Abstract
Inhalation is an important route of aldehyde exposure, and lung is one of the main targets of aldehyde toxicity. Octanal is distributed ubiquitously in the environment and is a component of indoor air pollutants. We investigated whether octanal exposure enhances the inflammatory response in the human respiratory system by increasing the expression and release of cytokines and chemokines. The effect of octanal in transcriptomic modulation was assessed in the human alveolar epithelial cell line A549 using oligonucleotide arrays. We identified a set of genes differentially expressed upon octanal exposure that may be useful for monitoring octanal pulmonary toxicity. These genes were classified according to the Gene Ontology functional category and Kyoto Encyclopedia of Genes and Genomes analysis to explore the biological processes related to octanal-induced pulmonary toxicity. The results show that octanal affects the expression of several chemokines and inflammatory cytokines and increases the levels of interleukin 6 (IL-6) and IL-8 released. In conclusion, octanal exposure modulates the expression of cytokines and chemokines important in the development of lung injury and disease. This suggests that inflammation contributes to octanal-induced lung damage and that the inflammatory genes expressed should be studied in detail, thereby laying the groundwork for future biomonitoring studies.
Introduction
Air pollution adversely affects the respiratory tract, causing symptoms of bronchial hyperactivity and asthma, while the health effects of indoor air pollutants have not been investigated extensively. 1 This is a critical topic in lung health since the indoor–outdoor air exchange rates are being decreased to increase the energy efficiency of buildings, which causes indoor accumulation of volatile organic compounds (VOCs).
Aldehydes are VOCs that are associated with health risks. Aldehydes are emitted from building materials, furnishings, paints, varnishes, waxes, solvents, detergents, cleaning products, cooking oil, and fried foods. 2 –4 Despite their usefulness, aldehydes can cause lung and respiratory diseases, as well as exert mutagenic and carcinogenic effects. Aldehyde exposure leads to inflammation, oxidative stress and immune modulation in the airways and is associated with airway inflammatory disorders, such as asthma and chronic obstructive pulmonary disease (COPD). 5,6 Formaldehyde, the best-known aldehyde, has been linked to various types of toxicity, particularly inflammation. 7 Acrolein is associated with COPD 8 and is linked to cancer of the lung, 9 and acetaldehyde causes inhalation toxicity. 10 Despite intensive investigation of the mechanisms underlying the effects of these aldehydes on inhalation and pulmonary toxicity over several decades, toxicity data are limited primarily to formaldehyde. Insufficient information is available regarding toxicities of other aldehydes, such as low-molecular weight saturated aliphatic aldehyde (LSAA, e.g. propanal, pentanal, hexanal, heptanal, and octanal), which are distributed ubiquitously in the environment and are components of indoor air pollutants. 2 –4,11 –15
The unsaturated aldehydes are particularly irritating. The mucus membranes of the nasal and oral passages and the upper respiratory tract are affected, producing a burning sensation, an increased ventilation rate, bronchial constriction, choking, and coughing. 16 Among these aldehydes, in the current study, we focused on octyl aldehyde (octanal), which is an aldehyde with the chemical formula C8H16O. It is used commercially as a component in perfumes and as a flavoring agent in the food industry. Occupational exposure to octanal may occur through inhalation and dermal contact with this compound at workplaces where octanal is produced or used. Monitoring data and use information indicate that the general population may be exposed to octanal via inhalation of ambient air, ingestion of food and drinking water, and dermal contact with products containing octanal. 17
Airway inflammation plays a central role in the pathophysiology of pulmonary disease. Exposure to environmental toxicants, injury, or infection induces the recruitment of inflammatory cells in the airways, which in turn results in the production of various cytokines, chemokines, proteases, and proinflammatory mediators, ultimately leading to increased oxidative stress, a protease/anti-protease imbalance and progressive lung tissue injury. 18 –20 Maintenance of the cytokine/chemokine microenvironment is critical for pulmonary homeostasis. Proinflammatory cytokines are produced predominantly by monocytes and macrophages; however, other cell types, such as epithelial cells and fibroblasts, can also produce such mediators. 21 –23 Many of these cytokines are important in the development of inflammation within the respiratory tract. Chemokines also play a key role in the recruitment of inflammatory cells to the lung from the circulation. 24 As several chemokines can activate a single receptor, their role is best described through their receptors, which are divided into those for CC chemokine receptors (CCRs) and those for CXC chemokine receptors (CXCRs).
Modulation of inflammatory cytokines, such as interleukin 6 (IL-6) and IL-11, has been demonstrated in formaldehyde-exposed human lung BEAS-2B cells and in the mouse respiratory system, potentially reflecting the effects of aldehydes on pulmonary inflammation by increasing inflammatory cytokines. 7,25 However, the LSAA-induced inflammatory responses in an in vitro system relevant to lung are largely unknown. Although saturated aldehydes are less toxic than unsaturated aldehydes such as formaldehyde, LSAA accumulation in indoor spaces causes discomfort and can have adverse effects on the health of residents, such as sensory irritation. 26 In this study, we focused on the pulmonary inflammatory responses induced by cytokines and chemokines following exposure of A549 cells to the LSAA octanal. This human lung adenocarcinoma cell line is derived from epithelial cells important in the development of lung injury 27,28 and is, therefore, a useful model for studying cytokine responses related to inflammation. Octanal-responsive genes in A549 cells were screened using microarray analysis, focusing on the expression of various cytokines and chemokines. To confirm that changes in cytokine expression are reflected in the protein levels, the level of IL-8 and IL-6 released were assessed. The results clarify the relationship between indoor pollution and inflammatory lung diseases by providing transcriptomic data and in vitro evidence.
Materials and methods
Chemicals and reagents
Octanal, dimethylsulfoxide (DMSO), and 3-(4,5-dimethylthaizol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) were purchased from Sigma-Aldrich Co. (St Louis, Missouri, USA). Culture media and buffer solutions were purchased from GIBCO™ (Big Cabin, Oklahoma, USA): Roswell Park Memorial Institute (RPMI) 1640, Dulbecco’s phosphate-buffered saline, fetal bovine serum (FBS), and antibiotics (penicillin and streptomycin). All chemicals used were of analytical grade or the highest grade available.
Cell culture
The human adenocarcinoma lung epithelial cell line, A549, was purchased from Korea Cell Line Bank (Seoul, Korea) and was cultured under conditions of a humidified atmosphere of 5% carbon dioxide and 95% air at 37°C. The culture medium was RPMI 1640 (Gibro-BRL, Grand Island, New York, USA) added 10% FBS, sodium bicarbonate (Sigma-Aldrich), 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) (Gibco), and penicillin/streptomycin (Gibco). The medium was renewed every 2–3 days.
Chemical treatment
The cells were treated with each compound at 0.1 and 1.0% of final solvent (i.e. DMSO) concentration in well plates. Due to the volatility of the compound, well plates were sealed with sealing films after chemical treatment. A549 cells were seeded at a density of 7 × 104 cells/mL per well in 500-μL media for MTT assay and seeded in a 6-well plate at a density of 25 × 104 cells/mL for RNA extraction. After incubation for 24 h at 37°C, the cells were treated with octanal for 48 h.
Determination of cell viability
To determine the cytotoxicity and effects on cell growth of octanal, the MTT cell proliferation assay was performed using the modifications described by Mosmann. 29
Microarray analysis and data processing
Messenger RNA (mRNA) expression analysis was conducted on the RNA samples using 4 × 44 K whole human genome microarray (Agilent Technologies, Santa Clara, California, USA). For control and test RNAs, the synthesis of target complementary RNA (cRNA) probes and hybridization were performed using low RNA input linear amplification kit (Agilent Technologies) according to the manufacturer’s instructions. Labeled cRNA target was quantified using a spectrophotometer (NanoDrop-1000, Wilmington, Delaware, USA). After checking labeling efficiency, fragmentation of cRNA was performed by adding 10× blocking agent and 25× fragmentation buffer and incubating at 60°C for 30 min. The fragmented cRNA was resuspended with 2× hybridization buffer and directly pipetted onto assembled Agilent’s Human Oligo Microarray (4 × 44 K). The arrays hybridized at 65°C for 17 h using hybridization oven (Agilent Technologies). The hybridized microarrays were washed as the manufacturer’s washing protocol (Agilent Technologies). The hybridized images were scanned using DNA microarray scanner and quantified using feature extraction software (Agilent Technologies).
After log2 transform, interarray normalization of expression levels was performed with quantiles for gene expression profiling to correct possible experimental distortions. Normalization function was applied to expression data of all experiments and then values of spot replicates within arrays were averaged. Furthermore, feature extraction software provides spot quality measures in order to evaluate the goodness and the reliability of hybridization. In particular flag, “glsFound” (set to 1 if the spot has an intensity value significantly different from the local background, 0 otherwise) was used to filter out unreliable probes: flag equal to 0 will be noted as “not available (NA)”. So, in order to make more robust and unbiased statistical analysis, probes with a high proportion of “NA” values were removed from the dataset.
GO category and pathway analysis
In order to classify the selected genes into groups with a similar pattern of expression, each gene was assigned to an appropriate category according to its main cellular function. To determine significantly overrepresented gene ontology (GO) analysis, the DAVID (http://david.abcc.ncifcrf.gov/home.jsp) functional annotation clustering tool was used by choosing the default option. 30 Pathway analysis was used to determine significant pathways for differentially expressed genes using microarray gene pathway annotations downloaded from Kyoto Encyclopedia of Genes and Genomes (KEGG) (http://www.genome.jp/kegg/). A Fisher exact test was used to find significant enrichment for pathways and the resulting p values were adjusted using the Benjamini–Hochberg False Discovery Rate (FDR) algorithm, 31 ; pathway categories with an FDR < 0.05 were reported.
Quantitative real-time RT PCR
mRNA expression levels for the genes of interest were analyzed via quantitative real-time reverse transcriptase polymerase chain reaction (RT-PCR) using an iCycler system (Bio-Rad Laboratories, Hercules, California, USA). Total RNA was reverse transcribed into cDNA using an Omniscript RT kit (Qiagen, Venlo, Limburg, Netherlands). Primer specificity was tested by performing PCR for 40 cycles (95°C for 20 s and 60°C for 1 min). Real-time RT-PCR was performed using an SYBR Supermix Kit (Bio-Rad Laboratories). Samples were subjected to 45 cycles at 95°C for 20 s and 60°C for 1 min. PCR efficiency was determined by running serial dilutions of template cDNA, and melting curve data were collected to ensure PCR specificity. Each cDNA sample was analyzed in triplicate and a corresponding no-RT mRNA sample was included as a negative control. A glyceraldehyde-3-phosphate dehydrogenase (GAPDH) primer was included in every plate as an internal loading control. The mRNA level of each sample for each gene was normalized against that of GAPDH mRNA. The relative mRNA level was determined as
Primer sequences for the quantitative real time RT-PCR used in this study.

RT-PCR: reverse transcriptase polymerase chain reaction.
ELISA for cytokines (IL-6 and IL-8)
The IL-6 and IL-8 concentration in A549 cell supernatants were measured using a commercially available Quantikine human IL-6 and Quantikine human CXCL6/IL-8 enzyme-linked immunosorbent assay (ELISA) kit (R&D Systems, Minneapolis, Minnesota, USA), according to the manufacturer’s protocol. All samples were assayed in duplicate and measured with a FLUOstar OPTIMA (BMG Labtech, Ortenberg, Germany).
Statistical analysis
The data were analyzed with SigmaStat (SPSS Inc., Chicago, Illinois, USA) using one-way analysis of variance followed by Dunnett’s method. The treatment groups were compared individually with the control group; p < 0.05 was considered significant.
Results
Octanal cytotoxicity in A549 cells
The relative survival of A549 cells following exposure to a range of octanal concentrations was determined using MTT assay. The survival percentage, relative to the DMSO solvent control, was determined as a percentage of the control optical density, measured after treatment. A dose-dependent A549 cell viability curve was obtained after a 48-h octanal exposure. Based on the results of the MTT assay, cell viability inhibitory concentrations (IC) of octanal at 5 (IC5 = 0.15 mM), 10 (IC10 = 0.33 mM), and 20% (IC20 = 0.58 mM) were calculated. The IC20 value was used to select genes less affected by cell cytotoxicity and to identify an appropriate number using microarrays.
Octanal regulates cellular mRNA transcript levels
A549 cells were treated with the IC20 concentration of octanal (0.58 mM) for 48 h, and total RNA was subjected to microarray analysis. Gene expression changes were analyzed by comparing the treated and control groups using statistical criteria of ≥1.5-fold changes with the value of p < 0.05 using three independent samples.
We initially evaluated the molecular profile after exposure to octanal. Generally, transcriptomic regulation occurs through dynamic intracellular responses to multiple, apparently unrelated, stimuli and stresses; thus, characteristic gene expression profiles reflect these stimuli. 32 To investigate whether a large-scale gene expression profile in a cellular system reflects ectopic stress, we performed an unsupervised hierarchical clustering analysis. Expression profiling demonstrated that the octanal-treated group was more sensitive to gene expression changes than the vehicle control group. Octanal exposure resulted in upregulation of 408 genes and downregulation of 291 genes; no such effects have to our knowledge been reported previously. These genes may be useful for monitoring octanal-induced pulmonary toxicity. The relative expression levels of these genes are illustrated as a heat map in Figure 1. Of the 699 differentially expressed genes, 687 (403 upregulated and 284 downregulated) were annotated with a gene name and subjected to further analysis (Table SI1).

Hierarchical clustering of significantly expressed genes in A549 cells exposed to an IC20 dose of octanal. Rows represent mRNA expression profiles indicating the octanal exposure. Columns represent probe measurements. Color saturation reflects differences in expression between sample RNA and reference RNA. IC: inhibitory concentration.
To identify the mechanism underlying octanal-induced pulmonary toxicity, differentially expressed genes were classified according to the GO terms of biological processes. Expression analysis systematic explorer was performed using http://david.abcc.ncifcrf.gov/ to identify the biological processes significantly overrepresented in the gene lists and to identify any biological themes in the response to octanal. As shown in Figure 2, the differentially expressed genes in octanal-treated cells included those involved in the inflammatory response, immune response, cell proliferation, chemical homeostasis, apoptosis, and chemotaxis (Benjamini test, p < 0.05). Also, according to the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, most of the genes were related to cytokine–cytokine receptor interaction and the mitogen-activated protein kinase (MAPK) signaling pathway (Table 2). Recently, our research group reported that octanal modulates MAPK signaling-related genes and stimulates p38 MAPK phosphorylation in A549 cells. 33 Thus, in the present study, we focused our analysis on the 31 genes involved in cytokine–cytokine receptor interaction (Figure 3; KEGG figure). These genes are CCL11, CCL14, CCL15, CCL19, CCL21, CCL23, CCL25, CCL3L3, CCL4, CCL8, CXCL13, CXCL9, CSF3, IFNG, IL1R2, IL1A, IL1B, IL17A, IL2RA, IL25, IL6ST, IL6, IL8, TNFRSF12A, TNFRSF14, TNFRSF1B, TNFRSF25, BMP2, CXCR4, TNFSF4, and TNFRSF11B.

Identification of overrepresented annotation terms of genes differentially expressed in octanal-exposed A549 cells. Gene-term enrichment analysis based on GO categories–biological process was carried out on the differentially expressed genes using the DAVID functional annotation chart tool. Biological processes with an EASE score < 0.05 are shown. GO: Gene Ontology; EASE: expression analysis systematic explorer.

Differentially expressed genes associated with the cytokine–cytokine receptor interaction pathway. This figure was adapted from: http://www.genome.jp/kegg/pathway.html.
Octanal-altered genes related to cytokine–cytokine receptor interaction and the MAPK) signaling pathway.

MAPK: mitogen-activated protein kinase; IL: interleukin; TAO: thousand-and-one amino acid; TNF: tumor necrosis factor.
Validation by quantitative real-time RT-PCR
Quantitative real-time RT-PCR (qRT-PCR) was performed on the nine genes involved in the cytokine–cytokine receptor interaction: CCL15, CCL4, CSF3, IL1A, IL1B, IL6, IL8, TNFRST12A, and BMP2, using the fluorescent dye SYBR green. GAPDH was used as an endogenous control for all genes, and similar data were obtained from all samples. As shown in Figure 4, the expression of these genes corresponded to the microarray data. The correlation coefficient (r 2 ) for the microarray and qRT-PCR data was 0.96.
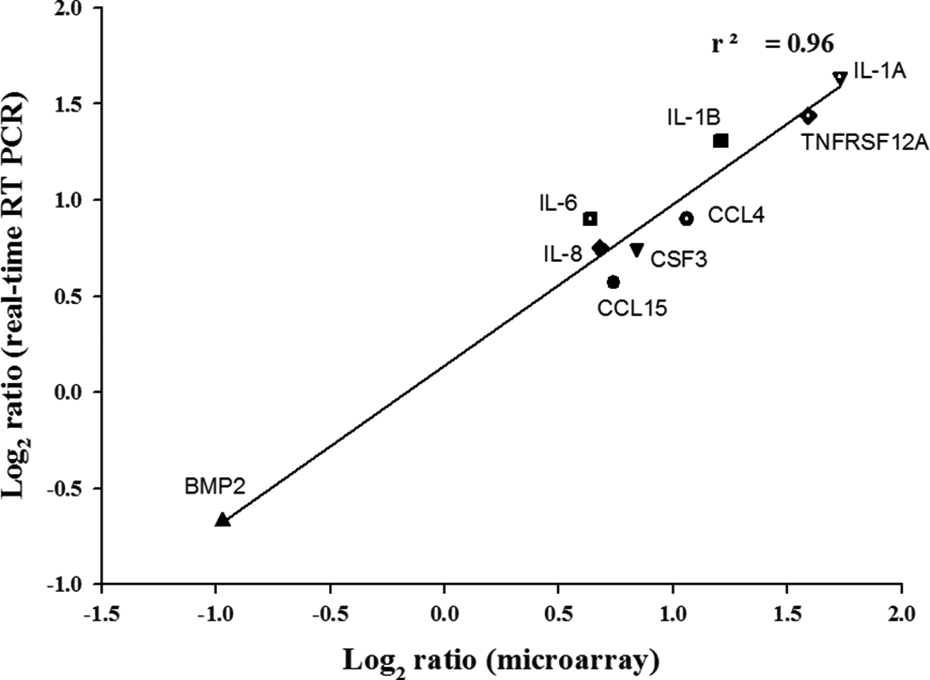
Comparison of the DNA microarray and quantitative real-time RT-PCR data. After 48 h treatment with IC20 dose of octanal, total RNA was isolated and microarray and quantitative real time RT-PCR was performed. The fold change in the mean mRNA ratio for differentially expressed genes related to the cytokine–cytokine receptor interaction pathway was log2 transformed and plotted as microarray versus real-time RT-PCR (r 2 , correlation coefficient). RT-PCR: reverse transcriptase polymerase chain reaction; IC: inhibitory concentration; mRNA: messenger RNA.
Octanal induces IL-6 and IL-8 secretion
IL-6 and IL-8 either initiate the inflammatory cascade in the lungs or that they are early markers of the inflammatory process. 34 Prior to this study, however, no investigation of the impact of other ubiquitous aldehydes, such as octanal, on inflammatory cytokine secretion has been published. We performed an ELISA to determine the effect of octanal on cytokine secretion. We assessed four groups to identify the effect of octanal concentration; the vehicle control group, and the low- (IC5, 0.15 mM), medium-, (IC10, 0.33 mM), and high- (IC20, 0.58 mM) octanal concentration groups. Although IC5 value of octanal exposure did not show significant effect on IL-6 secretion, other doses of octanal significantly increased IL-6 and IL-8 secretion compared to controls (Figure 5). This supports the hypothesis that an increase in these cytokines contributes to the octanal-induced inflammatory response.

Effects of octanal on IL-6 and IL-8 secretion by A549 cells. Cells were stimulated with three doses of octanal (IC5, IC10, and IC20). IL-6 and IL-8 concentrations in cell culture supernatants after 48 h were assayed by ELISA. The results are expressed as means ± SD of three independent experiments. *p < 0.05. IL: interleukin; IC: inhibitory concentration; ELISA: enzyme-linked immunosorbent assay.
Discussion
Characterization of the relevant toxicity pathways and the identification of biomarkers of key event parameters that can be monitored in human studies of chemical exposure are required. The combination of these data with distributional data on population characteristics of exposure and dose (magnitude, frequency, and duration) would provide a scientifically based approach for reducing the uncertainties associated with current risk assessments. The utilization of existing human data from epidemiological studies and clinical trials to retrospectively and prospectively demonstrate that the approach successfully and adequately predicts human toxicological responses was proposed. 35 Also, data from genome-wide association studies are envisioned to provide additional support for the pathway-based models.
Toxicogenomic study has been widely used to characterize toxicological properties of disease such as cancer. Through previous studies in our laboratory, we confirmed toxicogenomic effects of several hazardous chemicals involved in inducing cancer or other diseases. 32,36 The aim of this study was to identify genes involved in the biological process in order to elucidate the cellular response to the well-known air pollutant aldehyde, in mammalian cells, using toxicogenomic technology.
The lung is a key target of aldehyde toxicity, and pulmonary inflammation contributes to the development of many lung diseases. In the present study, we show that octanal affects the expression of several cytokines and chemokines by A549 cells. We exposed A549 cells to octanal using an in vitro system. Although we recognize that A549 cells may not completely mimic the normal lung cell response, there are several advantages in using these cells for air toxin studies. Epithelial cells are present throughout the respiratory tract as nasal, tracheal, bronchial, and alveolar cells. Cell lines of all pulmonary cell types have been developed, 37 but the classical model in respiratory toxicology is the unique human alveolar cell line. In particular, a large majority of in vitro studies have determined the effects of environmental pollutants on the lung using these cells. 38,39
The airway epithelium is at the interface between the environment and the respiratory tract. Maintenance of its integrity is therefore of prime importance in the defense of the lung against atmospheric pollutants. Previous study by Kastner et al. showed that a very short-term exposure to aldehyde resulted in a limited effect on cell death in bronchial epithelial cell lines. 40 They suggest that long-term exposure to high concentrations of aldehyde—at least 200 times higher than the physiological concentrations can lead to a loss of airway epithelium integrity. Persoz et al. reported a limited loss of viability only, and Pariselli et al. did not observe any lactate dehydrogenase release in airway epithelial cells exposed for a very short period of time (30 min and 1 h, respectively) to concentration classically found indoors and concentration far beyond the ones found indoors aldehyde, respectively. 41,42 The exposure conditions used in our current experiment are based on long-term (48 h) and subcytotoxic dose levels (>80% cell viability). Octanal concentrations were considered appropriate for microarray analysis due to the fact that such an exposure level gives meaningful outcomes in in vitro toxicogenomic studies on carcinogenesis and other toxicity in our research group. 36,43
Our results focused on microarray data, which indicated differential expression of inflammatory response genes after octanal exposure. Aldehyde-induced alterations in stress-responsive genes in the human respiratory system have been studied, 44 but the data regarding changes in cytokine expression are limited. Actually, our research group recently examined global transcriptomic responses upon seven LSAA compounds exposure (propanal, butanal, pentanal, hexanal, heptanal, octanal, and nonanal) in A549 cells to identify significant molecular signature for aldehyde toxicity. Although all these compounds altered many types of genes involved in pulmonary toxicity, only octanal altered a large number of inflammatory response genes that reveal the impact on cytokine secretion.
In this study, the expression of 31 cytokines and chemokines were dysregulated more than 1.5-fold by exposure to an IC20 dose of octanal. Extensive knowledge of these genes has impacted many inflammation-associated diseases. Among these, IL-1, IL-6, and IL-8/CXCL8 are likely the most important chemotactic factors in the lung. 27,45 These proinflammatory cytokines are synthesized by various inflammatory and pulmonary cells upon stimulation by hyperoxia, microorganisms, endotoxin, other bacterial cell wall constituents, and biophysical factors, such as volutrauma and barotrauma. 46 They play a crucial role in the initiation and evolution of the inflammatory response. 47
The IL-1 gene family is a group of cytokines involved in a variety of acute and chronic lung diseases. Several studies have described the important role of IL-1 in acute lung injury. 48,49 Furthermore, IL-1 induces the production of other proinflammatory cytokines. In the present study, we found significantly increased mRNA and protein levels of IL-6 and IL-8 in octanal-exposed A549 cells, which likely act in concert with IL-1 to perpetuate inflammation and subsequent events.
IL-6 and IL-8 play central roles in inflammatory responses and pulmonary disease. Serum IL-6 levels are elevated in a number of inflammatory diseases. 50 Increased IL-6 gene expression was identified in epithelial cells from bronchial biopsy specimens of asthmatic subjects, and IL-6 levels were increased in the bronchoalveolar lavage fluid of symptomatic asthmatic subjects. 51 Several lung pathologies are associated with enhanced IL-8 levels. 46 For example, elevated IL-8 levels are present in the lung of COPD and cystic fibrosis patients. 27,52 The importance of IL-8 in lung inflammatory processes has been reported, as IL-8 antibodies partly inhibited inflammation induced by toxic agents. 53 Increased IL-8 protein levels after exposure to formaldehyde in lung cells have been reported. 54 This suggests that IL-8-signaling-mediated inflammatory responses in lung cells exposed to aldehydes are altered. Interestingly, IL-8 levels are also increased in formaldehyde-exposed lung cells after presensitization to tumor necrosis factor α (TNF-α). 41 TNF-α is a proinflammatory mediator, the levels of which are increased upon exposure to formaldehyde. 55 However, few investigations have addressed the effects of LSAA exposure on cytokine secretion and the inflammatory response.
BMP2, CXCR4, TNFSF4, and TNFRSF11B were among the other cytokines downregulated in our microarray data. Bone morphogenetic proteins (BMPs) directly modulate the pathophysiology of various inflammatory/fibrotic conditions. The composition of BMP ligands produced in allergen-challenged animals is markedly altered during the progression of airway inflammation. 56 Recent studies have reported increased levels of CXCL12 in injured lungs and increased levels of its receptor, CXCR4, on accumulated neutrophils; furthermore, these studies showed that the CXCL12/CXCR4 signaling pathway participates in neutrophil accumulation during lung injury. 57 We did not evaluate CXCL12 expression in the A549 cells used in this study, but expression of the CXCR4 gene was downregulated. The effect of such downregulation of cytokine expression on octanal-induced lung toxicity is difficult to interpret, since the inflammatory outcome is dependent on the total cytokine network.
Although many studies have investigated the cellular mechanisms that mediate the toxic effects of aldehydes, the toxicity data have to date been limited to the effects of formaldehyde; few studies focused on the involvement of LSAAs in inflammatory processes, particularly lung toxicity. We provide evidence that inflammatory mediator expression is a mechanism underlying the cellular effects of octanal. We identified a set of 31 inflammatory mediators dysregulated in human lung cells exposed to octanal. Further studies on computational systems biology pathway models and in vitro to in vivo extrapolation are necessary to evaluate dose–response relationships and low-dose risks in biologically relevant in vitro assays. Although our data are insufficient to identify the mechanism(s) underlying octanal toxicity, we provide here a valuable mechanistic overview that will facilitate future investigations and a follow-up of functional analysis. In future, we will investigate whether the expression levels of these inflammatory mediators can serve as biomarkers of aldehyde exposure in humans. Such biomarkers can be used to monitor human exposure to environmental toxicants and elucidate their relationships with health effects.
Footnotes
Conflict of interest
The authors declared no conflicts of interest.
Funding
This work was supported by the Korea Research Foundation grants from Korea Ministry of Environment as “The Ecoinnovation Project (412-111-010)” and Korea Institute of Science and Technology Program, Republic of Korea, to J.C.R.
