Abstract
To explore potential therapeutic targets and underlying mechanisms of apigenin (API) for papillary thyroid carcinoma (PTC) treatment.
Keywords
Introduction
Despite improving living standards, cancer is still one of the most serious life-threatening health problems and is the leading cause of death worldwide. 1 The number of new cancer cases in 2020 was 19.3 million, but the number is expected to be over 28.4 million by 2040, 2 which is an extreme burden on the whole society. 3
Thyroid cancer (TC) is one of the most malignant endocrine system diseases. 4 Papillary thyroid cancer (PTC) is the most prevalent type of thyroid carcinoma accounting for more than 85% of all diagnosed TC cases. 5 Recently, with the wide application of high-resolution thyroid ultrasonography, the number of diagnosed PTC cases has increased significantly. 6 Several studies indicated that genetic alterations, environmental factors, radiation exposure and family history of PTC may contribute to its development. 7 In general, surgical treatment is still the first-line therapy for diagnosed PTC. Other clinical treatments include radiotherapy, chemotherapy, immunotherapy and combination therapy. 8 However, excessive toxicity, 9 radiation-related thyroid dysfunction 10 and chemoresistance 11 of the existing treatment all remind us that we need to find more efficient and safer therapeutic approaches to PTC.
Phytochemicals can exert immense beneficial effects to our body. 12 Until now, researchers have identified nearly 10,000 phytochemicals, though many more are yet to be discovered. 13 Polyphenols are secondary metabolites which account for the majority of phytochemicals. 14 Recently, more and more attention has been paid to identify polyphenol's health beneficial properties. 15 Apigenin (API; 4′,5,7-trihydroxyflavone), a bright yellow flavone, is abundant in many plants, such as celery, parsley and chamomile flowers. 16 Recently, accumulating studies in vitro and in vivo have shown API's anti-inflammatory, anti-tumor, anti-apoptosis and cardioprotective effects. 17 Sunanda Panda et al found that 0.78 mg/kg API significantly increased serum concentrations of thyroid hormone (T3 and T4) in alloxan-induced diabetic mice, and stimulated synthesis and secretion of thyroid hormone thus regulating thyroid function. 18 Zhen Sun et al revealed that API played a chemotherapeutic role in human PTC BCPAP cells and significantly induced autophagy and ROS production in BCPAP cells. 19 All these findings suggest that API may act as a promising drug in treating PTC. However, few studies have been carried out to identify the underlying mechanisms of API on PTC.
Network pharmacology is a newly emerging method that uses network analysis to identify a drug's biological ingredients and underlying therapeutic mechanisms. 20 Recently, in-silico techniques have become more and more powerful in drug research.21–24 The application of network pharmacology and in-silico techniques gives us better insights into the effect of drugs in specific diseases.25–28 In the present study, network pharmacological methods combined with bioinformatic tools were used to search for molecular interactions between API and PTC. Furthermore, an interaction network between API and PTC was constructed based on publicly available databases. Our study aimed to elucidate potential therapeutic targets and detailed therapeutic mechanisms of API for treating PTC and to provide a theoretical basis for PTC research.
Materials and Methods
Database Search for API-Interacting Targets
In the present study, API (Pubchem CID:5280443) was used as the ligand and was then retrieved from PubChem Database (https://pubchem.ncbi.nlm.nih.gov/). 29 The structure of API was downloaded in SDF format and was then converted to PDB format by using Open Babel software(http://ope-nbabel.org/wiki/Main_Page). 30 A 2D structure of API was presented by using ChemDraw software (Version 18.0). 31 We searched for potential API-interacting targets in SwissTargetPrediction database (http://www.swisstargetprediction.ch/index.php), Traditional Chinese Medicine Systems Pha-rmacology Database (TCMSP, https://old.tcmspe.com/tcmsp.php), and Encyclopedia of Traditional Chinese Medicine database (ETCM, http://www.tc-mip.cn/ETCM/index.php).25,32,33 The search results were pooled, and duplicates were removed.
Database Search for PTC-Related Targets
We searched for PTC-related targets in GeneCards database (https://www.genecards.org/), DisGeNET database (https://www.disgenet.org/) and Online Mendelian Inheritance in Man database (OMIM, https://omim.org/), using keywords ‘papillary thyroid carcinoma’ and ‘papillary thyroid cancer’ and species ‘Homo sapiens’.34–36 The search results were pooled, and duplicates were removed.
Protein-Protein Interaction (PPI) Network Construction and Core Targets Identification
PPI networks are effective internet tools that are used for discovering biological interaction among proteins. Firstly, potential API-interacting targets and PTC-related targets were overlapped in a Venn diagram (http://bioinformatics.psb.ugent.be/webtools/Venn/) to find targets in common. 37 The intersected targets were considered potential therapeutic targets of API against PTC. STRING database (https://cn.string-db.org/), an online database, currently covers more than 24,000,000 proteins from 5000 organisms and enables researchers to construct functional associations for the proteins.38,39 We then investigated functional association of these targets by using STRING database. The minimum interaction score was set as high confidence >0.7. Furthermore, cytoHubba plugin was used to predict core targets. Three different methods, namely MCC, MNC and Degree method, were applied in core target prediction. Cytoscape software (Version 3.8.0) was used to visualize the network analysis. 40
Functional Enrichment Analysis
In the present study, the Database for Annotation, Visualization and Integrated Discovery (D-AVID database, https://david.ncifcrf.gov/) was used to investigate the Gene Ontology(GO)-Biological Processes (BP) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. 41 Intersected targets were uploaded to DAVID database and the species was selected as Homo sapiens. KEGG analysis revealed crucial pathways involved in the predicted targets. A P value less than 0.05 was considered statistically significant. Additionally, Bioinformatic database (http://www.bioinfor-matics.com.cn/) was used to illustrate a target's bioinformatic annotation.
Construction of Gene-Pathway Network
To understand better the molecular mechanism involved in API-related PTC treatment
Verification of Expression Patterns and Prognostic Values of Predicted Core Targets
The Human Protein Atlas (HPA database, https://www.proteinatlas.org/) is an open access database aimed at mapping all the human proteins in cells, tissues and organs. 42 To increase further the credibility of our prediction, the HPA database was used to detect expression of predicted core targets in normal tissues and PTC tissues. Furthermore, the Gene Expression Profiling Interactive Analysis (GEPIA2) database was used to investigate potential interaction between disease-free survival (DFS) and predicted core targets. 43 In the present study, group cutoff was set as median and percentage of cutoff-high and cutoff-low was set as 50%. Furthermore, log-rank P < 0.05 was considered as statistically significant.
Molecular Docking
Molecular docking analysis was used to identify potential interaction between API and predicted targets. API(CID:5280443) was retrieved from PubChem database. Entry number of predicted proteins was obtained from Uniprot Database (https://www.uniprot.org/). 3D crystal structures of predicted macromolecules were obtained from Protein Data Bank database (PDB, https://www.pdbus.org/), whereas selection criterion was selected as Homo sapiens with refinement resolutions ranging from 1.0A to 3.0A. The collected original pdb format files were uploaded into Autodock and MGL tools for further analysis. Furthermore, ligand and macromolecules were prepared by using Pymol software (https://pymol.org/2/) including removal of water molecules, and charging addition and extraction of unwanted hetero-atoms. Additionally, Autodock (https://autodock.scripps.edu/) and Autodock tools 1.5.7 software (https://ccsb.scripps.edu/m-gltools/) were used to perform docking procedures between ligand and macromolecule using default parameters and the grid box was laid for the whole protein. 44 In the present study, the generated conformation with the best binding affinity was selected and visualized in Pymol and Discovery Studio 2019 software.45,46 The ligand binds to proteins freely when the binding energy is less than 0. A lower binding energy score indicates a higher affinity of the ligand to target macromolecules. 47
Results and Discussion
Screening for Potential Therapeutic Targets of API for Treating PTC
The 2D structure of API is shown in Figure 1. A search for API-interacting targets was carried out in three databases, namely TCMSP, SwissTargetPrediction, and ETCM. The search yielded 80, 104, and 41 targets, respectively. These molecules were pooled together to obtain a total of 204 targets. A search for PTC-related targets was carried out in three online databases, namely GeneCards, DisGeNET, and OMIM. The search yielded 2918, 1348, and 634 targets, respectively. These targets were pooled together to get a total of 3826 targets. We then compared API-interacting targets with PTC-related targets and found 110 targets in common (Figure 2A). These were considered as potential therapeutic targets for API against PTC (Table 1). Gene network analysis of potential therapeutic targets of API for PTC treatment are shown in Figure 2B.
Chemical structure of API.

Network diagram of the overlapping targetsbetween API and PTC. (A)Gene network analysis of potential therapeutic targetsof API for treating PTC. The PPI network contains 96 nodes and 475 edges. (B)Network diagram of drug-disease target genes. The green V node represents API, the orange V node represents PTC, purple triangles represent unique API-related targetsand cyan nodes represent API-related therapeutic targets for treating PTC.
Putative Targets of API for Treating PTC.

Drug-Disease-Interaction Network Construction
STRING database was used to construct a protein-protein interaction (PPI) network among 110 potential therapeutic targets. The resulting network was visualized by using Cytoscape software (Figure 3A). After removing outliers, the final PPI network contained 96 targets and 475 interactions, denoted by nodes and edges, respectively. Nodes with larger size and deeper color were more essential elements in the network. To identify the most essential API targets, we used three prediction methods, namely Maximal Clique Centrality (MCC), Maximum Neighborhood Component (MNC), and Degree. MCC, a newly proposed topological method, has a better performance in identifying essential proteins from complex PPI networks. 48 To ensure the accuracy of prediction, by overlapping prediction results from the three methods on a Venn diagram (Figure 3B), we obtained 9 targets in common. These were tumor protein p53(TP53), heat shock protein HSP90-alpha member 1(HSP90AA1), RAC-alpha serine/threonine-protein kinase (AKT1), epidermal growth factor receptor (EGFR), roto-oncogene tyrosine-protein kinase Src (SRC), vascular endothelial growth factor A (VEGFA), actin, cytoplasmic 1 (ACTB), transcription factor Jun (JUN) and estrogen receptor1 (ESR1) (Figure 3C). The degree of the top 9 targets is shown in Table 2 and they were considered core pharmacological targets and used for further analysis.

Network analysis of core targetsof API for treating PTC. (A)Cytoscape analysis representing potential therapeutic targetsof API in the treatment of PTC. (B)Venn diagram of analysis results from CytoHubba(MCC, MNC, Degree). The overlapped parts representing core genes. (C) Network diagram of predicted core targets. Darker and redder represents higher degree and diamond nodes represent predicted core targets.
Degree of Top 9 Core Targets.

Functional Enrichment Analysis
To understand better the functional mechanism of API in the treatment of PTC, DAVID database was used to perform enrichment analysis. We found a large number of enrichment terms including 371 BP terms and 112 KEGG terms that were significantly enriched. The top 20 BP and KEGG terms were visualized in a chord diagram (Figure 4A and 5A), bubble chart (Figure 4B and 5B) and histogram chart (Figure 4C and 5C) (P < 0.05). BP enrichment results suggested that the overlapping targets were involved in regulation of apoptotic process, positive regulation of phosphatidylinositol 3-kinase signaling, regulation of cell proliferation, and response to drug and protein phosphorylation. According to KEGG pathway analysis, the potential therapeutic targets were mostly enriched in hsa05200 (Pathways in cancer), hsa04151 (PI3K-AKT signaling pathway) and hsa04115 (p53 signaling pathway). The PI3K-AKT signaling pathway plays an important role in tumor apoptosis and the p53 signaling pathway is known to be involved in regulation of the apoptotic process, cell proliferation and DNA repair. Our KEGG enrichment analysis results suggested that API might exert positive effects on PTC by regulating the PI3K-AKT and p53 signaling pathways.

Biological processes enrichment analysis of core targets. (A)The chord plot circle represents the top 20 enriched BP terms. The left grey part represents enriched targets and the colored ribbons represent enriched GO terms. (B)The bubble chart represents the top 20 enriched BP terms. The X-axis represents gene ratio, the Y-axis represents description and node sizes are represented as enriched gene counts. (C)The histogram represents the top 20 enriched BP terms. The X-axis represents enriched gene counts, the Y-axis represents description and -log10(pvalue) are represented as intensities of colors.
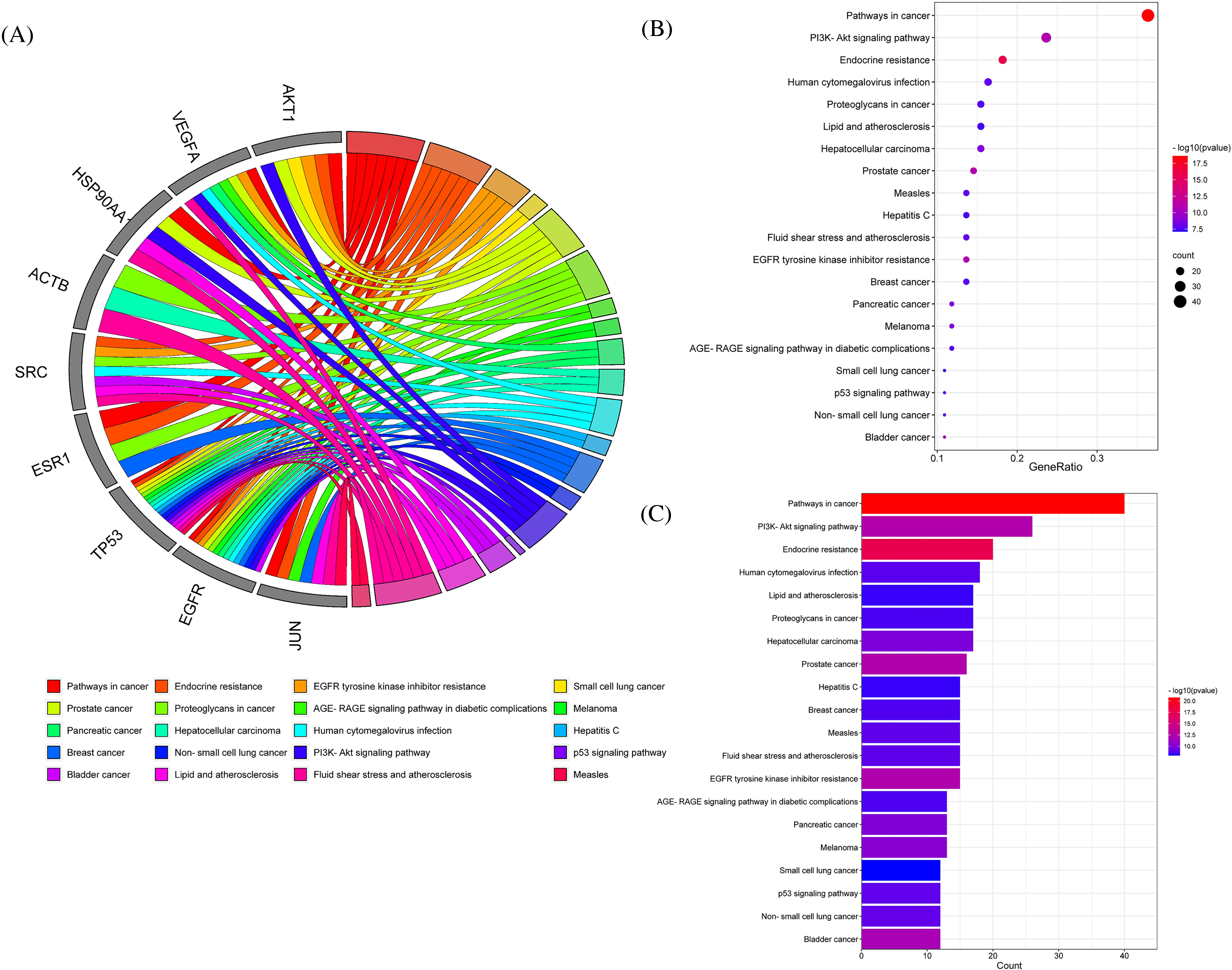
KEGG enrichment analysis of core targets. (A)The chord plot circle represents the top 20 enriched KEGG terms. The left grey part represents enriched targets and the colored ribbons represent enriched KEGG terms. (B)The bubble chart represents the top 20 enriched KEGG terms. The X-axis represents gene ratio, the Y-axis represents description and node sizes are represented as enriched gene counts. (C)The histogram represents the top 20 enriched KEGG terms. The X-axis represents enriched gene counts, the Y-axis represents description and -log10(pvalue) are represented as intensities of colors.
Gene-Pathway Network Construction
The gene regulatory pathway network was constructed to identify the distribution of potential therapeutic targets and relevant pathways. The gene-pathway network is shown in Figure 6. In this network, predicted core targets and the related top 20 GO BP terms and KEGG signaling pathways were carried out with degree and visualized using Cytoscape software. Our results demonstrated that AKT1 had the most maximum degree and was considered as the core target gene. Other genes such as EGFR, SRC, VEGFA, TP53, and JUN also had larger degree, which further indicated that they might be core targets for the regulatory pathways of API against PTC. Enriched terms and corresponding contained genes are shown in Table 3 (BP) and Table 4 (KEGG).
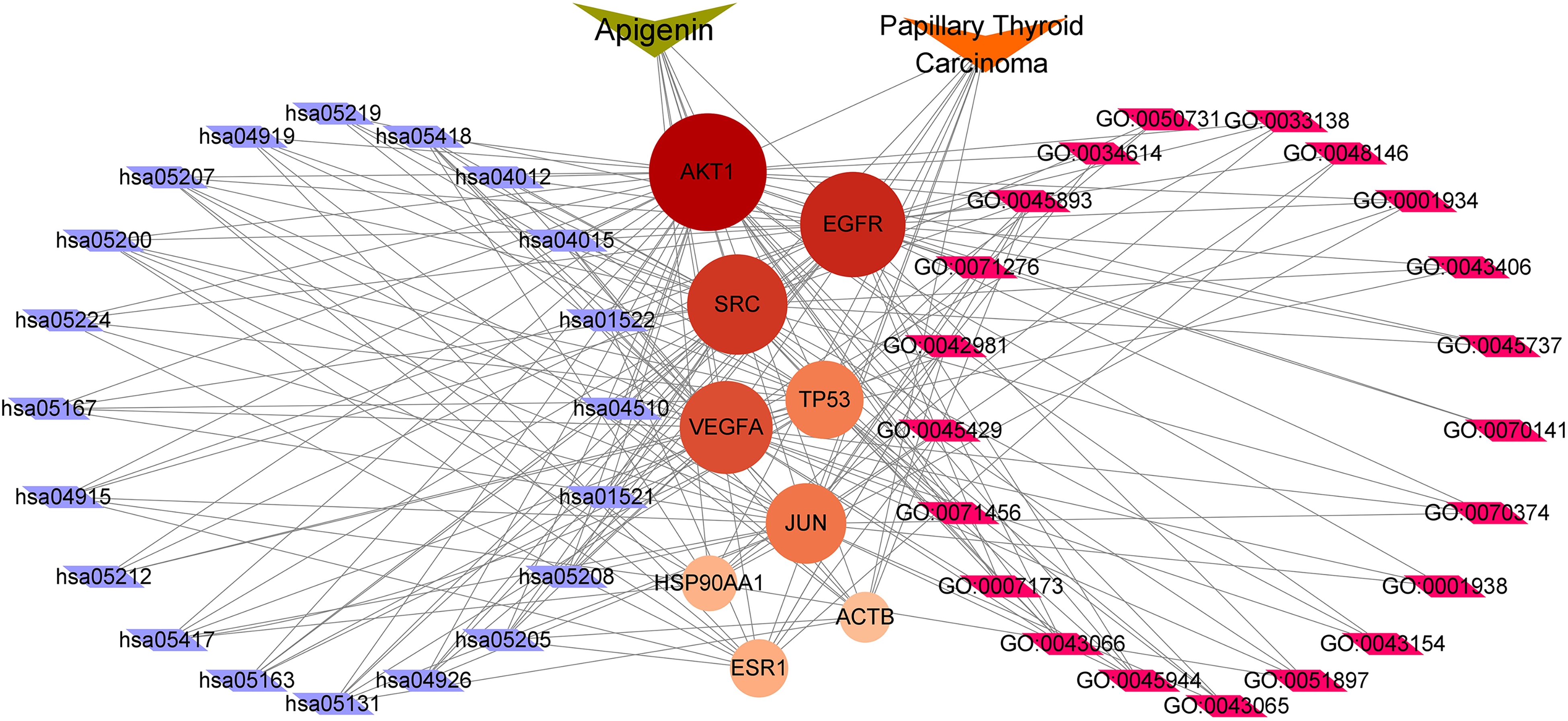

Gene-Pathway network of API for treating PTC. The orange V node represents PTC, the brown V node represents API, red circles represent core targets, purple parallelograms represent top 20 predicted KEGG pathway and pink rectangles represent top 20 predicted GO biological processes. Larger represents higher degree.
Enriched Biological Processes Relating to Core Targets.

Enriched Pathways Relating to Core Targets.

Expression Patterns and Survival Analysis of Predicted Core Targets
HPA and GEPIA databases were used to examine the expression pattern of API core targets in PTC patients and their relationship with survival rates. The group cutoff was set as median and the axis units were set as months. Significantly higher protein expression of ACTB, EGFR, HSP90AA1, AKT1, SRC, and VEGFA were observed in PTC tissues compared with normal tissues, whereas lower expression of JUN was observed in PTC tissues (Figure 7). Interestingly, lower expression of JUN was also correlated with lower survival rate of PTC patients (Figure 8F). Our findings suggested that JUN might serve as a potential biomarker for PTC treatment.

Differential expressed analysis of predicted core targets from HPA database. Representative IHC results of (A)ACTB, (B)AKT1, (C)TP53, (D)EGFR, (E)ESR1, (F)VEGFA, (G)SRC, (H)JUN and (I)HSP90AA1 in normal and PTC tissues from HPA database.

DFS analysis of predicted core genes from GEPIA2 database. Representative prognostic results of (A)ACTB, (B)AKT1, (C)EGFR, (D)ESR1, (E)HSP90AA1, (F)JUN, (G)SRC, (H)TP53 and (I)VEGFA in patients. The patients are divided into low(green) and high(red) group based on the median expression of predicted genes. Log-rank p < 0.05 is considered as statistically significant.
Molecular Docking Analysis
In this study, in silico analysis was used to identify biomolecule interaction between API and predicted proteins. Crystal structures of HSP90AA1(PDB ID: 7lt0), VEGFA (PDB ID: 4kzn), Akt1 (PDB ID: 7nh5), SRC (PDB ID: 1a09), Jun (PDB ID: 5t01), TP53 (PDB ID: 1aie), ESR1 (PDB ID: 1err), EGFR (PDB ID: 1m17) and ACTB (PDB ID: 6ict) were used as receptors for molecular docking.49–57 Binding energy of API with predicted proteins ranged from −7.52 to −4.47 (Table 5) suggesting API had strong affinity with these proteins (Figure 9). Previous studies indicated that the affinity between curcumin and TP53 and VEGFA was −6.5 and −5 kcal/mol and the binding affinity between resveratrol and AKT1 was −6.2 kcal/mol.58,59 Our study revealed that API showed better affinity with VEGFA (−5.27) and AKT1 (−6.83) in in silico studies. The co-crystalized ligand of the chosen macromolecule was re-docked using the same docking parameter as a test (Table 6). As for TP53, our results have shown that API possessed three conventional hydrogen bonds with ILE332, THR329 and ASN345, one carbon hydrogen bond with ILE332, and one pi-alkyl interaction ARG342; the residues LEU330, PHE341, GLU346, PHE338 and GLN331 were involved in van der Waals’ interactions with API. Additionally, API also showed strong binding affinity with AKT1 (−6.83) by forming one conventional hydrogen bond with THR211, SER205 and LYS268; three pi-pi stacked interactions with TRP80; one pi-anion interaction with ASP292; one pi-sigma interaction with LEU210; one pi-alkyl interaction with VAL270; the residues, including ASN 204, ALA212, ILE290, LEU264, and ASN53, took part in van der Waals’ interaction.

Molecular view of best docking complexes between API and (A)TP53, (B)AKT1, (C)ACTB, (D)SRC, (E)HSP90, (F)VEGFA, (G)EGFR, (H)JUN, (I)ESR1. The API was hosted in the active site and was represented as green color. The macromolecule is represented as purple color and the binding sites are represented as cyan color. 2D intermolecular interactions of API with macromolecules are shown in the left.
Molecular Docking Score.

Molecular Docking Simulation Results.

Discussion
Thyroid cancer is the most prevalent endocrine system disease. PTC, the most common type of thyroid cancer, has gained attention in recent years due to its high incidence. 60 Though surgical removal is still the choice of treatment for PTC, its unexpected complications and side effects make it imperative to search for more reliable therapeutic strategies. The beneficial role that phytochemicals play in thyroid function has been fully discussed in recent years. 61 As one of the most common type of phytochemicals, API has received much attention due to its low toxicity and potent biological activity. 62 Though API is reported to be effective in the treatment of PTC, the central therapeutic targets and mechanisms of API against PTC still remain unclear.
In the present study, we created a network prediction of API-associated pharmacological targets and molecular mechanisms for treating PTC. To the best of our current knowledge, this is the first study revealing potential pharmacological targets and mechanisms of API against PTC by using network pharmacological methods. By using bioinformatic analysis, we obtained 110 targets which were related to PTC pathology and API treatment. Through the drug-disease interaction network, the top 9 targets with higher degree (TP53, HSP90AA1, AKT1, EGFR, SRC, VEGFA, ACTB, JUN and ESR1) were identified as core pharmacological targets related to PTC. Meanwhile, JUN was found to be closely related to the prognosis of PTC patients. According to our results, these core targets also showed better affinity with API in in silico studies, suggesting that API exerted its pharmacological effects by treating PTC via these targets. The enrichment analysis results demonstrated that regulation of the apoptotic process, positive regulation of phosphatidylinositol 3-kinase (PI3K) signaling, modulation of cell proliferation, and regulation of the PI3K-Akt/p53 signaling pathway might also be involved in the API treatment process. It appears that API could function against PTC by inducing apoptosis, cell death and immunological reactions.
TP53 displays the highest degree of protein-protein interaction among the identified core targets, which suggests that it could be the central target of API against PTC. This finding agrees with the existing observation that tp53 plays an essential role in regulating cell proliferation, apoptosis and DNA damage response. Previous studies have established a strong correlation between p53 expression and PTC progression. Elevated p53 expression was observed in diagnosed PTC patients. 63 Activation of the p53 signaling pathway was closely associated with the progression of PTC. 64 These findings are in agreement with our current results. A therapeutic approach targeting p53 and its associated signaling pathway has achieved preliminary success in preclinical studies. 65 Furthermore, API has been reported to induce apoptosis through a p53-independant signaling manner. 66 As an effective drug for treating PTC, API increases the stability of p53 at the same time and plays a vital role by targeting p53. A recent study reported that API induced G2/M phase cell cycle arrest and apoptosis in MCF-7 cells by regulating the p53 signaling pathway. 67 Our results imply that the regulation could be mediated by direct API-p53 interaction.
Aberrant activation of apoptosis-associated signaling pathway is central in PTC development. The PI3k-Akt signaling pathway is a well-known intracellular apoptotic pathway which is closely associated with the occurrence of PTC. 68 AKT1 is a key regulator in cell motility and the apoptotic process, the activation of which promotes metastasis in thyroid cancer. 69 Four out of 9 core targets identified in this study, namely VEGFA, AKT1, EGFR, and HSP90AA1, are in the PI3k-Akt signaling pathway. PI3K belongs to the lipid kinase family 70 ; activated PI3k induces phosphorylation of Akt and leads to the translocation of Akt from cytoplasm to the nucleus further regulating its downstream cellular process. 71 Emerging evidence shows that inhibition of PI3k and its related pathway can be a novel target against cancer. 72 Studies have shown that vitamin C dose-dependently inhibited the PI3k-Akt signaling pathway in BRAF thyroid cancer cells. 73 Thyroid hormone receptor beta (TRβ), a known PI3K inhibitor, exerts an anti-tumor effect in anaplastic thyroid cancer (ATC). 74 A study by Lu et al found that LncRNA HOTTIP deficiency inhibited cell proliferation, invasion and migration in PTC cells by targeting AKT1. 75 The pro-apoptosis effect of API in vitro was extensively verified. Furthermore, API has been shown to be effective in regulating the PI3K-AKT signaling pathway. 76 Yang et al reported that API could inhibit cell proliferation and induced apoptosis in hepatocellular carcinoma cells by inhibiting the PI3k/Akt/mTOR signaling pathway. 76 Thus, by regulating both p53 and the PI3K-AKT signaling pathway, our study indirectly demonstrates that API appears to be a promising and multitarget drug in the fight against PTC.
HSP90 is an upstream regulator of the AKT-related signaling pathway. 77 HSP90AA1, a stress inducible isoform of Hsp90, is reported to be involved in cell survival, cell circle control and apoptosis. 78 Under heat shock conditions, structural changes in cell membrane contribute to cell death. In PTC cell lines, expression of HSP90 is extremely high. 79 White et al found that HSP90 inhibitor could effectively inhibit migration and invasion of thyroid cancer stem cells. 80 Besides, API was reported to overcome the chemo-resistance of pancreatic cancer by targeting the mTOR-HSP90-mutp53-p62-NRF2-antioxidant response axis. 81 The SRC oncogene has been proved to act as key regulator in tumor progression. 82 Functionally, SRC binds to focal adhesion kinase (FAK) forming FAK/SRC complex and phosphorylates various downstream signaling molecules. SRC is also reported to phosphorylate the serine site of AKT and further affects its activity. By using an immunohistochemistry approach, a recent study found that compared with benign thyroid lesion, the percentage of positively SRC-stained cells was significantly high. 83 A recent study revealed that inhibition of the SRC-mediated PI3K pathway improved the overall survival of PTC patients with BRAF and RAS mutations. 84 Furthermore, treatment with API significantly attenuated SRC expression and exerted a chemopreventive activity against ultraviolet-induced skin inflammation. 85
EGFR is a transmembrane glycoprotein receptor belonging to the ErbB family of tyrosine kinase proteins. ErbB family members are associated with apoptosis and cell proliferation. Compared with benign thyroid tissues, a significantly increased mRNA level of EGFR was observed both in PTC tissues and cancer-adjacent tissues. 86 The BRAF V600E mutation is one of the most frequent genetic alterations in PTC patients. Targeting BRAF V600E appears to be a promising strategy for PTC patients exhibiting BRAF V600E mutations. Previous study demonstrated that treatment with EGFR inhibitor effectively enhanced the anticancer-effect of PLX4032 (a selective BRAF V600E inhibitor) in vitro. 87 What's more, Wang et al revealed that treatment with API could enhance the antitumor effects of cetuximab in nasopharyngeal carcinoma by inhibiting the EGFR signaling pathway. 88 These findings indicated that EGFR was involved in PTC development and API treatment.
The fact that PTC is more prevalent in females than in males implies possible involvement of estrogen in PTC progression. The estrogen receptor α gene ESR1 mediates the biological activity of estrogens and is reported to play a key role in PTC progression. Previous study indicated that increased level of ESR1 was observed in PTC tissues than in adjacent normal thyroid tissues. 89 Besides, lower overall survival is observed for PTC patients with higher ESR1 expression. Researchers have found that API could function as an effective antiestrogen in various type of cancer. 90 For instance, Nephew et al reported that API could inhibit the ESR1-dependant signaling pathway in breast cancer cell line. 91 Furthermore, treatment with API significantly inhibited cervical cancer growth by reversing histamine-induced abnormal activation of ER signaling. 92 Our findings suggest that API might exert antiestrogen effect in PTC tissues by targeting ESR1 and its-related signaling.
VEGFA is a member of the vascular endothelial growth factor (VEGF) family that regulates angiogenesis and vascular permeability, both of which are involved in tumor progression and metastasis. Angiogenesis is reported to be involved in PTC progression 93 and VEGFA has been reported to induce MMPs secretion, which can degrade the basal membrane. 94 Dysregulated permeability may lead to tumor expansion. Recent research suggested that inhibition of VEGF-mediated angiogenesis could be an effective way of PTC treatment. Salajegheh et al found that miR-126 significantly reduced VEGFA expression and inhibited cell proliferation in the PTC cancer cell line. 95 To some extent, API is a potent angiogenesis inhibitor. In non-small cell lung carcinoma, API significantly suppressed VEGF-A/VEGFR2 and PDGF-BB/PDGFβR signaling pathways and further reduced microvessel density. 96 Additionally, Ko et al found that in breast cancer cells, API suppressed secretion of VEGF and significantly overcame drug resistance via inhibition of the STAT3 signaling pathway. 97
c-JUN, one of the most well-studied transcription factors of the activator protein-1 (AP-1) family, is reported to be involved in a various of physiological processes including cell proliferation, immune activity and tumorigenesis. The role that c-JUN plays in the tumorigenesis of papillary carcinomas is well studied. A recent bioinformatics analysis indicated that c-JUN was closely relevant to PTC genesis. 98 Furthermore, Li et al found that increased expression of phosphorylated c-Jun NH2-terminal kinase (p-JNK) was highly associated with the presence of lymph node metastases. 99 Therefore, targeting the JNK/c-JUN signaling pathway may be a promising therapeutic intervention in PTC treatment. In the present study, JUN was identified as a core pharmacological target involved in API-related PTC treatment and was deduced as a prognosis marker for PTC patients. What's more, API is more likely to play a role by targeting JUN. Previous studies revealed that treatment with API (30 μM) significantly attenuated phosphorylation of JNK associated with MAPK signaling pathway in RAW264.7 cells. 100 Jeong et al found that pre-treatment with API and luteolin in HaCaT cells suppressed ultraviolet irradiation-induced expression of c-JUN. 101 Thus, API targeting JUN may provide totally fresh insights and improve prognosis in the treatment of PTC.
In conclusion, API is expected to be a promising and effective drug for treating PTC. Despite all these novelties, there are several limitations to this study. On one hand, network pharmacological methods are based on the existing network databases, and in-depth mechanisms and relevant results still require constant updating. On the other hand, further pharmacological verifications are still required to verify our results and we will continue to investigate API's role in the treatment of PTC. Finally, much remains to be done to fully understand the pathogenesis of PTC and transfer our findings to clinical practice.
Conclusions
Taken together, in this study we systematically elucidated possible targets and potential therapeutic pathways of API in the treatment of PTC by using network pharmacology, bioinformatics analysis and molecular docking. Based on the network pharmacological results, 9 effective pharmacological targets (TP53, HSP90AA1, AKT1, EGFR, SRC, VEGFA, ACTB, JUN and ESR1) were identified as therapeutic targets of API against PTC, and molecular docking results also suggested that API could bind with these targets stably, further providing a theoretical basis to investigate the pathogenesis of PTC. Meanwhile, JUN served as a new prognosis of PTC. The functional enrichment results strongly suggest that therapeutic mechanisms of API against PTC might be associated with regulation of the apoptotic process, mediating cell proliferation, immunomodulation and regulating the PI3k-Akt/p53 signaling pathway. Our study provides valuable insights and a theoretical basis for further investigation of API for treating PTC.
Footnotes
Acknowledgements
The authors thank Dr Zhiyun Chen (Department of Microbiology and Immunology, University of Oklahoma Health Sciences Center, Oklahoma City, Oklahoma, USA) for the English editing of the manuscript.
Author's Contribution
All authors contributed to the revision of the study. Dongyu Li carried out the study and contributed to data collection and analysis. Lei Wang and Yuchen Jing contributed to manuscript writing. Bo Jiang, Lei Zhao and Yuxi Miao contributed to data analysis. Shijie Xin and Chunlin Ge contributed to manuscript corrections. All authors approved final manuscript to be published.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Availability of Data and Materials
The data supporting the findings of this study are available within the article.
