Abstract
The immune system is a complex set of physiological mechanisms whose general aim is to defend the organism against non-self-bodies, such as pathogens (bacteria, viruses, parasites), as well as cancer cells. Circadian rhythms are endogenous 24-h variations found in virtually all physiological processes. These circadian rhythms are generated by circadian clocks, located in most cell types, including cells of the immune system. This review presents an overview of the clocks in the immune system and of the circadian regulation of the function of immune cells. Most immune cells express circadian clock genes and present a wide array of genes expressed with a 24-h rhythm. This has profound impacts on cellular functions, including a daily rhythm in the synthesis and release of cytokines, chemokines and cytolytic factors, the daily gating of the response occurring through pattern recognition receptors, circadian rhythms of cellular functions such as phagocytosis, migration to inflamed or infected tissue, cytolytic activity, and proliferative response to antigens. Consequently, alterations of circadian rhythms (e.g., clock gene mutation in mice or environmental disruption similar to shift work) lead to disturbed immune responses. We discuss the implications of these data for human health and the areas that future research should aim to address.
Keywords
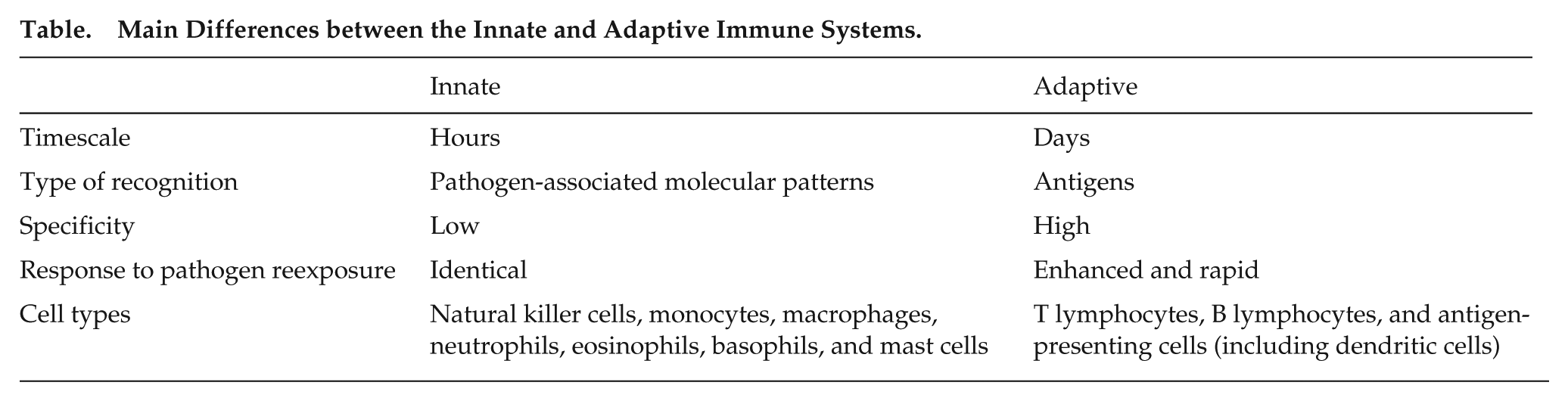
The immune system is a complex set of physiological mechanisms that defend the organism against infectious agents (bacteria, viruses, parasites) as well as cancer cells. The system can be divided in 2 arms: the innate and adaptive immunity (see Box 1 for a brief description of the immune system, its 2 arms, and major cell types). The innate immunity allows for a nonspecific but rapid response, which in most cases is sufficient to control the infection. This first line of protection includes physical barriers (e.g., skin, mucosa), phagocytosis, and secreted substances to kill the foreign body. The innate immunity will also alert the adaptive immune system, which can then mount a slower but antigen-specific response, allowing for the complete elimination or inactivation of the pathogens. Adaptive immunity also mediates generation of immune memory, responsible for specific and long-term protection against the infection.
Box 1: The Innate and Adaptive Immune Systems
Innate Immunity
The innate immune system is the first line of defense against infectious agents. It first depends on physical barriers, consisting of the skin and mucosal membranes. Past these first barriers, the innate immunity relies on a number of cell types. These cells can act on the invaders by different mechanisms. One of them is phagocytosis, through which the pathogen is engulfed within a vesicle. Another defense mechanism is the secretion of antimicrobial compounds that will kill the pathogen. The recognition of the pathogens is through conserved molecular patterns (called pathogen-associated molecular patterns, or PAMPs) that characterize group of pathogens (but do not discriminate small differences of particular pathogens). This recognition occurs via pattern-recognition receptors (PRRs) and allows the innate immune cells to detect the presence of infectious agents and to mount an inflammatory response. This leads to the secretion of cytokines and chemokines, which activate immune cells and attract them to the site of infection. In most of the cases, the response of the innate immune system is sufficient to eliminate the infectious agents. In addition, if the innate immunity has failed to eliminate the pathogen, the activation of innate immune cells will also lead to the induction of the adaptive immune response that will further fight the infection.
The following cell types participate in the innate immune responses:
Natural killer (NK) cells are large granular lymphocytes that can kill tumor cells and certain infected cells. They do not use antigen-specific receptors to recognize target cells. Instead, they recognize cell abnormalities (e.g., a decreased expression of major histocompatibility complex [MHC] class I molecules) via their NK cell receptors. They can also eliminate tumor and infected cells using antibody-dependent cell-mediated cytotoxicity.
Mononuclear phagocytes include monocytes circulating in the blood and macrophages in the tissues. Macrophages can be activated by phagocytosis of particulate antigens or via the recognition of pathogen molecular patterns via their PRRs. Activated macrophages have enhanced phagocytic activity, increased ability to kill the ingested infectious agents, and increased secretion of inflammatory molecules, and they produce antimicrobial compounds.
Granulocytes include neutrophils, eosinophils, basophils, and mast cells:
Neutrophils are usually the first innate cells being recruited at the site of infection. They have phagocytic activity, are able to kill the ingested microbes, and produce a variety of antimicrobial agents.
Eosinophils are also motile phagocytes, but their phagocytic activity is less important than that of neutrophils. However, they play an important role in the defense against large parasites: the secretion of their vesicular content can damage the parasite membrane.
Basophils are nonphagocytic granulocytes that release active substances contributing to allergic responses.
Mast cells have large numbers of cytoplasmic granules containing histamine and other active substances. As such, they play a key role during allergy.
Adaptive Immunity
T lymphocytes (or T cells) are key players in the induction of the adaptive immune response. T cells recognize, via their variable T-cell receptor (TCR), a foreign peptidic fragment of antigen in association with self-molecules of the MHC expressed at the surface of specialized antigen-presenting cells. CD4+ T cells (helper) recognize foreign antigen in association with MHC class II molecules, while CD8+ T cells (cytotoxic) see antigenic peptide bound to MHC class I molecules.
The initiation of the adaptive immune response to infectious agents critically depends on the activation of naive T cells by antigen-presenting cells, in particular dendritic cells (DCs). This occurs within secondary lymphoid organs (spleen and lymph nodes) and as such requires that the antigens from the pathogens infecting peripheral tissues be delivered to these organs. The presence of DCs within peripheral tissues and their ability to capture antigens for display on MHC I and II molecules are essential for this process. Infection of a tissue leads to the maturation of DCs that have taken up antigens. This maturation event only occurs during infection and involves sensing of pathogens by PRRs or by cytokines produced during the inflammatory response. Upon activation, DCs upregulate the expression of costimulatory molecules that are essential to prime naive T cells and migrate to the T-cell zone of the secondary lymphoid organ draining the site of infection. Following antigen recognition, naive T lymphocytes proliferate extensively and differentiate into effector cells able to eliminate the infectious agent. Effector CD4+ T lymphocytes secrete cytokines that are essential to help the response of B and CD8+ T cells and to activate macrophages. B cells that have received help from CD4+ T lymphocytes secrete antibodies that participate in the neutralization of pathogens. On the other hand, effector CD8+ T lymphocytes are able to kill infected cells. The coordinated action of T and B cells leads to the elimination of the infectious agent. After clearance of the pathogen, a subset of B and T lymphocytes differentiates into long-lived memory cells. These cells will confer long-term protection against reinfection by providing a rapid and very sensitive response.
Main Differences between the Innate and Adaptive Immune Systems.

Living organisms have developed adaptations to the changing environment they live in. In particular, endogenous circadian (~24-h) rhythms in physiology and behavior allow organisms and cells to anticipate and respond effectively to the daily variations in light, temperature, or food availability they experience (see Box 2 for a brief description of the circadian system). Importantly, the environmental daily variations may include the risk of infection. For example, some pathogens are transmitted by vectors (e.g., insects) that are active at particular times of day. Conversely, the organisms are active in the day or night and may be more prone to meet pathogens when active. As we present in this article, immune functions are controlled by circadian clocks. This circadian control of immune functions allows organisms to mount their most efficient response when it is most needed. The ability to anticipate the time of highest risk of infection is energy efficient, but also, a too high activity of immunocompetent cells might be detrimental (e.g., autoimmune and allergic reactions), so it is better to restrict highest activity to times when it is most needed. Another role of this circadian regulation is to allow different functions to be coordinated in time within the immune system (e.g., between cell types) and with other physiological systems.
It is well known that the symptoms of autoimmune or allergic diseases follow a daily rhythm. For example, asthma symptoms get worse in the night or early morning (e.g., dyspnea, cough), which is associated with higher levels of inflammation in the lungs (Durrington et al., 2014; Sutherland, 2005). Similarly, symptoms associated with rheumatoid arthritis, such as joint stiffness and pain, are stronger in the early morning (Gibbs and Ray, 2013). In addition, lethality in response to endotoxic shock (Halberg et al., 1960) or bacterial infection (Shackelford and Feigin, 1973) is known to be rhythmic in nocturnal rodents, with more deaths in the late rest period. At the cellular level, efforts to find an explanation for these clinical observations have mostly focused on the variation in the blood counts of various immune cell populations (Born et al., 1997; Depres-Brummer et al., 1997). However, over the past decade, an increasing number of studies have taken advantage of molecular and genetic approaches to studying circadian rhythms to address the mechanisms underlying circadian rhythms in the immune system and their implications for healthy physiology and development of diseases. This review focuses on the impact of circadian clocks on the function of various immunocompetent cell types. The reader is referred to other recent reviews for a more in-depth description of other aspects of the circadian-immune crosstalk, such as the circadian regulation of inflammation, the effect of immune responses on the circadian system, and clinical implications (Cermakian et al., 2013; Cermakian et al., 2014; Coogan and Wyse, 2008; Curtis et al., 2014; Scheiermann et al., 2013).
Circadian Influence on Innate Immunity
Rhythms in Monocyte and Macrophage Functions
Macrophages and monocytes have been the most studied cell types in the context of circadian rhythms so far. A reason for this might be that they have a robust intrinsic circadian clock, with a high amplitude of clock gene expression. This is seen, for example, when quantifying clock gene transcripts in peritoneal macrophages from mice sacrificed at different times over 24 h (Keller et al., 2009) and upon culturing of macrophages from PER2::Luciferase knock-in mice and recording bioluminescence rhythms in a luminometer (Gibbs et al., 2012; Keller et al., 2009).
Like in other cell types, the clock in macrophages and monocytes controls the rhythmic expression of hundreds of downstream genes (Keller et al., 2009). Consequently, different functions of macrophages and monocytes display circadian rhythms (Figure 1): the response of pattern-recognition receptors (PRRs), the expression of cytokines in response to an endotoxin challenge, recruitment to tissues, and phagocytosis.
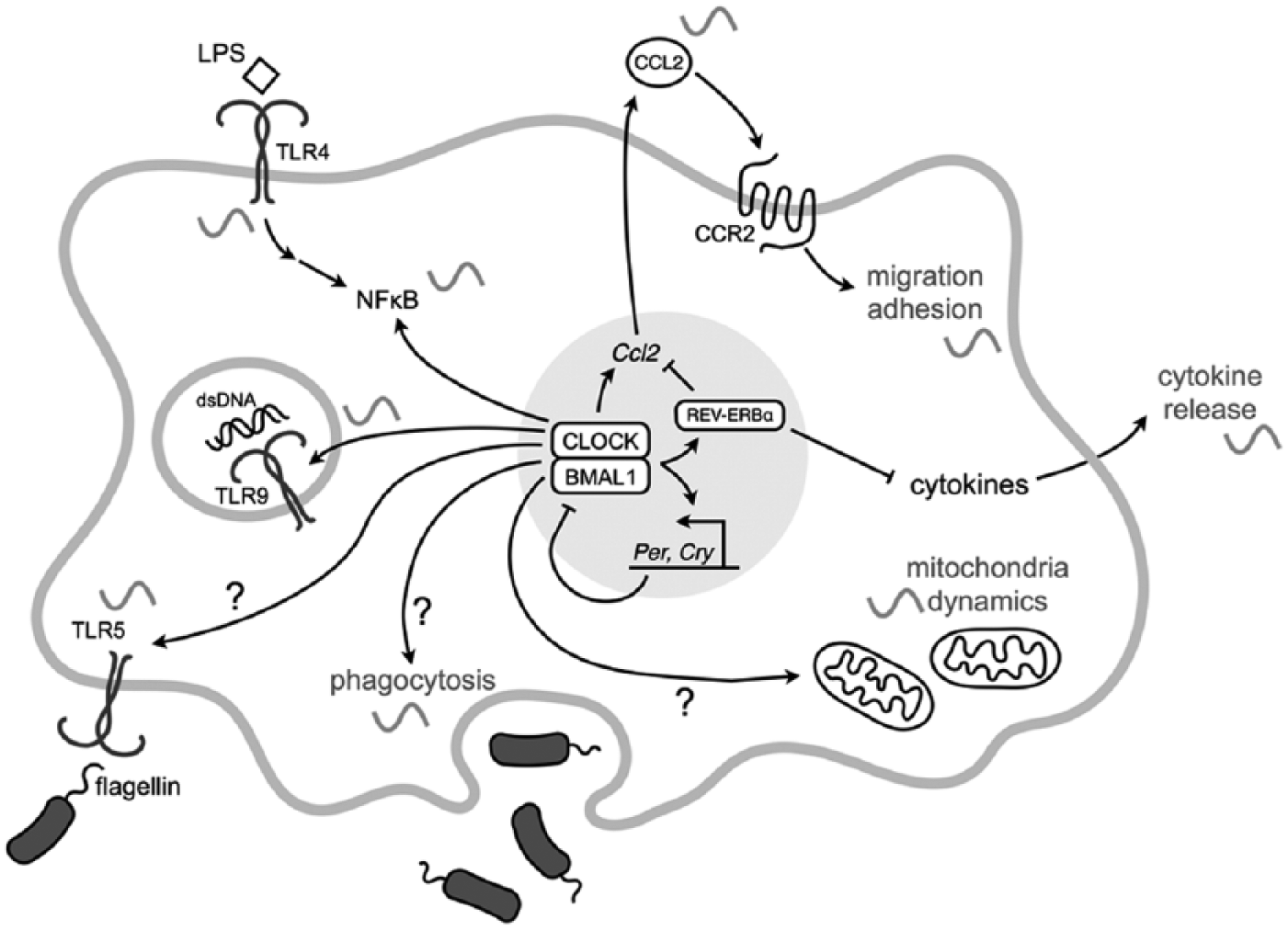
Circadian control of cellular functions in monocytes and macrophages. Schematic representation of the known aspects of monocyte and macrophage function that are regulated by the circadian clock. See text for details. Question marks indicate circadian regulation for which the exact mechanism is not known.
Gating of PRR responses
PRRs are activated by pathogen-associated molecular patterns (PAMPs), which are typically microbial or viral macromolecular components. Three such receptors were found to show a daily variation in the response to their ligand. Several studies have shown that this was the case for Toll-like receptor 4 (TLR4), which is the receptor for lipopolysaccharide (LPS) of the outer membrane of gram-negative bacteria. Treatment of mice at the end of their rest period or beginning of their active period leads to a stronger cytokine response than a treatment in the late active period or early rest period (Gibbs et al., 2012; Marpegan et al., 2009).
This rhythmic response to LPS can be traced down to the macrophages. Indeed, peritoneal macrophages taken from mice at different times of day and cultured in vitro also show a time of day-dependent response to LPS (Gibbs et al., 2012; Keller et al., 2009). This is even true for cultured macrophages whose clocks are synchronized in vitro and then stimulated at different times over several days (Keller et al., 2009). Although the abundance of TLR4 itself does not present a circadian rhythm, many components of the downstream pathways are rhythmic, which might explain the time-dependent response (Keller et al., 2009).
Box 2: The Circadian System and Clock Molecular Mechanisms
One of the key features of circadian rhythms is their endogenous origin (i.e., they occur even in the absence of external time cues). The persistence of these rhythms in a time-free environment is a consequence of circadian clocks located in various organs and cell types. A master pacemaker is located in a region of the anterior hypothalamus called the suprachiasmatic nucleus (SCN). Although SCN lesion and transplantation experiments had originally indicated a central role of the SCN for circadian rhythms generation, research in the past 15 years has shown that circadian oscillators are found in many other brain regions and in most peripheral tissues (Duguay and Cermakian, 2009). Importantly, circadian clock function can be traced down to the cellular level: many cell types exhibit cell-autonomous clock function. For example, isolated SCN neurons and immortalized fibroblasts like the NIH 3T3 cell line display 24-h rhythms in gene expression, metabolism, and other cellular functions. Cells associated with the immune system are not an exception, as most immunocompetent cell types were demonstrated to have circadian clock mechanisms (see main text for details) and, therefore, display cell-autonomous rhythms.
Within those cells, the clock mechanisms rely on so-called clock genes (see Figure). These genes are involved in autoregulatory feedback loops, in which clock factors regulate their own gene expression (Duguay and Cermakian, 2009). In the main loop, the CLOCK and BMAL1 transcription factors activate the expression of Period(Per)1 and 2 and Cryptochrome(Cry)1 and 2 genes, whose protein products then feed back on CLOCK/BMAL1 activity, thus repressing their own expression. In another loop, REV-ERBα, another gene target of CLOCK/BMAL1, represses the expression of the genes encoding these factors. These feedback mechanisms lead to 24-h variations in the abundance of the RNA and protein products from many of these genes.
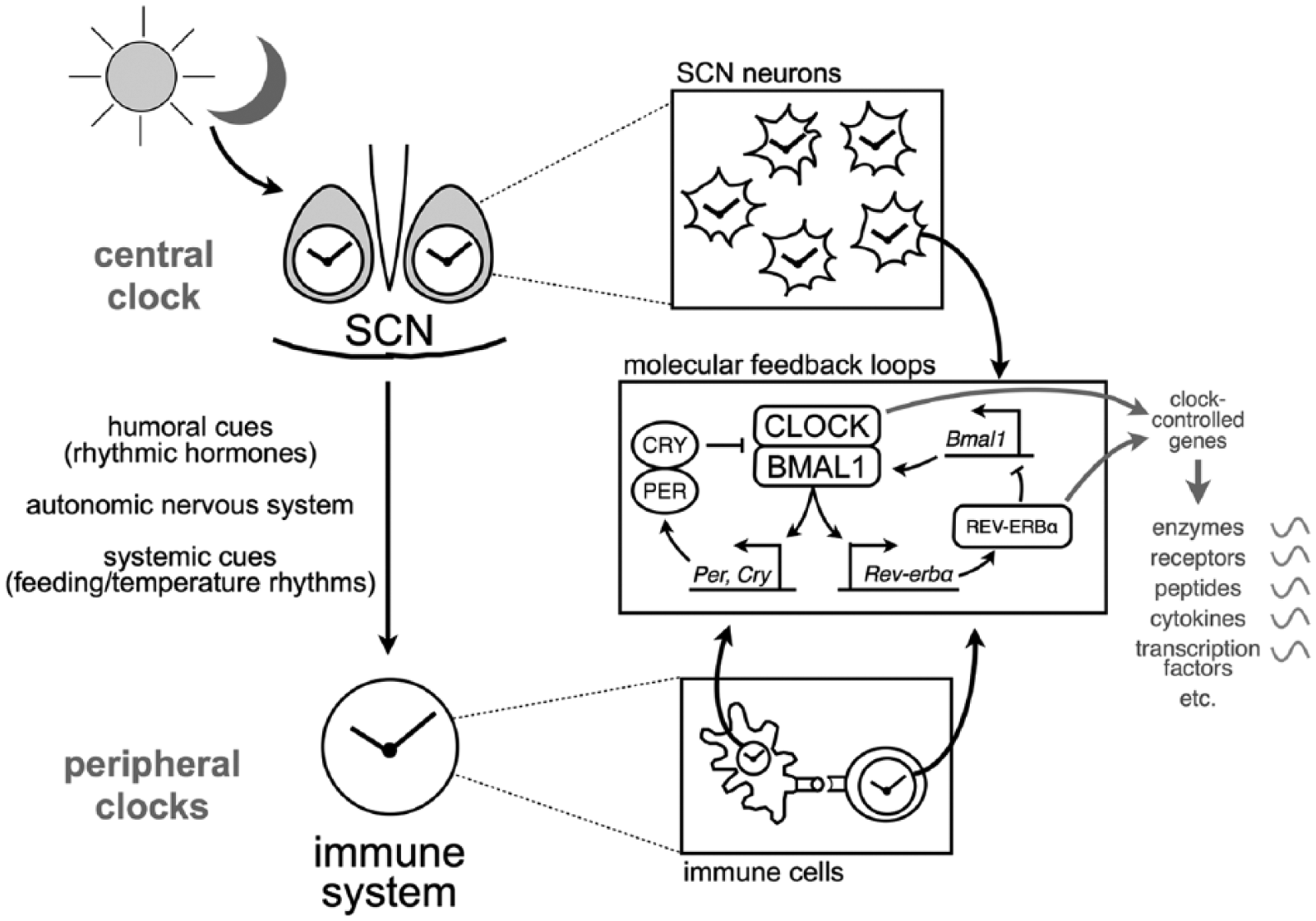
The circadian system and clock molecular mechanisms. Cells of the central clock in the SCN (top) and of peripheral clocks in many other tissues, including in the immune system (bottom), have a clock based on autoregulatory feedback loops (middle right inset; see Box text for details). This clock controls the rhythmic expression of various types of molecules. The SCN clock entrains peripheral clocks (including in immune cells) via humoral, neuronal, and systemic cues.
One way by which this molecular clock regulates cellular function is via transcription factors whose activity varies across the day. For example, transcriptional activation by the CLOCK/BMAL1 heterodimer will vary over the day, due to the rhythmic availability of their repressors (in particular CRY1 and CRY2). Likewise, transcriptional repression by REV-ERBα also varies in a circadian fashion, due to the rhythmic expression of the gene that encodes this nuclear receptor (see Figure). In any given tissue or cell type, hundreds of genes accounting for 10% to 20% of all expressed genes show circadian rhythms in their expression. These genes, whose expression is rhythmic but not part of the circadian clockwork, are called clock-controlled genes (ccgs) (Yan et al., 2008). As described in the main text, ccgs can encode a variety of proteins, including cytokines, chemokines, or receptors.
Another PRR that shows diurnal variation in its response is TLR9, which is bound by unmethylated repeats of the dinucleotide CpG within DNA (as found, for example, in bacterial genomes). In contrast to TLR4, TLR9 expression shows a daily variation. Accordingly, a rhythmic response is observed in the in vivo models of immunization and of sepsis that depend on TLR9 function (Silver et al., 2012b). Finally, a third PRR that shows a daily rhythm of its response is TLR5, the receptor for bacterial flagellin: the activation of nuclear factor (NF)–κB pathway and interleukin (IL)–1β expression in mice in response to a TLR5 agonist varies with the time of injection (Spengler et al., 2012).
Clock-controlled cytokine response
As noted above, stimulated macrophages express cytokines with a circadian rhythm. For example, macrophages stimulated with LPS at different times after synchronization of their clocks in vitro will show a rhythm of secretion of tumor necrosis factor α (TNFα) (Keller et al., 2009). The rhythmic regulation appears to be at least partly at the transcriptional or posttranscriptional level, because transcripts for different cytokines show a 24-h rhythm (Gibbs et al., 2012; Hayashi et al., 2007; Keller et al., 2009). Rhythms of cytokine secretion from monocytes and macrophages have also been observed in human subjects, both under light/dark sleep/wake conditions (Born et al., 1997) and under constant laboratory conditions (Rahman et al., 2014).
The exact mechanism behind this circadian control of cytokine secretion remains to be defined in most cases. Gibbs and colleagues addressed this question first by using mice with a Bmal1 gene deletion targeted specifically to the myeloid cells (macrophages, monocytes, granulocytes): these mice lacked the rhythm of cytokine response, indicating that the clock in macrophages and monocytes is required (Gibbs et al., 2012). The rhythm was also lost in Rev-erbα knockout (KO) mice, even though the macrophage clock seemed intact in these mice, suggesting that the rhythmic transcriptional repressor REV-ERBα is the mechanistic link between the macrophage clock and the circadian regulation of cytokine production in response to LPS (Gibbs et al., 2012). This was further confirmed by the use of a REV-ERBα agonist, which prevented the LPS-induced activation of genes such as Il-6 or Ccl2 in human macrophages (Gibbs et al., 2012).
Different groups have also shown that in models of clock dysfunction, cytokine secretion is altered. One study addressed the effect of an environmental disruption consisting of subjecting mice to a shift of the LD cycle every week for several weeks. This led to a strongly enhanced IL-6 response of peritoneal macrophages upon LPS challenge, compared to nonshifted mice (Castanon-Cervantes et al., 2010). Similarly, a lesion of the SCN, the site of the central circadian clock, strongly potentiated the IL-6 and TNFα response to LPS injection in rats (Guerrero-Vargas et al., 2014). The response of various proinflammatory cytokines is also altered in bone marrow–derived macrophages from ClockΔ19/Δ19 mutant mice (reduced cytokine expression in response to LPS or Salmonella bacterial infection) (Bellet et al., 2013) and from Cry1/Cry2 double KO mice (increased cytokine expression in response to LPS) (Narasimamurthy et al., 2012). It is interesting to note that the effect of the Clock mutation and the Cry gene deletion produces opposite effects, considering the opposite roles these genes play in the molecular clockwork. Of note, though, both CLOCK and CRY proteins may act on NF-κB independently of their classical roles within the clock: CLOCK by directly stimulating NF-κB–mediated transcription (Spengler et al., 2012) and CRYs by repressing the activity of adenylyl cyclase and, hence, indirectly inhibiting NF-κB phosphorylation and transcriptional activity (Narasimamurthy et al., 2012).
Clock-controlled recruitment to tissues
A number of studies also point toward a circadian regulation of the recruitment of macrophages and monocytes to tissues, for example, in situations of infection. Accordingly, the expression of adhesion molecules is controlled by the circadian clock (Sato et al., 2014; Scheiermann et al., 2012). This is also the case for chemokines. In particular, many groups identified Ccl2 (or MCP-1) as a clock-controlled gene. This gene encodes a chemokine that is promoting migration of monocytes (precursors of some macrophages subsets) out of the bone marrow and to the inflamed or infected tissues. The Ccl2 transcript presents a 24-h rhythm (in basal conditions or in response to LPS, depending on the reports) in peritoneal macrophages, blood monocytes, and endothelial cells of mice (Gibbs et al., 2012; Hayashi et al., 2007; Nguyen et al., 2013; Scheiermann et al., 2012) in the human blood (Rahman et al., 2014). Different mechanisms were proposed for this rhythm of Ccl2 expression. Two reports provided data supporting a regulation of this gene by REV-ERBα: one study showed that Ccl2 induction by LPS treatment of macrophages is blunted upon treatment of the cells with a REV-ERBα agonist (Gibbs et al., 2012), while the other study showed that REV-ERBα binds to DNA elements in the Ccl2 promoter and, hence, represses its expression (Sato et al., 2014). On the other hand, two other reports rather implicated a regulation by CLOCK/BMAL1: in one case, the expression of these transcription factors in RAW264.7 cells (a macrophage cell line) activated the Ccl2 expression (Hayashi et al., 2007), while in the other case, BMAL1 was shown to bind to this same promoter but this time causing its repression, presumably via the polycomb repressive complex 2 (Nguyen et al., 2013). This appears to be critical for the generation of a daily rhythm of the numbers of Ly6Chi inflammatory monocytes in the blood, spleen, and bone marrow and a time-dependent efficiency of the response to the bacteria Listeria monocytogenes (whose clearance depends on inflammatory monocytes) (Nguyen et al., 2013). Importantly, a deletion of Bmal1 specifically in myeloid cells (using the Lyz2-Cre driver) abolished both the rhythm of Ccl2 expression and that of inflammatory monocytes, implying a direct role of the clock in these cells for their trafficking between tissues. This loss of rhythmicity led to an increased inflammatory response, and accordingly, the mice became highly susceptible to death following infection with a dose of L. monocytogenes that was not lethal for control mice (Nguyen et al., 2013). Therefore, the circadian regulation of monocyte trafficking has great consequences for the ability to fight against infections.
Other functions
Other functions of macrophages and monocytes were found to vary as a function of time of day. This is the case of phagocytosis, which is higher in peritoneal macrophages harvested from mice in the late light period compared to other times of the day, under a light-dark cycle (Hayashi et al., 2007). Moreover, in cultured peritoneal macrophages, mitochondrial dynamics, mitochondria membrane potential, endocytosis of dextran, and phagocytosis of bacteria all appear to vary with time after synchronization of the cellular clocks (Oliva-Ramirez et al., 2014).
Rhythms in Natural Killer Cell Functions
Natural killer (NK) cells are cells important for the elimination of tumor cells and some infected cells, via secretion of cytolytic substances. It has been known for some time that NK cell cytotoxicity presents a daily rhythm in rats, with maximal activity in the dark phase (Esquifino et al., 1996; Fernandes et al., 1979). However, it is only in the past decade that studies have started to uncover the mechanisms underlying these rhythms and their physiological implications. Interferon γ (IFNγ) and the cytolytic factors granzyme B and perforin present a rhythm in the abundance of their messenger RNA (mRNA) and protein in spleens of rats held under a light-dark cycle (Arjona et al., 2004). The maximal levels occur around the late dark phase or early light phase, similar to the cytolytic activity of splenic NK cells. However, these factors are expressed by other spleen cell types. Therefore, a subsequent study used an enriched preparation of NK cells from spleen (>70% NK cells) and showed again a rhythm of cytokines and cytolytic factors (Arjona and Sarkar, 2005). The same study was also the first to show that NK cells express clock genes. The rhythms of expression of clock genes, cytokines, and cytolytic factors persist in constant darkness, suggesting an endogenous nature of this rhythmicity (Arjona and Sarkar, 2006b). Furthermore, knockdown of the clock gene Per2 in RNK16 cells (an NK cell line) reduced the levels of granzyme B and perforin proteins expressed (but not of IFNγ or TNFα). On the other hand, in Per2 mutant mice, while IFNγ mRNA and protein levels lose their rhythmicity, there is little or no alteration of the rhythms of granzyme B, perforin, and TNFα mRNAs (Arjona and Sarkar, 2006a). This does not necessarily mean that these factors are not clock regulated, as this strain of Per2 mutant mice retains some rhythmicity, although with a shorter period (Zheng et al., 1999). Another study showed that the same Per2 mutant mice were resistant to endotoxic (LPS) shock, which was paralleled by a bluntening of cytokine responses, including IFNγ (Liu et al., 2006). Interestingly, a clear reduction of the percentage of IFNγ-positive cells was observed only in cells positive for the NK cell marker DX5 (Liu et al., 2006).
So far, no studies have demonstrated a specific role for the NK cell-intrinsic clock in the function of these cells. Rhythms in NK cells could also be the consequence of rhythmic cues from other body clocks (e.g., the SCN; see also Box 3). This central clock appears to have a role, via the sympathetic nervous system. Logan and colleagues used guanethidine injections in the spleen to achieve a chemical splenic sympathectomy in rats (Logan et al., 2011). This led to alteration in the expression of clock, cytokine, and cytolytic factor genes in the spleen (but not a loss of the day-night difference). Another systemic circadian disruption used by the same group was to subject the rats to repeated 6-h shifts of the light-dark cycle. Under these repeated jetlag conditions, enriched NK cell preparations showed phase changes of clock gene and protein expression, as well as highly disturbed rhythms of IFNγ, granzyme B, and perforin mRNAs and proteins (Logan et al., 2012). Interestingly, this was paralleled with a loss of the nighttime peak in NK cell cytotoxic activity and lower protection against inoculated tumors (Logan et al., 2012). Altogether, these data demonstrate the importance of the circadian system in controlling NK cell function.
Box 3: Central or Local Rhythmic Control?
When studying circadian rhythms of physiological functions, including in the immune system, one important question to address is the source of the circadian regulation. On one hand, the rhythms could have their origin in the clock within the immune cells themselves. On the other hand, the circadian control could originate from other cells in the organism, particularly from the central clock of the SCN. It is well known that the SCN can control peripheral rhythms via different pathways, including endocrine mechanisms (rhythmic hormones that affect peripheral tissue function) and neuronal pathways, via the autonomic nervous system.
A recurrent feature in the literature on the circadian rhythms in the immune system is that in many cases, there is a control both by the local clock and by the central clock. For example, the local clock can gate the integration of the central signal (i.e., prevent or limit its effect at particular times of the day) or reinforce it when both regulatory mechanisms are in phase. Here are a few examples:
The recruitment of leukocytes to tissues (in situations of inflammation or under homeostasis) shows a circadian rhythm, which depends on the sympathetic nervous system (Scheiermann et al., 2012). However, the clock intrinsic to the leukocytes also plays a role, as illustrated by the rhythmic transcriptional regulation of the genes encoding chemokines involved in tissue recruitment (e.g., the rhythmic expression of the Ccl2 gene in monocytes and macrophages) (Sato et al., 2014).
The inflammatory response to LPS in the lung presents a daily rhythm (Gibbs et al., 2014). This is due to a rhythmic expression of the chemokine CXCL5 and recruitment of neutrophils. Interestingly, the Cxcl5 expression rhythm is dependent on the adrenal corticosterone. However, the response to corticosterone within the lung epithelial cells is gated by a rhythmic occupancy of the Cxcl5 promoter by the glucocorticoid receptor, a rhythm that is controlled by the local circadian clock in these cells.
NK cells have an intrinsic clock, which appears to play a role in the circadian rhythm of expression of cytolytic factors (granzyme B, perforin) (Arjona and Sarkar, 2005). However, chemical splenic sympathectomy leads to alteration of the rhythms of expression of clock genes, cytokines, and cytolytic factors in NK cells, which suggests that in addition to a local clock control, input from the central SCN clock also has a role to play (Logan et al., 2011).
Passive cutaneous anaphylactic reaction presents a circadian rhythm, which is lost in mice with clock function abolished specifically in mast cells (Nakamura et al., 2014b). However, the reaction is also controlled by the central SCN clock (Nakamura et al., 2014a) and by glucocorticoids (Nakamura et al., 2011; Nakamura et al., 2014b). As for the lung inflammation, a role of the local mast cell clock might be to gate the response to corticosterone.
Rhythms in Neutrophil Functions
Neutrophils are important cells of the innate immune system, attracted early at inflamed tissues, and acting both via phagocytosis and secretion of antimicrobial agents. As for other immunocompetent cells, neutrophils express clock genes (Haimovich et al., 2010). Importantly, some recent studies have started to uncover daily rhythms in the function of neutrophils. Gibbs and colleagues administered LPS to mice as an aerosol to study rhythms in the lung inflammatory response (Gibbs et al., 2014). Neutrophil recruitment to the lungs varied as a function of time of day, with more cells in the early rest period. This rhythm was not dependent on a clock in the neutrophils themselves. Instead, the clock in lung epithelial cells, called club cells, was shown to be required for the rhythmic recruitment of neutrophils, via the daily gating of the response of these cells to glucocorticoids, and the rhythmic production of the glucocorticoid receptor-regulated chemokine CXCL5, which is known to attract neutrophils to inflamed tissues (Gibbs et al., 2014).
Rhythmic neutrophil function may also be important for normal homeostasis, in the absence of inflammation. Casanova-Acebes and colleagues have identified a population of aged neutrophils, which show a daily rhythm of abundance in the blood (Casanova-Acebes et al., 2013). They showed that the clearance of these aged neutrophils triggers a reduction in the numbers of hematopoietic niche cells in the bone marrow. Since this hematopoietic niche acts to retain hematopoietic stem cells (HSCs) in the bone marrow, such a neutrophil-controlled rhythm might underlie the previously shown daily variations in HSC release in the bloodstream (Mendez-Ferrer et al., 2008). It is unclear which circadian clock is involved in the daily variations of neutrophil clearance and hematopoietic niche, but the authors showed an involvement of phagocytosis by macrophages, whose clock might have a role to play (Casanova-Acebes et al., 2013). In addition, prior work has shown that the circulation of HSCs is regulated by the autonomic nervous system (and thus, presumably, the central SCN clock) (Mendez-Ferrer et al., 2008). The authors also suggested that the rhythm of neutrophil clearance might also regulate other immune functions outside the bone marrow (Casanova-Acebes et al., 2013).
Circadian Influence On Adaptive Immunity
A pioneer study by Fernandes and colleagues in 1976 reported a rhythm of the adaptive immune response, measured as antibody production, following immunization with sheep red blood cells at different times of the day (Fernandes et al., 1976). These results suggested that either the B- or the T-lymphocyte response was influenced by the circadian system. More recent investigations have shown that the molecular clockwork is expressed within the adaptive immune system and that T-cell responses shows circadian rhythmicity.
Expression of the Clock Genes in Lymphocytes
Very few studies have evaluated whether T and B lymphocytes express the different clock genes and if their expression is rhythmic. In mouse lymph nodes (LNs), which mostly contain B and T cells, several clock genes are rhythmically expressed (Fortier et al., 2011; Keller et al., 2009), and this rhythm is blunted in LNs of ClockΔ19/Δ19 mutant mice (Fortier et al., 2011). Using PER2::Luciferase knock-in mice, it was shown that CD4+ T cells purified from spleen or thymus had a rhythm of luciferase activity, indicating that mouse CD4+ T cells express the clock gene Per2 in a rhythmic manner (Bollinger et al., 2011). Similarly, CD4+ T cells, purified from blood drawn from human subjects at different times over 24 h, showed a rhythm of expression of several clock genes (Bollinger et al., 2011). Moreover, clock gene expression was sustained following in vitro culture of human CD4+ T cells (Bollinger et al., 2011). Clock genes are also expressed in B lymphocytes enriched (80%) from mouse spleens (Silver et al., 2012a). Altogether, these results indicate that a functional molecular clockwork is present within the adaptive immune system and that this clock is at least present within CD4+ T cells and B cells. Further studies should reveal whether CD8+ T cells also express clock components.
Distribution of T-Cell Populations
It has been long recognized that blood counts of leukocytes, including lymphocytes, vary in a circadian manner. In human blood samples, T- and B-cell numbers are generally found to be high at night and then decline in the morning and remain low during the day (Abo et al., 1981; Besedovsky et al., 2014; Born et al., 1997; Dimitrov et al., 2009; Kirsch et al., 2012). Among the T lymphocytes, naive, central memory, and effector memory CD4+ and CD8+ T cells show similar rhythms (Besedovsky et al., 2014; Dimitrov et al., 2009; Kirsch et al., 2012). This morning decrease in T-cell numbers in the human blood coincides with the morning rise in cortisol and is associated with an increased expression of the chemokine receptor CXCR4 by T cells (Besedovsky et al., 2014; Dimitrov et al., 2009). Blocking the morning rise of cortisol in human subjects with the drug metyrapone prevents the morning decrease of T-cell number in the blood and inhibits the upregulation of CXCR4 (Besedovsky et al., 2014). Furthermore, ex vivo treatment of human T cells with cortisol leads to an increased expression of CXCR4 (Dimitrov et al., 2009). Therefore, the rhythm of T-cell counts in the blood is under the control of the circadian variation in cortisol production, which regulates the expression of CXCR4 on T cells, leading to their probable redistribution to the bone marrow, the site of production of the CXCR4 ligand (CXCL12). A circadian rhythm of T-lymphocyte numbers was also observed in rat blood samples, and this rhythm was maintained even when the central clock was inactivated (using extended exposure to constant light), suggesting the presence of an autonomous immune clock, possibly in T cells themselves (Depres-Brummer et al., 1997).
Although lymphocyte numbers displayed a rhythm in human and rat blood samples, no variations in B, CD4+, and CD8+ lymphocyte subsets were observed in the mouse LNs over a 24-h cycle under constant darkness (Fortier et al., 2011). Moreover, there was no circadian difference in the proportion of naive, regulatory, effector, and memory T cells (Fortier et al., 2011). In contrast, variation in B- and T-lymphocyte proportions was observed in LNs from rats kept in LD, with varying phases across studies (Bonacho et al., 2001; Esquifino et al., 2004).
In Vitro T-Cell Response
Initial experiments on rats suggested that B- and T-lymphocyte proliferation in response to B-cell (LPS) or T-cell (Concavalin A) mitogens was showing 24-h rhythms under a light-dark cycle (Esquifino et al., 1996).
More recently, the impact of the circadian clock on T-cell responses was reevaluated using more physiologically relevant T-cell stimulation assays. In humans, a rhythm of cytokine production (IL-2, IFNγ, IL-10, and TNFα) was observed when purified CD4+CD25– T cells, obtained at different times of the day, were stimulated in vitro with an anti-CD3 antibody, which mimics recognition of the antigen by the TCR (Bollinger et al., 2010). However, in these experiments, the authors did not measure T-cell proliferation, which could also show a daily rhythm, and therefore the amount of cytokine detected by enzyme-linked immunosorbent assay (ELISA) could have been influenced by the number of T cells present in the wells. In another report, the same group reported a 24-h variation of the response of human CD4+ T cells purified from blood at different time points (Bollinger et al., 2011). In this report, CD4+ T cells were stimulated for a short period (6 h) with PMA and ionomycin, and cytokine production by the T cells was measured using intracellular staining. It is important to note that only activated and memory CD4+ T cells will be able to produce cytokines in such a short period of stimulation. In this system, the authors observed a circadian variation of cytokine production (IL-2, IFNγ, and IL-17), both in the percentage of CD4+ T cells producing the cytokine and the amount of cytokine per cell (Bollinger et al., 2011). Furthermore, a rhythm of cytokine production was also observed over a 24-h period of in vitro culture (Bollinger et al., 2011). Using a microarray approach, the authors were able to identify genes that may be responsible for this circadian rhythm of cytokine production by CD4+ T cells. Among promising candidates are 2 genes, IκBα and SGMS2, involved in the regulation of the NF-κB pathway, which becomes activated in T cells following TCR stimulation (Bollinger et al., 2011). Another group observed a similar circadian variation of the human CD4+ T-cell response. A rhythm of IFNγ production by antigen-specific CD4+ T cells was observed after a short in vitro stimulation with adenovirus antigens or Staphylococcus aureus enterotoxin B, with a lowest response in the morning (Kirsch et al., 2012). While these results indicate that the circadian clock has an influence on human CD4+ T cell responses, additional work will be needed to determine whether CD8+ T-cell responses are also influenced.
In mouse LNs, both CD4+ and CD8+ T-cell proliferation in response to anti-CD3 stimulation showed a circadian rhythm in constant conditions, with stronger T-cell proliferation during the late subjective day and during the subjective night (Fortier et al., 2011). Furthermore, this T-cell proliferation rhythm was blunted when LNs were obtained from ClockΔ19/Δ19 mutant mice (Fortier et al., 2011). This T cell proliferation rhythm was not the result of variations in expression of the TCR/CD3 complex (Fortier et al., 2011). However, a circadian variation in the level of expression of ZAP-70, a downstream tyrosine kinase involved in TCR/CD3 signaling, could be correlated with the T-cell proliferation rhythm (Fortier et al., 2011). These results demonstrate that the circadian clock influences T-cell proliferation. Altogether, the data obtained with human and mouse T cells strongly suggest that several aspects of T-cell response, proliferation and cytokine production, are under control of the circadian clock. However, the mechanisms by which the clock regulates T-cell responses need to be defined.
In Vivo T-Cell Response
Very few studies have evaluated the impact of the circadian clock on in vivo T-cell responses. Fortier et al. have observed a rhythm of the in vivo CD8+ T-cell response to immunization of mice with dendritic cells loaded with the ovalbumin antigen. T-cell expansion and production of IFNγ were stronger when mice were immunized in the middle of the light phase than at night (Fortier et al., 2011). The same preparation of DCs was used to immunize mice at the different time points. So although DCs express clock genes (Silver et al., 2012a), the variation in T-cell responses is probably intrinsic to the CD8+ T cells (Fortier et al., 2011). However, it cannot be excluded that soluble factors or other variations within the lymphoid organ itself are contributing to the rhythm of in vivo CD8+ T-cell response. Further studies using conditional deletion of clock genes specifically in T cells should unravel whether the rhythmicity is intrinsic to T cells.
Another study has shown a day-night difference in the in vivo response of mouse CD4+ T cells to immunization with ovalbumin in the presence of CpG oligodeoxynucleotides (a ligand for the PRR TLR9). As a day-night difference in TLR9 expression was observed, the enhanced response of T cells during the night was believed to occur due to higher activation of antigen-presenting cells expressing TLR9 (Silver et al., 2012b). However, it is also possible that the rhythm of the CD4+ T cell response to the immunization was a direct consequence of an intrinsic rhythm of T-cell responses.
Recently, a diurnal regulation of the differentiation of mouse CD4+ T cells into IL-17–producing effector cells (Th17 subset) was observed (Yu et al., 2013). Mouse Th17 cells play a very important role for the control of bacterial and fungal infections (Korn et al., 2009). Following infection, the differentiation of Th17 cells from naive CD4+ T lymphocytes requires expression of the orphan nuclear receptor RORγt (Korn et al., 2009). RORγt is a short isoform of a nuclear receptor, RORγ, which is involved in the circadian clock and is rhythmically expressed in several tissues (Mongrain et al., 2008). A link with the circadian clock of T cells was revealed by studying Nfli3–/– mice, which have a higher frequency of Th17 cells within their small intestine. This increased frequency of Th17 occurs because NFIL3 (or E4BP4), a basic leucine zipper transcription factor, acts as direct repressor of Rorγt transcription. The Nfil3 gene is also a direct target of the circadian clockwork via the rhythmic transcription factor Rev-Erbα, which represses Nfil3 transcription. As a consequence, fewer Th17 cells are present in Rev-Erbα–/– mice (Yu et al., 2013). Consistent with this regulation by the clock, naive CD4+ T cells differentiated more into Th17 cells in vitro when CD4+ T cells were collected during the day than in the night (Yu et al., 2013). Given the long time course of the differentiation process, the Th17 cells did not show a rhythm in vivo. Nevertheless, the authors were able to show an in vivo role of the clock in Th17 differentiation by using a repeated phase advance experiment: following four 6-h shifts imposed every 3 days, a nearly 2-fold increase in Th17 frequency was observed in wild-type (WT) mice but not in Rev-Erbα–/– mice (Yu et al., 2013).
Involvement Of Circadian Clocks In Immune Disease Models
Rhythms in Allergic Reactions
Patients with allergic diseases such as asthma have a daily pattern of symptoms (Seery et al., 1998). Similarly, in mouse models, a circadian regulation of immediate-type skin allergic reactions has been observed. A passive cutaneous anaphylactic (PCA) reaction was induced by subcutaneous administration of anti-TNP IgE, followed by intravenous injection of DNP-BSA. The extent of the allergic skin reaction in this model showed a variation over the day-night cycle (Nakamura et al., 2011). This rhythmic response was shown to be lost in Per2 mutant mice and in adrenalectomized mice. This suggested a circadian influence via the rhythmic secretion of corticosterone (Nakamura et al., 2011). Furthermore, the response of mast cells to glucocorticoids was altered in Per2 mutant mice, suggesting that the clock gates this response in some way, which then regulates the daily variation in PCA reaction (Nakamura et al., 2011). However, since the Per2 mutation is present in all tissues in these mice, it was not possible to determine the source of the circadian regulation of PCA reaction and response to glucocorticoids. Mast cells express clock genes with a 24-h rhythm (Baumann et al., 2013; Nakamura et al., 2014b; Wang et al., 2011), so the circadian control could originate from the mast cell–intrinsic clock. To address this, Nakamura and colleagues used a mast cell–deficient mouse strain (the W/Wv mice) and injected them subcutaneously with bone marrow–derived cultured mast cells. These reconstituted mast cells were either WT or with the ClockΔ19/Δ19 mutation. The daily variation in PCA reaction was abolished in the mice with the Clock-mutant mast cells, implying a role for the clock intrinsic to these cells (Nakamura et al., 2014b). They further showed that this mast cell clock controls the rhythm of IgE-triggered mast cell degranulation. The mechanism appears to involve a CLOCK-dependent rhythmic regulation of the expression of the receptor for IgE in mast cells (Nakamura et al., 2014b). However, firm conclusions were hard to make, due to the very quick dampening of the rhythms in cultured mast cells, which prevented the authors to study the variations over a full 24-h cycle. They proposed that in vivo, the mast cell clock is likely to be entrained by rhythmic glucocorticoids, and they showed that indeed, glucocorticoids affect Per gene expression in mast cells in vitro and in vivo. Therefore, both the mast cell–intrinsic clock and rhythmic system cues seem to be at play.
The same group also studied the passive systemic anaphylactic reaction by measuring the drop in rectal temperature in response to the treatment. This was also dependent on the time of treatment (Nakamura et al., 2011). Interestingly, the day-night difference was lost upon SCN lesion (Nakamura et al., 2014a). This adds weight to the notion that allergic reaction in mice undergoes dual control by a local clock but also by systemic (SCN-driven) signals.
The direct influence of the circadian rhythm has not been evaluated for contact hypersensitivity (CHS), a T-cell–mediated allergic response. However, one study has shown an exacerbation of the inflammatory response during CHS in ClockΔ19/Δ19 mutant mice (Takita et al., 2013), suggesting a possible circadian regulation of this allergic reaction. Further studies should reveal whether the circadian clock also controls T-cell–mediated allergic reactions and if this control depends on a rhythmic response of T cells or other immune effectors involved in CHS.
Rhythms in Models of Rheumatoid Arthritis
Several studies have explored the involvement of circadian clock mechanisms in the pathogenesis of rheumatoid arthritis. To do this, Hashiramoto and colleagues used a mouse model of arthritis: intraperitoneal injection of anti–type II collagen antibody, followed by LPS injection the next day (Hashiramoto et al., 2010). This leads to arthritis symptoms in joints of the injected mice. In the synovial cells of arthritic foot joints, PER2 protein levels did not show the usual morning-evening difference. In the spleen, clock gene expression was also strongly altered. Interestingly, the arthritis symptoms were exacerbated in Cry1/Cry2 double KO mice (whose clock is nonfunctional), which was paralleled by increased serum levels of TNFα. Conversely, treatment with an anti-TNFα antibody reversed the phenotype of Cry double KO mice upon experimental arthritis (Hashiramoto et al., 2010). These results can be put in relation with those of Narasimamurthy and colleagues (already mentioned above), showing increased release of TNFα (and other cytokines) by Cry1/Cry2 KO macrophages (Narasimamurthy et al., 2012). Another study using a distinct model of mouse arthritis (injection of type II collagen emulsified in Freund’s adjuvant) suggested a link between melatonin, Cry1, and arthritis symptoms (Bang et al., 2012). Finally, work on human synovial cells, harvested upon joint surgery of patients with rheumatoid arthritis, uncovered a possible mechanism for the alteration of clock gene expression observed in the model described above. Treatment of the synovial cells with TNFα suppressed the expression of Per2 and increased that of Bmal1 and Cry1 (Yoshida et al., 2013). The changes in Per2 expression could be due to the strong changes in the expression of the transcription factors DBP, HLF, TEF, and E4BP4/NFIL3, which are known to bind D-boxes, DNA elements that are found in the Per2 promoter region (Yoshida et al., 2013). The reader is referred to a recent review by Gibbs and Ray (2013) for an in-depth discussion of the possible links between circadian clocks and rheumatoid arthritis. The authors present a model whereby not only the core clock mechanisms but also neuroendocrine pathways (including melatonin, glucocorticoids, adipokines) and nuclear receptor pathways (glucocorticoid receptor, ROR factors, PPAR factors) would affect the development of arthritis. They also advocate for these different regulatory pathways as possible therapeutic targets for the disease.
Conclusion
Even if the research to decipher the impacts of circadian clocks on immune cell function is in its infancy, it is clear that this circadian regulation is profound and widespread. All immunocompetent cell types studied so far show some kind of circadian regulation. In many cases, the clock intrinsic to these cells was shown to be required for the circadian regulation. Of note, though, a dual regulation by both the local clock and systemic cues can occur (see Box 3). However, a lot remains to be done to evaluate the consequences and physiological roles of the immune system clocks. Most studies so far have focused on acute responses such as the cytokine response to LPS stimulation, the proliferative response of T cells to antigen, or the expression of chemokines involved in recruitment of leukocytes to inflamed tissues. Future studies will need to measure the deeper circadian regulation of immune functions (e.g., the long-term acquisition of distinct phenotypes by the cells) and their development over the course of infection or cancer progression. To this end, mouse models of tissue-specific clock dysfunction coupled with advanced tools to dissect immune cell phenotypes will be of great help. Furthermore, the circadian regulation of immune cells will need to be integrated in a broader physiological context. In this regard, research will need to link the circadian-immune regulation to metabolic, neuroimmune, and cardiovascular systems. Finally, it will be important to address the significance of this clock regulation in the immune system for infection, autoimmune diseases, and cancer, both for the understanding of the etiology of these diseases and for the use of a chronobiological component in designing more efficient treatments.
Footnotes
Acknowledgements
The authors thank Dr. David Morse for proofreading the manuscript and members of the Labrecque and Cermakian laboratories for discussion. This work was supported by the Canadian Institutes of Health Research (grant number MOP-119322).
