Abstract
PRP-Exos are nanoscale cup-shaped vesicles that carry a variety of proteins, mRNAs, microRNAs, and other bioactive substances. PRP-Exos can be formed through several induction pathways, which determine their molecular profiles and facilitate their tailormade participation in intercellular communication. Currently, little is known on how the PRP-Exos activation method influences the quality and quantity of PRP-Exos. The present study aims to observe and analyze the number, profile, and growth factors of PRP-Exos through TEM, Nanoflow, and WB after PRP activation and compare the difference in function of PRP-Exos on HUVECs, with different stimuli (calcium gluconate, thrombin, or both). We found that PRP activated with both thrombin and calcium gluconate harvested the highest concentration of exosomes [(7.16 ± 0.46) × 1010 particles/ml], compared to thrombin group [(4.87 ± 0.15) × 1010 particles/ml], calcium gluconate group [(5.85 ± 0.43) × 1010 particles/ml], or saline group [(7.52 ± 0.19) × 109 particles/ml], respectively (P < 0.05) via Nanoflow analysis. The WB analysis showed that cytokines (VEGF, PDGFBB, bFGF, TGF-β) are differentially encapsulated in PRP-Exos, depending on the PRP stimulus, in which the mixture-PRP-Exos yielded the highest concentration of cytokines. In the function assay of PRP-Exos on HUVECs, the mixture-PRP-Exos promoted HUVECs proliferation, increased HUVECs migration, promoted the formation of vessel-like by HUVECs via the AKT ERK signal pathway more dramatically, compared with other groups. In summary, our studies showed that PRP activated by the mixture of calcium gluconate and thrombin harvested the best quality of exosomes which had the top biological functions. This study provides a protocol for selecting appropriate PRP activators to obtain high-quality exosomes for future applications.
Introduction
Platelet-rich plasma (PRP) is a mixture of highly concentrated platelets obtained from centrifugation of fresh whole blood. PRP releases a large number of autologous growth factors and cytokines after activation 1 . PRP has been widely used in the field of repair and regeneration, including in chronic tendinitis 2 , osteoarthritis 3 , chronic wounds 4 , and plastic surgeries 5 . A series of our previous studies on PRP treatment has revealed positive effects on wound healing in patients with diabetic foot ulcers 6 –9 . The basic cytokines identified in platelets include transforming growth factor-β (TGF-β), platelet-derived growth factor (PDGF), basic fibroblast growth factor (bFGF), vascular endothelial growth factor (VEGF), and endothelial cell growth factor. These natural cytokines are present in “normal” biological proportions and play important roles in cell proliferation, chemotaxis, cell differentiation, and angiogenesis 10 . Despite these advantages, the limitation of PRP applications is the requirement for autologous platelets.
Exosomes are a kind of extracellular vesicles (EVs), originating from intracytoplasmatic multivescicular bodies (MVBs), and are directly released into the extracellular space upon the fusion of the MVB membrane with the plasma membrane 11 . Platelet-derived exosomes (PLT-Exos) are stored in MVBs and α-granules 12 . After activation, the outer membrane of MVBs is invaded and released out of the platelets through extracellular secretion. EVs are a means of communication between long-distance cells in the body 13 . At present, EVs are mainly divided into three categories: exosomes, microvesicles, and apoptotic bodies 14 . Given that exosomes have no obvious adverse effects such as immunogenicity or tumorigenicity 15 , the potential of exosomes in wound healing has attracted increasing attention.
In 2014, Torreggiani et al. 16 first isolated exosomes from PRP and demonstrated their potential effects on the proliferation and migration of mesenchymal stem cells, which was the first described role of PRP-Exos in tissue regeneration. Furthermore, Tao et al. 17 demonstrated that PRP-Exos efficiently promote the re-epithelialization of chronic cutaneous wounds via activation of Yes-associated protein (YAP) in a diabetic rat model. Although PRP-Exos have been extensively studied in recent years 17,18 , few studies have assessed the method of activating PRP to release exosomes.
The current standards for preparing PRP are not uniform because of consensus regarding PRP preparation techniques and the complicated PRP activation methods. In this study, we aimed to develop a standard protocol for preparing exosomes for clinical treatment or research purposes, with consideration given to cytokines being predominantly stored in PRP-Exos after PRP activation 16,19 . Previous studies have shown that PRP-Exos can be obtained through a four-step method 20 , but the steps are cumbersome, the agonists cannot be widely used in clinical practice, and the platelets may be prematurely activated to release exosomes during the separation and purification steps.
Materials and Methods
PRP Preparation
Venous blood (400 mL) was gained from three healthy adults recruited from the Department of Blood Transfusion, Southwest Hospital, Army Medical University, Chongqing, China. Volunteers did not take any drugs that would have affected platelet number or function within the 2 weeks before blood donation. All volunteers provided written informed consent following the approval of the Ethical Committee Board of Southwest Hospital. Citrate glucose (22 g/L sodium citrate, 24.5 g/L glucose monohydrate, and 8 g/L citric acid monohydrate) was used as an anticoagulant for blood samples at a ratio of 1:10, and 100 ng/mL of prostaglandin E1 (PGE1, Sigma-Aldrich, St. Louis, MO, USA) was added to prevent platelet activation during isolation. PRP (40 mL) was isolated by a fully-automatic blood separator (CS-3000 plus; Baxter International, Inc., Deerfield, IL, USA), which has a leukocyte reduction system to avoid leukocyte contamination. The concentration of apheresis platelets in PRP and venous blood were counted by the automatic hematology analyzer (KX-21 N; Sysmex Corporation, Kobe, Japan). The prepared PRP was stored at 24°C while shaking or stored at -80°C until ready to use.
PRP Activation
For the present study, each PRP was divided into four aliquots of 10 mL and stimulated in the presence of (A) 0.1 U/mL thrombin (Sigma Aldrich, St. Louis, MO, USA); (B) 2 mm/L calcium gluconate solution (Sigma Aldrich, St. Louis, MO, USA); (C) a mixture with an equal volume of 2 mm/L calcium gluconate solution and 0.1 U/mL thrombin; or (D) an equal volume of 0.9% saline. These mixtures were incubated with agonist or with saline control for 90 min at 37°C in a water bath.
Histological Examination of PRG
Platelet-rich gel (PRG) was collected from activated PRP, and samples were fixed with 4% paraformaldehyde and embedded in paraffin. Embedded samples were sectioned at 5 µm thickness to perform hematoxylin and eosin (H&E) staining. The staining was performed according to the manufacturer’s protocol (Sigma Aldrich, St. Louis, MO, USA). Finally, the paraffin section was sealed using resinene, and pictures were taken under an Olympus BX51 optical microscope (Olympus America, Inc., Center Valley, PA).
Exosome Separation
Exosomes were isolated and purified by the gradient ultracentrifugation method created by Thery et al. 21 . Briefly, PRP was transferred to a 15 mL sterile polypropylene tube, and different agonists were added to activate platelets to release exosomes. All centrifugation steps were carried out at 4°C. Activated PRP was sequentially centrifuged at 300 × g for 10 min, 1500 × g for 15 min, and 3200 × g for 20 min to remove cellular debris. Then, the supernatant was successively centrifuged at 20,000 × g at 4°C using an SW26 rotor (Beckman Instruments, Inc., Fullerton, CA, USA) for 30 min to separate the exosomes from the MVs. After each centrifugation, the pellet was discarded, and the supernatant was used in the following steps. Then, the supernatant was filtered through a 0.22-µm filter (Millipore, Burlington, MA, USA). Exosomes were collected by ultracentrifugation using an SW26 rotor (Beckman Instruments, Inc., Fullerton, CA, USA) at 100,000 × g for 70 min at 4°C, washed in sterile 1 × PBS (no calcium, no magnesium, and no phenol red; Gibco; Thermo Fisher Scientific, Waltham, MA, USA), and pelleted by ultracentrifugation at 100,000 × g for 70 min. The pellets were carefully resuspended in 200 µL of sterile 1 × PBS containing protease inhibitor cocktail (Thermo Fisher Scientific, Waltham, MA, USA) and phosphatase inhibitor cocktail (Thermo Fisher Scientific, Waltham, MA, USA) and frozen at −80°C for future use. In the study, the supernatant of activated PRP was named PRP-AS. The exosomes isolated from thrombin-activated PRP were named thrombin-PRP-Exos. The exosomes isolated from calcium gluconate-activated PRP were named Ca2+-PRP-Exos. The exosomes isolated from saline-activated PRP were named saline-PRP-Exos. The exosomes isolated from mixture-activated PRP were named mixture-PRP-Exos.
TEM Analysis
Exosome suspensions (10 μl) were fixed by the same volume of glutaraldehyde (filtered twice through the 0.22-µm filter). Then, 10 µl of fixed exosomes were coated on 400 mesh grids (formvar/carbon-coated, glow-discharged) for 90 s. The grids were negatively stained with uranyl acetate (2%) for 30 s, and excess uranyl acetate was carefully removed using filter paper. The samples were examined immediately. The size and shape of the exosomes were observed, and micrograph images were taken by TEM (HITACHI-H7650, Hitachi, Tokyo, Japan) operating at 100.0 kV with an AMCT XR80 CCD sensor allowed to dry at room temperature (RT).
Particle Size Statistics of TEM Images
ImageJ 1.42q software (National Institutes of Health) was used to calculate the diameter of all exosomes from the TEM images. The graphs and data analyses were generated by Origin 2019 (OriginLab Corporation, Northampton, MA, USA).
Nanoflow Analysis
Exosome samples (10 μl) were diluted (1:100) and analyzed using the Flow Nano Analyzer (NanoFCM Inc. China), according to the manufacturer’s protocol 22 . Briefly, calibration of the instrument was performed using 200 nm control beads (NanoFCM Inc. China), which were analyzed as a reference for particle concentration. Furthermore, a mixture of beads with different sizes (NanoFCM Inc. China) was provided as a reference for size distribution. PBS was used as a background signal and subtracted from other measured values during analysis. The number of particles after the sample was diluted was in the optimal range of 4000–14,000. Particle concentration and size distribution were analyzed using the NanoFCM software (NanoFCM Profession V1.0) and normalized for the starting volume of exosome resuspension and the dilution factor that was necessary for proper NanoFCM reading.
Cell Cultures
Human umbilical vein endothelial cells (HUVECs) were obtained from ScienceCell and cultured in endothelial cell medium (ECM, ScienceCell Research Laboratories, Carlsbad, CA, USA), supplemented with 10% FBS and EC growth supplement (ECGS, ScienceCell Research Laboratories, Carlsbad, CA, USA). All experiments were conducted with HUVECs in passages 3-6. Cells were incubated in a humidified atmosphere of 5% CO2 at 37°C.
Exosomes Labeling, Internalization, and Confocal Microscopy
Exosomes were labeled using PKH-26 (red, Sigma-Aldrich) and added into the HUVECs culture at a concentration of 50 μg/ml. After incubation for 6 hours, HUVECs were washed twice with diluted PBS. Then cells were fixed with 4% paraformaldehyde for 20 min at RT. The nuclei of HUVECs were stained with 4′,6-diamidino-2-phenylindole (DAPI). The uptake of exosomes was observed by the laser scanning confocal microscope (Leica Microsystems, Wetzlar, Germany).
Cell Proliferation Assay of HUVECs
HUVECs suspension was seeded in 96-well plates (5 × 103 cells per well) and were co-cultured with PRP-Exos for 24, 48, 72, and 96 h, respectively. The cells of each group were treated accordingly, and three parallel wells were set up in each group. Cell proliferation was analyzed using a Cell Counting Kit-8 (CCK-8, Beyotime, Jiangsu, China).
Transwell Migration Assay of HUVECs
HUVECs were seeded to the upper well of transwells in a 6-well plate with an 8-µm pore size (Corning-Costar, Corning, NY, USA) at 5 × 104cells per well, and PBS or PRP-Exos was added to the medium containing 5% serum of the lower chamber. After incubation at 37°C for 24 h, the cells in the upper chamber were gently wiped off with cotton swabs. The Transwell chamber was fixed with 4% paraformaldehyde for 30 min, stained with crystal violet dyeing solution for 1 min. Images were obtained on an inverted microscope (Olympus America, Inc., Center Valley, PA). Each experiment was repeated three times, and each time three wells were tested for a sample.
Matrigel Tube Formation Assay of HUVECs
HUVECs were seeded onto a Matrigel (BD, New Jersey, USA) precoated 96-well plate at the cell density of 2 × 104 cells/well, and cultured in ECM for 6 h, in the presence of PRP-Exos (50µg /mL) or not. The tube-like structures were pictured using an inverted microscope (Olympus America, Inc., Center Valley, PA). The angiogenic property was assessed by measuring the total branching length from three random microscopic fields using Image J 1.42q software (National Institutes of Health). Each assay was repeated at least three times. One representative of three independent experiments was shown.
Western Blot Assay
The total protein was extracted using a Total Exosome RNA & Protein Isolation Kit (Thermo Fisher Scientific, Inc. Waltham, MA, USA). The protein concentration was determined using the Pierce™ BCA Protein Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). Cell or exosome lysates were diluted at a ratio of 1:4 with protein loading buffer (5×) and denatured at 95°C for 5 min. Samples (60 µg of total protein) were separated by SDS-polyacrylamide gels (5% stacking gel and 12% separation gel) at 120 V for 1 h and blotted onto a polyvinylidene difluoride (PVDF) membrane (Millipore, Jaffrey, NH, USA) for 90 min at 210 mA. The membrane was blocked with 5% weight/volume non-fat milk powder (BSA, Sangon Biotech, Shanghai, China) in TBST (0.1 M Tris-HCl PH 8, 1.5 M NaCl, and 1% Tween-20) for 1 h at RT. The membrane was probed with primary antibodies overnight at 4°C [anti-CD9, anti-CD63, anti-CD41, anti-CD81, anti-TSG101, anti-flotillin, anti-VEGF, anti-PDGFBB, anti-Calnexin (1:1,000 dilution; Abcam, Cambridge, MA, USA); anti-TGF-β, anti-bFGF, anti-AKT, anti-p-AKT, anti-p-ERK1/2, anti-ERK1/2, anti-β-actin (1:1000 dilution, Cell Signaling Technology, Beverly, MA, USA)]. After rinsing three times using TBST, we incubated the immunocomplexes with horseradish peroxidase (HRP)-conjugated secondary antibodies (1:4000 dilution, Cell Signaling Technology, Beverly, MA, USA) for 1 h at RT. We detected immunoreactivity via the chemiluminescence method using a ChemiDoc™ MP Imaging System (Bio-Rad, Hercules, CA, USA), processed using an enhanced-chemiluminescence system (ECL, Pierce, Rockford, USA). The densitometric quantification on the protein bands was performed using Image J 1.42q software (National Institutes of Health). The proteins encapsulated into exosomes were analyzed by classical western blotting to test for specific exosome markers, such as CD9, CD63, CD81, TSG101, flotillin, and the source marker CD41. Calnexin was used as a negative control. β-actin served as the internal reference. For detailed antibody information, please see Supplementary Table S1.
Statistical Analysis
All data were analyzed using Origin Pro 2020 (OriginLab Corp., Northampton, MA, USA). The data are presented as the means ± standard deviation (SD) of at least three independent simples. Two-tailed Student’s t-test or one-way analysis of variance was used for evaluating the significance of differences between subgroups. The difference between samples was considered significant when P < 0.05.
Results
In the present study, an efficient and simple protocol of activation PRP to release exosomes has been established. What’s more, exosomes disengaged utilizing calcium gluconate, thrombin, the mixture of calcium gluconate and thrombin were successfully compared on morphologic features, particle size distribution, surface charge, and encapsulated protein level using electron microscopy, Nanoflow, and WB analysis, individually. The average platelet count was (1532 ± 81) × 109 cell/L in PRP, while the average platelet count was (236 ± 22) × 109 cell/L in donor venous blood. By using our protocol, those numbers for nucleated cells, such as leukocytes or monocytes, did not surpass one in the 1 × 106 platelet tests.
Different Agonists Influenced PRP Activation
In the thrombin- and calcium gluconate-activated PRP group, the region of fibrin staining by HE in the PRG shaped by PRP activation after 90 min has been extensive and thickly dispersed. The area of fibrin in the PRG shaped by PRP activation after 90 min in the group of the thrombin actuation assembly may have been smaller (P < 0.05), yet the area in the PRG shaped by PRP actuation after 90 min in the group of calcium gluconate activation assembly may have been small, even dispersed. Nevertheless, the control group (saline treatment) did not form a fibrin clot (Supplemental Fig. S1).
PRP-Exos Harvested by Gradient Centrifugation
A protocol for the production of PRP-Exos starting in vein blood is shown in Fig. 1A,B. After separating the exosomes, the total protein concentration in all groups was measured using the BCA colorimetric protein assay, in which exosomes were lysed with RIPA buffer, and the results are shown in Fig. 1C. The protein concentration of PRP-Exos obtained in the three stimuli groups was significantly higher than that of the control group (saline treatment). The protein concentration obtained in the calcium-activated group was higher than that in the thrombin-activated group (P < 0.01). Although the protein concentration obtained in the mixture group (calcium combined with thrombin) was higher than the calcium-ionophore group, there was no significant statistical difference (P > 0.05).
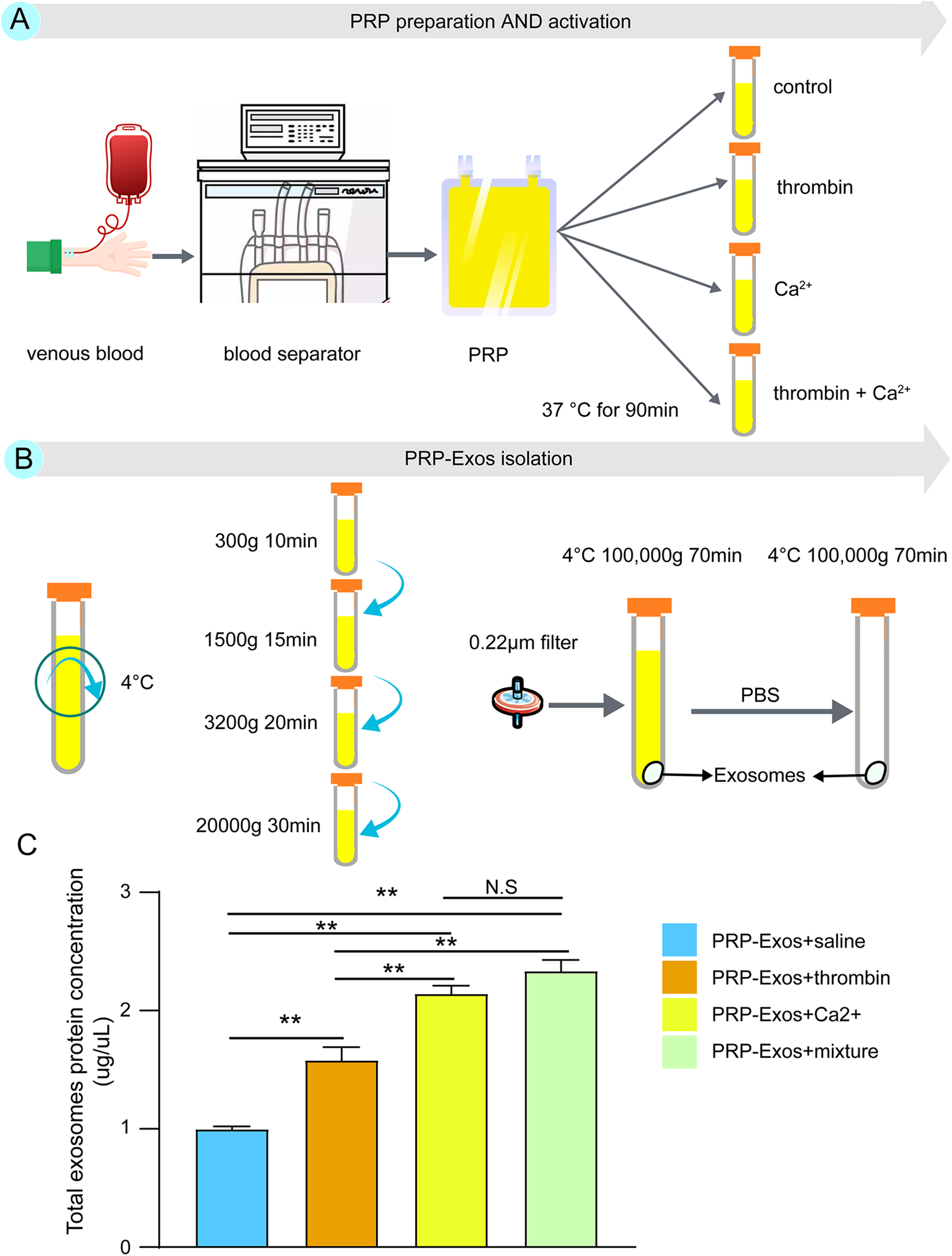
The workflow of PRP-Exos isolation protocol, starting from PRP and comparison of the total exosome protein concentration. Exosomes from PRP were isolated using UC. (A) The recommended protocol for PRP preparation and activation. (B) The protocol used in this study for PRP-Exos extraction. The precipitates in the final step were the obtained PRP-Exos, which were dissolved in sterile PBS. (C) The total protein levels were estimated in intact exosomes by the BCA protein assay. The highest PRP-Exos yield was detected for the mixture of calcium gluconate and thrombin, while saline yielded the lowest concentration of exosomes. Statistical significance was determined using an independent-sample t-test. Data are shown as mean ± SD of values of three measurements in each group. n = 3 independent samples. N.S., not significant. *P < 0.05, and **P < 0.01. UC, ultracentrifugation; PRP, platelet-rich plasma; PRP-Exos, exosomes derived from platelet-rich plasma; PBS, phosphate-buffered saline.
The Difference of PRP-Exos Structure with Different Agonists
As shown in Fig. 2, the exosomes were isolated from PRP without membrane breakage. The exosomes were ascertained with typical cup-shaped vesicles, measuring 30–150 nm in diameter (Fig. 2). However, the size and shape of exosomes are different from PRP activated by different agonists. Through TEM observations, a few of MVBs were present in the calcium-activated group, which contains exosomes. The surfaces of PRP-Exos were also rougher in the calcium-activated group than that of the other groups. The size of PRP-Exos (Fig. 2A, C) was significantly larger than the other two groups (Fig. 2B, D). The number of PRP-Exos activated by saline (Fig. 2A) was the lowest, but the number of other groups could not be identified under the microscope.
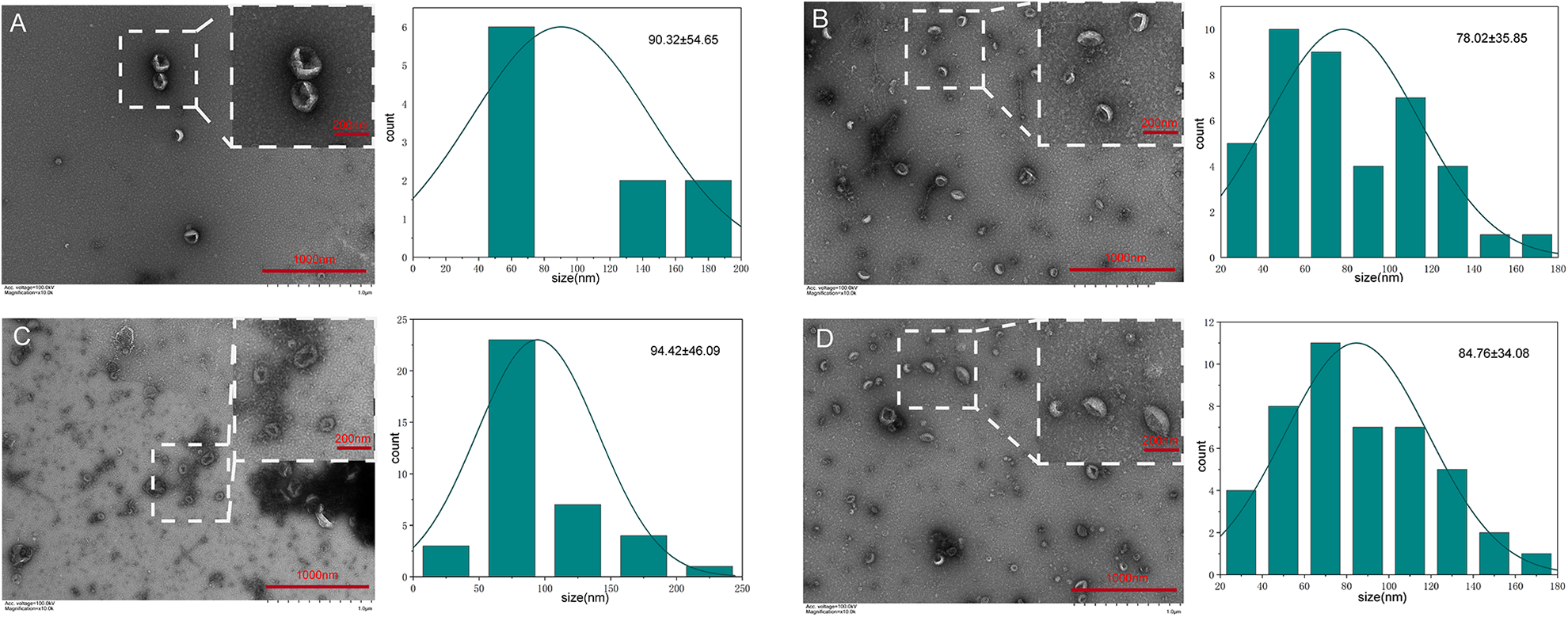
Representative transmission electron microscopy (TEM) images of PRP-Exos. TEM reveals the structure of exosomes. Images obtained from the exosome population generated by platelets exposed to different agonists: (A) saline; (B) thrombin; (C) calcium gluconate; and (D) mixture of calcium gluconate and thrombin. The left side of each group are TEM images and the right side is the particle size distribution diagram of the exosomes, which were counted in the TEM images by Image J. Date presented as mean ± SD; n=3 independent samples. Scale bar as shown. PRP-Exos, exosomes derived from platelet-rich plasma.
The Size Distribution and Concentration of PRP-Exos Significantly Vary Among Preparations from Different Agonists
To further determine the size and concentration of PRP-Exos obtained by different stimuli, the Nanoflow assay was performed for PRP-Exos. As shown in Fig. 3, the average size of exosomes is (78.98 ± 4.62) nm. The size of calcium-activated PRP-Exos (85.16 ± 0.59 nm) was larger than the thrombin-activated group (79.81 ± 0.43 nm) and the mixture-activated group (76.00 ± 0.37 nm), which showed broader size distribution with a shift toward the larger size. Furthermore, PRP activated by the mixture released the highest concentration of exosomes, followed by the calcium-activated group and then the thrombin-activated group. The concentration of particles obtained by different activators varies among preparations. The highest concentration of particles was observed when PRP-Exos were obtained by the mixture [(7.16 ± 0.46) × 1010 particles/mL; P < 0.05 compared to the calcium-, thrombin-, and saline-activated groups]. PRP treated with saline yielded the lowest concentration of particles [(7.52 ± 0.19) × 109 particles/mL]. The calcium-activated group harvested a higher concentration of particles [(5.85 ± 0.43) × 1010 particles/mL] than the saline-activated group (P < 0.01), as did the thrombin-activated group [(4.87 ± 0.15) × 1010 particles/mL; P < 0.01]. Also, the concentration of particles in the calcium-activated group was slightly higher than the thrombin-activated group (P < 0.05).
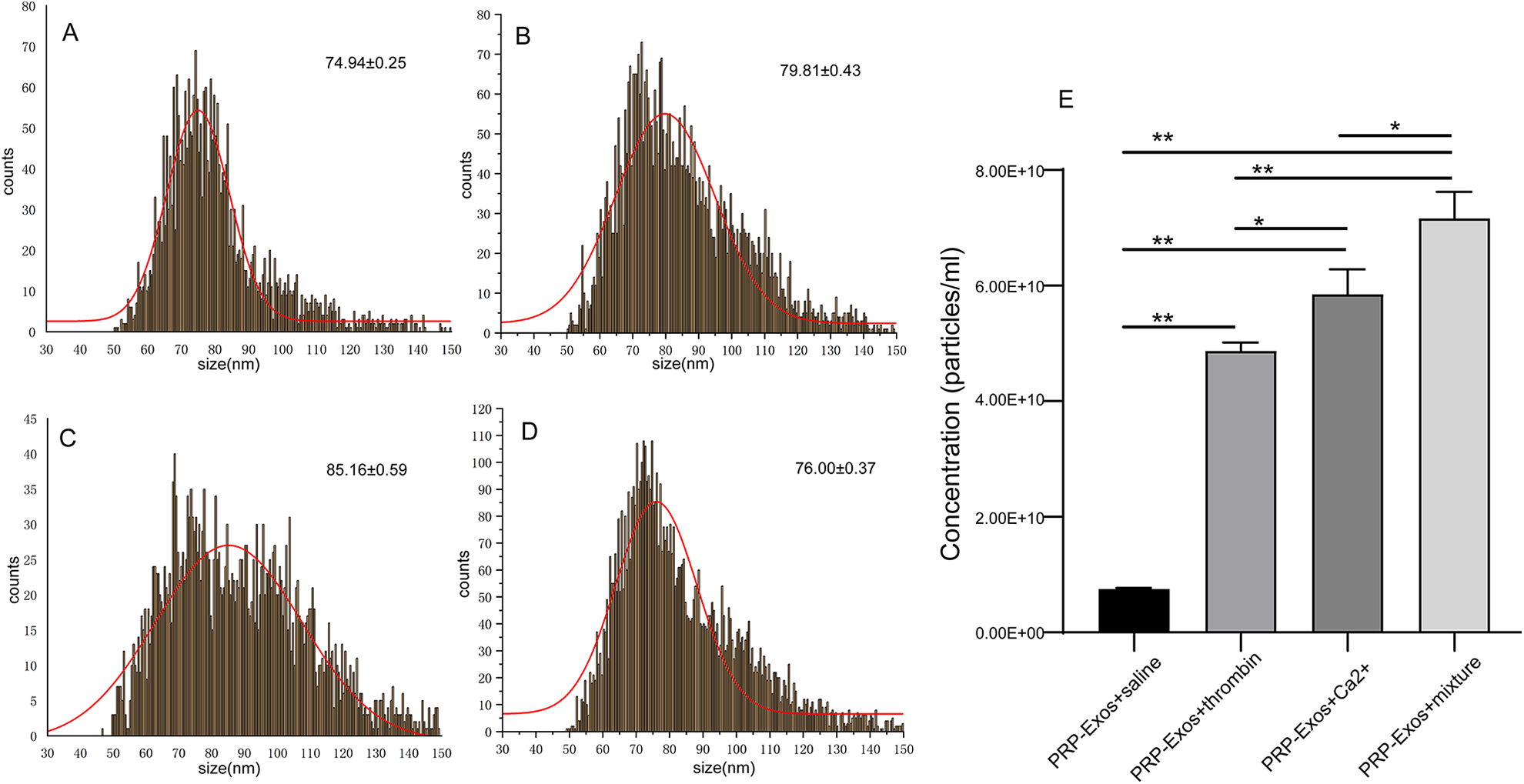
Size and concentration of exosomes from Nanoflow analysis. The size and concentration of the exosome population derived from platelets exposed to different agonists: (A) saline; (B) thrombin; (C) calcium gluconate; and (D) mixture of calcium gluconate and thrombin. (E) Concentration (particles/mL) of exosomes was determined by Nanoflow analysis. Statistical significance was determined using an independent-sample t-test. Data are presented as mean ± SD; n = 3 independent samples. N.S., not significant. *P < 0.05, and **P < 0.01. PRP-Exos, exosomes derived from platelet-rich plasma.
The Differences of Exosomal Surface Biomarker and the Encapsulated Proteins with Different Agonists
To further investigate the difference between exosomal surface proteins and the encapsulated proteins, the purity of exosomes was examined by immunoblotting for specific marker proteins. CD81, TSG101, CD63, Flotillin, CD9 are specific markers for exosomes. The source marker (CD41) and negative control (calnexin) proteins were also detected by immunoblotting. PRP-Exos activated by different stimuli showed different signal intensities for specific marker proteins, with the highest intensity detected for prep from the mixture group (Fig. 4). In contrast, only faint signals for specific marker and negative control proteins in the saline treatment group were identified, which indicated lower purity and fewer numbers of exosomes. The signal intensities of the calcium-activated group were stronger than that of the thrombin-activated group (P < 0.01). The platelet marker CD41 confirmed the exosome identity. Negative control (calnexin), which was found to be present in cell lysates, acted as the negative control. These data suggested that the nanoparticles were exosomes and the WB results were consistent with the results of the Nanoflow analysis.
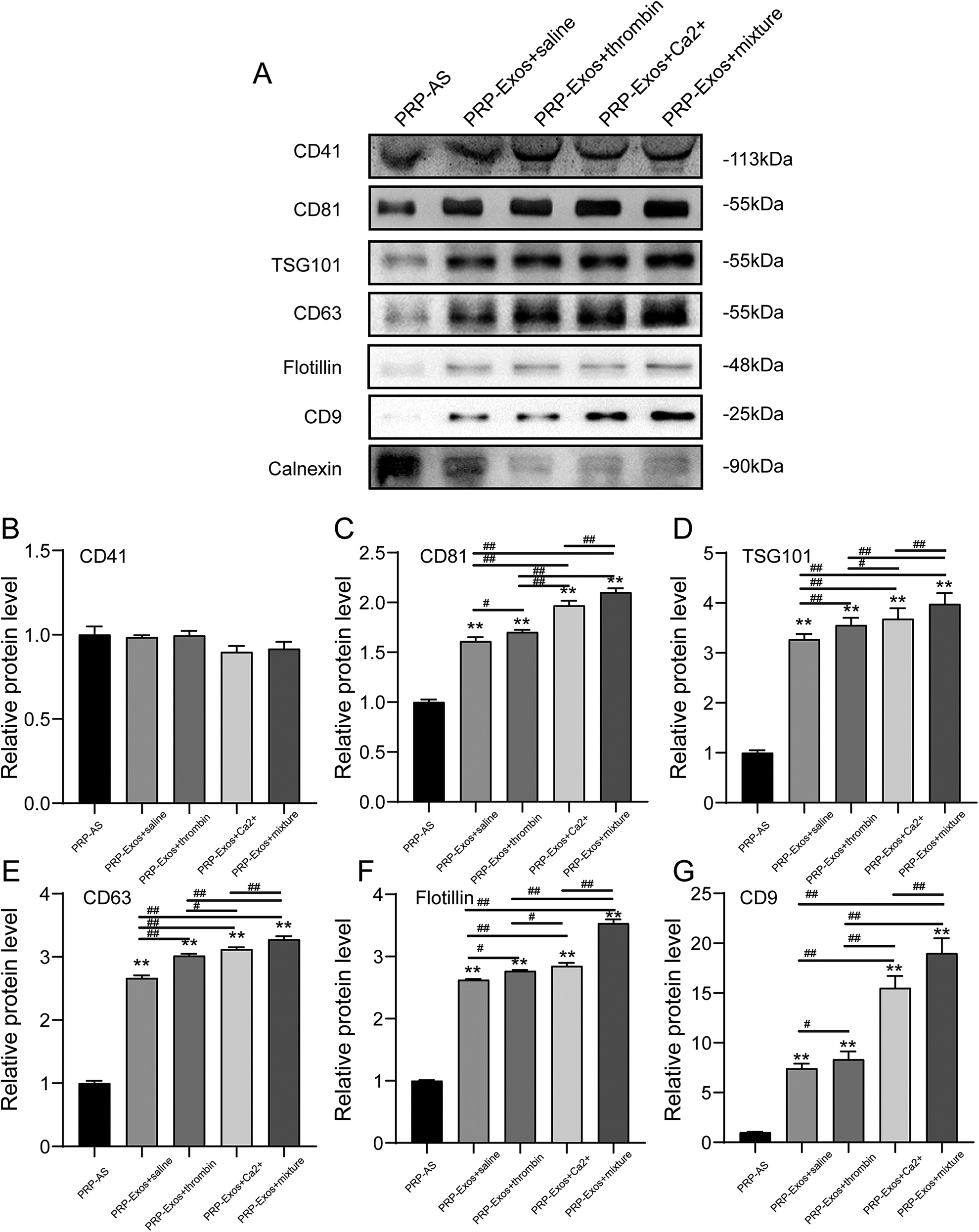
Western blot analysis of the surface biomarkers: CD81, TSG101, CD63, Flotillin, CD 9 on PRP-Exos; the source marker CD41; and the negative control (calnexin), which were found to be present in cell lysates. Densitometric analysis of western blot results: (A) representative western blot images and (B–E) densitometric analysis. Each lane was loaded with 60 µg of total exosomal protein. Statistical significance was determined using an independent-sample t-test. Data are shown as mean ± SD of values of three measurements in each group. n = 3 independent samples. N.S., not significant. *P < 0.05, and **P < 0.01 versus PRP-AS; # P < 0.05 and ## P < 0.01.
Considering that platelet basic cytokines are mainly present in exosomes 16,19 . WB analysis was conducted to further confirm the differences between cytokine-encapsulated proteins in each group. The results are presented in Fig. 5. The results from the WB show that VEGF, PDGFBB, bFGF, and TGF-β mainly exist in PRP-Exos, not PRP-AS (P < 0.05). The protein levels of VEGF, PDGFBB, bFGF, TGF-β in the mixture-activated PRP-Exos group were higher than that of the other groups (P < 0.05). There were no statistical differences in the protein content of VEGF, bFGF, and TGF-β between the calcium- and thrombin-activated groups (P > 0.05). The protein content of PDGFBB in the thrombin-activated group was higher than that of the calcium-activated group but lower than that of the mixture group (Fig. 5B).
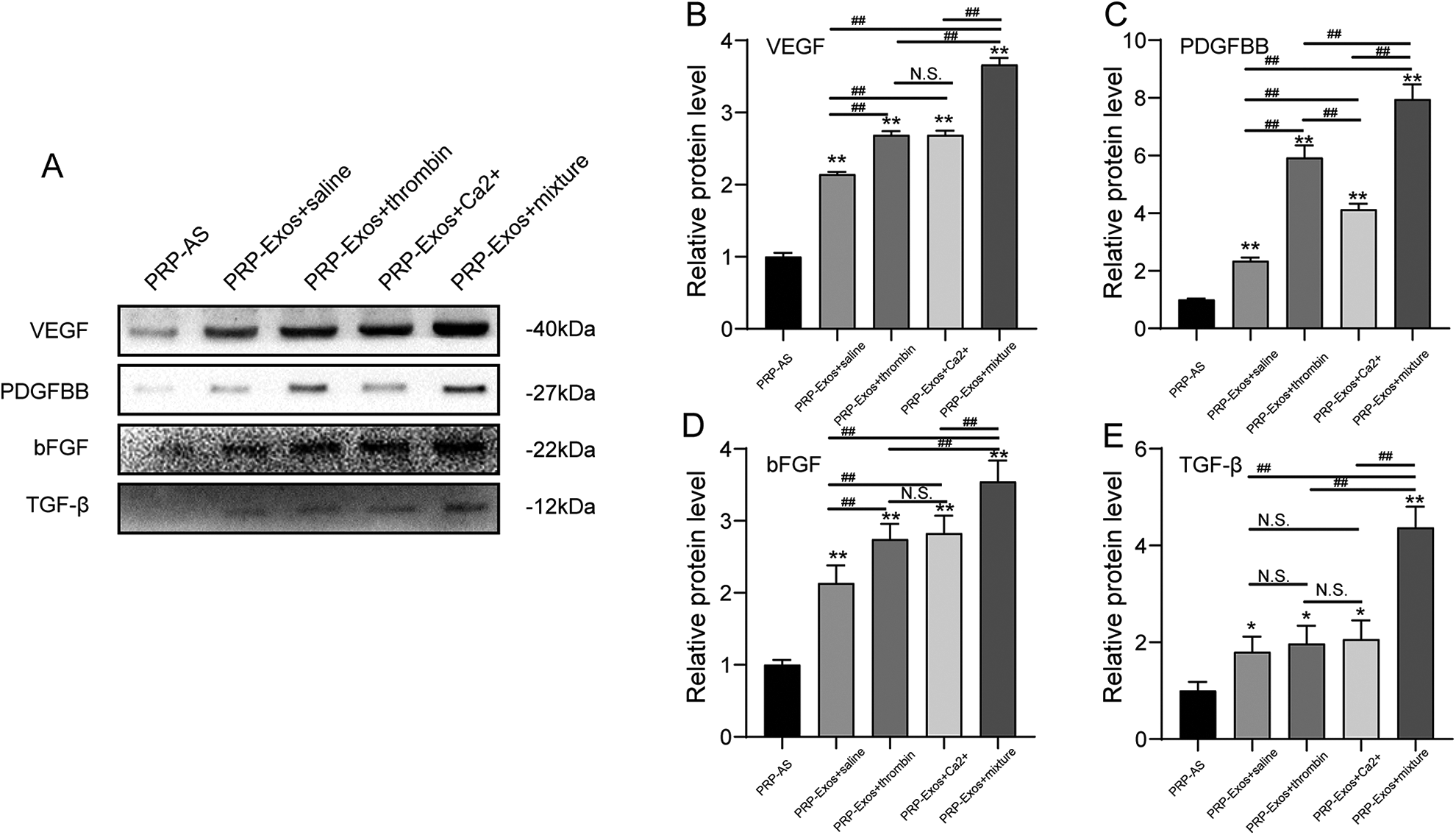
Western blot analysis of PRP-Exos-encapsulated proteins. Representative western blot showing VEGF, PDGFBB, bFGF, TGF-β, and densitometric analysis of western blot results: (A) representative images; (B–E) densitometric analysis. Each lane was loaded with 60 µg of total exosomal protein. Statistical significance was determined using an independent-sample t-test. Data are shown as mean ± SD of values of three measurements in each group. n = 3 independent samples. N.S., not significant. *P < 0.05 and **P < 0.01 versus PRP-AS; # P < 0.05, and ## P < 0.01.
The Function Assays of PRP-Exos on HUVECs
Considered that PRP-Exos promote tissue wound repair and endothelial cells play an important role in both angiogenesis and tissue regeneration, the effect of PRP-Exos on HUVECs in vitro was examined.
PRP-Exos were internalized by HUVECs, stimulated their proliferation, promoted migration, stimulated in vitro formation of vessel-like structures. PRP-Exos were internalized by HUVECs, which were mainly located in the cytoplasm of endothelial cells (Fig. 6A). The proliferation of HUVECs cultured in different concentrations of PRP-Exos and PRP-AS is shown in Fig. 6B, C. The PRP-Exos had a concentration-dependent effect on cell proliferation (Fig. 6B), in which HUVECs cultured in 50 μg/mL PRP-Exos activated by saline proliferated significantly better than other groups at 24 h (P < 0.05). 50 μg/mL PRP-Exos activated by the mixture have a stronger proliferation capacity, compared to the other groups (Fig. 6C) at all time points tested (P < 0.05). Transwell and tube formation assays showed that the PRP-Exos activated by the mixture more significantly promoted migration, formation of vessel-like, compared to control, PRP-AS, calcium-activated PRP-Exos, thrombin-activated PRP-Exos at the same total protein level (P < 0.05) (Fig. 6D–F). However, there was no statistically significant difference between the calcium-activated PRP-Exos group and the thrombin-activated PRP-Exos group enhancing the formation of vessel-like structures. Figure 5 indicated that VEGF, bFGF, and PDGFBB were enriched in the PRP-Exos, suggesting that AKT and ERK pathways can be activated by PRP-Exos. To further evaluate the performance differences between the different stimulus types, we examined the levels of AKT and ERK phosphorylation by Western Blot following treatment with PRP-Exos or PRP-AS for 6 h (Fig. 6G–I). Compared to PRP-AS, incubation with PRP-Exos resulted in a significant increase in phosphorylation of AKT and ERK (Fig. 6G). The mixture PRP-Exos group has the highest level of AKT and ERK phosphorylation compared with all other groups (Fig. 6G–I).
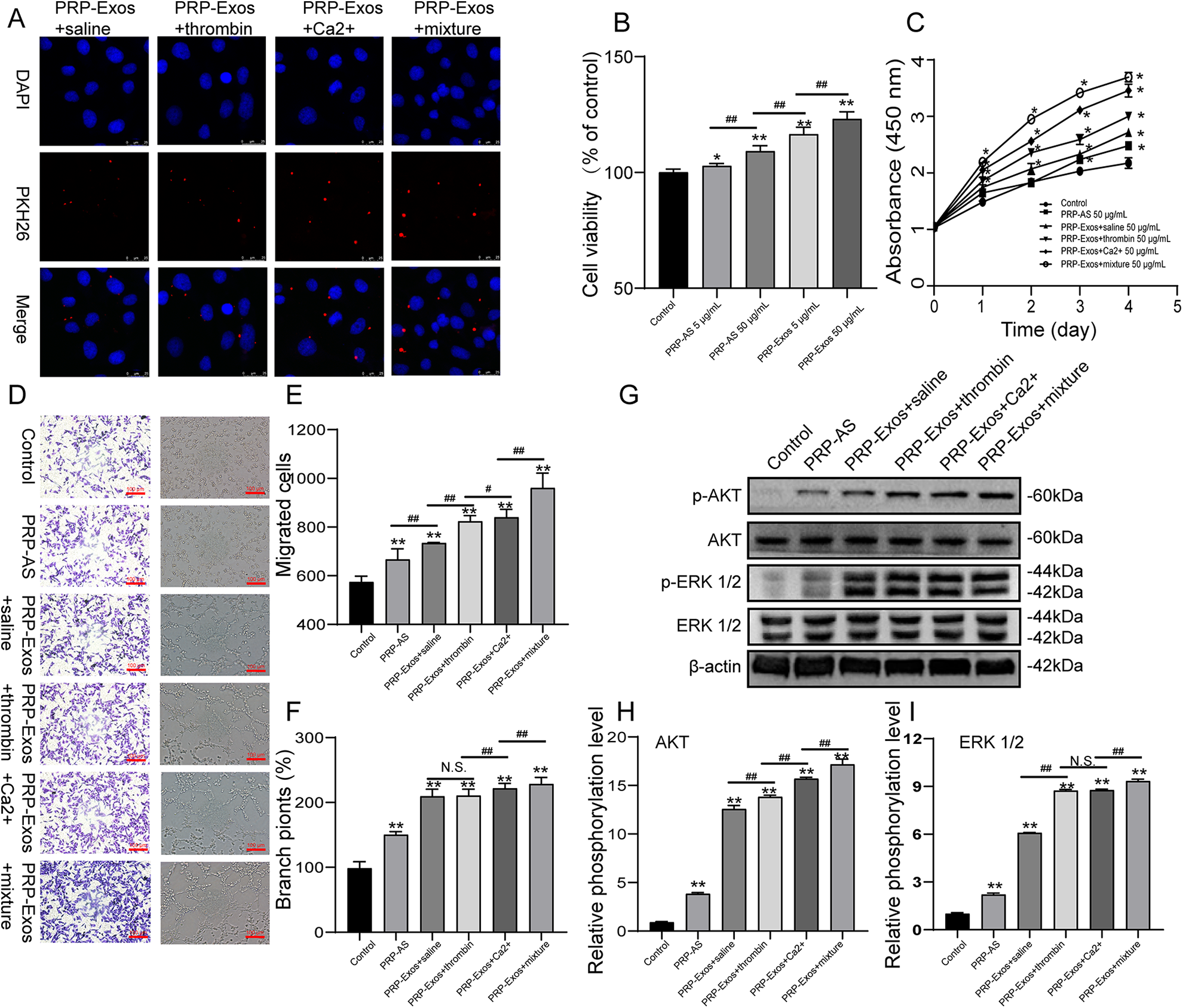
PRP-Exos effect on HUVECs. (A) representative confocal microphotography of PKH-26 labeled exosomes internalized by HUVECs; (B-C) CCK8 results of HUVECs treated by PRP-Exos and PRP-AS; (D–F) Transwell migration assay and tube foramtion results of HUVECs with different treatments. HUVECs were treated with PBS (control), 50 μg/mL PRP-AS, 50 μg/mL PRP-Exos+saline, 50 μg/mL PRP-Exos+thrombin, 50 μg/mL PRP-Exos+Ca2+, 50 μg/mL PRP-Exos+mixture; (G–I) Western blot analysis of HUVECs with different treatments. HUVECs were treated with PBS (control), 50 μg/mL PRP-AS, 50 μg/mL PRP-Exos+saline, 50 μg/mL PRP-Exos+thrombin, 50 μg/mL PRP-Exos+Ca2+, 50 μg/mL PRP-Exos+mixture. Statistical significance was determined using an independent-sample t-test. Data are shown as mean ± SD of values of three measurements in each group. n = 3 independent samples. N.S., not significant. *P < 0.05, and **P < 0.01 versus Control; # P < 0.05 and ## P < 0.01.
Discussion
PRP has been widely used in the field of repair and regeneration because it offers many advantages, including easy applicability, low cost, and an autologous nature 23,24 . However, the life span of platelets is only around 7–10 days when circulating in the blood and they are not easy to store 25 . In general, autologous platelets do not often meet clinical application, and allogeneic platelets have the risk of immune rejection. The mechanism by which PRP produces improvements in the field of repair and regeneration remains subject to debate 5 . With the continuous growth of research on cellular exosomes, the potential of exosomes has gained increasing attention. Exosomes have the following characteristics: low immunogenicity, easy storage, stability, easy mass production, and controllability 11,14,26 . Exosomes have a high degree of biocompatibility and immune inertness and can cross the body’s thick tissue barriers, such as the blood–brain barrier (BBB). Under normal circumstances, exosomes can be directly used as potential therapeutic agents for immune regulation and tissue repair.
Different platelet activations may affect PRP-Exos protein profiles and roles in intercellular communication 27 –30 . When being used in clinical applications, methods of activating PRP to release exosomes should be rapid and standardized, produce a high yield of pure exosomes from small PRP volumes, and be compatible with a large number of samples. A large number of comparative studies have been performed, which compare different activators to activate platelets to release exosomes 20,27,28,30,31 . However, these activation steps are cumbersome. It is necessary to isolate platelets from PRP and then activate the platelets to release the exosomes, and these activators are not commonly used in clinical applications. Therefore, it is necessary to compare the ability of common drugs (thrombin or calcium gluconate) to activate PRP to release exosomes. The methods studied here are straightforward and easy to perform and several samples can be processed simultaneously in short periods.
All the activation methods were suitable for PRP-Exos isolation, as demonstrated by TEM (cup-shaped vesicles), particle size, and WB detection of the exosome markers: CD81, TSG101, CD63, Flotillin, and CD9 15 . In particular, calcium gluconate alone was found to be weaker than thrombin or thrombin and calcium gluconate together in PRP activation, which was consistent with the previous studies that investigated the physiological strength of different agonists for platelet activation 20,27 . TEM showed that some exosomes activated by calcium had rough surfaces, which may be attributed to the formation of a calcium citrate chelate causing a partial effect of low pH on the exosome surface proteins 32,33 . However, they could be successfully isolated without membrane breakage, and their morphological features were similar in each group.
Moreover, the concentration of PRP-Exos activated by thrombin alone was found to be lower than that of calcium gluconate or thrombin and calcium gluconate together. To our knowledge, signaling originates from specific receptors followed by common vesiculation processes, such as Ca2+ entry, cytoskeletal remodeling, calpain/caspase activity, and mitochondrial depolarization during activation 34 . The reason may be that non-physiological agonists, such as Ca2+-ionophore or detergents, induce most microvesiculation, acting as relaxants 27,30 .
The protein content on the exosomal surface was detected by WB and was consistent with the concentration of exosomes detected by Nanoflow. Previous studies demonstrated that biologically-active growth factors are mainly present in PRP-Exos, which have concentration-dependent biological effects 16 –18 . Our study showed that the basic cytokines were almost exclusively present in PRP-Exos and their concentrations varied depending on the PRP activation method. PRP-Exos activated by thrombin and calcium gluconate together were found to contain more cytokines than groups activated by thrombin or calcium gluconate alone.
To further understand functional differences in angiogenesis between all groups, the effect of PRP-Exos on HUVECs was examined, which play an important role in both angiogenesis and tissue regeneration 35 . Results showed that 50 μg/mL PRP-Exos activated by thrombin and calcium gluconate together could significantly promote HUVECs proliferation, increase HUVECs migration, promote the formation of vessel-like by HUVECs via the AKT ERK signal pathway, compared with those in the other groups. While the downstream signaling kinases AKT and ERK were activated by PRP-Exos, it appeared that AKT was a major one since much higher levels of p-AKT than that of p-ERK were observed. Although PRP-Exos activated by thrombin or thrombin and calcium gluconate together contained more PDGFBB than the other groups, PRP-Exos activated by thrombin did not promote HUVECs proliferation, increase HUVECs migration, promote the formation of vessel-like by HUVECs, enhance activation of AKT and ERK signal pathway more dramatically than PRP-Exos activated by calcium gluconate. It is suggested that calcium might perform a cell-intrinsic role in regulating the expression of certain important molecules by activating PRP. Relevant studies showed that the lipid growth factors of platelet microparticles may be the main active factor of promoting angiogenesis, while the protein component may be the secondary factor 36,37 . For instance, Sphingosine 1-phosphate (S1P), one of these platelet-derived lipid growth factors, has been demonstrated that mediates cytoprotection, proliferation, migration, and tube formation via activating PI3K/AKT 38,39 .
In summary, we compared the effects of different agonists on the quality and quantity of PRP-Exos and the effect of PRP-Exos on HUVECs. We found that all agonists could activate platelets to release exosomes, but the activation efficiency and cytokine content are different. Calcium gluconate activation can release more exosomes, and thrombin activation can obtain exosomes containing high concentrations of PBDGBB. The effects from activating PRP with thrombin and calcium gluconate together are better than using thrombin or calcium alone, both in quality and quantity of exosomes. 50 μg/mL PRP-Exos activated by thrombin and calcium gluconate together could more significantly promote HUVECs proliferation, increase HUVECs migration, promote the formation of vessel-like by HUVECs via the AKT ERK signal pathway, compared with other groups. Our studies showed that the number of PRP-Exos and the content of cytokines in the PRP-Exos, PRP-Exos biological effects on HUVECs are different when PRP is activated by different agonists. This study provides a protocol for selecting the appropriate PRP activator to obtain high-quality exosomes for future applications. However, the mechanism behind the difference in the number of exosomes and cytokines the effect of PRP-Exos on HUVECs, generated by different activators, are not clear, and further exploration should be performed.
Supplemental Material
Supplemental Material, sj-docx-1-cll-10.1177_09636897211017833 - Comparison and Investigation of Exosomes Derived from Platelet-Rich Plasma Activated by Different Agonists
Supplemental Material, sj-docx-1-cll-10.1177_09636897211017833 for Comparison and Investigation of Exosomes Derived from Platelet-Rich Plasma Activated by Different Agonists by Shunli Rui, Yi Yuan, Chenzhen Du, Peiyang Song, Yan Chen, Hongyan Wang, Yahan Fan, David G. Armstrong, Wuquan Deng and Ling Li in Cell Transplantation
Supplemental Material
Supplemental Material, sj-docx-2-cll-10.1177_09636897211017833 - Comparison and Investigation of Exosomes Derived from Platelet-Rich Plasma Activated by Different Agonists
Supplemental Material, sj-docx-2-cll-10.1177_09636897211017833 for Comparison and Investigation of Exosomes Derived from Platelet-Rich Plasma Activated by Different Agonists by Shunli Rui, Yi Yuan, Chenzhen Du, Peiyang Song, Yan Chen, Hongyan Wang, Yahan Fan, David G. Armstrong, Wuquan Deng and Ling Li in Cell Transplantation
Footnotes
Authors Contribution
SR, YY, WD, CD, and LL conducted and collected the study. SR, WD, YY, HW, YF and PS analyzed the data and contributed to the discussion. SR, DGA, and WD wrote the manuscript, contributed to the discussion, reviewed, and edited the manuscript. All authors read and approved the final manuscript. WD and LL took full responsibility for this study.
Ethical Approval
The ethical approval to report this work was obtained from the Ethical Committee Board of Southwest Hospital.
Statement of Human and Animal Rights
All procedures in this study were conducted following the protocol approved by “the Ethical Committee Board of Southwest Hospital” and complied with the recommendations of the Declaration of Helsinki.
Statement of Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Natural Science Foundation of Chongqing Municipal Science and Technology Bureau (cstc2018jcyjAX0335; cstc2020jcyj-msxmX0298); the Fundamental Research Funds of Central Universities at Chongqing University (2019CDYGYB020) awarded to Dr. Wuquan Deng. This work was funded by the Natural Science Foundation of Science, and Basic Research and Frontier Exploration of Science and Technology Commission by Chongqing Municipality (Grant No.cstc2018jcyjAX0788) awarded to Dr.Yan Chen. This study is partially supported by National Institutes of Health, National Institute of Diabetes and Digestive and Kidney Diseases Award Number 1R01124789-01A1 awarded to Prof. DG Armstrong.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
