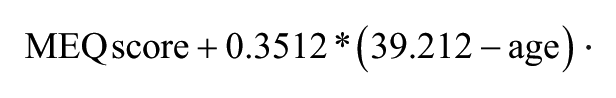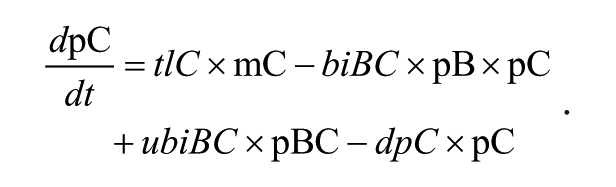Abstract
Anxiety and other mood disorders, such as major depressive disorder (MDD) and seasonal affective disorder (SAD), affect nearly one-fifth of the global population and disproportionately affect young adults. Individuals affected by mood disorders are frequently plagued by sleep and circadian problems, and recent genetic studies provide ample support for the association of circadian and sleep syndromes with depression and anxiety. Mathematical modeling has been crucial in understanding some of the essential features of the mammalian circadian clock and is now a vital tool for dissecting how circadian genes regulate the molecular mechanisms that influence mood. Here, we model the effect of five clock gene polymorphisms, previously linked to mood disorders, on circadian gene expression and, ultimately, on the period length and amplitude of the clock, two parameters that dictate the phase, or alignment, of the clock relative to the environment. We then test whether these gene variants are associated with circadian phenotypes (Horne-Ostberg Morningness-Eveningness scores) and well-established measures of depression (Beck Depression Inventory) and anxiety (State-Trait Anxiety Inventory) in a population of undergraduates (n = 546). In this population, we find significant allelic and/or genotypic associations between CRY2 and two PER3 variants and diurnal preference. The PER3 length polymorphism (rs57875989) was significantly associated with depression in this sample, and individuals homozygous for the PER3 single nucleotide polymorphism (SNP) (rs228697) reported significantly higher anxiety. Our simple model satisfies available experimental knockdown conditions as well as existing data on clock polymorphisms associated with mood. In addition, our model enables us to predict circadian phenotypes (e.g., altered period length, amplitude) associated with mood disorders in order to identify critical effects of clock gene mutations on CRY/BMAL binding and to predict that the intronic SNPs studied represent gain-of-function mutations, causing increased transcription rate. Given the user-friendly structure of our model, we anticipate that it will be useful for further study of the relationships among clock polymorphisms, circadian misalignment, and mood disorders.
Anxiety and mood disorders, such as major depressive disorder (MDD) and seasonal affective disorder (SAD), affect nearly 20% of the global population (Steel et al., 2014). In addition to exacting personal costs, such disorders have a substantial economic cost, as do sleep and circadian disorders (Fineberg et al., 2013; Hafner et al., 2017). Further understanding of these disorders may allow us to provide adequate therapy, dramatically improving quality of life for many individuals.
Recent studies highlight numerous and diverse associations between genes involved in the circadian clock and mood disorder phenotypes (Antypa et al., 2015; Kim et al., 2015; McCarthy and Welsh, 2012; Soria et al., 2010; Zhang et al., 2016). Large genome-wide association studies (GWAS) link multiple circadian genes with depression and other psychological disorders (McCarthy and Welsh, 2012; McCarthy et al., 2012; Soria et al., 2010; Terracciano et al., 2010). Experimental data from studies of individuals with severe circadian disorders, like those with Familial Advanced Sleep Phase (FASP) and Delayed Sleep Phase (DSP) phenotypes, support the high frequency of comorbidity between circadian disorders and negative affect (Courtet and Olié, 2012; Lee et al., 2011; Mendlewicz, 2009; Robillard et al., 2013; Sivertsen et al., 2015; Zhang et al., 2016). Murray et al. (2017) used melatonin profiles to disentangle circadian factors from sleep deprivation and other confounding factors in individuals with DSP phenotypes. Their results suggest that depression is closely linked to circadian misalignment (the degree to which the circadian phase of an individual is advanced or delayed relative to his or her social environment or sleep-wake cycle). Furthermore, studies on self-reported diurnal preference, the behavioral phenotype associated with advanced (morningness) or delayed (eveningness) circadian phase, have shown that individuals with extreme types are more likely to show symptoms of depression and/or anxiety (Kitamura et al., 2010; Levandovski et al., 2011; Antypa et al., 2015; Fares et al., 2015). Not surprisingly, a number of circadian clock polymorphisms are also associated with the diurnal preference (Archer et al., 2003; Hida et al., 2014; Kim et al., 2015; Lavebratt et al., 2010a, 2010b; Partonen et al., 2007; Soria et al., 2010; Zhang et al., 2016). Overall, the results from these studies provide support for the hypothesis that circadian misalignment, where the phase of the intrinsic clock is not synchronized with environmental cues, influences negative affect.
One possible framework for understanding the connections between the circadian clock and human mood is becoming clear (Figure 1). There is strong evidence linking circadian gene polymorphisms with extreme diurnal preference as well as with depression. Circadian misalignment is associated with both depression and extreme diurnal preference. The most parsimonious hypothesis is that polymorphisms in circadian clock genes alter the period length or amplitude of the clock oscillations, thus causing an advance or delay in clock phase that results in circadian misalignment. The desynchrony of clock oscillations with environmental cues disrupts the downstream signals regulating behavioral phenotypes, including sleep-wake pathways (diurnal preference and/or chronotype) (Archer and Oster, 2015; Cuesta et al., 2017; Fuller et al., 2006), hormone regulatory pathways (mood) (Bedrosian et al., 2016; Ben-Hamo et al., 2016; Salgado-Delgado et al., 2011), and likely many others. There are multiple potential pathways by which circadian misalignment may affect mood by altering cellular processes in the brain and in peripheral tissues (McClung, 2013).
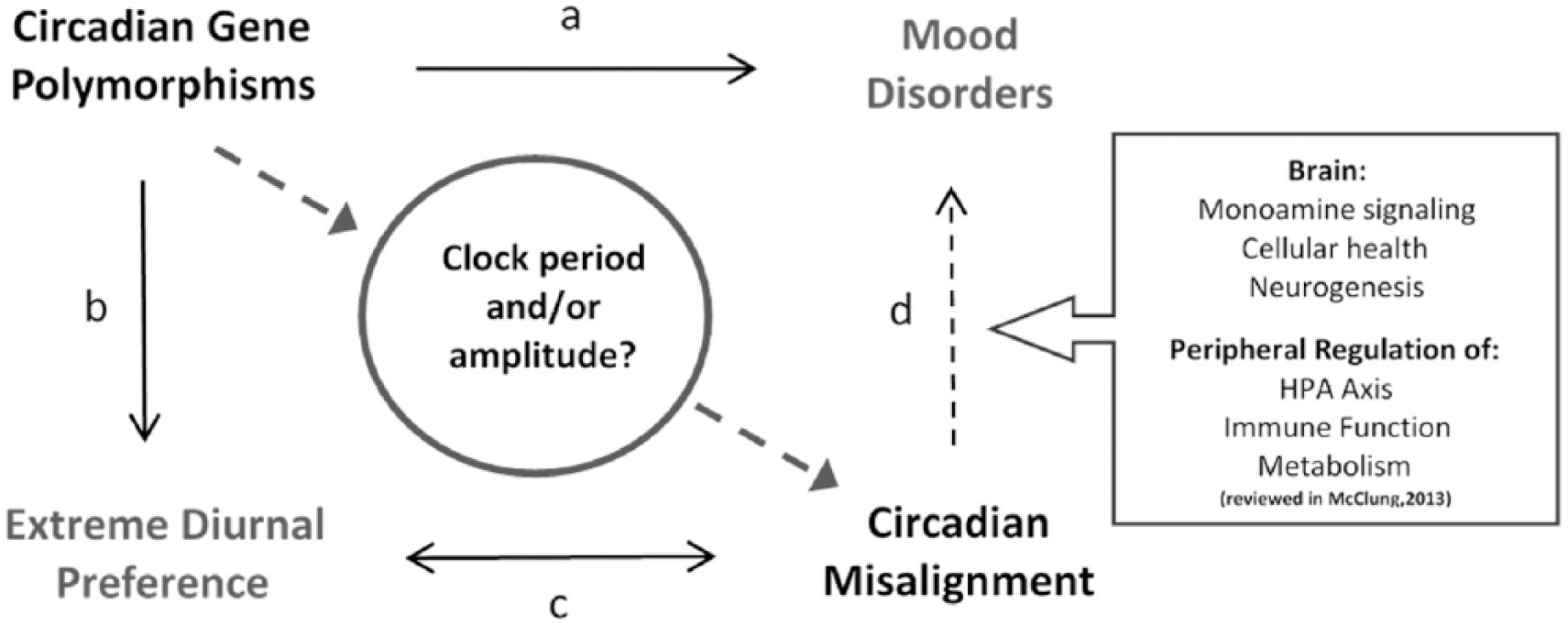
A framework for understanding the connections between the circadian clock, diurnal preference, and human mood. Strong associative evidence links circadian gene polymorphisms with both (a) mood disorders and (b) extreme diurnal preference. Circadian misalignment is also associated with both (c) extreme diurnal preference and (d) mood disorders. One hypothesis is that polymorphisms in circadian clock genes alter the period length and/or amplitude of the clock oscillations, thus causing an advance or a delay in clock phase that results in circadian misalignment. The desynchrony of clock oscillations with environmental cues disrupts the downstream signals regulating behavioral phenotypes, including sleep-wake pathways (diurnal preference and/or chronotype) and hormone regulatory pathways (mood). There are multiple potential pathways by which circadian misalignment may affect mood by altering cellular processes in the brain and in peripheral tissues (black box) (reviewed in McClung, 2013). To build a more complete picture, we modeled the molecular mechanisms responsible for how single-nucleotide changes in clock genes cause circadian misalignment (black circle). Black arrows are evidenced-based links; dashed lines represent outstanding questions in the field.
To build a more complete picture, we take a mathematical modeling approach to understand the molecular mechanisms responsible for how single-nucleotide changes in clock genes cause circadian misalignment, a missing link between clock polymorphisms and mood disorder phenotypes in humans. Mathematical modeling has been crucial in understanding some of the essential features of the mammalian circadian clock gene network and its associated mutations (Bordyugov et al., 2013), motivating novel experiments to validate model predictions. For example, a model developed by Forger and Peskin (2003) contradicted earlier understanding of the role of the casein kinase I epsilon mutation, CKItau, in vivo, and later experimental findings verified the model’s predictions (Gallego et al., 2006; Virshup et al., 2007). Meanwhile, a circadian clock model developed by Becker-Weimann et al. (2004) was later used to explain why increased stabilization of PER2 leads to a disruption of rhythmicity in mice (Reischl et al., 2007).
We have developed a simple deterministic differential equation model to explain how polymorphisms in clock genes might influence circadian gene expression and changes in clock period and amplitude, two circadian parameters that dictate circadian alignment. Our previous, more comprehensive model considered only the role of period length and did not include light as a parameter (Liberman et al., 2017). In this model, we simplified the molecular network and incorporated a seasonal light parameter, allowing us to test for the effects of amplitude in circadian gene expression and to consider the role of amplitude in SAD conditions.
To test the predictions of our model, we selected five polymorphisms from four genes previously linked to anxiety, depression, and/or SAD: PER2, PER3, CRY1, and CRY2 (Archer et al., 2003; Hida et al., 2014; Lavebratt et al., 2010a, 2010b; Soria et al., 2010). We investigated the relationships between the frequency of these polymorphisms, diurnal preference, and depression and anxiety in a population of undergraduates, where depression is likely to be widespread due to cofactors such as alcohol abuse, social jetlag, and high levels of stress. We measured depression using the Beck Depression Inventory (BDI), a widely used self-report instrument of depression severity (Beck and Beamesderfer, 1974). We measured dispositional (trait) anxiety using part of the State-Trait Anxiety Inventory (STAI) (Spielberger et al., 1983). The BDI and STAI are well-supported instruments for quantifying levels of negative affect. We tested the hypothesis that depression and anxiety are associated with allelic or genotypic variants of the clock genes. To explore how negative mood intersects with diurnal preference, we measured individual scores on the Horne-Ostberg Morningness-Eveningness Questionnaire (MEQ), a self-assessment of diurnal preference (Horne and Ostberg, 1976).
Of the 5 polymorphisms tested, two are linked to alterations in phosphorylation and degradation rates, respectively (Archer et al., 2003; Hida et al., 2014), while 3 are intronic single nucleotide polymorphisms (SNPs) (Lavebratt et al., 2010a, 2010b; Soria et al., 2010). By altering rates of phosphorylation or degradation, or by altering circadian gene transcription within the model consistent with the proposed roles of intronic SNPs (Chorev and Carmel, 2012), we test two potential mechanisms for circadian influence on mood disorder: shifts in phase of oscillating genes and decreased amplitude in circadian oscillations (Li et al., 2013; Wehr et al., 2001). Additionally, by changing light intensity in the winter, we test whether the model predicts decreased amplitude in clock gene oscillations in the winter, consistent with seasonal changes in circadian gene expression (Wehr et al., 2001).
Methods
Experimental Data Collection
Students from the Colgate University undergraduate program volunteered to participate in the study (n = 528; 190 males, 338 females, age range 18-20 years). All methods adhered to the principles of the Declaration of Helsinki; the Institutional Review Board at Colgate University approved all procedures and consent forms (#FR-F13-07, #ER-F14-12, #F15-13, #ER-F16-19). All participants gave written informed consent.
Each participant completed a computer-based survey that included the STAI, BDI, and Horne-Ostberg MEQ (Beck and Beamesderfer, 1974; Horne and Ostberg, 1976; Spielberger et al., 1983). MEQ scores, which range from 30 (extreme morning preference) to 70 (extreme evening preference), were age-adjusted using the following formula (Hida et al., 2014):
Individuals were assigned the following designations: MEQ scores 41 or lower were designated evening types (ET), MEQ scores 59 or higher were morning types (MT), and MEQ scores between 42 and 58 were neither type.
From the STAI, we measured the Trait Anxiety Scale, which evaluates aspects of “anxiety proneness,” including general states of calmness, confidence, and security. We averaged the STAI scores across genotype to test for an effect of genotype on anxiety. BDI scores were treated as quantized values; scores ranged from 0 to 63, with any score of 14 or above indicating the presence of mild to severe symptoms of depression.
Hair samples were collected from each participant to determine genotypes for PER3 SNP rs228697 and PER3 variable number of tandem repeats (VNTR) rs57875989. Following digestion of hair at 37 °C for 24 h, DNA was extracted and purified with the Qiagen DNAeasy Micro Kit. Genotyping for SNPs was performed using a TaqMan SNP Genotyping assay (Applied Biosystems, Foster City, CA) on an ABI 3700HT real-time qPCR instrument. Participants were identified as homozygous or heterozygous for the major and minor alleles.
To measure the VNTR length polymorphism of 54 base pairs in exon 18 of the PER3 gene, we used a fragment length analysis on an ABI 3100 sequencer. The following PCR primers were used with the forward primer fluorescently labeled with 6-FAM : forward, 5′-CAAAATTTTATGACACTACCAGAAT-GGCTGAC-3′, and reverse, 5′-AACCTTGTACTTCC-ACATCAGTGCCTGG-3′ (Ebisawa et al., 2001). The PCR was performed in a 25-µL volume using Qiagen PCR Mastermix. Positive and negative DNA controls were included with each PCR plate of samples. The PCR cycling conditions were 3 min at 94 °C, followed by 35 cycles of 45 sec at 94 °C, 45 sec at 58 °C, and 45 sec at 72 °C, with a final step at 72 °C for 3 min. PER3 alleles were separated by capillary electrophoresis and sized using ABI ROX standards. Participants were identified as PER34/4, PER34/5, or PER35/5.
Odds ratios were used to test differences in allele frequencies and frequency of depressed individuals (BDI ≥ 14) among genotypes. Chi-square tests were used to test for Hardy-Weinberg equilibrium in our population sample. One-way analyses of variance (ANOVAs) were used to test for differences in genotype frequencies between morning and evening types and for differences in STAI scores among VNTR and SNP genotypes.
Mathematical Model Structure
We have developed a deterministic differential equation model, in which each equation represents the rate of change of an mRNA, protein, or protein complex. The genes included in our model are PER1, PER2, PER3, CRY1/2, BMAL1, CLOCK/NPAS2, and REV-ERBα/β (Kondratov et al., 2006). As has been done in previous studies (Becker-Weimann et al., 2004; Kim and Forger, 2012; Relógio et al., 2011), we treat the gene pairs CLOCK and NPAS2, CRY1 and CRY2, and REV-ERBα and REV-ERBβ as one gene each for simplicity. Together, all modeled genes and their translated protein counterparts comprise 30 ordinary differential equations and 93 parameters.
We use the following naming convention: all mRNAs are represented with a lowercase “m,” while all proteins are represented with a lowercase “p.” PER1, PER2, and PER3 are written as P1, P2, and P3, respectively; CRY genes and proteins are indicated with “R,” BMAL1 with “B,” CLOCK with “C,” and REV-ERB genes and proteins with “E.” Additionally, activation by phosphorylation is represented with a lowercase “a.” For instance, the compound pP1a represents a phosphorylated PER1 protein molecule. A representative equation is given below:
In the above equation, CLOCK protein (pC) is generated by the translation (tlC) of Clock mRNA (mC). Since CLOCK can bind to BMAL1 (biBC), this would decrease the amount of available CLOCK protein, while unbinding from the BMAL1-CLOCK complex (ubiBC) would increase the amount of available CLOCK. Finally, the protein may degrade (dpC). All parameters, variables, and equations are described in full in Supplementary Tables S1, S2, and S3.
Mathematical Model Assumptions
To develop a simplified circadian model, we made several assumptions. Most prominently, the model ignores the effects of nuclear translocation and combines several circadian clock genes (CLOCK and NPAS2, CRY1 and CRY2, and REV-ERBα and REV-ERBβ). Other assumptions include the following:
The model assumes that the repressive effect of REV-ERBα/β is more important than the activatory effects of RORα/β/γ for BMAL1 transcription, an assumption made in a previous circadian clock model (Kim and Forger, 2012).
The model assumes that PER3 must be in complex with PER1 to be phosphorylated (Lee et al., 2004). The PER1-PER3 complex is able to bind to the BMAL1-CLOCK complex but does so less frequently than do the PER-CRY complexes (Kume et al., 1999).
CLOCK is constitutively expressed in the cell (Ko and Takahashi, 2006).
CRY inhibits BMAL1-CLOCK phosphorylation when not in complex with PER1 or PER2 (Matsumura et al., 2014; Tamaru et al., 2015).
Changes in light affect transcription of PER1 and PER2 (Hughes et al., 2015).
Numerical Simulation
Our deterministic model is solved numerically using ode15s, a built-in variable-step-size solver for stiff systems in MATLAB. To simulate a knockout mutation, we set the transcription rate of the corresponding clock gene to zero (see Suppl. Table S5). To simulate an SNP or a VNTR condition, we parameterized the appropriate biochemical rate, as described in Table 1.
Model representation of circadian gene variants.
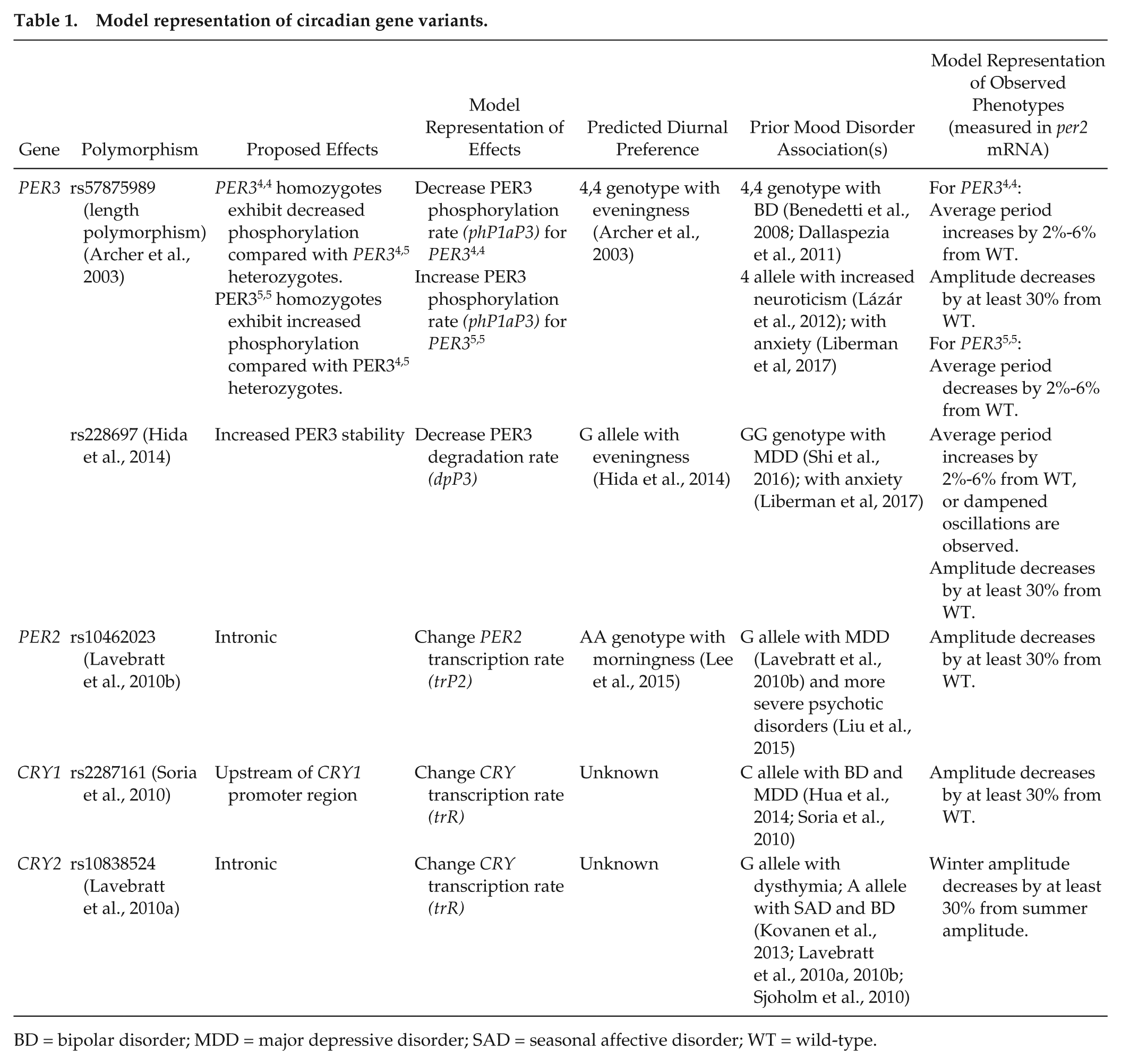
BD = bipolar disorder; MDD = major depressive disorder; SAD = seasonal affective disorder; WT = wild-type.
Oscillation Features and Model Conditions
The experimental observations used in this study were related to the amplitude and period of circadian gene transcript oscillations (Archer et al., 2003; Baggs et al., 2009; Hida et al., 2014; Kim et al., 2015; Lavebratt et al., 2010a, 2010b; Liu et al., 2007; Partonen et al., 2007; Ramanathan et al., 2014; Soria et al., 2010; Zhang et al., 2016). To calculate the amplitude of oscillations in our model, we used the average difference of peaks and troughs observed in Per2 mRNA oscillations from 48 to 168 h of simulation, since the system takes some time to stabilize. (In the case of PER2 knockdown, Bmal1 mRNA oscillations were used instead of Per2 mRNA.) Period was calculated as the average time difference between all peaks from 24 to 148 h of simulation. Both BMAL1 and PER2 are circadian clock genes known to be both essential and cyclical and therefore were used as markers of period (Cheng et al., 2007; Lee et al., 2004). The periods calculated from these 2 genes differed by less than 0.8% (mean ± 2 standard errors: 0.730% ± 0.037%). However, using other necessary clock genes did not appear to significantly change our findings; the maximum difference in period calculated from BMAL1, PER1, PER2, and CRY was less than 2% (1.51% ± 0.14%).
All polymorphism features were calculated using Per2 mRNA. Extreme morningness and advanced sleep phase (ASP) were each defined as a 2% to 6% decrease in average period. Extreme eveningness and delayed sleep phase (DSP) were each defined as a 2% to 6% increase in average period. MDD was represented as a decrease in amplitude of at least 30% from wild-type (Li et al., 2013), while SAD was represented as a decrease in amplitude of at least 30% occurring only in the winter, with a summer mutant amplitude within 15% of wild-type summer amplitude. Further details of the model conditions are specified in Supplementary Tables S4 and S5.
Parameter Estimation
Most of the reaction rates in the human circadian clock network have not been measured experimentally due to technical limitations. We used a parameter search algorithm to identify parameter values that could reproduce experimental observations. Experimental knockout conditions were obtained from studies performed in transgenic mice and in isolated human cells; all mouse conditions were chosen based on their resemblance to gene expression patterns in human cell lines (Suppl. Table S5) (Baggs et al., 2009; Liu et al., 2007; Ramanathan et al., 2014). All SNP and VNTR data were obtained from peer-reviewed sources linking human PER3 to specific variations in sleeping patterns and mood disorders (Archer et al., 2003; Hida et al., 2014; Lavebratt et al., 2010a, 2010b; Lázár et al., 2012; Soria et al., 2010). For a full list of experimental conditions used, see Table 1 and Supplementary Tables S4 and S5.
To identify points at which the system could oscillate, a search for Hopf bifurcations was performed. Hopf bifurcations indicate the point at which a system switches from stability to instability, at which point oscillations may be observed. Once sustained oscillations were confirmed, parameters were allowed to vary up to 1000% around Hopf points, and a global parameter search commenced to identify parameter sets that could satisfy all available experimental conditions.
Parameter search was performed using the stochastic ranking evolutionary strategy (SRES) algorithm (Runarsson and Yao, 2000), which looks for suitable parameter sets that demonstrate an approximately 24.2-h period and fit all experimental conditions described in Supplementary Tables S4 and S5. Previous research indicates that SRES performs better than other global parameter estimation algorithms, such as simulated annealing, in large-scale biological systems (Fakhouri et al., 2010; Moles et al., 2003). To produce the biologically feasible parameter ranges described in Supplementary Table S1, parameter ranges were refined based on the results of initial parameter searches.
Coding
The codes for the study were implemented in MATLAB R2017a (with the Parallel Processing Toolbox) because of the program’s user-friendliness and advanced data processing features. Each simulation took less than 1 min (performed on a desktop computer with 32 gigabytes of RAM). Each SRES parameter estimation run with 2000 generations, 15 parents, and a population size of 100 took approximately 8 h (using 2 nodes of a computer cluster containing 19 nodes, 248 processors, and 48 gigabytes of RAM per node).
Results
Associations with Diurnal Preference
The undergraduate population sample (n = 528) consisted of 190 males and 338 females. Of the 242 individuals who scored as evening or morning types (MEQ scores ≤41 or ≥59, respectively), 28 males (12%) and 50 females (21%) scored as MT, and 60 males (25%) and 104 females (43%) scored as ET. Forty-four of 169 (26%) participants indicated mild to severe depression (i.e., BDI score ≥14), representing 21% of males and 27% females. Average BDI scores were higher in females (Mf = 11.25, SE = 0.61; Mm = 8.61, SE = 1.01; F(1,224) = 2.07, p = 0.039) (Fig. 2a). Females had higher anxiety levels than males (Mf = 39.10, SE = 0.64; Mm = 42.29, SE = 0.74; F(1,356) = 4.51, p = 0.034) (Fig. 2b).
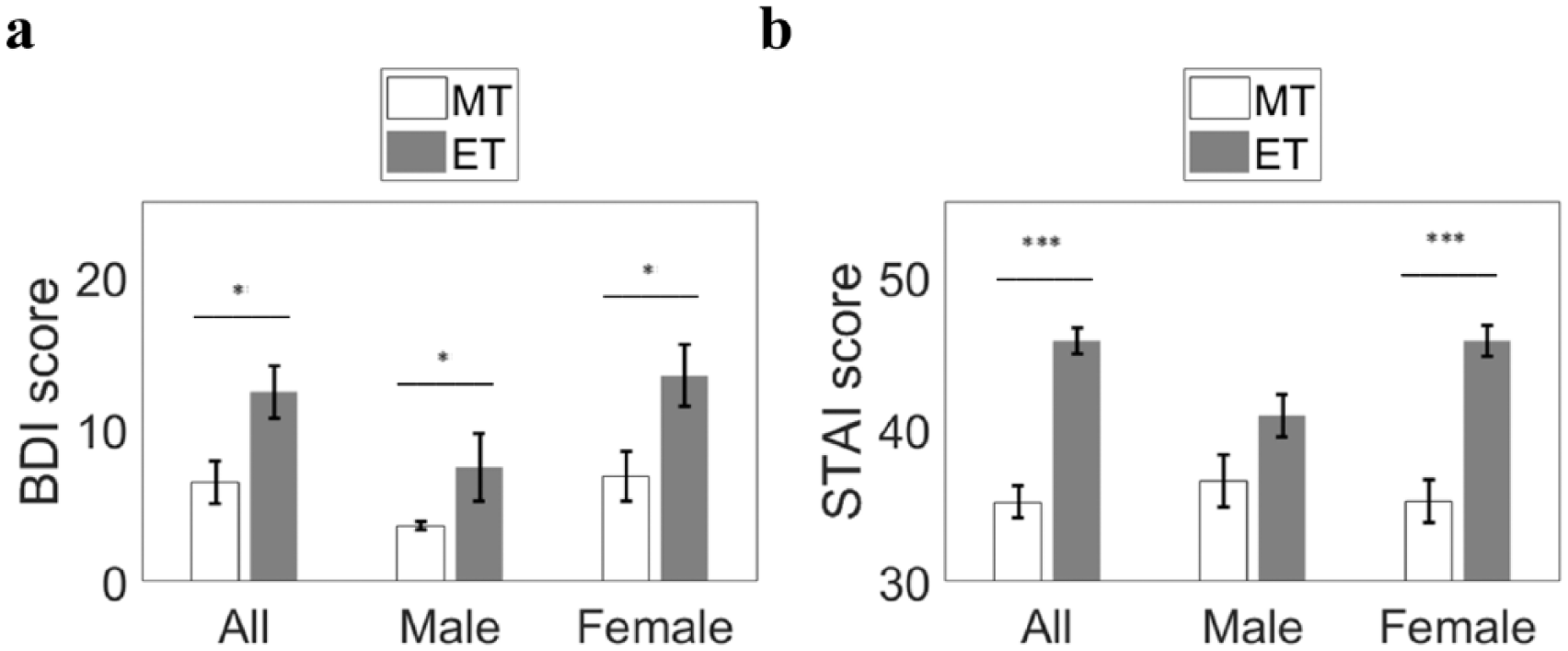
Average anxiety (STAI) and depression (BDI) scores in morning types (MT) and evening types (ET). Evening types had higher BDI scores (MET = 12.47, SE = 1.71) than MT (MMT = 7.52, SE = 1.41; F(2,172) = 3.24, p = 0.042). Morning types were significantly less anxious (MMT = 35.23, SE = 1.07) than ET (MET = 45.83, SE = 0.83; F(2,355) = 8.85, p = 0.000). Average BDI scores were higher in females (Mf = 11.25, SE = 0.61; Mm = 8.61, SE = 1.01; F(1,224) = 2.07, p = 0.039), and females had higher anxiety levels than males (Mf = 39.10, SE = 0.64; Mm = 42.29, SE = 0.74; F(1,356) = 4.51, p = 0.034). *p < 0.05, ***p < 0.001.
Evening types had higher BDI scores (MET = 12.47, SE = 1.71) than MT (MMT = 7.52, SE = 1.41; F(2272) = 3.24, p = 0.042). Morning types were significantly less anxious (MMT = 35.23, SE = 1.07) than ET (MET = 45.83, SE = 0.83; F(2,355) = 8.85, p = 0.000).
Genotype Associations
Of the 5 clock polymorphisms studied, only the CRY1 (rs2287161) polymorphism deviated significantly (p < 0.01) from Hardy-Weinberg expectations in this population (Table 2). The deviation may be due to miscalling of alleles or the relatively small sample size for this gene.
Allele and genotype frequencies of circadian clock genes in an undergraduate population.
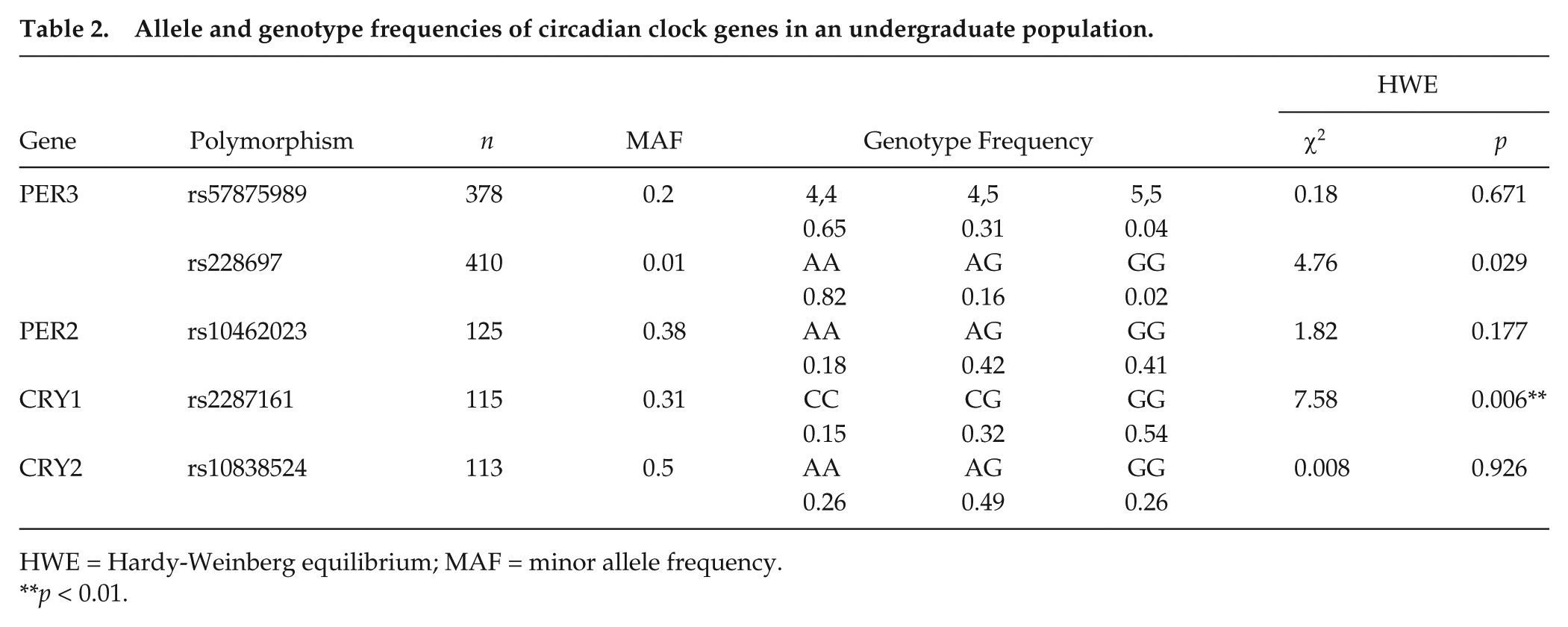
HWE = Hardy-Weinberg equilibrium; MAF = minor allele frequency.
p < 0.01.
The two PER3 polymorphisms had significant genotypic associations with MEQ scores under a recessive model (Table 3, Fig. 3a); 4,4 genotypes (rs57875989) were 3 times more likely to be ET, and GG genotypes (rs228697) were 8-fold more likely to be ET. CRY2 (rs10838524) had both an allelic association (individuals with the G-allele were 3-fold more likely to be ET) (Fig. 3a) and a genotypic association (GG homozygotes and AG heterozygotes were 5-fold more likely to be ET than AA homozygotes) (Fig. 3b).
Association of circadian clock genes with diurnal preference.
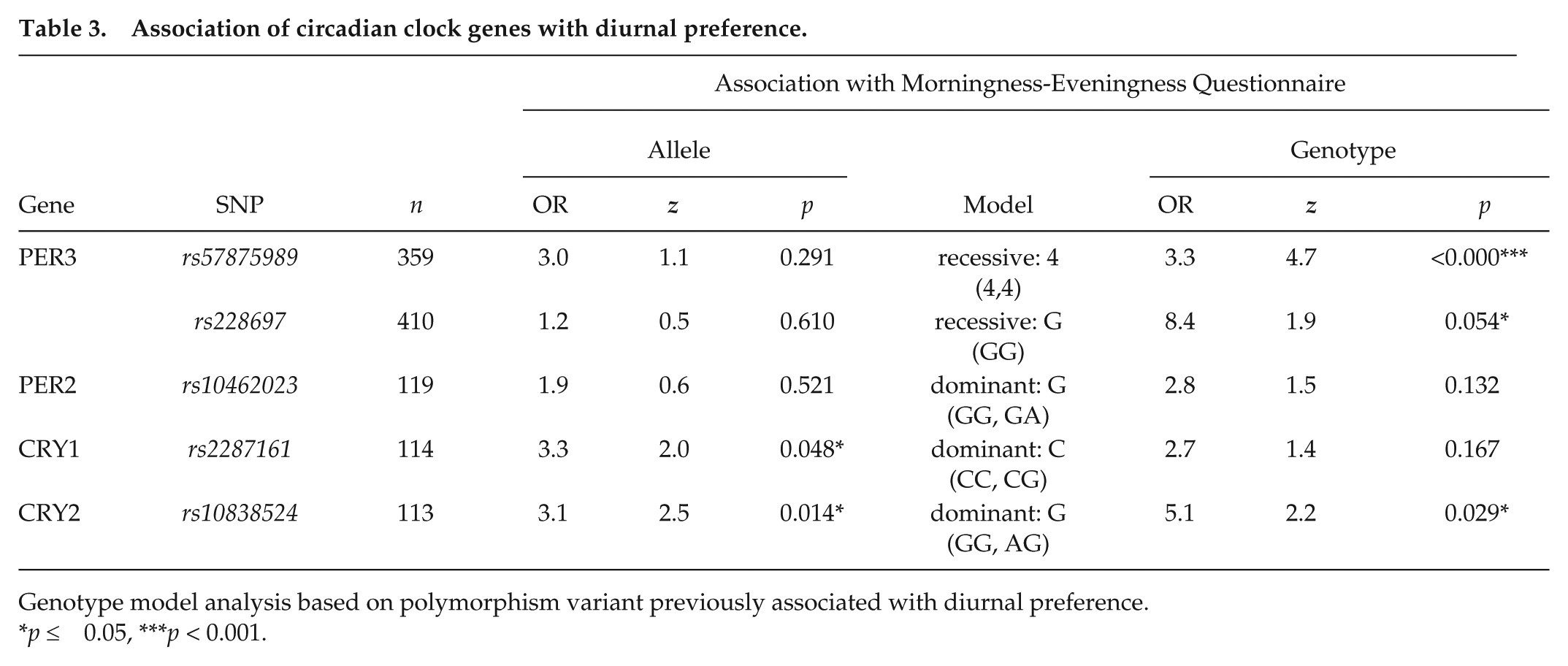
Genotype model analysis based on polymorphism variant previously associated with diurnal preference.
p ≤ 0.05, ***p < 0.001.
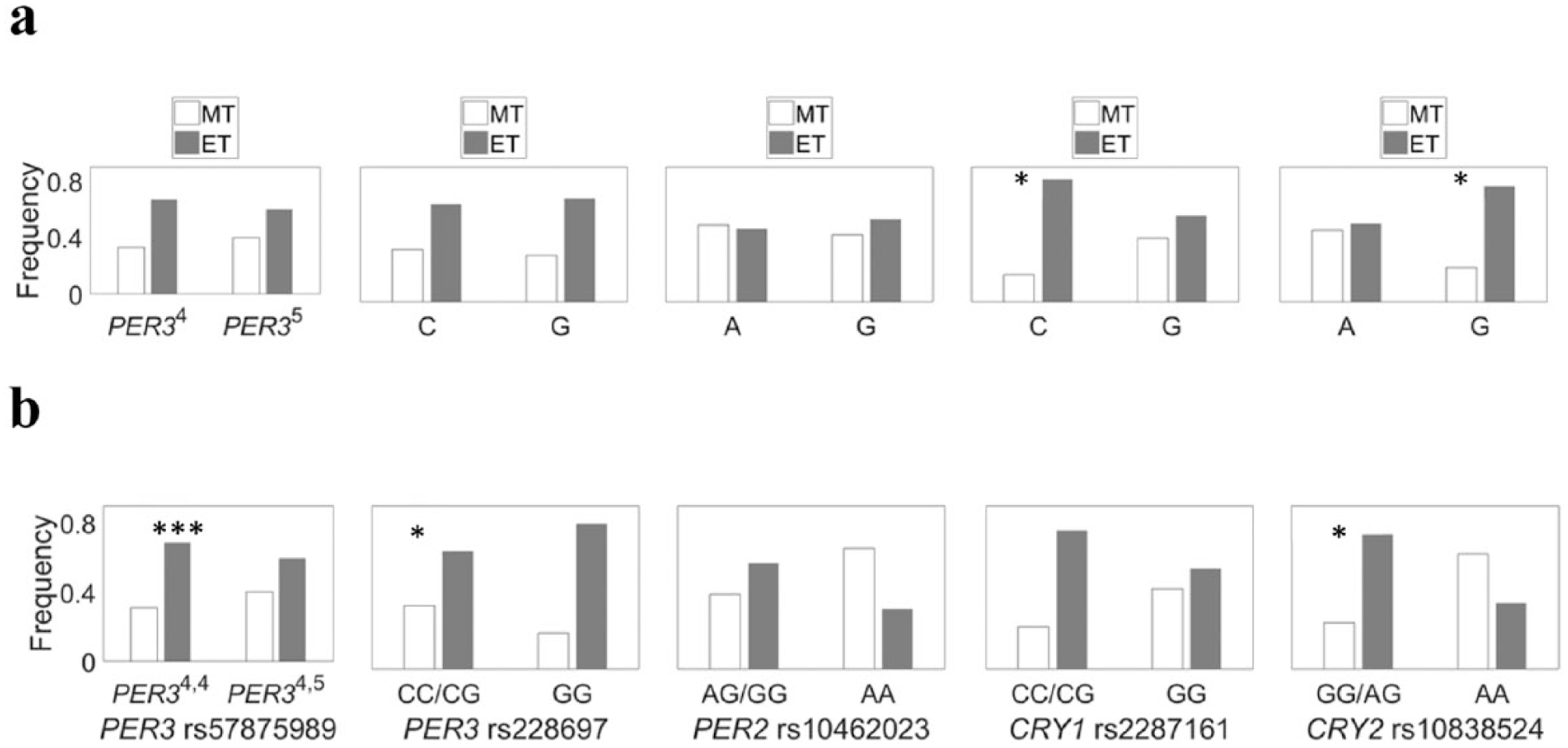
Association of selected clock genes with Horne-Ostberg Morningness-Eveningness Questionnaire scores in undergraduates for (a) allele frequencies and (b) genotype frequencies. For PER3, individuals homozygous for 4,4 (rs57875989) were 3 times more likely to be evening types (ET), and those homozygous for GG (rs228697) were 8-fold more likely to be ET. CRY2 (rs10838524) had both an allelic association (individuals with the G-allele were 3-fold more likely to be ET) and a genotypic association (GG homozygotes and AG heterozygotes were 5-fold more likely to be ET than AA homozygote). *p ≤ 0.05. ***p < 0.001.
Of the 4 polymorphisms studied for association with depression, only the PER3 length polymorphism (rs57875989) was associated with BDI scores in this population (Table 4, Fig. 4a). Individuals homozygous for the 4-repeat length variant were 3 times more likely to be mildly or severely depressed. The PER3 SNP, (rs228697), was significantly associated with anxiety, with GG homozygotes experiencing greater anxiety (F(1405) = 4.30, p = 0.039) (Table 4, Fig. 4b). Across the 5 polymorphisms previously linked to mood disorders, individuals with only 1 polymorphism did not report symptoms of mild or severe depression as measured with the BDI. The frequency of individuals with mild to severe depression differed with the number of polymorphisms present in individuals (Fig. 5) (χ21 = 5.43, p = 0.018).
Association of circadian clock genes with Beck Depression Inventory and State Trait Anxiety Index scores.
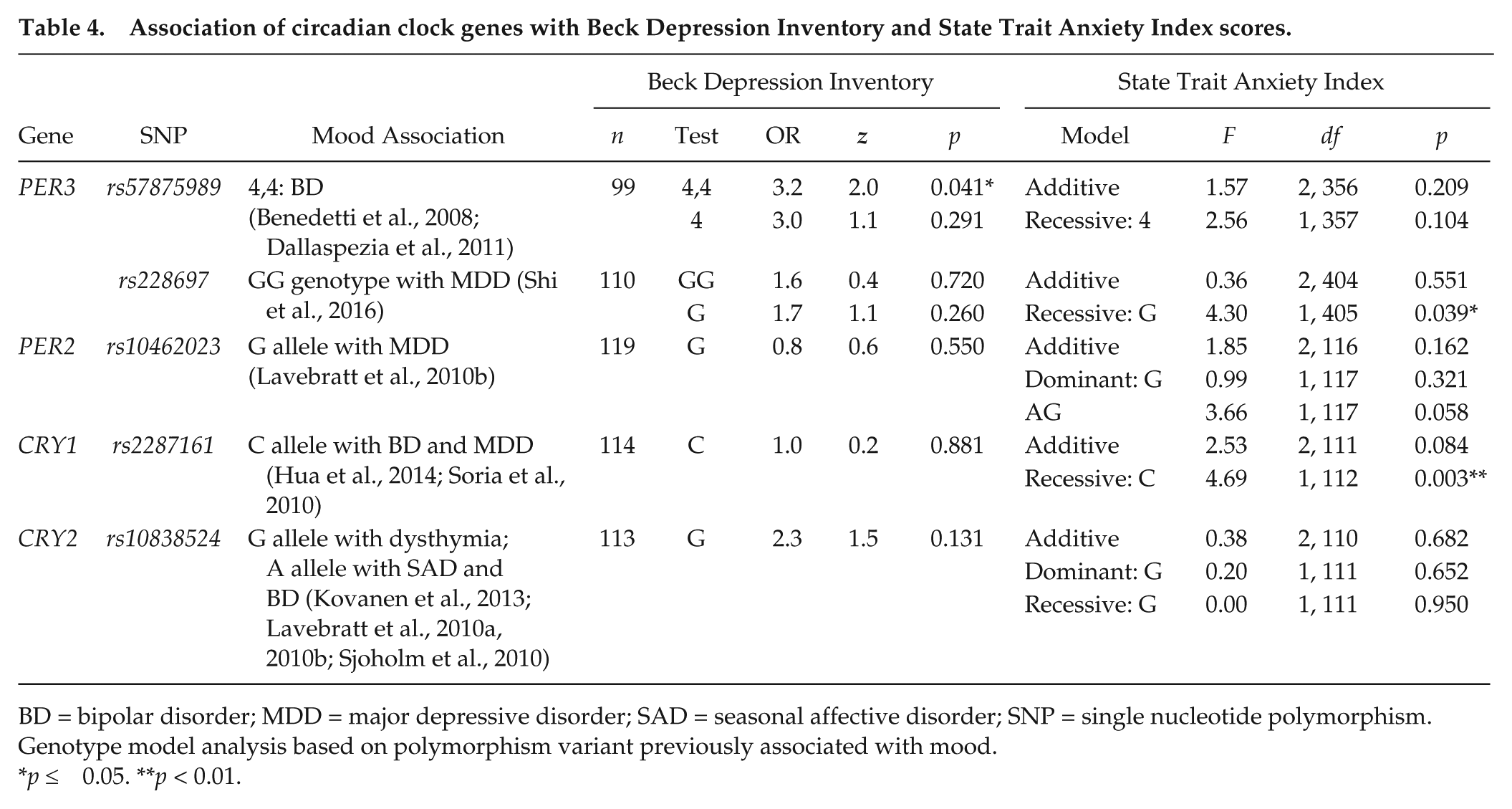
BD = bipolar disorder; MDD = major depressive disorder; SAD = seasonal affective disorder; SNP = single nucleotide polymorphism. Genotype model analysis based on polymorphism variant previously associated with mood.
p ≤ 0.05. **p < 0.01.
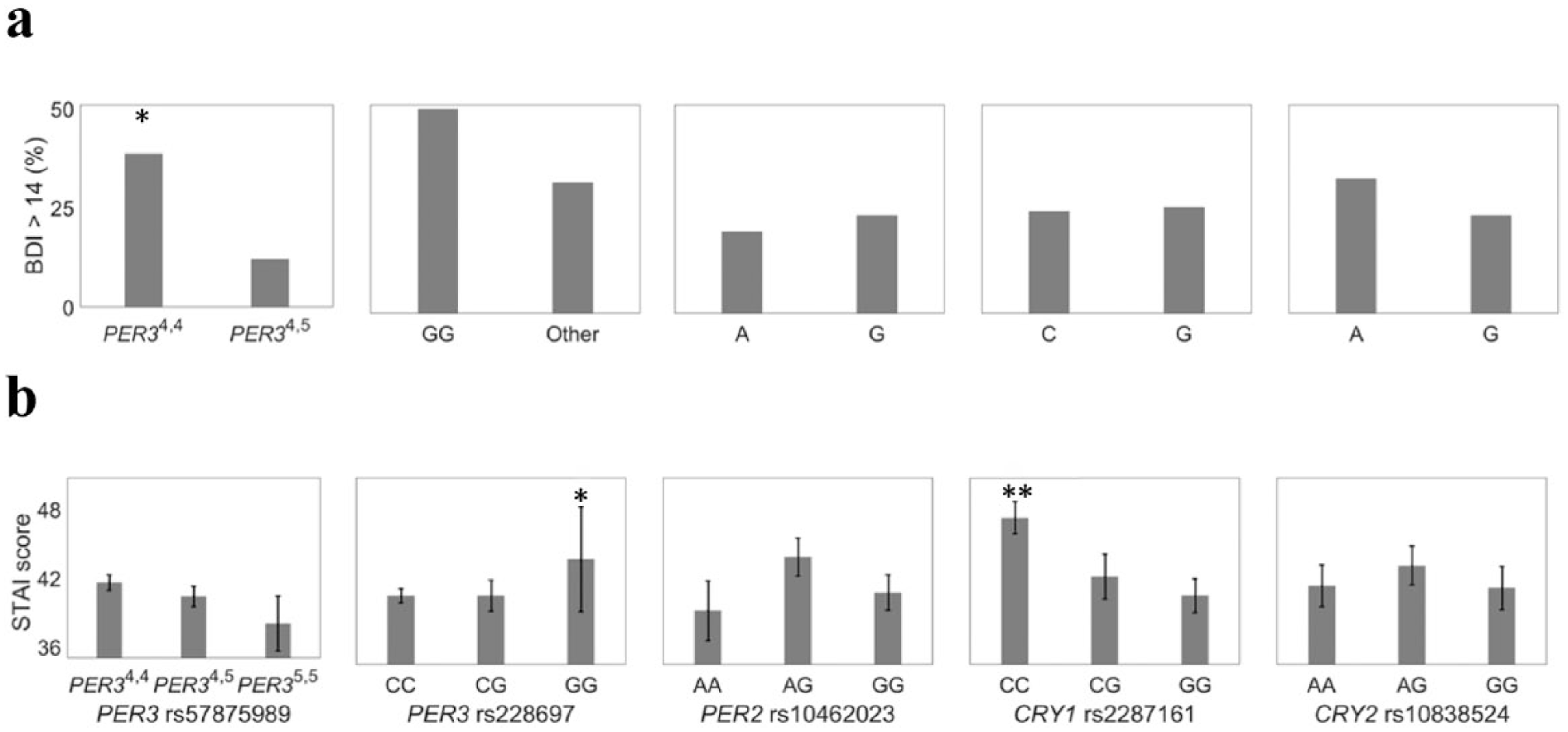
The association of clock gene polymorphisms with (a) Beck Depression Inventory (BDI) and (b) State-Trait Anxiety Inventory (STAI) scores. Only the PER3 length polymorphism (rs57875989) was significantly associated with BDI scores in this population. Individuals homozygous for the 4-repeat length variant were 3-fold more likely to be mildly or severely depressed. The PER3 SNP, (rs228697), was significantly associated with anxiety, with GG homozygotes experiencing greater anxiety (F(1,405) = 4.298, p = 0.039). *p ≤ 0.05. **p < 0.01.
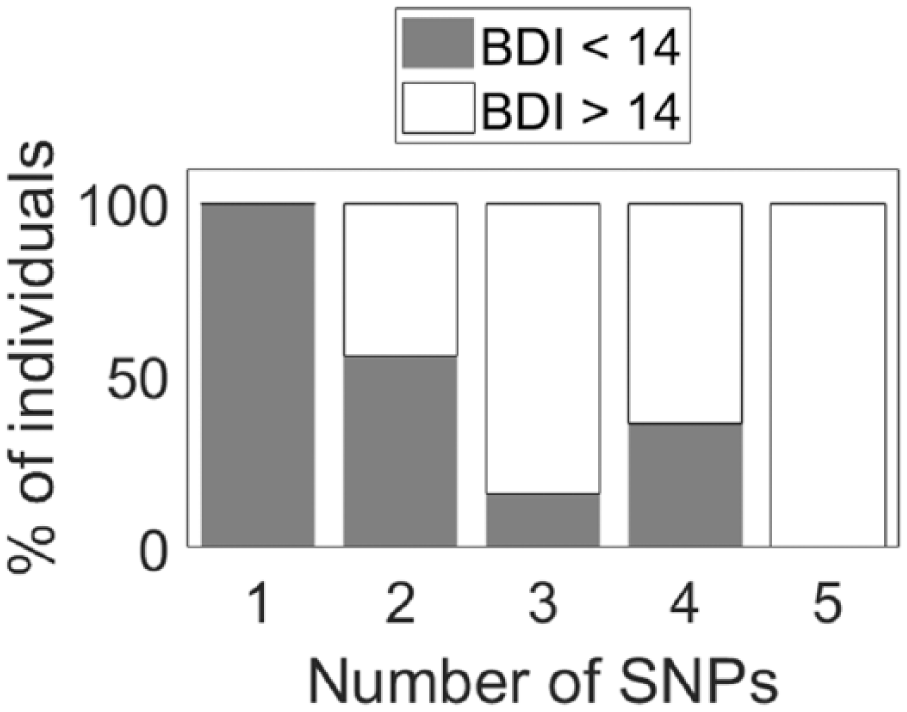
The proportion of depressed individuals and number of circadian clock polymorphisms. No single polymorphism conferred a depressed phenotype in our sample, but the frequency of individuals with depression differed with the number of clock polymorphisms present in an individual (χ21 = 5.43, p = 0.018).
Modeling
To investigate the mechanisms by which these circadian clock genes lead to mood and circadian disorders, we built a mathematical model containing 30 ordinary differential equations with 93 parameters (Fig. 6, Suppl. Table S3). This model was trained using genetic knockout conditions previously identified in mice and in various cell lines (Baggs et al., 2009) (Suppl. Table S5). Once developed, this model system allowed us to apply mutations linked with 9 different polymorphisms previously identified in CLOCK, PER, and CRY genes (Archer et al., 2003; Hida et al., 2014; Lavebratt et al., 2010a, 2010b; Soria et al., 2010) (Figs. 7 and 8). All mutant conditions are described in Table 1.
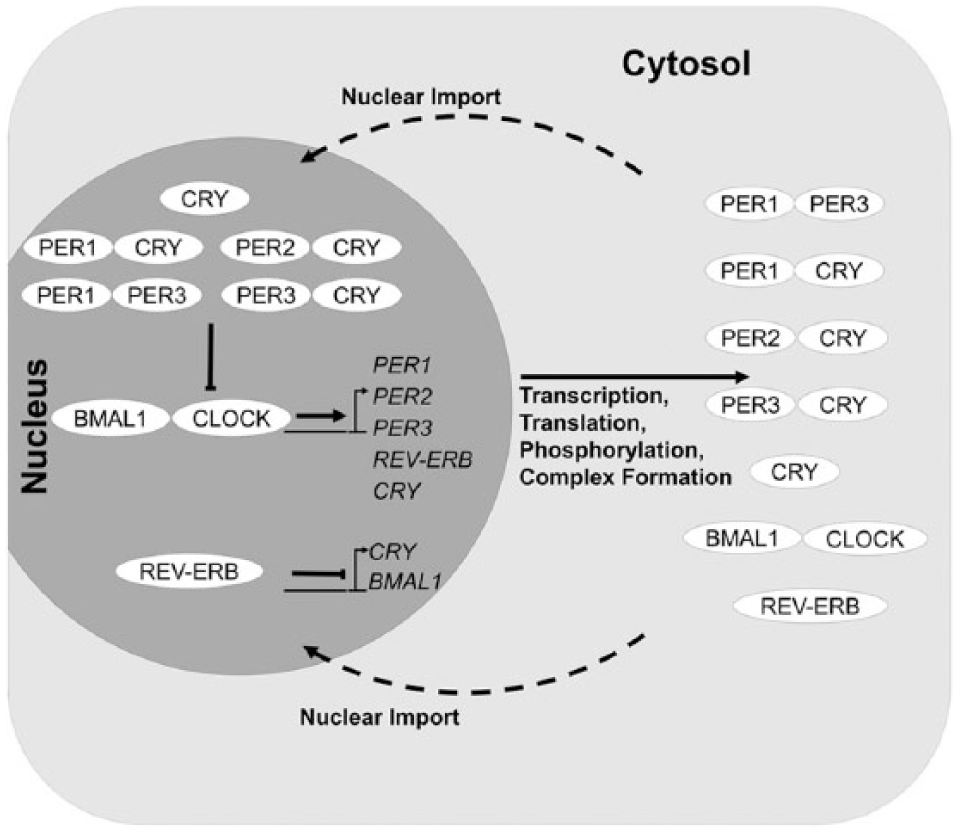
Schematic of model structure. The model structure entails a double negative feedback loop. In the first negative feedback loop, the BMAL1-CLOCK complex activates transcription of the PER and CRY genes, which, once translated, feedback to inhibit their own transcription. In the second negative feedback loop, the BMAL1-CLOCK complex activates the transcription of the REV-ERB genes, which inhibit the transcription of BMAL1 (Shearman et al., 2000).
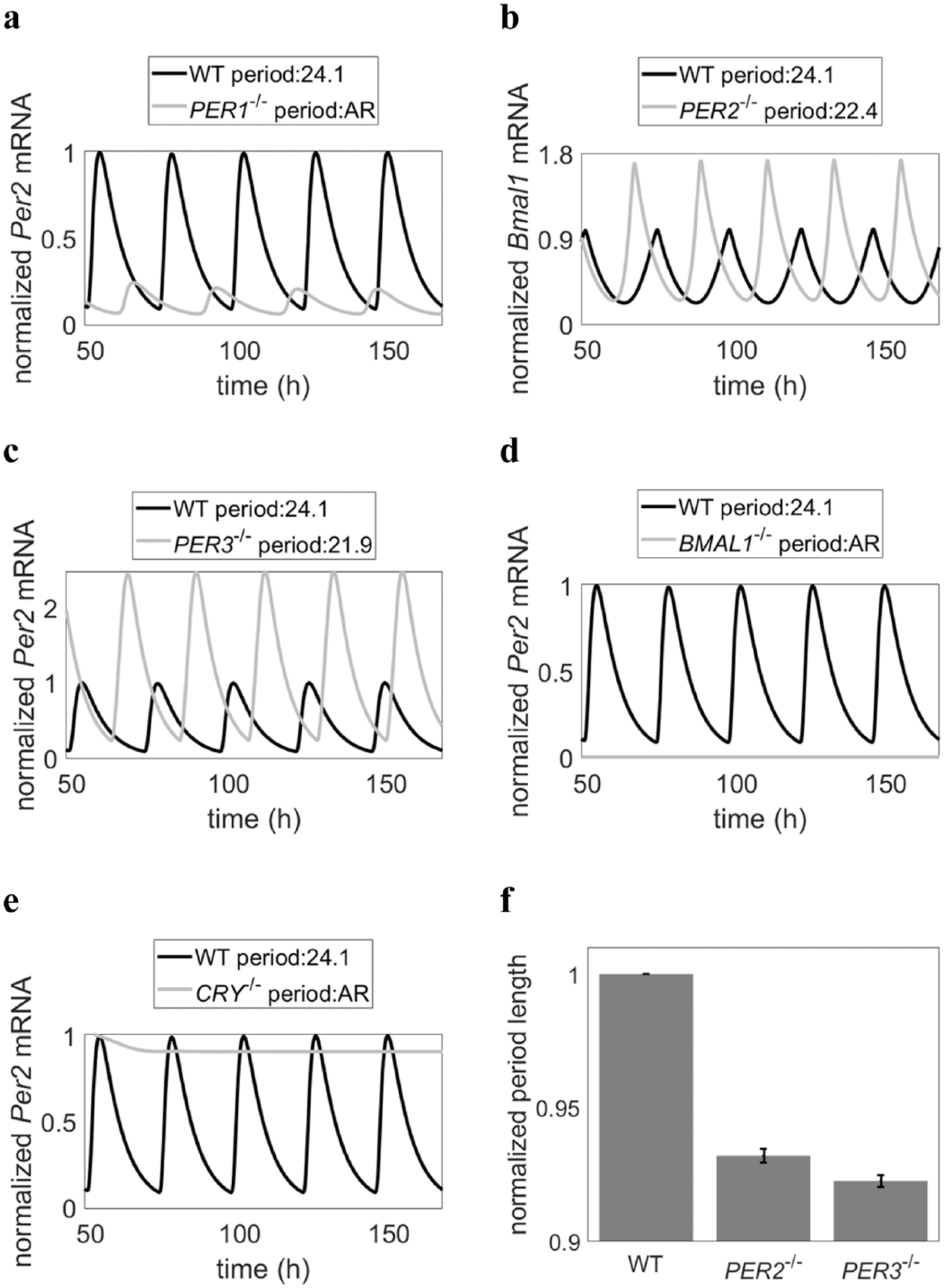
Mathematical model recapitulates experimentally observed knockdown conditions. (a-e) Mathematical model successfully simulates knockdowns in (a) PER1, (b) PER2, (c) PER3, (d) BMAL1, and (e) CRY1/2 as reported experimentally (Baggs et al., 2009; Martynhak et al., 2017). To simulate each mutation, the transcription rate of the corresponding gene was set to 0. (f) Comparison of average knockdown periods using 100 parameter sets. Arrhythmic conditions (BMAL1-/-, PER1-/-, CRY-/-) not reported. Error bars represent 2 standard errors from the mean.
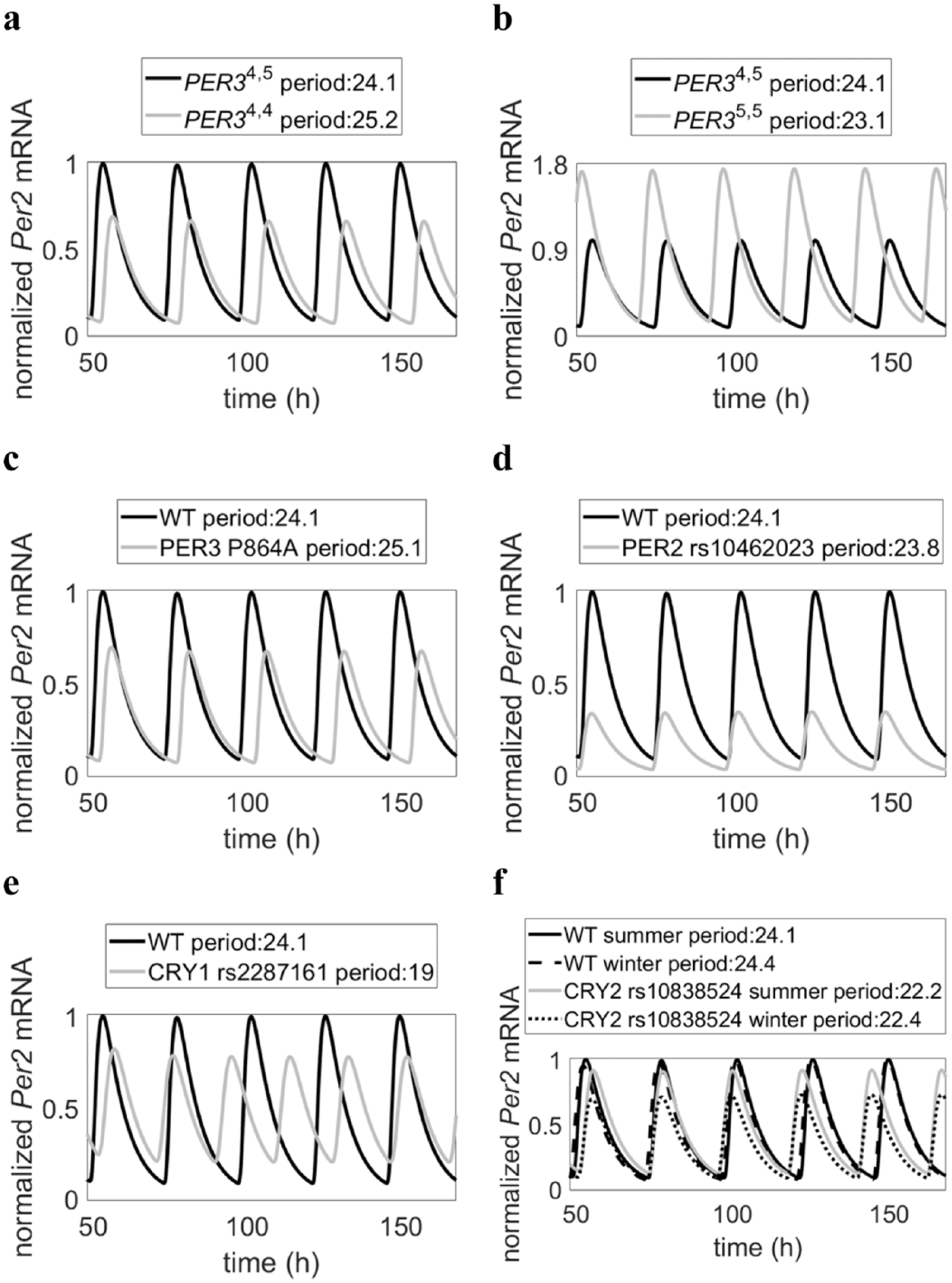
Mathematical model successfully simulates circadian variants experimentally linked to major depressive disorder (MDD), seasonal affective disorder (SAD), and/or diurnal preference. (a, b) A PER3 length polymorphism (rs57875989) is linked to diurnal preference, with 5,5 homozygotes being more prone to morningness (shorter period), while 4,4 homozygotes are prone to eveningness (longer period) (Archer et al., 2003). Additionally, the shorter allele has been associated with mood disorder MDD, as represented by a decreased amplitude in Per2 mRNA expression. (c) PER3 SNP rs228697 is linked to MDD and eveningness, as represented by a decreased amplitude and longer period in Per2 mRNA expression (Hida et al., 2014). (d) PER2 SNP rs10462023 is linked to MDD (Lavebratt et al., 2010b), as represented by a decreased amplitude in Per2 mRNA levels. (e) CRY1 SNP rs2287161 is linked to MDD (Soria et al., 2010), as represented by a decreased amplitude in Per2 mRNA levels. (f) CRY2 SNP rs10838524 is linked to SAD (Lavebratt et al., 2010a), as represented by a decreased amplitude in Per2 mRNA levels during the winter season.
The model was able to recapitulate experimental data on SNPs that are associated with diurnal preference and/or mood as well as predict potential downstream molecular pathways influencing mood. Figure 7 shows the model’s representation of knockdown conditions, indicating model fitness to experimental data from mice and from human cell lines. In Figure 8, we were able to simulate the changes in period and amplitude associated with extreme diurnal preference and mood disorders. For example, Figure 8a shows one of the model’s proposed effects of PER34,4, lengthened period and decreased amplitude of PER2 gene expression, indicative of eveningness preference and increased risk of depression. Intrinsic circadian period has long been used as a marker of morningness-eveningness preference (Duffy et al., 2001), and PER2 transcriptional rhythms have specifically been shown to be necessary for intrinsic clock oscillation (Yamamoto et al., 2005), with period lengths similar to those of intrinsic circadian period (Yamaguchi et al., 2003). Additionally, decreased amplitude of PER2 has previously been associated with depression (Logan et al., 2015), and lithium, an antidepressant and antibipolar medication, has been shown to increase the amplitude of PER2 gene expression (McCarthy et al., 2013). Thus, the model findings successfully recapitulate experimental data, indicating model fitness.
Novel insights into clock mechanisms affecting human mood offered by the model include the following: the potential mechanism by which intronic SNPs influence mood-related molecular pathways, the differential effects of light on PER1 and PER2 transcription, and the critical role of CRY protein binding in inhibiting the phosphorylation of BMAL1 and CLOCK. Of the five polymorphisms investigated, three are intronic. Although all have previously been linked with mood disorders (Lavebratt et al., 2010a, 2010b; Soria et al., 2010), the mechanism by which these SNPs lead to mood disorders is poorly understood. Our model suggests that all are associated with transcriptional increases. When given significant flexibility, the model suggests that in order to satisfy all experimental conditions provided, the three SNPs rs10462023 (in PER2), rs2287161 (in CRY1), and rs10838524 (in CRY2) must lead to increases in transcription rate. Additionally, our model allows flexibility for seasonal changes in PER1 and PER2 transcription, since expression of these genes is affected by light (Challet et al., 2003; Yan, 2009). Model results indicate that seasonal changes in light may lead to increases in PER1 transcription in the winter, while PER2 transcription generally decreases. Our results from 200,000 parameter sets show an 8.47-fold increase in PER1 transcription in the winter and a 0.43-fold decrease in PER2 transcription (Suppl. Table S1).
One critical interaction highlighted by the model was the importance of CRY as an inhibitor of BMAL1 and CLOCK phosphorylation (Matsumura et al., 2014; Tamaru et al., 2015), an interaction not included in previous mathematical models, to our knowledge. The results from our model show that the binding and unbinding rates of CRY to the unactivated BMAL-CLOCK complex are about 10 times higher than any other protein or protein complex binding rates, indicating its indispensability for the biological system as represented in our model. Additionally, the model highlighted several other interactions. The Hill coefficient in the model was estimated to be approximately 1, suggesting that transcriptional activation by the BMAL-CLOCK complex does not have significant cooperative or noncooperative binding effects. Furthermore, the model predicted slow Cry mRNA degradation relative to other mRNAs and fast phosphorylation of the model’s BMAL-CLOCK complex (representative of both BMAL1 phosphorylation and BMAL1-dependent CLOCK phosphorylation) relative to other proteins and complexes in the model. Finally, the model predicted PER3 expression to be higher than that of PER1 or PER2 in the SCN, a finding observed in a previous experimental study (Hamada et al., 2001). We highly encourage other researchers to verify these findings experimentally.
Discussion
Circadian Misalignment and Mood
Our model predicts period length changes and/or decreased amplitude in circadian oscillations in all polymorphisms tested. Changes in period length and amplitude both cause phase shifts in the timing of circadian oscillations (Brown et al., 2008), which in turn cause misalignment between the downstream functions regulated by the clock and the external environment, leading to depression symptoms by yet unknown processes. Delayed or advanced phases in clock gene oscillations are also characteristic of individuals with extreme diurnal preference, and this may help explain why depression and SAD have been previously linked to extreme morning- or evening-type individuals (Antypa et al., 2015; Fares et al., 2015; Kitamura et al., 2010; Lee et al., 2011; Levandovski et al., 2011).
If individuals with advanced or delayed clock oscillations are more likely to suffer from negative affect, individuals with alleles or genotypes associated with period length or amplitude changes and/or those with extreme diurnal preferences should be more depressed. We tested these associations in an undergraduate population and found that college-age students with allelic or genotypic variants that affected period length or amplitude tended to have a higher frequency of depression, and this relationship was significant in those individuals with the PER3 VNTR length polymorphism (rs57875989) allele. Individuals homozygous for this 4-repeat allele were three times more likely to be mildly or severely depressed and were significantly more likely to be evening types. These associations with diurnal preference are consistent with the longer period and decreased amplitude predicted for this genotype by our model. Similarly, the PER3 single nucleotide polymorphism (rs228697) is significantly associated with both anxiety and eveningness in this population, with PER3G,G individuals exhibiting higher levels of anxiety, also consistent with the model prediction of increased period length and decreased robustness of oscillations in individuals homozygous for the G-allele at this locus.
The links between PER3 polymorphisms, diurnal preference, and mood lend support to the “phase-shift hypothesis” (Stephenson et al., 2012) that depression is caused by a misalignment of circadian phase with the sleep-wake cycle. However, our model also predicts a concordant decrease in amplitude of circadian oscillations under these genetic backgrounds. Thus, PER3 polymorphisms may confer multiple functional changes to circadian rhythms that affect downstream processes, including misalignment of rhythms, decreased robustness of circadian signaling, or both. Why does PER3 have such a robust association with diurnal preference and mood relative to other circadian clock genes? In this study, this may be an artifact of the larger sample size of individuals sampled for this gene, relative to the other candidate genes studied. The lack of identified associations between PER2 and CRY SNPs could represent false negatives; further genotyping studies may be needed to test these gene associations and the model predictions fully. However, it should be noted that PER3 has been implicated in several large, genome-wide studies on diurnal preference (Archer et al., 2003; Hida et al., 2014) and mood (Benedetti et al., 2008; Dallaspezia et al., 2011; Shi et al., 2016), suggesting that polymorphisms in this gene may indeed have a robust effect on circadian misalignment. Perhaps because it is not an essential core clock gene (Lee et al., 2004), PER3 may have the flexibility to exert functional changes that alter both period length and amplitude.
It has long been recognized that circadian clock genes likely affect mood as a polygenic trait. In this study, we observed that none of the individuals in our population possessed a single polymorphism that alone produced depressive symptoms. Instead, the frequency of individuals with depressive symptoms tended to increase with the number of clock polymorphisms per individual, suggesting that the circadian system possesses multiple compensatory mechanisms sufficient to counteract some functional mutations in clock genes.
One limitation of our behavioral results is that we did not examine the role of sleep in this study. Sleep quality and duration are also likely to be linked to the degree of circadian misalignment (Barclay et al., 2011) and may exacerbate these effects independently of the circadian system. Another limitation is that we relied on the use of questionnaires to measure depression and anxiety, rather than a diagnostic interview. However, these surveys have been validated in numerous studies and thus represent well-established criteria for understanding depressive or anxious symptoms. A limitation of our genetic results is that our candidate gene approach was highly selective. We selected only polymorphisms that had been previously linked to mood disorders in order to limit the number of genes we needed to model. In doing this, we did not have the opportunity to discover novel polymorphisms associated with mood disorder. In addition, our analysis was restricted to one population of undergraduates and may not be generalizable to other age groups or populations.
Modeling Seasonal Affective Disorder
By adding seasonal light parameters to the model, we were able to consider the seasonal effects of light on circadian oscillations. Previous research has indicated the necessity of light in circadian modeling; in mice, constant light-dark conditions change period lengthening effects of genetic knockouts (van der Veen and Archer, 2010), and in humans, certain genotypes show increased sensitivity to light (Chellappa et al., 2012). While some studies demonstrate transcriptional increases associated with light intensity (Yokota et al., 2000), others find that these increases are mitigated with increased duration of daily light exposure (Steinlechner et al., 2002). In peripheral human immune tissues, seasonal effects on circadian gene expression vary by gene (Dopico et al., 2015). Therefore, our model allowed seasonal effects to cause transcriptional increases, decreases, or no change in PER1 and PER2 gene expression, two circadian genes commonly thought to be regulated by light inputs (Challet et al., 2003; Yan, 2009). After range refinement performed on initial parameter sets, our model indicates that PER1 transcription increases in the winter, while PER2 transcription generally decreases. While this finding is surprising, it is not unusual for PER1 and PER2 to display different effects. Studies in mutant mice found that PER1 mutants exhibited lengthened period with increased light intensity, while PER2 mutants showed the opposite effects (Steinlechner et al., 2002). Additionally, prolonged light exposure has been found to increase PER2, but not PER1, gene expression (Challet et al., 2003), so it is possible that limited light exposure, as is seen in the wintertime, may have the opposite effect.
Once seasonal light parameters had successfully been incorporated into the model, we investigated the mechanism by which CRY2 rs10838524, an intronic SNP, may be linked to SAD (Lavebratt et al., 2010a). After applying this mutation through transcriptional variation in CRY, our model demonstrated at least 30% amplitude decrease in the winter light condition, while the amplitude in the summer condition remained within 15% of wild-type. These results strengthen the previously observed connection between SAD and circadian gene expression and suggest that circadian-based therapies aimed at increasing the robustness of circadian oscillations may improve the condition of patients with mood disorders. In fact, bright light therapy, a common treatment for circadian-linked disorders, has previously been shown to reduce both anxiety and depression-like behaviors in mice (Ashkenazy et al., 2009).
Possible Function of Intronic SNPs in Circadian Genes
Very little is known about the role that intronic polymorphisms play in regulating mood and behavior. However, introns play important transcriptional roles, including involvement in transcription initiation, termination, and elongation (Chorev and Carmel, 2012). By using a stochastic ranking evolutionary strategy for global parameter estimation, our model is able to propose functional explanations for intronic SNPs linked to some of these disorders. Our model suggests that intronic SNPs rs2287161, rs10462023, and rs10838524 are gain-of-function mutations, causing increases in transcription rate.
Previous research suggests that CRY2 SNP rs10838524 is located in a regulatory region (Kovanen et al., 2013), which our model suggests leads to increased transcription of CRY2, leading to a decrease in amplitude of circadian gene expression in the model’s “winter” condition, indicative of winter depression or SAD. Next, PER2 SNP rs10462023 is believed to occur in the 5′ untranslated region (UTR) (Lavebratt et al., 2010b), which may influence mRNA stability and lead to increased nuclear export of mRNA (Chatterjee et al., 2010), eventually leading to decreased winter amplitudes of circadian gene expression representative of SAD. Finally, CRY1 SNP rs2287161 occurs upstream of the CRY1 promoter region (Soria et al., 2010), leading to increased transcription of CRY1 and an eventual decrease in amplitude of circadian gene expression representative of MDD. These predictions should be verified experimentally in future studies.
The Unstudied Outstanding Question: What Are the Downstream Links between Clock Genes and Mood?
Desynchrony in circadian-related gene networks is likely to disrupt many downstream physiological processes, as nearly one-third to one-half of all human transcripts are regulated by the circadian clock. In fact, circadian dysfunction is implicated in diverse disorders from obesity and diabetes to cardiac disease and cancer (McClung, 2013; Morris et al., 2012), and identifying the molecular pathways involved in circadian-related disorders is the next frontier of chronobiology. Our computational model is simplistic and therefore does not involve downstream molecular pathways. Nevertheless, we can predict molecular interactions that may influence these pathways. In a previously published paper, using a comprehensive model, we noted that PER3 binding rates with CKIδ/ε and CRY were significantly larger than PER1/2-CRY binding rates, suggesting that the potential mechanism by which PER3 SNP variants influence anxiety occurs via the interactions of PER3 with CRY and CKIδ/ε proteins (Liberman et al., 2017). While the simplified model does not include CKIδ/ε, PER3-CRY findings coincide with those of the previous paper. Additionally, the new model suggests that PER3 expression levels might be slightly higher than those of the other two PER genes. These findings reiterate the importance of PER3 and PER3-CRY binding in the circadian clock network and to mood and behavioral disorders.
Another result from our current model suggests a critical function for CRY genes in the circadian clock network, with downstream effects on mood. CRY genes play a role in the clock mechanism both as part of the PER-CRY complexes and as inhibitors of BMAL1 and CLOCK phosphorylation (Matsumura et al., 2014; Tamaru et al., 2015). The results from our model show that the binding and unbinding rates of CRY to the inactive BMAL-CLOCK complex (representative of this inhibition) are about 10 times higher than any other protein or protein complex binding rates, indicating that CRY inhibition of BMAL1 and CLOCK phosphorylation is critical to the model.
In humans, CRY genes have been associated with bipolar and related disorders as well as depression and related disorders (Hua et al., 2014; Kovanen et al., 2013; Lavebratt et al., 2010a; Melo et al., 2017; Soria et al., 2010), and CRY knockout mice (lacking either or both CRYs) have been shown to have elevated anxiety-like behavior (De Bundel et al., 2013). At the molecular level, both BMAL and CRY proteins play important roles in monoamine signaling and neurogenesis, connecting the clock mechanism to diurnal regulation of dopamine levels (Albrecht, 2017). BMAL phosphorylation and binding are known to affect downstream physiological changes in the dopamine pathway; BMAL1 and NPAS2 are known to regulate dopamine levels via activation of monoamine oxidase (MAO) enzymes that act to degrade dopamine, and these genes have been linked to seasonal affective mood disorder (Partonen et al., 2007). Depression and related disorders, including anxiety, are associated with an imbalance in dopamine levels (Kacprzak et al., 2017; van der Wee et al., 2008), suggesting that altered levels of BMAL phosphorylation via CRY binding could influence the dopaminergic control of anxiety and depression-related behaviors. Interestingly, microarray experiments also link the expression of CRY2 to bipolar and related disorders; valproate, a mood stabilizer, decreased CRY2 expression in the amygdala, and cotreatment with the stimulant methamphetamine prevented this change, suggesting that pathways involving CRY may also influence manic-depressive states and related mood disorders (Ogden et al., 2004). Additionally, CRY proteins are involved in hypothalamus-pituitary-adrenal (HPA) axis regulation (Lamia et al., 2011), acting as repressors of glucocorticoid receptor activity. CRY dysfunction could be linked with the onset of mood disorders via this pathway (McClung, 2013).
Conclusions
Here, we have investigated the relationships between circadian polymorphisms and mood disorders. We have experimentally verified links between PER3 variants and extreme diurnal preferences as well as depressive and anxious phenotypes. We built a simple mathematical model capable of simulating these mutations and their effects on circadian misalignment. The model satisfies experimental knockdown and genetic conditions, and it is written in MATLAB, a user-friendly coding platform. Our model enables us to predict circadian phenotypes (e.g., altered period length and amplitude) associated with mood disorders and to identify critical effects of clock gene mutations on CRY/BMAL binding. Our model also predicts that the three intronic SNPs studied are gain-of-function mutations, causing increases in transcription rate, predictions that should be verified experimentally in future studies. Thus, the model explains potential mechanisms underlying the compelling links between clock genes, circadian rhythms, and mood. However, our model provides only one possible framework for this system. We encourage other researchers to use our computational structure to include future experimental findings.
Supplemental Material
Supplementary_material – Supplemental material for Modeling Strengthens Molecular Link between Circadian Polymorphisms and Major Mood Disorders
Supplemental material, Supplementary_material for Modeling Strengthens Molecular Link between Circadian Polymorphisms and Major Mood Disorders by Amanda R. Liberman, Lumbardh Halitjaha, Ahmet Ay, and Krista K. Ingram in Journal of Biological Rhythms
Footnotes
Acknowledgements
We thank Dan Schult for assistance with writing the code for the Hopf bifurcation analysis and for his support of the project and Kerri Woods for assistance with genotyping. This work was supported by a grant from the Picker Interdisciplinary Science Institute (KKI) and the Beckman Foundation as part of the Beckman Scholars program (ARL). The code and README files are available on request. All data are available in the DRYAD repository.
Conflict of Interest Statement
The author(s) have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.