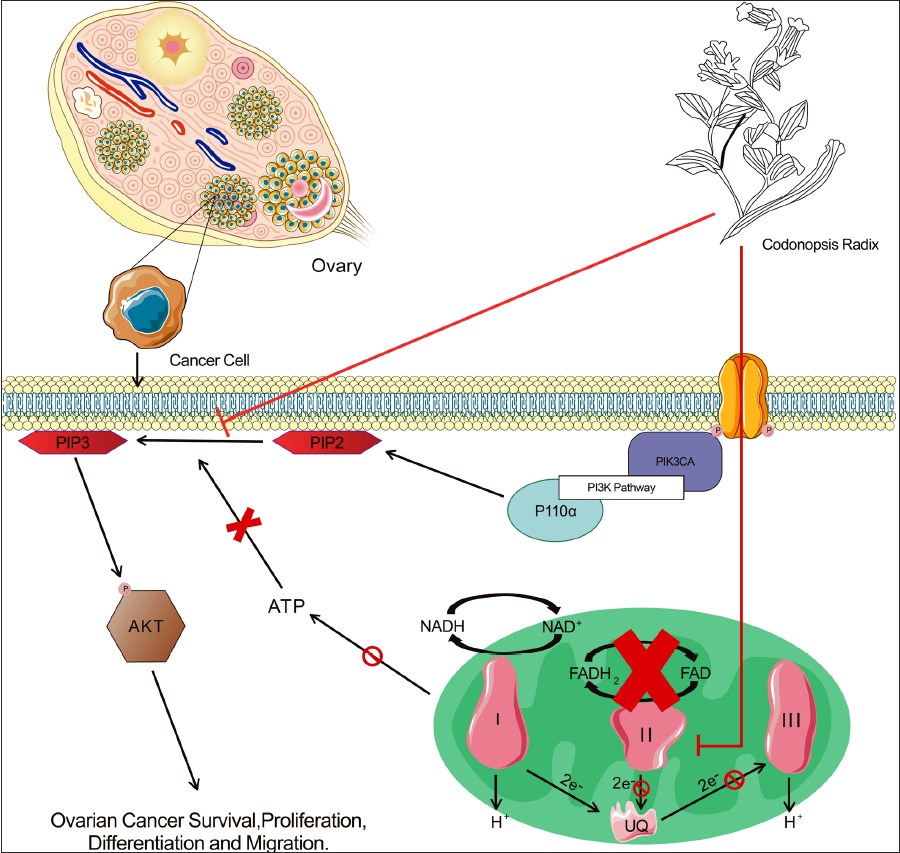
Therefore, we concluded that a potential mechanism of action of CR in OC treatment may be that CR inhibits the activity of mitochondrial complex II, reduces ATP production by mitochondrial respiration, inhibits phosphorylation of the PI3K/AKT pathway, and promotes the apoptosis of OC cells (Figure 5). However, our study had some limitations. All data were obtained from databases, and we explored the therapeutic role of CR in OC only at the bioinformatics level, with no subsequent in vivo or in vitro trials to support our conclusions. Additionally, since CR has many active ingredients, other ingredients may also play a role in OC inhibition, which might have been overlooked by our research and screening methods. Therefore, additional research is required to further explore the potential molecular mechanisms of CR in OC treatment in vitro and in vivo.
Conclusion
This study used bioinformatics methods to identify target genes and active components in CR for OC treatment. We identified a highly significant PI3K/AKT pathway and screened out the key biological process of mitochondrial electron transport and the key target PIK3CA through enrichment analysis. Finally, we confirmed our findings with molecular docking, genetic changes, and survival analyses of key targets. Therefore, we can conclude that CR is a potential therapeutic drug for OC treatment.
Abbreviations
OC: Ovarian cancer; CR: Codonopsis radix; PPI: Protein-protein-interaction; KEGG: Kyoto Encyclopedia of Genes and Genomes; ESR1: Estrogen receptor 1; PIK3CA: Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha; PGR: Progesterone receptor; TNF: Tumor necrosis factor; ATM: ATM Serine/Threonine Kinase; PIP2: Phosphatidylinositol-(4,5) bisphosphate; PIP3: Phosphatidylinositol 3,4,5-trisphosphate; UQ: Ubiquinone; PI3K: Phosphoinositide 3-kinase; NAD+: Nicotinamide adenine dinucleotide; NADH: Nicotinamide adenine dinucleotide, reduced; FAD: Flavin adenine dinucleotide; FADH2: Flavine adenine dinucleotide, reduced; ATP: Adenosine triphosphate; AKT: Protein kinase B; HGSC: High-grade serous carcinoma; LGSC: Low-grade serous carcinoma; CCC: Clear cell carcinoma; EC: Endometrioid carcinoma; MC: Mucinous carcinoma; TCM: Traditional Chinese Medicine; GO: Gene ontology; HGSOC: High-grade serous ovarian cancer; ETC:Electron transport chain ; TCGA: the Cancer Genome Atlas; MSK: Memorial Sloan Kettering; MSKCC: Memorial Sloan Kettering Cancer Center.
Footnotes
Acknowledgments
The authors would like to thank the Editage team for polishing the English language.
Authors’ Contributions
ZHC and YRY proposed and designed this study, and YZ performed the analyses. YRY and YS revised the manuscript. All the authors read and approved the final manuscript. ZHC and YZ contributed equally to this work.
Availability of Data and Materials
The authors confirm that the data supporting the findings of this study are available within the article (and/or) its supplementary materials.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Not applicable.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Supplementary Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
