Abstract
Objective
To explore potential multi-target anti-ovarian cancer mechanisms of Panax notoginseng (PN) using an integrated computational–experimental approach.
Methods
Network pharmacology was used to identify PN constituents, predict targets, and intersect them with ovarian cancer (OC) genes. Core targets were screened via a PPI network, followed by GO/KEGG enrichment. Predicted binding was evaluated by molecular docking and 100-ns MD simulations. In vitro validation was performed in A2780 cells to assess viability and migration, including combination with olaparib.
Results
A total of 127 putative anti-OC targets were identified for 8 PN components, and 7 core targets (HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1, AKT1) were highlighted. Enrichment suggested involvement of lipid metabolism/atherosclerosis and xenobiotic metabolism pathways. Docking suggested favorable predicted interactions, and MD supported stable behavior of representative complexes. PN-derived components, alone or with olaparib, reduced A2780 cell viability and migration under the tested conditions.
Conclusion
PN may modulate OC through multi-target mechanisms supported by computational predictions and in vitro observations. Further mechanistic and in vivo studies are needed to assess translational relevance.
This is a visual representation of the abstract.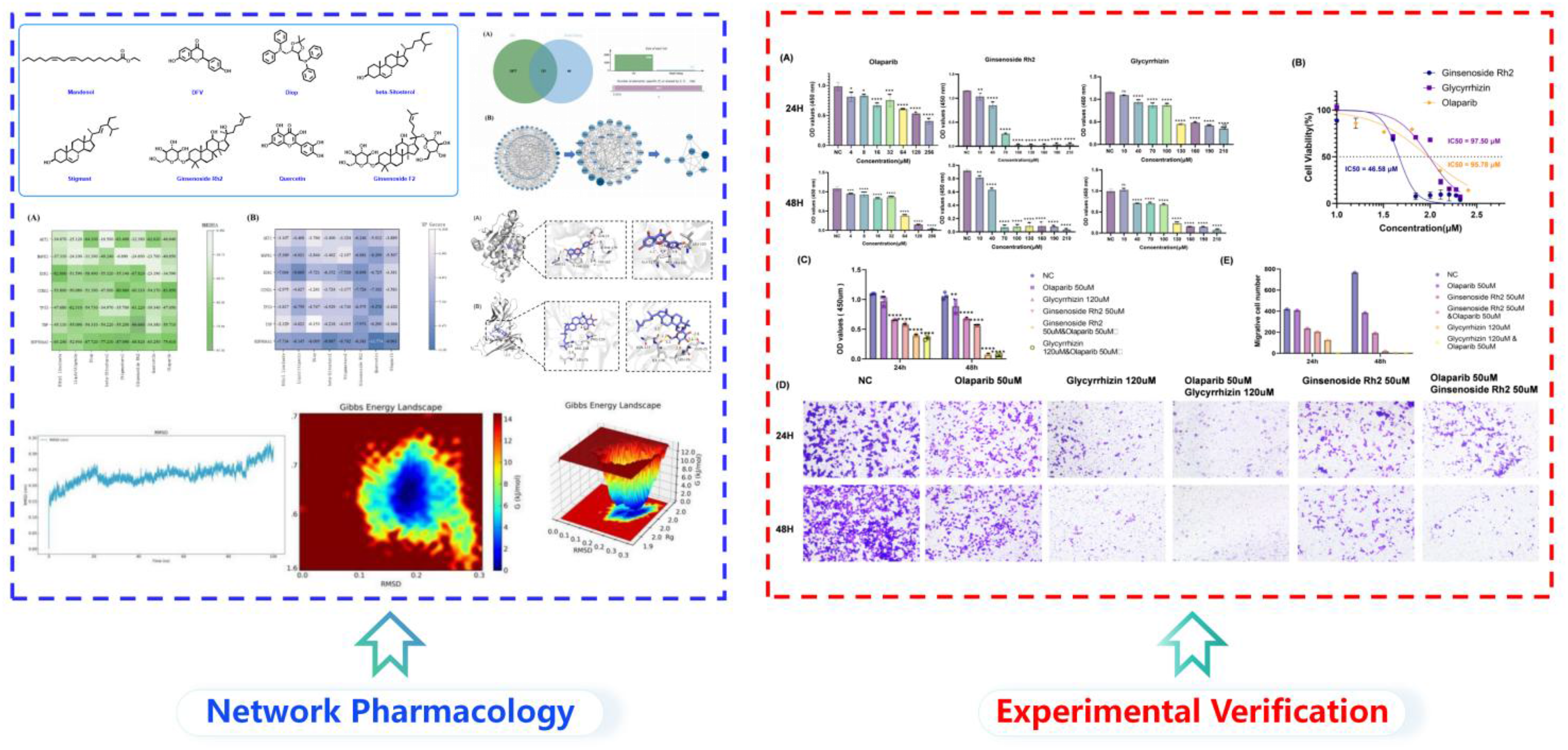
Introduction
Ovarian cancer (OC) represents a molecularly heterogeneous disease encompassing epithelial (EOC), stromal, and germ cell malignancies, with high-grade serous Ovarian carcinoma (HGSOC) constituting 70% of EOC cases.1,2 Globally, Ovarian cancer constitutes 4.7% of cancer-associated mortalities among females, characterized by delayed clinical diagnosis—60%–70% of cases are detected at FIGO stage III/IV—and dismal long-term prognosis, with a 5-year survival rate of only 49% for advanced-stage patients.2,3 Residual disease post-cytoreduction and chemoresistance-driven recurrence remain critical barriers to long-term remission.2,4 Therefore, exploring drugs that can effectively treat Ovarian cancer and overcome drug resistance is of great significance.
Panax notoginseng (PN), traditionally called Sanqi in Chinese, is renowned for its historical medicinal use. The root of Panax notoginseng (Burk.) F.H. Chen, a perennial herbaceous plant in the Panax genus of the Araliaceae family, has been used for more than 500 years in clinical treatments for a variety of diseases. Emerging evidence supports PN's applicability in Ovarian cancer (OC). Network pharmacology studies identify OC-related targets regulated by PN components, suggesting roles in apoptosis induction and chemoresistance reversal. 5 Clinical data from herbal formulations containing PN demonstrate improved survival in OC patients, correlating with enhanced immune modulation and reduced inflammation. 5 For example, a retrospective study of OC patients receiving PN-based therapies reported prolonged median survival, attributed to p53 pathway activation and NF-κB suppression. 5 Additionally, ginsenoside Rd exhibits blood-brain barrier protection, potentially mitigating neurotoxicity from OC therapies. 6 While preclinical data are promising, rigorous clinical validation is needed. Current trials on PN focus on cardiovascular and metabolic diseases, 7 underscoring the necessity for OC-specific trials.
Traditional Chinese herbal medicines are characterized by multi-target and multi-pathway actions. Network pharmacology, a computational framework integrating systems biology and polypharmacology, facilitates: Comprehensive network analysis of biological systems, Rational selection of druggable nodes within signaling cascades, Mechanistic interpretation of herbal medicine efficacy across multiple therapeutic axes. This integrative methodology systematically resolves the molecular basis of herbal pharmacodynamics. This study employed network pharmacology to predict PN's anti-OC targets and pathways, molecular docking, molecular dynamics to verify component-target binding stability, and in vitro experiments using the human OC cell line A2780 to validate functional effects. This prediction-simulation-validation framework uncovers the targets and mechanisms of PN in treating OC, providing both theoretical and experimental evidence for its clinical application.
Materials and Methods
Potential Targets of PN in Treating OC
The Traditional Chinese Medicine Systems Pharmacology Database (TCMSP, https://www.tcmsp-e.com/#/database) 8 was queried to identify potential targets of PN's main components. Candidate constituents of PN were screened in TCMSP using the ADME-related criteria of oral bioavailability (OB) ≥ 30% and drug-likeness (DL) ≥ 0.18. Accordingly, this database–and threshold–based screening strategy may miss potential active compounds that do not meet the above criteria or are incompletely documented in the database. The target genes corresponding to the main components were batch-processed to retrieve their full names using the Practical Extraction and Report Language (Perl, https://www.perl.org/). Subsequently, the Universal Protein database (UniProt, https://www.uniprot.org/) 9 was employed to batch-convert the full names of the genes into protein species names.
The disease-related genes for Ovarian cancer (OC) were retrieved from the human gene database (GeneCards, https://www.genecards.org/), 10 DisGeNET database (https://disgenet.org/), 11 Therapeutic Target Database (TTD, https://db.idrblab.net/ttd/), 12 Pharmacogenetics and Pharmacogenomics Knowledge Base (PharmGKB, https://www.pharmgkb.org/), 13 and OMIM-Gene Map database (https://omim.org/) 14 using “ovarian cancer” as the search term. The potential targets collected from various databases were subjected to data organization, duplicates across databases were removed after merging the target lists, and all gene symbols were standardized to UniProt official gene names to ensure consistency. All drug target and disease gene names were standardized using the Universal Protein database (UniProt, https://www.uniprot.org/). Genes were considered candidates only if they met both criteria: a minimum score of 1.5 in GeneCards and ≥0.1 in DisGeNET, ensuring stringent primary selection and robust secondary confirmation. Database-specific scoring systems are not directly comparable; therefore, threshold filtering may introduce selection bias. The common targets of PN and OC were screened via jvenn (https://jvenn.toulouse.inrae.fr/app/example.html) 15 and shown in a Venn diagram.
Establishment of PPI Network
The common targets were uploaded imported into the Multiple proteins module of the STRING database (https://cn.string-db.org/cgi/input?sessionId = b6TaSUmWAuv0&input_page_show_search = on). 16 The protein-protein interaction (PPI) network was subsequently constructed with the organism parameter specified as “Homo sapiens”. This two-step process ensures systematic integration of target data and species-specific network generation for subsequent topological analysis. In light of the extensive intersection genes, a stringent interaction score threshold (0.900) was applied to ensure high-confidence protein-protein associations. Meanwhile, nodes without connections in the network were hidden to improve visual clarity and analytical focus. Subsequently, Cytoscape 3.9.1 software 17 was utilized to visualize the imported PPI data. The CytoNCA plugin was used to calculate multiple topological parameters between nodes, including betweenness, degree, eigenvector, closeness, and local average connectivity (LAC). Hub genes were defined as nodes with key topological parameters exceeding the median values, a commonly used heuristic for identifying influential nodes in PPI networks. We acknowledge that alternative hub-selection methods could yield partially different hub sets. Given the large number of common targets, we applied a two-step filtering procedure using the same median-based criteria to reduce network complexity and obtain a parsimonious hub set. We acknowledge that the final hub list is method-dependent.
GO and KEGG Analysis
By leveraging the BiocManager package in the R programming language, the symbol IDs of the intersection genes were translated into gene IDs. Subsequently, KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway and GO (Gene Ontology) enrichment analyses were performed using R language to explore the potential mechanisms underlying the treatment of OC by PN.
For the GO analysis, the parameters of ‘species’, ‘list type’, and ‘identifier’ were set to ‘Homo sapiens’, ‘official gene symbol’, and ‘gene list’, respectively. The GO enrichment analysis comprised three components: CC (cellular component), BP (biological process), and MF (molecular function). Statistical significance was defined as p < 0.05, while q < 0.05 was applied to adjust for potential false discovery rates (FDR) in multiple hypothesis testing.
Molecular Docking of 7 PN Components with Hub Genes
The ligand files in SDF format for the 7 PN components were acquired from the PubChem database (https://pubchem.ncbi.nlm.nih.gOC/). The selected targets (ie hub genes) were associated with the 3D (three-dimensional) structure of HSP90AA1 (PDB ID: 1UY6; resolution: 1.90 Å), 18 TNF (PDB ID:5M2J; resolution: 1.90 Å), 19 TP53 (PDB ID: 3ZME; resolution: 1.35 Å), 20 CCND1 (PDB ID:6P8E; resolution: 2.30 Å), 21 ESR1 (PDB ID: 1SJ0; resolution: 1.90 Å), 22 MAPK1 (PDB ID:2Y9Q; resolution: 1.55 Å), 23 AKT1 (PDB ID: 3CQW; resolution: 2.00 Å). 24 These structures were obtained from the RCSB - PDB database(https://www.rcsb.org/) and the UniProt database (https:// www.uniprot.org/). Molecular docking experiments were carried out by means of Maestro 13.5 software (Schrodinger Suite 2023 version 2). Subsequently, the docking results underwent further analysis and visualization using Discovery Studio 2019 software. To minimize methodological variability, all receptors and ligands were prepared using the same workflow and identical docking settings within Maestro/Glide.
MD Simulation
To determine the dynamic patterns and binding stability of protein-ligand complexes, three molecular systems were subjected to 100-ns molecular dynamics (MD) simulations: HSP90AA1 and Quercetin, TNF and Ginsenoside Rh2, ESR1 and Liquiritigenin (with an additional fourth complex explicitly named if applicable), selected to represent top-ranked docking pairs, different target classes, and those most relevant to subsequent in vitro validation.
78 simulations were executed using GROMACS 2024.5 under the AMBER99SB-ILDN all-atom force field with explicit TIP3P solvent. 25 The thermodynamic ensemble was maintained at physiological temperature (300 K) through Langevin dynamics coupling. An initial energy minimization phase (5000 steps) was implemented via the steepest descent algorithm to resolve steric clashes, achieving convergence at a maximum force threshold of 1000 kJ/mol/nm. Subsequent equilibration protocols were performed using sequential NVT/NPT ensembles. Positional constraints on heavy atoms were gradually relaxed over a 200 ps period, with the restraint release implemented in a stepwise manner. Production runs employed a 2-fs integration time step with LINCS-constrained bond lengths and Particle Mesh Ewald (PME) electrostatics (12-Å real-space cutoff). Trajectories were recorded at every 10 ps for subsequent analysis of backbone RMSD, residue fluctuations, radius of gyration, and hydrogen bond occupancy using standard gmx utilities.
Cell Counting kit-8 (CCK-8) Assay
Methods of CCK-8 Assay Cells in the logarithmic growth phase were harvested, digested with trypsin, centrifuged, and resuspended in the corresponding culture medium. The human ovarian cancer cell line A2780 was used as previously reported (PMID: 30485824). After counting with a hemocytometer under a microscope, cells were diluted to an appropriate density based on their growth characteristics for seeding. Blank wells around the 96-well plate were filled with PBS to minimize errors caused by medium evaporation in peripheral wells, with 6 replicate wells per group. Each well was added with 100 μL of cell suspension and incubated for 24-48 h until cell adhesion. Old medium was aspirated, and media containing different concentrations of Glycyrrhizin, Ginsenoside Rh2, and Olaparib (either alone or in combination) were added. The control group was treated with solvent-containing medium (containing DMSO at the same final concentration as the drug-treated group; final DMSO concentration ≤ 0.1%), followed by incubation for 24 or 48 h. After discarding the culture medium, 100 μL of medium supplemented with 10% CCK-8 was added to each well and incubated at 37 °C with 5% CO₂ for approximately 1 h. The optical density (OD) at 450 nm was measured using a microplate reader, and the effects of the drugs on A2780 cell viability, as well as the corresponding cell survival rate and half-maximal inhibitory concentration (IC50) values, were calculated. The concentration ranges (0-160 μM) and exposure times (24 and 48 h) were selected based on preliminary dose–response screening experiments and published in vitro studies of these compounds in ovarian or other cancer cell models, aiming to cover sub-cytotoxic to cytotoxic levels for IC₅₀ determination rather than to directly reflect clinically achievable plasma concentrations. All compounds were freshly prepared from concentrated stock solutions, filtered when appropriate, and stored according to manufacturer recommendations. Working solutions were prepared immediately prior to each experiment and protected from light. The final DMSO concentration was kept at ≤0.1% to minimize solvent-related instability or cytotoxicity.
Cell Migration and Invasion Assay
Logarithmic-phase cells were divided into six experimental groups, each with three replicates. Vehicle-only controls were included in parallel with the same solvent concentration used for compound preparation. After 24 or 48 h of drug treatment, the cells were harvested by trypsinization, washed twice with PBS, and resuspended at a final density of 1 × 105 cells/mL. For the Transwell migration assay (performed without Matrigel coating), 200 μL of the cell suspension (containing 2 × 104 cells) was introduced into the upper chamber, while the lower chamber was filled with 600 μL of DMEM supplemented with 10% fetal bovine serum (FBS). Following a 48-h incubation period, the migrated cells were fixed with 4% paraformaldehyde for 30 min at room temperature and stained with crystal violet solution in the dark for another 30 min. Non-migrated cells remaining on the upper surface of the membrane were carefully removed using a cotton swab. The migrated cells were then visualized under an inverted light microscope and quantified with ImageJ software. For migration assays, drug concentrations were selected based on CCK-8 results to avoid excessive cytotoxicity (>50% cell death), minimizing confounding effects of reduced proliferation on migration outcomes.
Statistical Analysis
All quantitative data are expressed as the mean ± standard deviation (SD). Comparisons among multiple groups were performed using one-way ANOVA followed by Tukey's post hoc test. A two-sided p-value < 0.05 was considered statistically significant. Statistical analyses were conducted using GraphPad Prism 10.1.2. All experiments were independently repeated at least three times.
Results and Discussion
Targets of PN Components
The structures of PN's main components are presented in Figure 1. The TCMSP database identified 165 targets corresponding to 8 components: ethyl linoleate, DFV (database annotation; full chemical identity provided in Supplementary Table S1 with PubChem CID), Diop (as annotated in TCMSP; detailed chemical name and PubChem CID provided in Supplementary Table S1), β-sitosterol, stigmast, ginsenoside Rh2, quercetin and ginsenoside F2. Following duplicate removal, 165 targets remained (Figure 2).

Structural characterization of PN's primary components.

Depicts interactions among PN's components (orange ovals denote), oc, and targets (while green rectangles).
Potential Targets of PN in Treating OC
Within the GeneCards database, 2649 genes associated with OC were detected, having a score exceeding 1.5 and a clear UniProt ID. The TTD database provided 172 OC - related genes, the PharmGKB database yielded 4, and the DisGeNET database was used to screen 587 genes. These genes not only had scores of ≥ 0.1 but also possessed definite UniProt IDs. 484 OC-related genes were retrieved from the OMIM database, and removing duplicates, 3698 OC-related genes were finally selected. To identify shared molecular targets, we imported the curated drug - target dataset and OC - associated genes into jvenn. Set - based analysis (Figure 3A) yielded 121 common targets, which were visualized via a Venn diagram to delineate the intersection of PN's therapeutic mechanisms and OC pathophysiology.
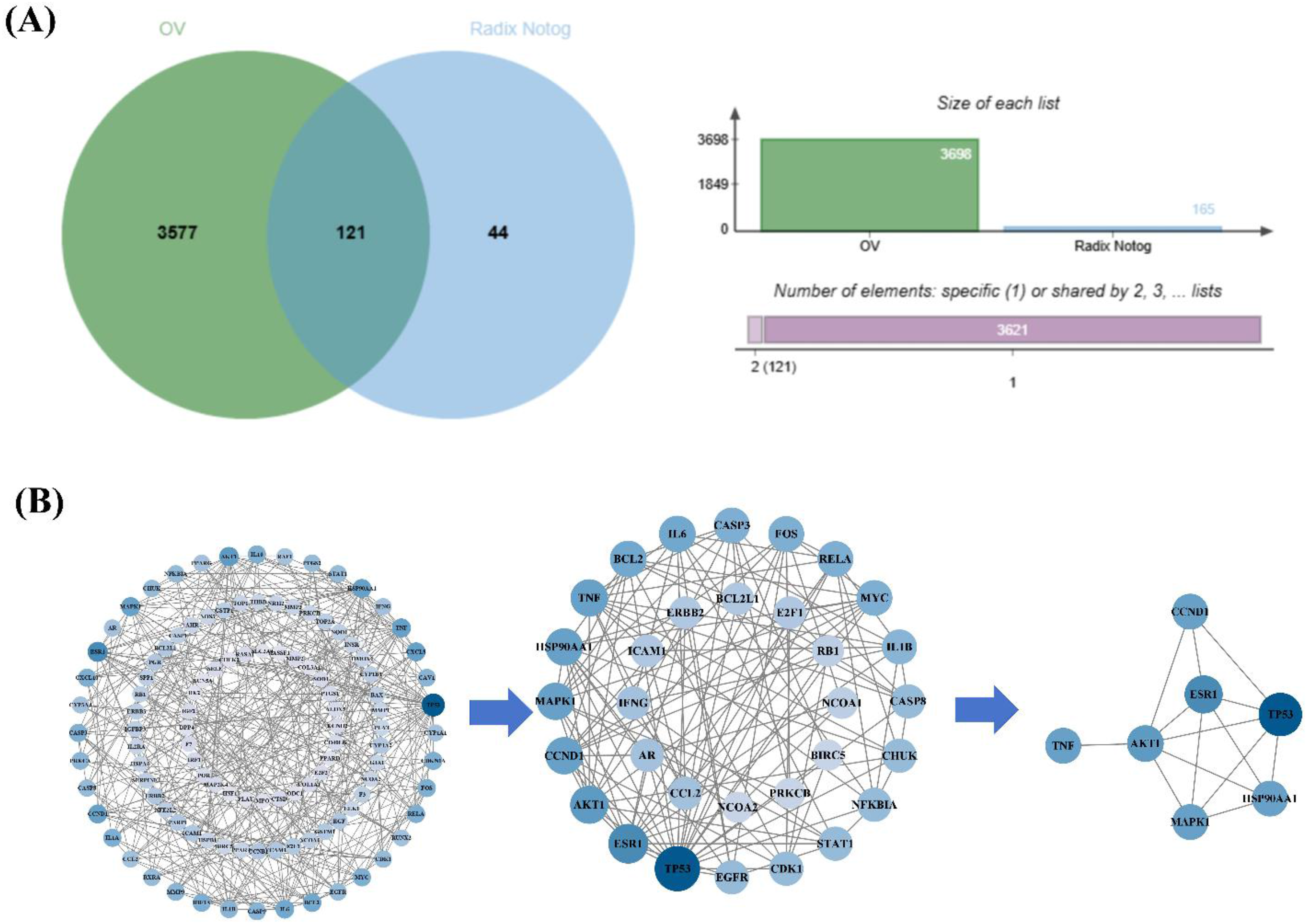
Network pharmacology analysis. (A) To exhibit intersections of PN and OC, a Venn diagram is used; (B) For network analysis of common targets, a PPI network of these genes is presented, with node size denoting the degree value.
PPI Network Analysis of PN in Treating OC
To construct a PPI network visualization (node size proportional to degree value; Figure 3B), we first analyzed 121 common targets in STRING and then imported the resultant interaction data into Cytoscape for graphical rendering. From the first - round screening, the values covering closeness, LAC, degree, betweenness, eigenvector, and network were displayed in Supplemental Table 4. The second round screening's values were shown in Supplemental Table 5. According to the screening criteria for hub genes, after the first round of screening, 112 nodes, 335 edges, and 32 key genes were obtained. By repeating the above operation for the second round screening, 32 nodes, 123 edges, and the 7 core targets were finalized, HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1 and AKT1, which were regulated by 7 active ingredients (Eethyl linoleate, Liquiritigenin, tert-dodecylmercaptan, beta-Sitosterol, Stigmasterol, Ginsenoside Rh2 and Quercetin) of PN (Table 1).
List of Major Protein Abbreviations.

GO and KEGG Analysis of the Core Targets
Both bar and bubble charts in Figure 4 showed the terms from GO enrichment analysis, in which BP, CC, and MF BP, CC, and MF each included 10 enriched terms meeting the significance criteria (FDR-adjusted q < 0.05).. PN treating OC might be associated with the following BP: response to xenobiotic stimulus, response to molecule of bacterial origin, response to lipopolysaccharide, epithelial cell proliferation, response to peptide hormone, regulation of apoptotic signaling pathway, response to oxygen levels, response to decreased oxygen species, response to oxidative stress, and response to reactive oxygen levels. The CC terms focused on membrane microdomain, RNA polymerase II transcription regulator complex, peptidase inhibitor complex, plasma membrane raft, membrane raft, caveola and external side of plasma membrane. Meanwhile, the MF terms with the highest enrichment predominantly encompass DNA−binding transcription factor binding, factor binding transcription coregulator binding, RNA polymerase II−specific DNA−binding transcriptionfactor binding, ubiquitin protein ligase binding, transcription coactivator binding, nuclear receptor activity, protein kinase regulator activity, kinase regulator activity, ubiquitin−like protein ligase binding, and ligand−activated transcription factor activity. Notably, several enriched KEGG terms are labeled as infectious diseases (eg, Hepatitis B, Influenza A). In enrichment analyses, such pathway labels often capture broadly shared host signaling nodes (eg, TNF-, MAPK-, and PI3K–AKT-related modules) and therefore may reflect overlapping stress/inflammatory and survival programs relevant to cancer biology rather than infection-specific mechanisms

GO enrichment analysis.
KEGG pathway analysis yielded 56 terms meeting the significance criteria (adjusted q < 0.05, FDR-corrected). Among these, Figure 5A, using bar and bubble charts, depicted the top 30 pathways ranked by enrichment values. The findings indicated that the mechanisms by which PN treats OC predominantly centered around regulating lipid and atherosclerosis, Hepatitis B, Chemical carcinogenesis−receptor activation, Fluid shear stress and atherosclerosis, Human cytomegalovirus infection, Kaposi sarcoma−associated herpesvirus infection, the AGE−RAGE signaling pathway in diabetic complications, Hepatitis C, Prostate cancer, Influenza A, Proteoglycans in cancer, Hepatocellular carcinoma, and Epstein−Barr virus infection. A Sankey diagram (Figure 6A) was constructed to further clarify the association between core genes and enriched pathways, illustrating that 10 core genes—with ALB excluded—were involved in pathway enrichment. Moreover, the ‘pathview’ tool was employed to visualize the highly enriched OC pathway, highlighting its crucial role in OC (Figure 5B).

KEGG analysis of the core targets. (A) KEGG analysis. (B) Visualization of signaling pathways.

Optimal binding energy of core targets and prominent components in molecular docking. (A) MM-GBSA; (B) XP Gscore.
Molecular Docking of PN Ingredients and OC Targets
Molecular docking simulations were performed to explore predicted interactive between PN bioactive compounds and OC-related targets. This approach assessed predicted binding score for seven PN components (Eethyl linoleate, Liquiritigenin, Diop, beta-Sitosterol, Stigmasterol, Ginsenoside Rh2 and Quercetin) and OC's clinical medicine Olaparib with seven key target genes (HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1, and AKT1).
The docking results indicated that the XP Gscore ratings of the seven PN components and Olaparib with seven targets ranged from −0.153 to −11.774, while the MM - GBSA ratings were within the interval of −6.69 kcal/mol to −96.66 kcal/mol, presenting a relatively large range(Figure 6). This wide range reflects heterogeneity in predicted binding energies across targets and ligands; docking scores in this study are only used for comparative ranking and hypothesis generation, rather than for the absolute quantification of binding affinity.
The binding dynamics between the ligand and receptor, as delineated in Figure 7, encompass multiple interaction modes: classical hydrogen bonds, carbon-hydrogen bonds, halogen bonding, alkyl/π-alkyl hydrophobic forces, and aromatic π-π stacking interactions. Overall, HSP90AA1 tended to yield relatively lower predicted binding energy values than several other targets in this workflow.

Representative docking complex of components and targets. (A) Interaction diagram between HSP90AA1 and Quercetin. (B) Binding mode of TNF with Ginsenoside Rh2. (C) Interaction profile of ESR1 and Eethyl linoleate. (D) Interaction diagram between HSP90AA1 and Olaparib.Hydrogen bonds (yellow line), hydrophobic interactions (pink line) and halogen (blue line).
Figure 7 illustrates the binding mechanisms of key components in Notoginseng (quercetin, ginsenoside Rh2, liquiritigenin) and the reference drug olaparib with three critical targets: HSP90AA1, TNF, and ESR1. For the HSP90AA1, quercetin showed the most favorable predicted docking scores (XP Gscore −11.774 ; MM-GBSA −63.250 kcal/mol) within this set. In the HSP90AA1 binding site, quercetin established hydrogen bonds with ASN51 (1.7 Å) and LEU103 (1.6 Å), while forming hydrophobic interactions with LEU107 (3.5 Å, 2.8 Å, 3.6 Å), PHE138 (3.4 Å), TRP162 (5.8 Å), PHE170 (5.9 Å), and ALA111 (4.1 Å) (Figure 7A).
Regarding TNF, ginsenoside Rh2 showed favorable predicted docking scores (XP Gscore −7.973; MM-GBSA −96.660 kcal/mol) within this set. In the TNF complex, ginsenoside Rh2 formed hydrogen bonds with GLN25 (3.2 Å, 2.6 Å), LEU26 (3.3 Å), ILE136 (3.4 Å), and ASN137 (1.8 Å), alongside hydrophobic interactions with LEU75 (2.5 Å), ARG138 (2.6 Å), and PRO139 (4.0 Å) (Figure 7B). For ESR1, liquiritigenin showed favorable predicted docking scores (XP Gscore −9.669; MM-GBSA −51.590 kcal/mol) within this set. In the ESR1 complex, liquiritigenin formed a hydrogen bond with GLU353 (2.9 Å), and established hydrophobic interactions with ALA350 (2.8 Å), LEU349 (2.5 Å), PHE404 (2.2 Å), LEU346 (2.4 Å), VAL533 (3.0 Å), LEU536 (2.9 Å), and TRP383 (2.9 Å) (Figure 7C).
The reference drug olaparib exhibited XP Gscore of −9.961, −3.364, and −3.381 for HSP90AA1, TNF, and ESR1 respectively, with corresponding MM-GBSA energies of −75.610 kcal/mol, −55.710 kcal/mol, and −34.590 kcal/mol. In the HSP90AA1 complex (Figure 7D), olaparib formed hydrogen bonds with ASN51 (2.8 Å), SER52 (2.7 Å), THR184 (1.8 Å, 2.9 Å), GLY97 (2.5 Å), LEU103 (2.5 Å), and LYS58 (2.4 Å), while establishing a fluorine interaction with ASP93 (3.7 Å) and hydrophobic interactions with PHE138 (3.9 Å, 4.1 Å), MET98 (4.1 Å, 3.2 Å, 4.0 Å), LYS58 (2.4 Å), ARG138 (2.6 Å), and PRO139 (4.0 Å). These results suggest that several Notoginseng components exhibit comparable or lower predicted binding energy values than olaparib in the docking analysis.
Molecular Dynamics Simulation Results Analysis
The backbone RMSD analysis served as a principal metric for evaluating system equilibration, with values below 1 Å indicating stable protein-ligand interactions under physiological conditions. As illustrated in Figure 8A, all three complexes achieved rapid stabilization: HSP90AA1 and Quercetin (0.25 ± 0.05 Å), TNF and Ginsenoside Rh2 (0.23 ± 0.03 Å), and ESR1 and Liquiritigenin (0.15 ± 0.10 Å). Notably, the ESR1 and Liquiritigenin complex exhibited the minimal RMSD fluctuation, suggesting relatively stable conformational behavior under the simulated conditions.

Molecular dynamics simulations results. (A) RMSD values of HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. (B) RMS Fvalues of HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. (C) Radius of gyration (Rg) values of HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. (D) Solvent-accessible surface area (SASA) values of HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. (E) Number of hydrogen bonds in HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. (F) The free energy landscape of HSP90AA1 with Quercetin, TNF with Ginsenoside Rh2, and ESR1 with Liquiritigenin. All MD metrics are reported descriptively as single-trajectory summaries and are not intended as statistically significant comparisons between complexes.
Residue-specific RMSF analysis (Figure 8B) characterized the flexibility and dynamic behavior of protein residues throughout the simulation trajectory. The HSP90AA1 and Quercetin complex displayed the most stable architecture with minimal atomic fluctuations, followed by ESR1 and Liquiritigenin, while TNF and Ginsenoside Rh2 showed relatively higher mobility in specific structural elements.
The Radius of gyration (Rg) analysis (Figure 8C) provided quantitative assessment of structural compactness, with stable values maintained throughout the simulation: HSP90AA1 and Quercetin (2.00 ± 0.02 nm), TNF and Ginsenoside Rh2 (1.70 ± 0.02 nm), and ESR1 and Liquiritigenin (2.00 ± 0.02 nm). The sustained Rg profiles indicated maintained structural integrity, with TNF and Ginsenoside Rh2 exhibiting the most compact tertiary organization during the simulation period.
Solvent-accessible surface area (SASA) analysis (Figure 8D) yielded the following values: HSP90AA1 and Quercetin (152 ± 4.21 nm2), TNF and Ginsenoside Rh2 (113 ± 3.52 nm2), and ESR1 and Liquiritigenin (138 ± 2.10 nm2). The 25% reduction in SASA observed for TNF and Ginsenoside Rh2 compared to HSP90AA1-Quercetin suggested enhanced hydrophobic core formation and more efficient solvent exclusion.
Hydrogen bond analysis (Figure 9E) revealed distinct interaction patterns among the complexes. TNF and Ginsenoside Rh2 demonstrated the most stable hydrogen bonding interactions in terms of both density and persistence, followed by HSP90AA1 and Quercetin, while ESR1 and Liquiritigenin exhibited relatively transient hydrogen bonding characteristics.
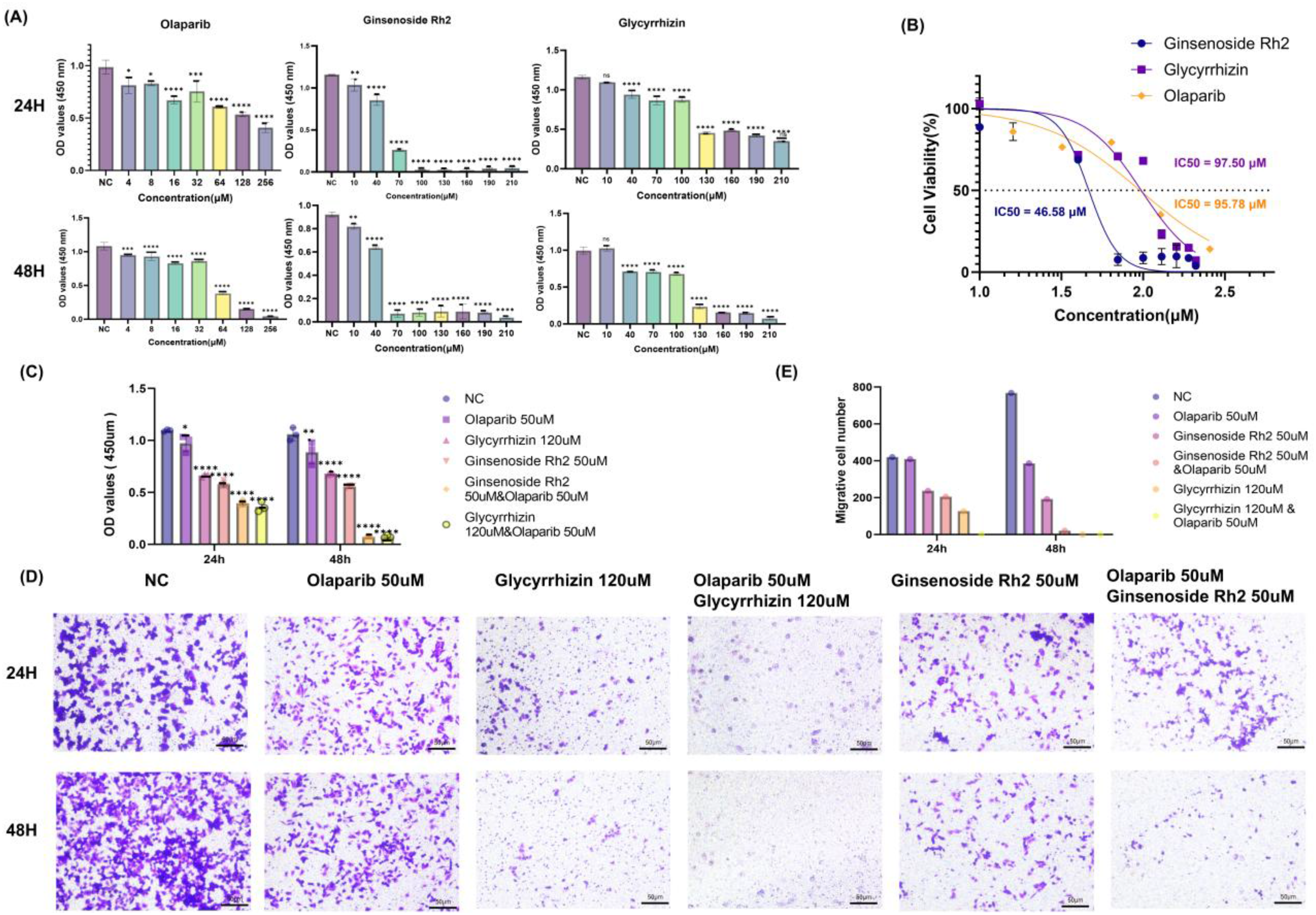
(A) Survival rates of Glycyrrhizin, Ginsenoside Rh2 or Olaparil monotherapy for 24 h and 48 h A2780 cell, compared with control group at the same time. (B) Anti-proliferative activity of Ginsenoside Rh2, Glycyrrhizin, and Olaparib. (C) Cell proliferation assessed by CCK-8 assay after 24 and 48 h of treatment with single agents or their combinations. (D) Cell migration ability after 24 and 48 h of treatment. (E) Statistical plots of the number of migrating cells in A2780 after 24 and 48 h of treatment.
The free energy landscape constructed from principal components of RMSD and Rg (Figure 8F) displayed pronounced energy minima corresponding to the bound conformations of each complex. This observation supports the structural stability under simulation of the protein-ligand complexes in their respective bound states within the simulated timeframe.
CCK8 Results Analysis
To assess the potential anticancer activity of Glycyrrhizin, Ginsenoside Rh2, and Olaparib—both individually and in combination—A2780 cells were treated with a range of concentrations of each drug for 24 h or 48 h. Cell viability was determined by CCK-8 assay, which revealed that all three agents significantly suppressed the proliferation of A2780 cells (Figure 9A). Notably, Olaparib, Ginsenoside Rh2, and Glycyrrhizin each inhibited ovarian cancer cell growth in a time- and concentration-dependent manner relative to the negative control (NC), consistent with the KEGG enrichment analysis indicating involvement of “cell proliferation regulation” pathways. The time-dependent enhancement of inhibitory effects (48 h > 24 h) suggested that the compounds may accumulate in cells to exert sustained regulatory effects on core targets such as AKT1 and MAPK1.
The half-maximal inhibitory concentration (IC₅₀) values at 48 h were calculated as 97.50 μM for Glycyrrhizin, 46.58 μM for Ginsenoside Rh2, and 95.78 μM for Olaparib (Figure 9B).Among the single agents, Ginsenoside Rh2 exhibited the lowest IC₅₀, which aligned with its strong binding affinity to TNF in molecular docking, highlighting a potential structure-activity relationship.
For combination studies, 16 μM Olaparib was selected based on its moderate monotherapeutic inhibition rate (20-30%) in both cell lines. As shown in Figure 9C, the NC group maintained consistently high cell viability across time points, confirming system stability and attributing observed reductions to drug treatments. Compared to single-agent groups, the combination treatments led to further decreases in cell viability, suggesting an enhanced combined antiproliferative effect. Statistical analysis (one-way ANOVA followed by Tukey's post-hoc test) confirmed that the cell viability of the Glycyrrhizin (120 μM) + Olaparib (50 μM) group was significantly lower than that of the Glycyrrhizin or Olaparib monotherapy groups (p < 0.01 at both 24 h and 48 h). In particular, the combination of Glycyrrhizin (120 μM) with Olaparib (50 μM) exhibited the strongest suppression of cell viability at both 24 h and 48 h, outperforming not only all monotherapies but also the Ginsenoside Rh2 (50 μM) plus Olaparib (50 μM) combination. The enhanced combined effect may involve coordinated modulation of overlapping pathways (eg, PI3 K/AKT and MAPK) by the two drugs, as revealed by KEGG enrichment analysis.
Cell Migration and Invasion Assay
To evaluate the effect of drug treatments on the metastatic potential of ovarian cancer cells, we performed Transwell migration assays using A2780 cells. As illustrated in Figure 9D (microscopic images) and quantified in Figure 9E, all single-agent and combination treatments significantly suppressed cell migration compared to the NC group after both 24 h and 48 h of exposure. Notably, the inhibitory effect on migration became more pronounced over time, indicating a time-dependent response. Among the treatments, the combination of Glycyrrhizin (120 μM) with Olaparib (50 μM) exhibited the strongest anti-migratory effect, resulting in the fewest migrated cells at both time points. This combination also appeared to be more effective than Ginsenoside Rh2 (50 μM) combined with Olaparib (50 μM), suggesting a stronger combined inhibitory effect between Glycyrrhizin and Olaparib in impairing cell motility under the tested conditions. Given that ESR1 is involved in regulating epithelial-mesenchymal transition (EMT) in ovarian cancer, the strong binding of Liquiritigenin to ESR1 may contribute to the anti-migratory effect, which is further enhanced by Olaparib's regulation of DNA damage response pathways.
Discussion
Ovarian cancer, although not the most prevalent malignant tumor of the female reproductive tract, remains one of the most common malignancies in women. Characterized by high malignancy, invasiveness, and recurrence rates, it exhibits the highest mortality among gynecological cancers. This poor prognosis is largely attributable to its insidious onset and the frequent absence of characteristic early symptoms. While numerous network pharmacology studies have investigated ovarian cancer, the precise mechanisms of action underlying many clinically effective agents remain incompletely elucidated. This study employed an integrated network pharmacology approach—leveraging its “multi-pathway, multi-target, multi-component” paradigm—combined with computational simulations and in vitro cellular experiments. This strategy was utilized to identify the principal molecular targets and elucidate the potential mechanisms by which Panax notoginseng exerts its anti-ovarian cancer effects.
The PN-target-OC network indicated that HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1, and AKT1 had high degree values. The therapeutic efficacy of PN's key components against OC was subjected to further analysis, given the findings above. It has been reported that Eethyl linoleate, Liquiritigenin, Diop, beta-Sitosterol, Stigmasterol, Ginsenoside Rh2, and Quercetin possess various biological and pharmacological activities. Ginsenoside Rh2 in Panax notoginseng can promote the transformation of cancer cells into non-cancer cells and has reported anti-tumor activity in the treatment of ovarian cancer. Quercetin and Liquiritigenin have been reported to exhibit antioxidant, anti-inflammatory, and anti-tumor effects in other experimental systems; however, these biological effects were not directly evaluated in the present study and therefore should not be interpreted as demonstrated mechanisms here
PPI network analysis revealed that seven potential key targets (HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1, and AKT1) were significantly associated with the treatment of ovarian cancer by Panax notoginseng (PN). These genes may play a role in the therapeutic mechanism of PN in treating ovarian cancer. Mechanistic studies have confirmed that HSP90AA1 upregulation drives metastatic dissemination in ovarian cancer cell models, implicating this chaperone protein in the invasive cascade of high - grade serous subtypes and validating its therapeutic relevance. Mna induces T cells and promotes the secretion of TNF - α to regulate the immune response. In HGSOC, TP53 gene mutations interact with adipocyte-derived IL-8, leading to joint upregulation of FABP4 expression. This mechanism further enhances fatty acid uptake and absorption, thereby facilitating cancer cell growth and metastasis within the omental microenvironment. The transition from G1 to S phase is mediated by Cyclin D1, which is encoded by the CCND1 gene. After the formation of the IgH/CCND1 fusion gene, the overexpression of Cyclin D1 leads to cell cycle disorders and uncontrolled cell growth. ESR1 gene mutations are rare in primary tumors but can have a mutation rate as high as 60% in patients with tumor resistance. The PRDX6 - induced upregulation of NNMT triggers the activation of the MAPK signaling cascade, consequently driving ovarian cancer cell proliferation and metastasis. AKT is usually abnormally activated in cancer, affecting multiple processes such as survival, metabolism, angiogenesis, and cell proliferation. Therefore, HSP90AA1, TNF, TP53, CCND1, ESR1, MAPK1, and AKT1 are widely implicated in OC progression, suggesting that PN's therapeutic effects may be exerted via interactions with these core target proteins, however, network pharmacology predictions may contain false positives and thus require further targeted experimental validation.
Central to PN's anti-OC effect were the pathways uncovered by GO and KEGG enrichment analyses. GO analysis showed that Panax notoginseng's key targets primarily participated in response to xenobiotic stimulus, localized to membrane raft and membrane microdomain, and functioned as DNA-binding transcription factor binding. The KEGG pathway analysis revealed that the anti-OC mechanisms of PN predominantly encompassed the modulation of lipid metabolism and atherosclerosis, chemical carcinogenesis-receptor activation pathways and Hepatitis B. KEGG pathway analysis revealed that catechins actively regulate multiple signaling axes critical for hypertension treatment, such as those governing vascular smooth muscle contraction, aldosterone - induced sodium reabsorption, and renal cell carcinoma progression. The experimental data from CCK-8 and Transwell assays are consistent with the possibility that the PN-derived components and Olaparib combinations may influence PI3 K/AKT and MAPK-related signaling, although direct molecular validation was not performed in this study. Accordingly, we focus our biological interpretation on the shared signaling components within these KEGG pathways rather than the disease labels themselves. This may contribute to the reduced cell proliferation and migration—two key hallmarks of OC progression.
Furthermore, molecular docking was used to assess binding affinities, whereas molecular dynamics simulations were conducted to explore the conformational changes and binding stability of PN - OC target complexes. Molecular docking results indicated that when 8 components of Panax notoginseng and Olaparib (OC's clinical medicine) were docked with 7 targets, and their MM - GBSA and XP Gscore were analyzed, the optimal Panax notoginseng - ligand complexes were HSP90AA1 and Quercetin, TNF and Ginsenoside Rh2, ESR1 and Eethyl linoleate, and HSP90AA1 and Olaparib, which were mainly governed by hydrogen bonding and hydrophobic forces. To further examine the kinetic characteristics and temporal stability of complexes formed between the primary components of Panax notoginseng and core targets, MD simulations were performed. The structural stability and compactness of the three complexes were corroborated by their RMSD, Rg, SASA values and hydrogen bond counts, suggesting reliable molecular integrity. In addition, RMSF accurately located the dynamic functional sites of proteins, revealing the relationship between conformational changes and biological functions. Through the dynamic analysis of the number, lifespan, and spatial distribution of hydrogen bonds, the mechanism of action of functional residues was revealed. The functional conformational states of the complexes were accurately identified through the free energy landscape, revealing the molecular mechanism of dynamic regulation. The MD simulations supported the stability favorable predicted binding of HSP90AA1 and Quercetin, TNF and Ginsenoside Rh2, and ESR1 and Liquiritigenin, consistent with the trends observed in molecular docking results.
Network pharmacology was applied to screen for OC-relevant targets and pathways modulated by eight PN constituents. Although computational simulations were carried out to explore the binding mechanisms of these PN components and OC core targets, this study acknowledges several limitations. Future studies are warranted to validate the molecular mechanisms underlying he anti-OC effects of PN using a wider range of ovarian cancer models that reflect disease heterogeneity, drug-resistant cell lines, and animal models, with a focus on bridging preclinical findings to translational therapeutic strategies. It should be noted that these computational simulation results only provide predictive insights into potential molecular interactions, and do not represent direct experimental evidence for the pharmacological superiority of PN components over clinical drugs. Additionally, in vivo xenograft models could verify the efficacy and safety of PN-derived components and their combinations, providing a foundation for clinical translation. Notably, some concentrations used in the in vitro assays may exceed physiologically achievable plasma levels; this dose selection was intended to ensure detectable biological effects within the constraints of short-term cell culture experiments and to explore potential mechanistic interactions at the cellular level. Accordingly, the observed antiproliferative and anti-migratory effects should be interpreted as proof-of-concept findings rather than direct reflections of clinical efficacy, and future investigations incorporating pharmacokinetic evaluation, bioavailability assessment, and in vivo validation are necessary to determine the translational relevance of these results. This approach will facilitate the development of safer, more precise targeted interventions for OC patients. Herbal compounds may exhibit different pharmacokinetic behaviors when administered as part of complex formulations compared to isolated monomers.
Conclusions
This study presents an integrative framework integrating network pharmacology, computational simulations, and in vitro cellular experiments to systematically delineate the therapeutic targets and molecular mechanisms underlying Panax notoginseng (PN)-mediated ovarian cancer (OC) suppression. Our analysis pinpointed Ginsenoside Rh2 and Liquiritigenin as the key bioactive constituents of PN showing inhibitory effects on OC cell proliferation and migration in vitro, while HSP90AA1, TNF, and ESR1 emerged as critical linchpins for targeted OC intervention. In silico investigations—encompassing molecular docking and molecular dynamics (MD) simulations—suggested that PN's bioactive compounds may engage with core OC targets via hydrogen bonding and hydrophobic interactions, under simulated conditions. Subsequent in vitro validation using the human high-grade serous OC cell line A2780 sugggested that these predicted molecular interactions may be associated observed cellular effects: PN-derived components, either as monotherapies or in combination with the clinical PARP inhibitor Olaparib, significantly suppress A2780 cell proliferation and migratory capacity. Collectively, these molecular-level insights not only delineate the mechanistic basis for PN's anti-OC efficacy but also provide candidate targets for further investigation in the development of potential OC treatment strategies—bridging computational predictions with experimental validation to advance translational research on PN in OC therapy.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X261435314 - Supplemental material for Unraveling the Multi-Target Anti-Ovarian Cancer Mechanisms of Panax Notoginseng: Computational and Experimental Insights
Supplemental material, sj-docx-1-npx-10.1177_1934578X261435314 for Unraveling the Multi-Target Anti-Ovarian Cancer Mechanisms of Panax Notoginseng: Computational and Experimental Insights by Danli Yuan, Liping Chen, Die Xia, Jiapan Li, Shiyu Zhang, Jinquan Zhang and Ziyan Meng in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X261435314 - Supplemental material for Unraveling the Multi-Target Anti-Ovarian Cancer Mechanisms of Panax Notoginseng: Computational and Experimental Insights
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X261435314 for Unraveling the Multi-Target Anti-Ovarian Cancer Mechanisms of Panax Notoginseng: Computational and Experimental Insights by Danli Yuan, Liping Chen, Die Xia, Jiapan Li, Shiyu Zhang, Jinquan Zhang and Ziyan Meng in Natural Product Communications
Footnotes
Ethical Approval
Ethical approval was not required for this study because it did not involve human participants, human tissue, or animal experiments. All experiments were performed using established commercial cell lines.
Author Contributions
D. Y. and L.C. contributed equally to this work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Youth Fund Project of Natural Science Foundation of Jiangxi Province (20242BAB21036), (20232BAB216135) and (20252BAC200103).
Youth Fund Project of Natural Science Foundation of Jiangxi Province, (grant number 20232BAB216135, 20242BAB21036, 20252BAC200103).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supporting Information
The Supporting Information is available free of charge on the ACS Publications website at DOI:
Data are included within the Supporting Information (PDF)
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
