Abstract
Background
Coptis chinensis Franch. (CCF) is a Traditional Chinese medicine known for its good anti-cancer effects. However, the bioactive compounds and mechanisms underlying its anti-colorectal adenocarcinoma (COAD) remains unclear.
Objectives
This study aims to reveal the bioactive compounds and mechanism of CCF against COAD by utilizing bioinformatics, molecular docking simulation and in vitro experiments.
Materials and Methods
Bioactive compounds and candidate targets of CCF against COAD were collected using the Traditional Chinese Medicine Systems Pharmacology Database, Analysis Platform (TCMSP), Gene Expression Omnibus (GEO), Gene Cards, and SwissTargetPrediction databases, respectively. The Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using the DAVID (v2022q3) database. The “Compounds-Targets-Pathways” (C-T-P) and “Protein-Protein Interaction” (PPI) networks were constructed using Cytoscape (v3.8.0) to identify the hub targets. AutoDock Vina (v1.2.0), UNLCAN, TCGA, MTT, plate cloning, and quantitative real-time PCR (qPCR) assays were used to confirm the results.
Results
The “C-T-P” and “PPI” networks revealed three hub targets: checkpoint kinase 1 (CHEK1), polo like kinase 1 (PLK1), and aurora kinase B (AURKB). Molecular docking simulation results showed that berberine, a candidate bioactive compound of CCF, had high affinity with the hub targets, comparable to that of positive drugs. Both CCF ethanol extracts and berberine significantly down-regulated CHEK1, PLK1, and AURKB, inhibited proliferation and promoted apoptosis of HCT-116 cells.
Conclusion
To summarize, CCF inhibits COAD by down-regulating CHEK1, PLK1, and AURKB, with berberine as its primary bioactive compound.
Introduction
Colorectal adenocarcinoma (COAD) is a prevalent and lethal form of digestive tract cancer with high incidence and mortality rates (Dang et al., 2020). Annually, millions of people are diagnosed with COAD, and many die from the disease (Qu et al., 2020). Although treatments like surgery, radiotherapy, chemotherapy, and medication have improved cure rates for early-stage COAD patients, the five-year survival rate remains low for those with locally advanced COAD. Endoscopy and blood screening are crucial in optimizing treatment and reducing COAD mortality (Koster et al., 2020). Unfortunately, most countries currently lack the capacity to conduct large-scale inspections. To tackle this global health issue, it is essential to develop highly effective, low toxicity, and affordable drugs.
In recent years, Traditional Chinese Medicine (TCM) has achieved remarkable curative effects in cancers prevention and treatment, attracting extensive attention worldwide (Wang et al., 2021). However, a gap between modern medicines and TCM is the lack of the bioactive compounds and mechanism of the latter, which has led to lower acceptance of TCM. Nonetheless, this has not affected their wide application in various fields. For instance, Coptis Chinensis Franch. (CCF), first recorded in Shennong’s Classic of Materia Medica, has been used to clear heat, purge fire, and detoxify for more than 3000 years (Zhang et al., 2021). Clinical epidemiology and modern pharmacology reveal that CCF is a highly promising anti-cancer herbal medicine, which shows an impressive inhibition effect on a variety of cancer cells. Gu, Song, et al. (2020) proved that berberine, an essential bioactive compound extracted from CCF, had inhibitory effects on cancer cells. However, the underlying bioactive compounds and mechanisms of CCF against COAD are not yet fully understood.
The emergence of bioinformatics has provided significant support for the in-depth study of TCM (Zhang et al., 2019), with the use of multiple algorithms to comprehensively analyze the internal relationship between drugs, diseases, and targets, which is very consistent with the overall concept of TCM in the treatment of diseases. Identifying the differentially expressed genes (DEGs) between case and normal samples is essential in this process (Guo et al., 2017), and public databases with large clinical samples provide convenience for researchers. Molecular docking simulation (MDS), a tool widely used in drug development, is effective in analyzing the affinity between drugs and targets, providing a very valuable reference for drug development (Pinzi & Rastelli, 2019). MDS has also achieved many substantive achievements (Khelfaoui et al., 2021). A plethora of studies have already proved that it is feasible to study TCM via these bioinformatics tools and in vivo and in vitro experiments synthetically (Gu, Xue, et al., 2020; Song et al., 2020).
This study utilized bioinformatics, public databases and in silico to gain a deeper understanding of the potential bioactive compounds and hub targets of CCF against COAD. After analyzing the function of hub targets and the pathways they participated in, in vitro experiments confirmed the above results. A graphical summary highlights the workflow of this study and is detailed in Figure 1.
The Workflow of this Study.
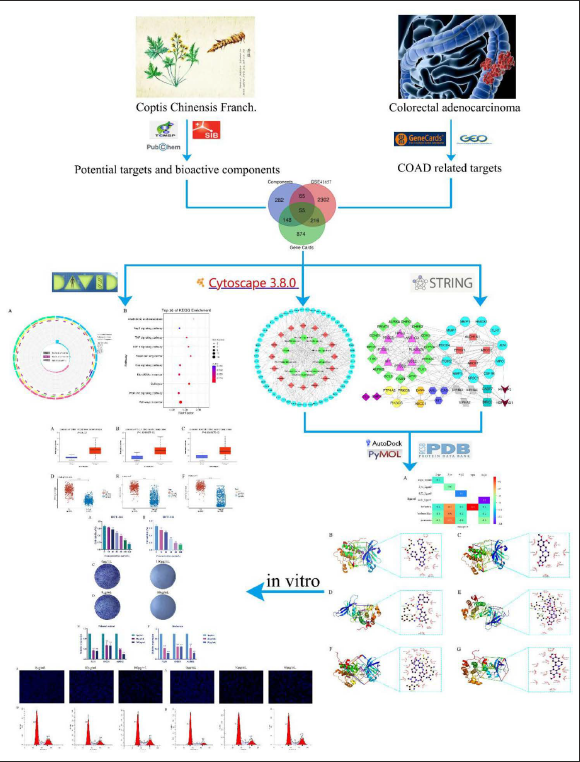
Materials and Methods
Bioactive Compounds Collection
To screen potential anti-COAD bioactive compounds in CCF, a crucial step is to collect as many compounds as possible and analyze their oral bioavailability (OB) and drug-likeness (DL). OB is a crucial standard to evaluate the quality of drugs, referring to the relative amount and rate of drugs absorbed by the body into circulation (Guo et al., 2019). DL is derived from the structures and properties of existing drugs and candidate drugs, including chemical stability, solubility, permeability, and metabolism (Wei et al., 2020). Both of them are commonly used to screen potential compounds in the early stage of drug development.
Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP,
Condidate Targets of CCF Against COAD
Swiss Target Prediction (
A wealth of clinical data can provide effective data support for researchers to mine the core targets of diseases. Gene Expression Omnibus (GEO,
Enrichment Analysis
The Database for Annotation, Visualization and Integrated Discovery (DAVID,
Acquisition of Hub Targets
Comprehensive analysis of the “Compounds-Targets-Pathways” (C-T-P) and “Protein-Protein Interaction” (PPI) networks is crucial for screening the potential bioactive compounds and core targets of CCF against COAD. A “C-T-P” network was constructed using Cytoscape (v3.8.0,
STRING (
Molecular Docking Simulation
MDS is a widely used computational method for screening drugs based on their affinity to proteins. AutoDock Vina, a molecular docking software based on semi-flexibility docking, is one of the best software for this purpose (Eberhardt et al., 2021). To ensure the accuracy of the results, we compared the affinity between positive drugs and proteins, bioactive compounds, and proteins.
The PDB formats of the five hub targets’ structureswere downloaded by RSCBPDB (
UNLCAN and TCGA Analysis
To further validate our experiments and establish a connection between research and the clinic, the public clinical data from UALCAN and TCGA were used. UALCAN (ualcan.path.uab.edu/home) is a free database that provides publicly available cancer omics data and allows users to identify biomarkers or to perform in silico validation of potential genes of interest (Chandrashekar et al., 2017). The Cancer Genome Atlas (TCGA,
Acquisition of CCF Ethanol Extracts and Berberine
An amount of 10 g CCF was purchased from Bozhou City (Anhui, China), crushed and passed through a sieve, and extracted twice using 75% ethanol for 30 min each time via ultrasonication. The ethanol solution of CCF was recovered using a rotary evaporator, and its dry paste yield was calculated. Berberine was purchased from Chengdu Pusi Biotechnology Co., Ltd., (Sichuan, China).
MTT Assay
HCT-116 cells in logarithmic growth phase were incubated in two 96-well plates at a density of 7.0×103 cells/well for 24 h until all cells were adherent. After that, they were treated with varying concentrations of CCF ethanol extracts (0, 10, 20, 40, 80, 160, and 320 µg/mL) and berberine (0, 10, 20, 40, 80, and 160 µg/mL) for 24 h, respectively. Subsequently, 10 µL MTT (Sigma, USA) was added to each well of both plates. After 4 h, the culture medium in each well was sucked away, and the absorbance of wells was read at 490 nm using a Spark 10 M microplate reader (Tecan, Mannedorf, Switzerland).
Plate Cloning Assay
HCT-116 cells were inoculated into two 6-well plates at 2.0×103 cells/well for 24 h until all cells were adherent. After culturing cells with different concentrations of CCF ethanol extracts (0 and 160 µg/mL) and berberine (0 and 80 µg/mL) for 24 h, photos were taken under fluorescence microscope (Olympus Corporation, Tokyo, Japan). Then, the culture medium in each well was replaced with normal culture medium and cultured for 14 days. The cells in 6-well plates were fixed with 4% paraformaldehyde, and reacted for 20 min at room temperature in the dark. The colony formation in each well was observed under a microscope and photographed.
Cell Cycle Detection by Flow Cytometry
After treating the cells with CCF ethanol extracts (0, 80, and 160 µg/mL) and berberine (0, 40, and 80 µg/mL) for 24 h, the cells were digested, and the cell suspension was collected. The cell suspension was centrifuged at 1000 rpm for 5 min, and the supernatant was discarded. After resuspension twice in PBS, cells were fixed overnight in 70% ethanol. PI dye was added to the cell samples, and they were incubated for 30 min at room temperature in the dark. The cell cycle of the samples was detected using a flow cytometer (BD, FACSCalibur, US).
Hoechst 33342 Staining Assay
Logarithmic growth phase HCT-116 cells were inoculated into two 6-well plates at 5.0×105 cells/well. After cells adhered to the walls, they were incubated for 24 h in different concentrations of CCF ethanol extracts (0, 80, and 160 µg/mL) and berberine (0, 40, and 80 µg/mL). The cells were fixed with 4% polyoxymethylene, washed twice with PBS, and incubated with 8 µg/mL Hoechst 33342 (Beyotime Biotechnology, Shanghai, China) for 10 min at 4℃. After washing the cells thrice with PBS, they were observed using a fluorescence microscope (Olympus, Nanjing, China).
Quantitative Real-time PCR Assay
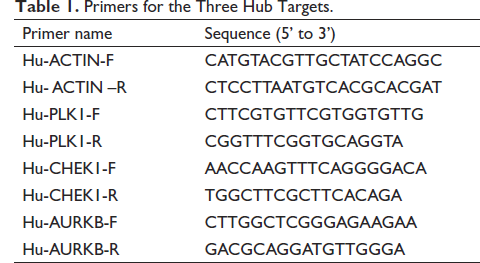
A qPCR was conducted on HCT-116 cells in the logarithmic growth phase, which were seeded in 6-well plates with 5.0×105 cells/well and treated with CCF ethanol extracts (0, 80, and 160 µg/mL) and berberine (0, 40, and 80 µg/mL) for 24 h. The primers for the three hub targets were designed and synthesized by Siwega Biotechnology Co., Ltd., (Wuhan, China), as shown in Table 1. The reverse-transcribed samples were detected on Bio-Rad CFX Manager software (Bio-Rad, CFX Connect, US) using a qPCR machine (Bio-Rad, CFX Connect, US).
Primers for the Three Hub Targets.

Statistical Analysis
Statistical significance was assessed using a two-tail, equal variance independent-samples t-test. GraphPad Prism 7.0 (GraphPad Software Inc., San Diego, CA) was used to perform data analysis and p < 0.05 was set as a significant threshold.
Results
Acquisition of Bioactive Compounds
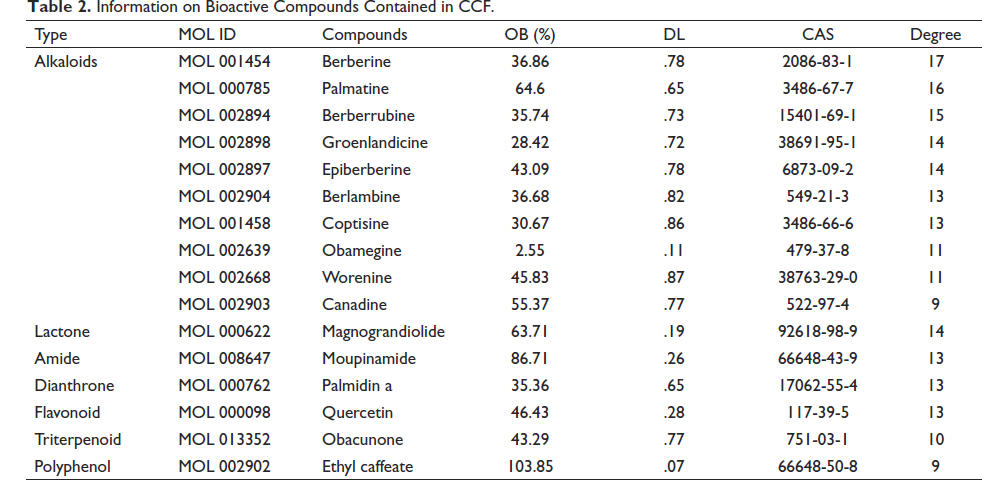
Based on OB ≥ 30%, DL ≥ 0.18 and literatures, a total of 16 potential bioactive compounds of CCF against COAD were collected. These compounds contained 10 alkaloids, 1 dianthrone, 1 flavonoid, 1 triterpenoid, 1 amide, 1 lactone, and 1 polyphenol. Numerous studies have reported that many alkaloids anti-inflammatory and anti-tumor bioactive properties, which supports the objectives of this study. Further information on the 16 compounds is shown in Table 2.
Information on Bioactive Compounds Contained in CCF.
Acquisition of Candidate Targets
The GES41657 dataset (12 normal and 76 COAD samples) in the GEO database was analyzed for DEGs, and the results were displayed as heat and volcano maps. A total of 2639 DEGs were obtained, including 1565 up-regulated genes and 1,074 down-regulated genes (Figure 2A and B). Additionally, 1,293 genes from Gene cards were collected. By comprehensively analyzing the targets of 16 compounds obtained from the SwissTargetPrediction database, and removing the redundancy between them, 556 targets were obtained. Finally, 55 candidate targets of CCF against COAD were obtained using the Venn 2.1.0 database (Figure 2C). The symbols of the 55 targets are listed in Table 3.
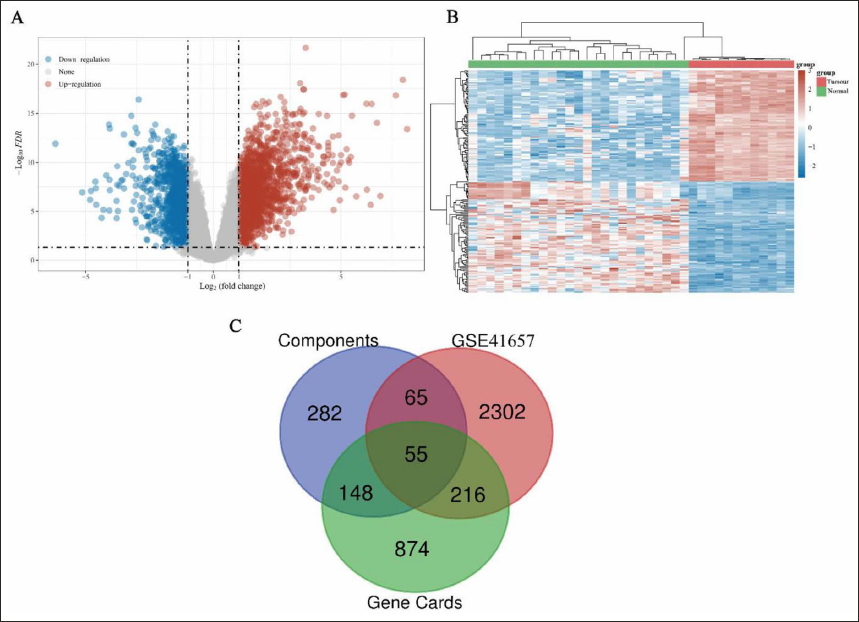
Candidate Targets of CCF Against COAD.
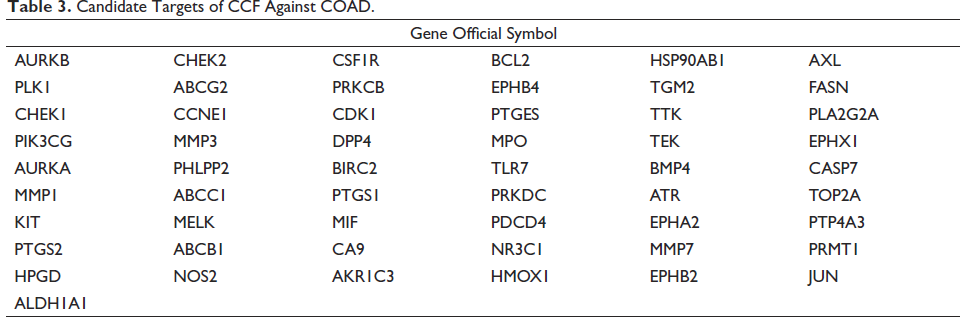
Functional Enrichment Analysis
Using the DAVID database, 18 KEGG pathways (52, 94.5%), 139 GO BP (55, 100%) entries, 28 GO CC (55, 100%) entries, and 28 GO MF (55, 100%) entries were obtained. Among them, 18 KEGG pathways, 98 GO BP entries, 21 GO CC entries, and 24 GO MF entries were significant (p < 0.05). The visualization results of the GO entries showed that the above candidate targets were widely distributed from the cell surface to the nuclear membrane (Figure 3A). And they regulated the activities of protein kinase, tyrosine kinase, and transmembrane receptor protein. Therefore, these targets affected COAD cell growth through multiple BP entries involved in cell apoptosis, proliferation, cell cycle, inflammation, and drug responses. These results were also confirmed in the KEGG pathways enrichment (Figure 3B). The top ten pathways, such as pathways in cancer, PI3K-Akt signaling pathway, Cell cycle, Ras signaling pathway, and other pathways had been widely reported to be significantly associated with the refractory tumor.
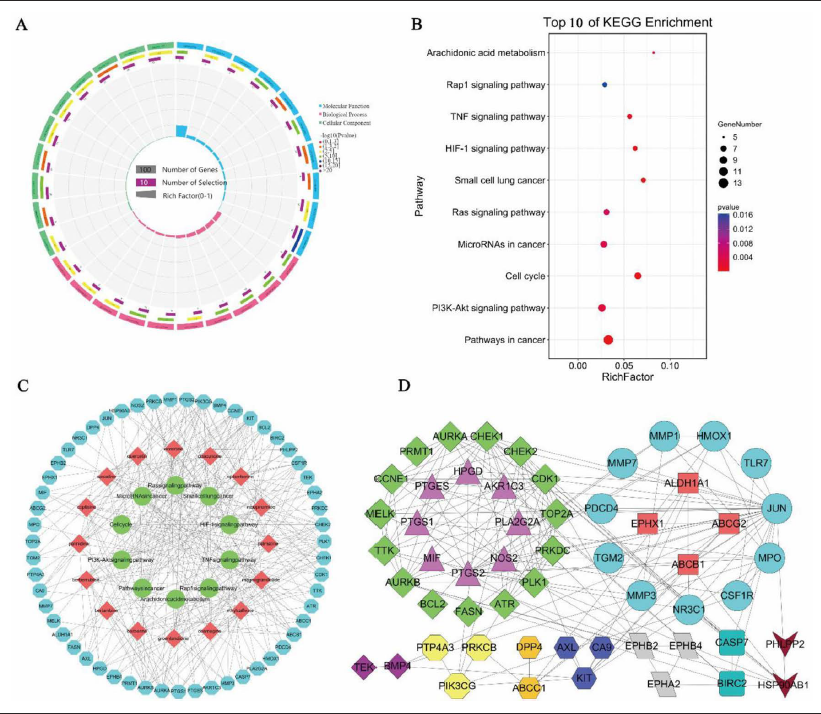
Construction and Analysis of “C-T-P” and “PPI” Networks
A “C-T-P” network consisting of 81 nodes and 280 edges was constructed using Cytoscape. The red nodes in the network represented bioactive compounds, greed nodes represented KEGG pathways, while blue nodes represented candidate targets (Figure 3C). Using the STRING and Cytoscape, a “PPI” network with 55 nodes and 174 edges was obtained. The module analysis of the 55 targets was carried out using the MCODE enhancement package of Cytoscape, and a total of 11 modules were obtained (Figure 3D). With comprehensive comparison and analysis of the degrees of nodes in both networks, we identified the top 5 targets based on the median principle (median = 11) as prostaglandin-endoperoxide synthase 2 (PTGS2, degree = 28), jun proto-oncogene (JUN, degree = 27), checkpoint kinase 1 (CHEK1, degree = 26), polo like kinase 1, (PLK1, degree = 26), and aurora kinase B (AURKB, degree = 25).
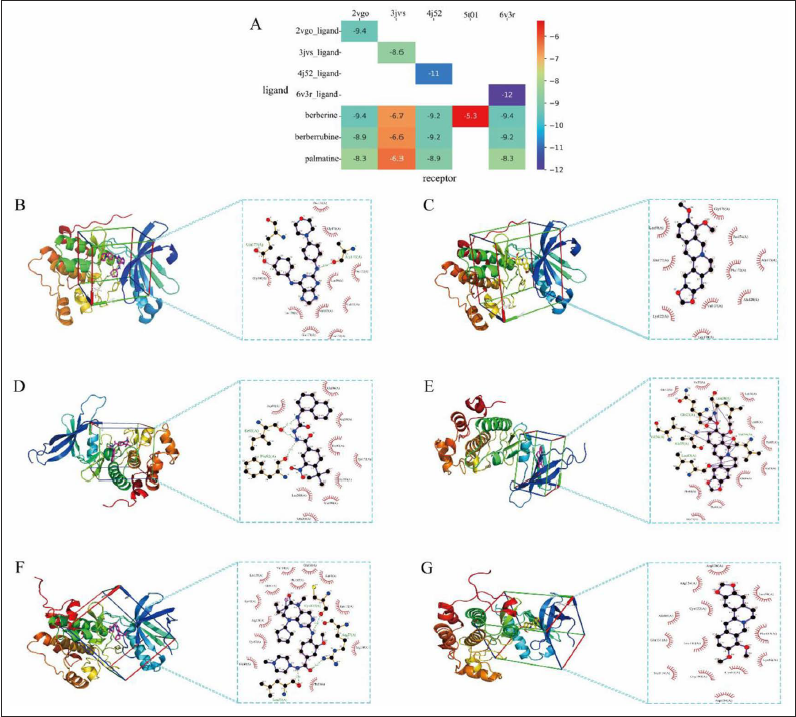
Molecular Docking Simulation
The docking affinity heatmap (Figure 4A) showed that berberine had the best potential bioactivity. The particularity of the structure of JUN (5t01) results in that it did not have a good binding pocket on its surface, so several bioactive compounds could not form a stable binding model with it. To consolidate the accuracy of the docking results, AURKB (2vgo), CHK1 (3jvs) and PLK1 (4j52) were considered as potential targets since berberine and positive drugs had similar affinities to these targets (affinity numerical difference was less than 2). The docking pockets and 2D modes of compounds with targets are shown in Figure 4. Undeniably, further validation of the expression of these three targets in COAD and normal colon tissues using clinical data was necessary to confirm their importance.
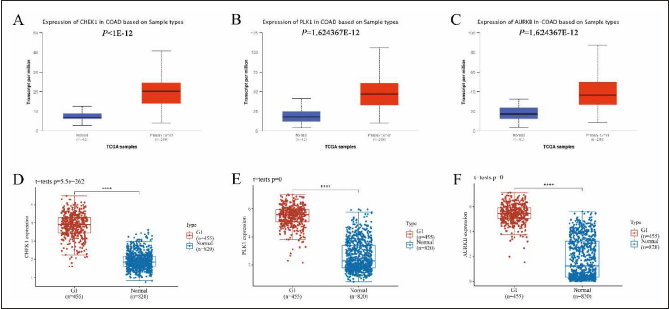
Clinical Dataset Validation
Analysis of 41 normal and 286 patient samples recorded in the UNLCAN database found that the expressions of CHK1 (p < 1×E-12), PLK1 (p = 1.62×E-12), and AURKB (p = 1.62×E-12) were significantly higher in tumor samples than in normal samples (Figure 5A–C). Simultaneous analysis of 455 normal and 820 patient samples found that the expression of CHK1 (p =5.5×E-262), PLK1 (p = 0), and AURKB (p = 0) in tumor was also significantly higher than that in normal samples (Figure 5D–F). These results were consistent with our previous findings. Multiple studies had shown that the high expression of CHK1, PLK1, and AURKB caused changes in the cycle of HCT-116 cells and promoted their proliferation.
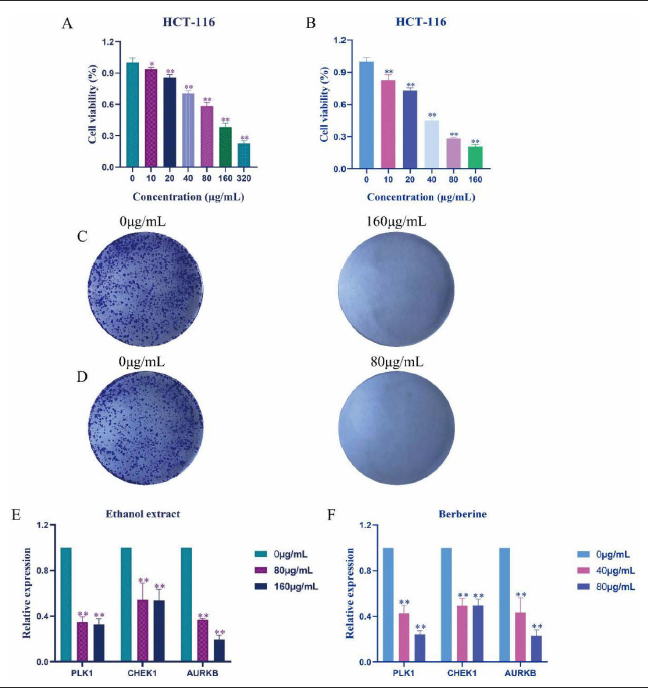
CCF Ethanol Extracts and Berberine Decreased the Proliferation of HCT-116 Cells
In vitro experiments revealed a significant decrease in cell viability of the treatment groups compared to the control group (*p < 0.05 or ** p < 0.01) as the concentrations of CCF ethanol extracts and berberine increased (Figure 6A and B). Notably, the inhibitory effect of berberine on HCT-116 cells was better than that of CCF ethanol extracts, with their half-maximal inhibitory concentrations (IC50) for HCT-116 cells at 33.61 µg/mL and 130.5 µg/mL. The results of plate cloning showed that HCT-116 cells treated with CCF ethanol extracts and berberine did inhibit the proliferation of HCT-116 cells in a concentration-dependent manner (Figure 6C and D).
CCF Ethanol Extracts and Berberine Blocked the Cell Cycle of HCT-116 Cells
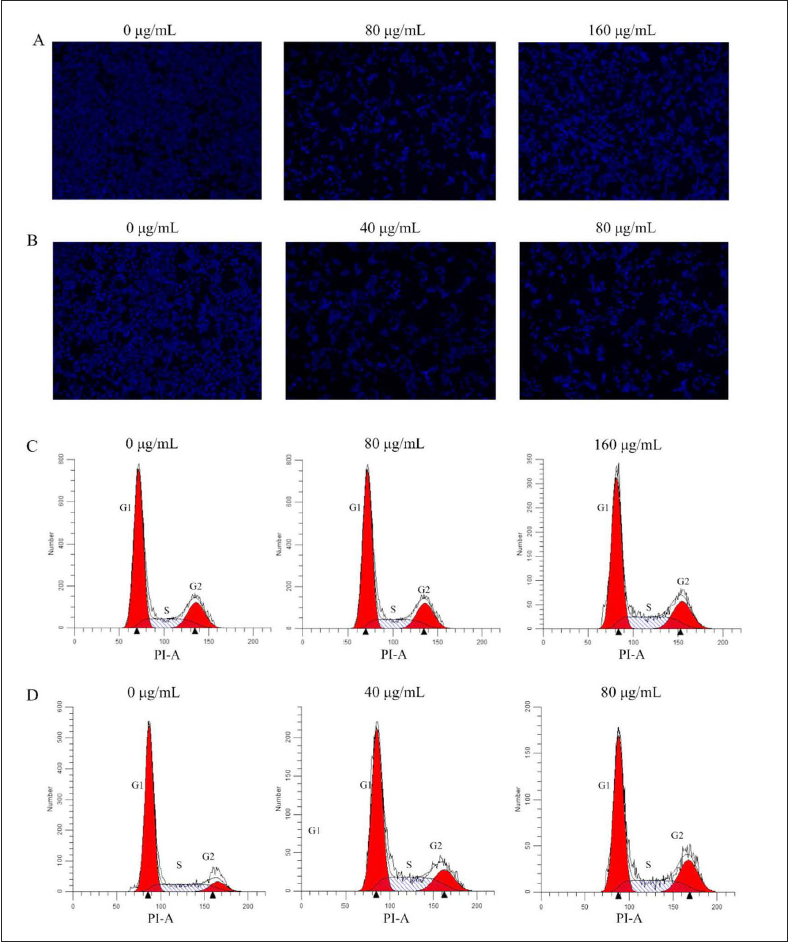
As shown in Figure 7D, the cell cycle of HCT-116 cells treated with different concentrations of berberine (40 and 80 µg/mL) was significantly inhibited in G2 phase in a dose-dependent manner compared with the control group. However, the effects of CCF ethanol extracts on the cell cycle of HCT-116 cells did not exhibit regularity (Figure 7C and D), possibly due to its extremely complex compounds, and the multi-targets and multi-pathways it acts on. Further analysis will be conducted to investigate the reasons for this phenomenon in future works.
CCF Ethanol Extracts and Berberine Promoted the Apoptosis in HCT-116 Cells
To determine whether CCF ethanol extracts and berberine promoted apoptosis in HCT-116 cells, the cells were preincubated with them for 24 h and stained with Hoechst 33342 for 10 min. The cells were then observed under the microscope, and with the increase in drug concentration, the number of cells decreased and the number of cells with bright blue nuclei increased (Figure 7A and B), confirming that different concentrations of CCF ethanol extracts and berberine inhibited the growth of HCT-116 cells and promoted their apoptosis.
CCF Ethanol Extracts and Berberine Down-regulated the Expression of the 3 Hub Targets
The expression of PLK1, CHK1, and AURKB was down-regulated in the CCF ethanol extracts and berberine-treated groups compared to the control group (**p < 0.01), as revealed by qPCR analysis, thus confirming the accuracy of the above results (Figure 6E and F).
Discussion
COAD is the leading cause of lethality globally, while the development of therapeutic drugs in this area remains a daunting challenge. Over the past few hundred years, TCMs have been explored as a potential source of anti-cancer drugs (Wang, Qi, et al., 2020). As a classic TCM, CCF is often used to clear heat, dry dampness, purge fire and detoxify, and has been proven to have anti-cancer effects by modern pharmacological studies (Yu et al., 2020; Zhang, Ma, et al., 2021). However, the bioactive compounds and mechanism of CCF against COAD are not well understood. With the advancement of science and technology, integrated bioinformatics has become an important way to identify core genes and pathways related to disease pathogenesis. As representatives of bioinformatics, network pharmacology and molecular docking have been widely used in the research and development of TCM (Taha et al., 2020; Xia et al., 2020). Therefore, this study aimed to explore the bioactive compounds and mechanism of CCF against COAD through comprehensive bioinformatics analysis and in vitro experiments.
Bioinformatics analysis of CCF revealed that it treated COAD by modulating the expression of 55 targets. The distribution of these targets in cells had no obvious regularity, but they were widely involved in the proliferation and apoptosis of cancer cells, which coincided with the results of the KEGG pathway analysis. Interestingly, in the “PPI” and “C-T-P” networks, the degree values of the above targets were different. In order to ensure the rationality of the experiments, the degree values of both networks were integrated and found that PTGS2 (degree=28), JUN (degree=27), CHEK1 (degree=26), PLK1 (degree=26), and AURKB (degree = 25) were the core targets of CCF anti-COAD. The semi-flexible docking results of Autodock Vina showed that the binding pocket of JUN could not fit well with small molecules. Berberine, a bioactive compound of CCF, was found to have the highest degree in the “C-T-P” network and a good affinity with the core targets, making it a promising candidate drug for further study. For the sake of accuracy, three targets (CHEK1, PLK1, and AURKB) whose binding affinity was closer to that of the positive drugs were selected as hub targets. More than 1,000 clinical samples in TCGA and UNLCAN databases also confirmed their significantly higher expression in COAD than in normal samples.
Numerous studies have hypothesized that CHEK1, PLK1, and AURKB are extensively involved in the regulation of tumor cell proliferation and apoptosis, which is consistent with GO and KEGG enrichment results (Liu et al., 2017; Õsz et al., 2019; Wang, Yu, et al., 2020). As hallmarks of tumor cells, rapid proliferation and resistance to apoptosis are major obstacles in the treatment of cancer (Wang et al., 2018; Wei et al., 2019). The plate cloning and morphological analysis of HCT-116 cells before and after administration indeed confirmed the inhibitory effect of CCF ethanol extracts and berberine on cell proliferation. From the results of Hoechst 33342 staining experiments, we found that the different concentrations of CCF ethanol extracts and berberine promote apoptosis and reduce the number of HCT-116 cells. Under the light microscope, the morphology of the cells after drug intervention did change and they were basically eliminated after washing with PBS. It cannot be ignored that the effects of CCF ethanol extracts and berberine on the cell cycle of HCT-116 were not completely consistent, and other chemical compounds in the extract might be the cause of this phenomenon. In HCT-116 cells, the three hub targets with high docking affinity to berberine, namely CHEK1, PLK1, and AURKB, were significantly downregulated after drug intervention, which verified the accuracy of our results.
Reckoning the study findings, CCF and its bioactive compounds berberine are potential adjuvant drugs for COAD. Nevertheless, further in vitro and in vivo experiments are necessary to confirm the efficacy and safety of these compounds.
Conclusions
In conclusion, this study demonstrated that CCF exhibits anti-COAD activity by down-regulating the expression of CHEK1, PLK1, and AURKB, with berberine identified as its main bioactive compound. The findings suggest that CCF and berberine have potential as adjuvant drugs for the treatment of COAD. However, further in vitro and in vivo experiments are needed to confirm these results and explore the clinical applications of CCF and berberine in COAD therapy.
Abbreviations
CCF: coptis chinensis franch.; COAD: colorectal adeno‑carcinoma; TCMSP: traditional chinese medicine systems pharmacology database, analysis platform; MDS: molecular docking simulation; GEO: gene expression omnibus; KEGG: kyoto encyclopedia of genes and genomes; GO: gene ontology; C-T-P: compounds-targets-pathways; PPI: protein-protein interaction; qPCR: quantitative real-time PCR; CHEK1: checkpoint kinase 1; PLK1: polo like kinase 1; AURKB: aurora kinase B; TCM: traditional chinese medicines; DEGs: differentially expressed genes; OB: oral bioavailability; DL: drug-likeness; DAVID: database for annotation, visualization and integrated discovery; GO BP: gene ontology biology; GO CC: gene ontology cellular compound; GO MF: gene ontology molecular function.
Footnotes
Acknowledgment
The authors declare that this work was done by the authors named in this article and all liabilities pertaining to claims relating to the content of this article will be borne by the authors.
The Statement of Informed Consent and Ethical Approval
Necessary ethical clearances and informed consent was received and obtained respectively before initiating the study from all participants.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
This study was supported by the research projects of clinical medicine in Wuhan in 2019 [Wuhan Municipal Health Commission, WZ19Q15]. Thanks to everyone who had contributed to this research.
Summary
CCF significantly down-regulated CHEK1, PLK1 and AURKB, inhibited proliferation, and promoted apoptosis of HCT-116 cells. Berberine is the main bioactive compound of CCF against COAD.
