Abstract
Purpose
As a usual malignant tumor in urinary system, renal cell cancer is regulated by microRNAs (miRNAs). This study revealed the prognostic value and regulatory effect of miR-190a-5p in renal cell cancer patients.
Methods
A total of 253 renal cell cancer patients were included for prognostic value analysis. The target gene of miR-190a-5p was detected by luciferase reporter assay. Cell Counting Kit-8 analysis and Transwell analysis were performed to explore the proliferation, removal capability, and invasiveness of 786–0 and A498 cells. Prognostic value was calculated by Kaplan–Meier curve and Cox regression analysis.
Results
miR-190a-5p was more down-regulated in tumor tissues than in adjacent tissues. Renal cell cancer cases were differed as low and high groups ground on mean miR-190a-5p expression in tumor tissues. Overall survival probability was obviously high in patients with high miR-190a-5p level (log-rank test
Conclusion
miR-190a-5p was reduced in renal cell cancer tissues, and predicted worse outcomes of renal cell cancer cases. Overexpressed miR-190a-5p could restrain the proliferation, removal capability, and invasiveness of renal cell cancer cells via suppressing GDF11.
Introduction
Renal cell cancer (RCC) is a common malignancy in urinary system, originating from the urinary tubular epithelia of the renal parenchyma. 1 Onset and death rates of RCC have been overstated in recent years, and the burden of RCC has continued to grow.2,3 Due to insensitivity to radiotherapy and chemotherapy, RCC is mainly treated by surgery, and the outcomes is poor.4,5 The prognostic factors of RCC patients include age, pathological characteristics of the tumor, growth rate, treatment method, surgical margin, etc.6,7 MicroRNAs (miRNAs), as a class of non-coding small RNA, regulate gene expression.8,9 MiRNA levels control the proliferation, differentiation, and apoptosis of cancer cells.10,11
miR-190a-5p is obtained from intron of the talin 2 gene. 12 Han et al. 13 suggested that miR-190a-5p was reduced in cervical cancers. Liang et al. 14 suggested that resveratrol could suppress the apoptosis of hepatocytes via reducing miR-190a-5p level. A negative relationship was discovered between ferroptosis and miR-190a-5p in rat cardiomyocytes; over-expressed miR-190a-5p could suppress reactive oxygen species (ROS). 15 Ferroptosis could induce the programmed cell death of renal parenchymal cells. 16 Sodium selenite could restrain the proliferation and migration, and promote apoptosis and ROS formation of 786-O cells. 17 Overexpressed miR-190 could restrain the proliferation, removal capability, and epithelial-mesenchymal transition (EMT) in breast cancer (BC). 18 Molecular data from The Cancer Genome Atlas indicated that miR-190 expression was correlated with the outcomes of kidney cancer. 19 Therefore, miR-190a-5p might participate in RCC development. However, there were no previous studies focused on the prognostic value and function role of miR-190a-5p in RCC patients.
Current research analyzed the prognostic value of miR-190a-5p in RCC patients. The regulatory role of miR-190a-5p on tumor progression also has been explored.
Methods
Participants
The sample size was calculated by GPower3.1 (http://www.gpower.hhu.de/), when the effect size = 0.3, α = 0.05, statistic power = 0.95, and the sample size was 253. Then 253 RCC patients were randomly selected from Tianjin Medical University General Hospital from June 2016 to May 2018. RCC patients were diagnosed by two pathologists according to the American Joint Committee on Cancer Staging. 20 Tumor tissues and adjacent tissues were obtained during the operations. Ratification was offered by the Institutional Review Board of the Tianjin Medical University General Hospital. Signed informed consent was provided by all patients before enrolling in this research. The follow-up time was 5 years.
Extraction of miRNA and quantitative real-time reverse-transcription polymerase chain reaction (qrt-PCR)
Total mRNA was separated by MagMAX™ mirVana™ total RNA extraction kit (Applied Biosystems, Thermo Fisher Scientific, Carlsbad, CA, USA) conforming with the direction. MiR-190a-5p and GDF11 were synthesized by SuperScript IV UniPrime One-Step RT-PCR System (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s instruction. Relative expression of miR-190a-5p and GDF11 were calculated by 2-ΔΔCt method. The reference genes were U6 and β-actin. The sequences are listed in Supplementary Table 1.
Cell culture
Human kidney epithelial cells (HKb-20, BNCC368130), RCC 786–0 (BNCC338472), and A498 (BNCC100609) cells were purchased from BeNa culture collection biotechnology Co., Ltd. HKb-20 and 786–0 cell lines were cultured in 90% DMEM-H medium with 10% fetal bovine serum (FBS), A498 was cultured in 90% EMEM medium with 10% FBS at 37 °C with 5% CO2.
Luciferase reporter assay
Target gene of miR-190a-5p was predicted by TargetScanHuman, StarBase, and miRDB databases. Venn diagram (http://www.bioinformatics.com.cn) obtained the candidate genes, then we selected GDF11 as the target gene.
Wild type (wt) or mutate type (mt) of GDF11 were transfected into miR-190a-5p mimic or miR-190a-5p mimic negative control (NC) cells by Lipofectamine 3000 (Invitrogen). Luciferase activity was tested by luciferase Reporter Assay System (Promega, Madison, WI, USA).
Cell transfection
miR-190a-5p mimic, miR-190a-5p mimic NC (negative control), GDF11 overexpression RNA (oeGDF11) and oe-NC sequences were conducted by Sangon Biotech (Shanghai) Co., Ltd (Shanghai, China). miR-190a-5p mimic and mimic NC were respectively mixed with Lipofectamine 3000 and transfected into 786–0 and A498 cell lines and incubated in serum-free DMEM-H medium at 37°C.
Cell counting kit (CCK)-8 analysis for proliferation
DMEM-H medium (100 μL) with 10% FBS (containing 106 cells) was cultured in 96-well plates. Cell proliferation of 786–0 and A498 cells in different transfected groups was detected by the CCK-8 kit (Shanghai Enzyme-linked Biotechnology Co., Ltd, Shanghai, China) following the directions. The OD values were detected at 450 nm after respectively cultured for 0, 12, 24, 48, and 72 h.
Migration and invasion analysis
Transfected cells (200 μL containing 2 × 104 cells) with serum-free DMEM-H medium were incubated in upper chamber of Transwell. Another 800 μL DMEM-H medium with 10% FBS was put into the lower chamber. After incubating at 37°C for 24 h, cells on the lower surface were fixed by paraformaldehyde and stained by 0.1% crystal violet. Migration cells were scanned and calculated under an inverted microscope in five randomly selected visual fields. The Matrigel matrix was precoated in the upper chamber, to perform the invasion assay.
Data analysis
GraphPad Prism 9.0 was used to plot the figures. All data were evaluated by SPSS 22.0. The differences between groups were calculated by χ2 test or independent sample
Results
miR-190a-5p was decreased in RCC tissues
Reduced miR-190a-5p level was discovered in the tumor tissues (0.593 ± 0.175) than in adjacent tissues (1.000 ± 0.144) (Figure 1(A),
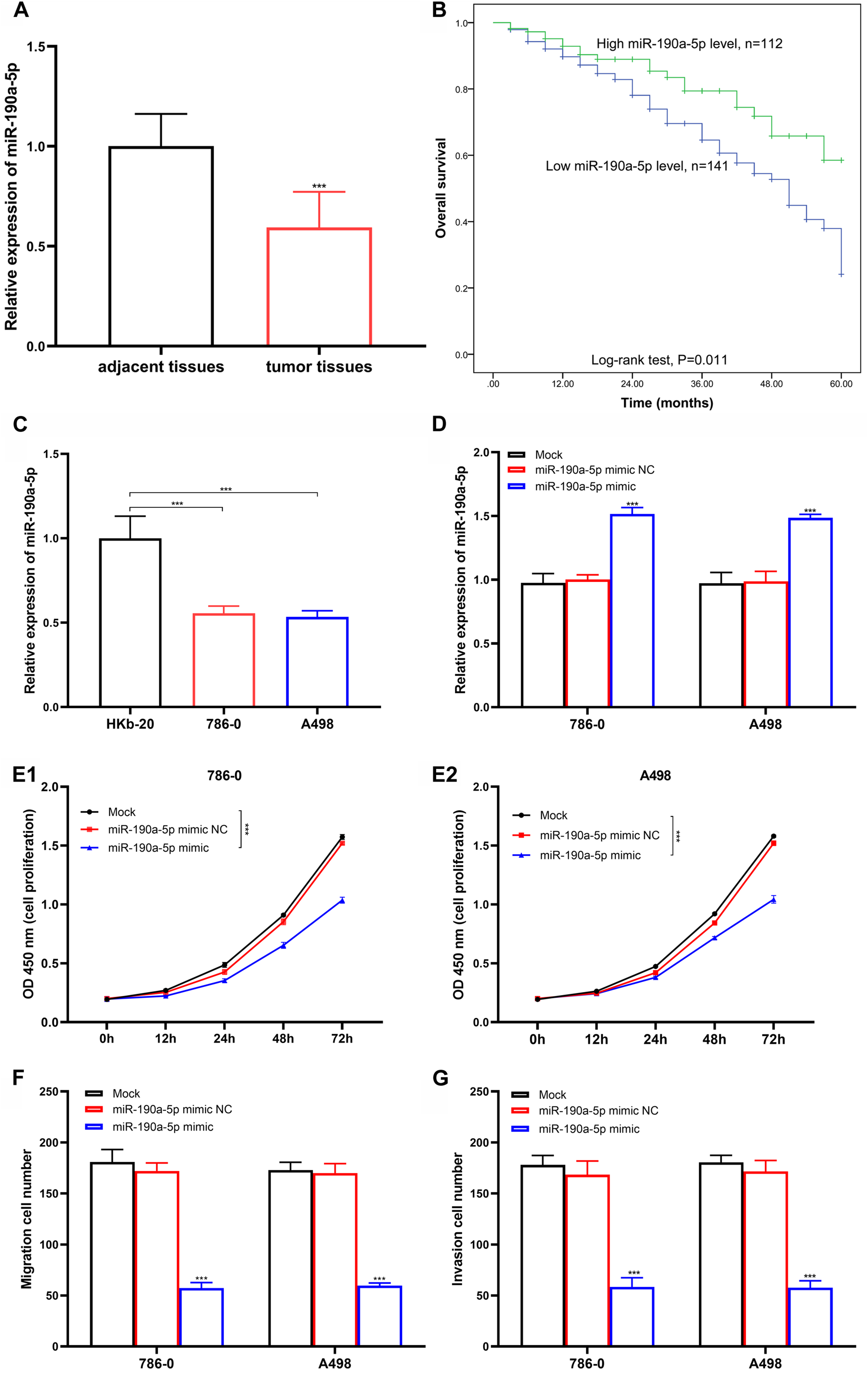
Influences of miR-190a-5p for RCC development. (A) miR-190a-5p levels in RCC tissues and adjacent tissues. ***
Baseline and clinical characteristics of RCC.
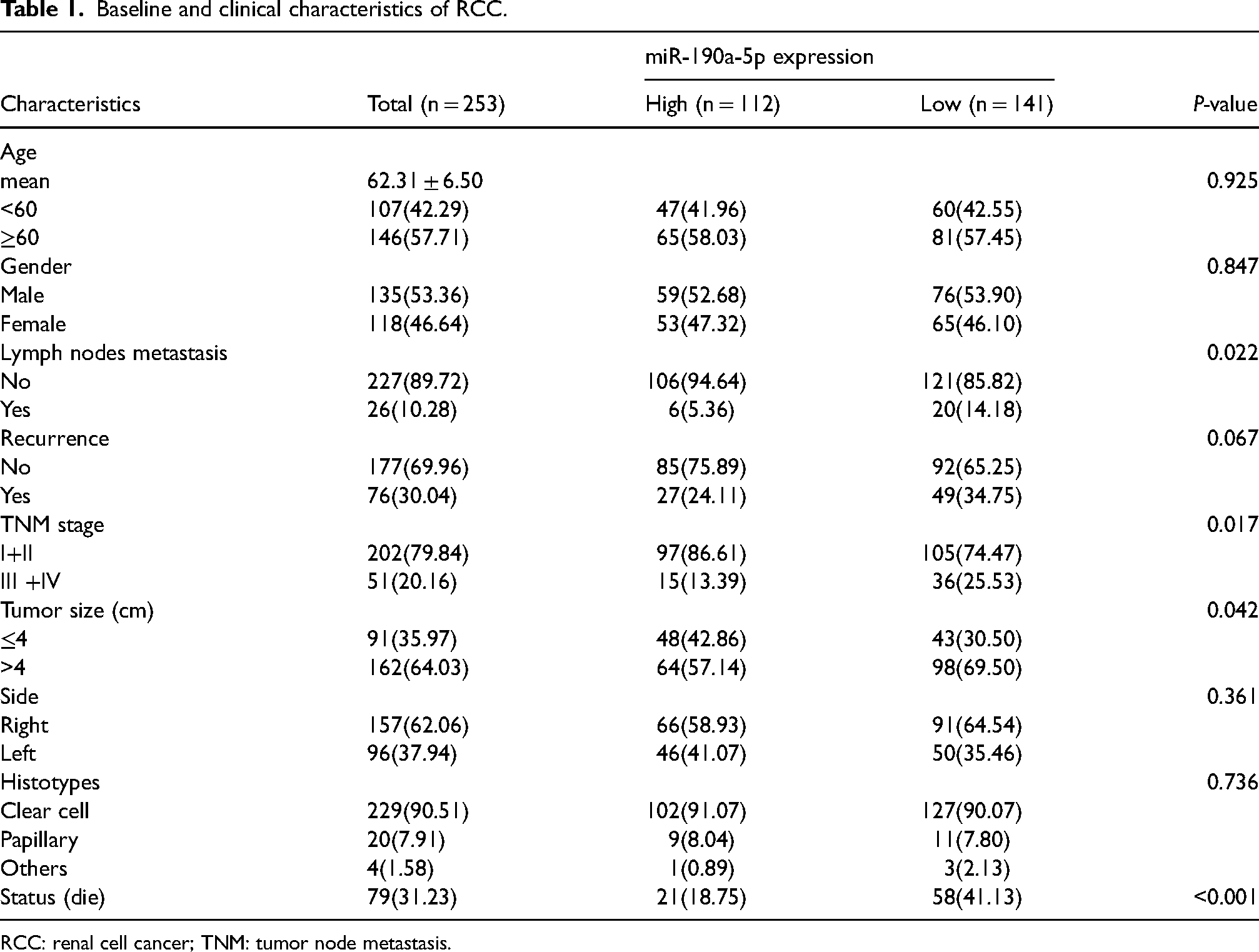
RCC: renal cell cancer; TNM: tumor node metastasis.
Prognostic value of miR-190a-5p for RCC
A total of 79 patients (31.23%) died during the 5-year follow-up (21 cases in the high group and 58 in the low group). The KM curve was performed to explore prognostic effects of miR-190a-5p for RCC and found that the overall survival probability was high in patients with a high miR-190a-5p level (Figure 1(B), log-rank test
Survival analysis of characteristics, which were significantly different between the high and low groups, was performed by the Cox regression model. In univariate analysis, low miR-190a-5p expression (relative risk (RR) = 1.870, 95% confidence interval (CI) = 1.134–3.083,
Univariate and multivariate survival analyses.
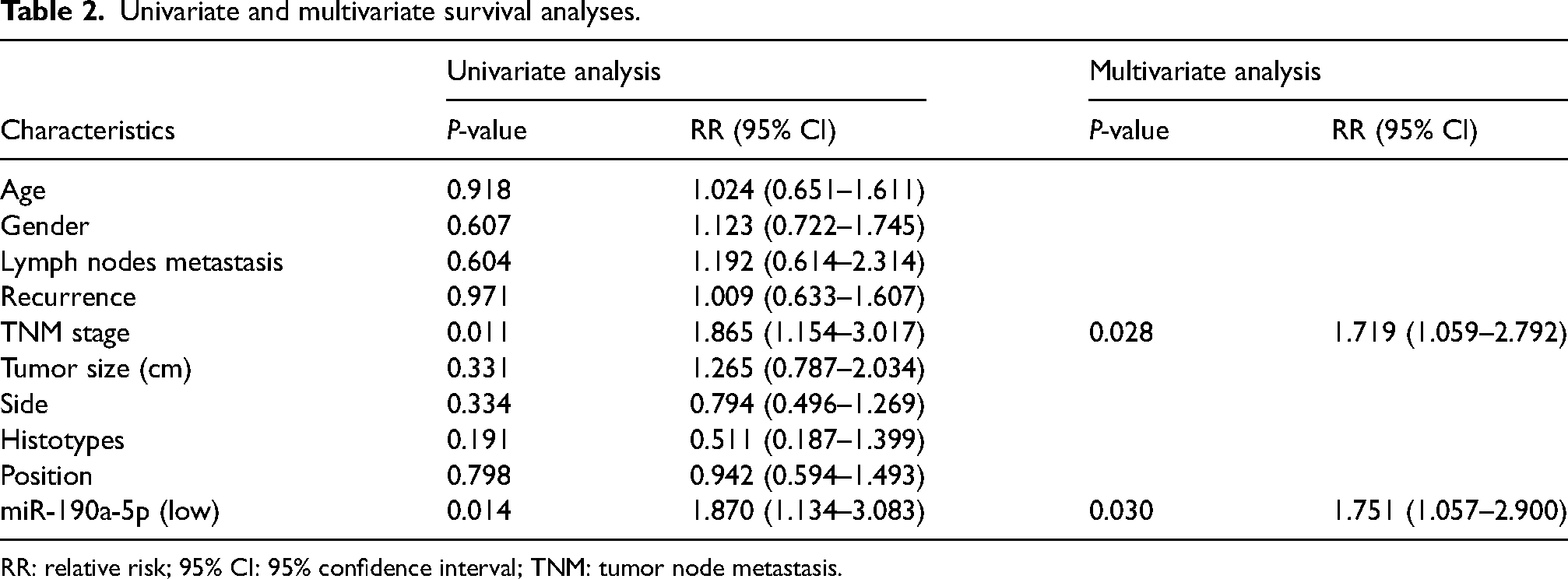
RR: relative risk; 95% CI: 95% confidence interval; TNM: tumor node metastasis.
Effects of miR-190a-5p for cell viability
Relative miR-190a-5p level was also reduced in 786–0 and A498 RCC cells when compared with HKb-20 (Figure 1(C),
GDF11 is a direct target gene of miR-190a-5p
Target gene of miR-190a-5p was selected from TargetScanHuman, StarBase, and miRDB databases. Common elements in three databases were gathered by the Venn diagram using bioinformatics (http://bioinformatics.com.cn). Five intersection genes have been found (Figure 2(A)), and GDF11 was selected for the following function analysis. miR-190a-5p directly combined with GDF11 mRNA at the position 2687–2694 of 3ʹUTR (Figure 2(B)). Relative GDF11 expression was significantly up-regulated in RCC cell lines than in HKb-20 cells (Figure 2(C),
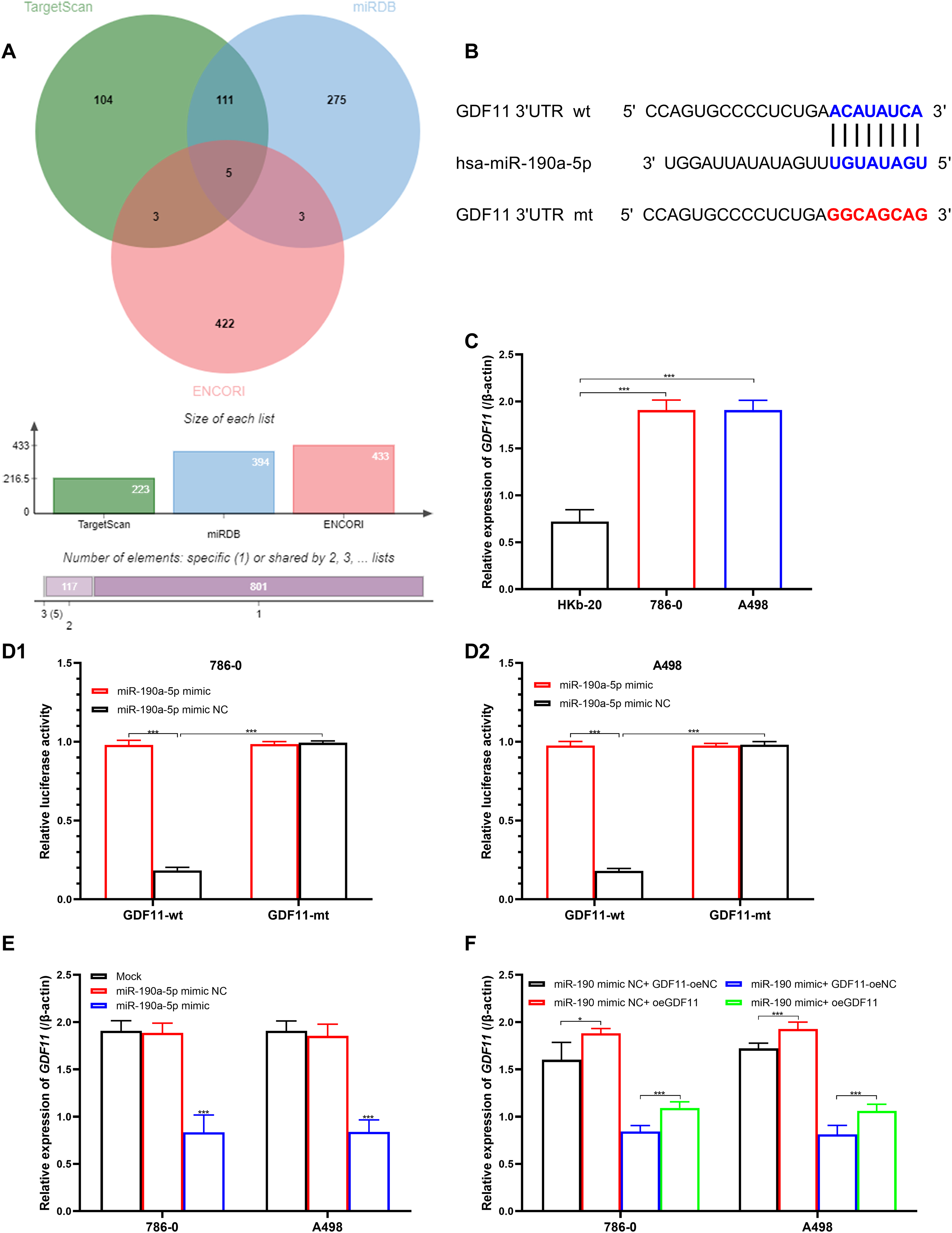
GDF11 is a target gene of miR-190a-5p. (A) Venn diagram for the selection of miR-190a-5p target genes. (B) Combined position between miR-190a-5p and GDF11 3′UTR. (C) Relative expression of GDF11 in HKb-20, 786–0, and A498 cell lines. (D1) Luciferase reporter assay between miR-190a-5p and GDF11 in 786–0 cells. (D2) Luciferase reporter assay between miR-190a-5p and GDF11 in A498 cells. (E) GDF11 levels in miR-190a-5p mimic transfected 786–0, and A498 cells. (F) Relative expression of GDF11 in 786–0, and A498 cells with the combined transfection of miR-190a-5p mimic and oeGDF11.
GDF11 could reverse the influences of miR-190a-5p for RCC cells
Cell proliferation of 786–0 (Figure 3(A1),
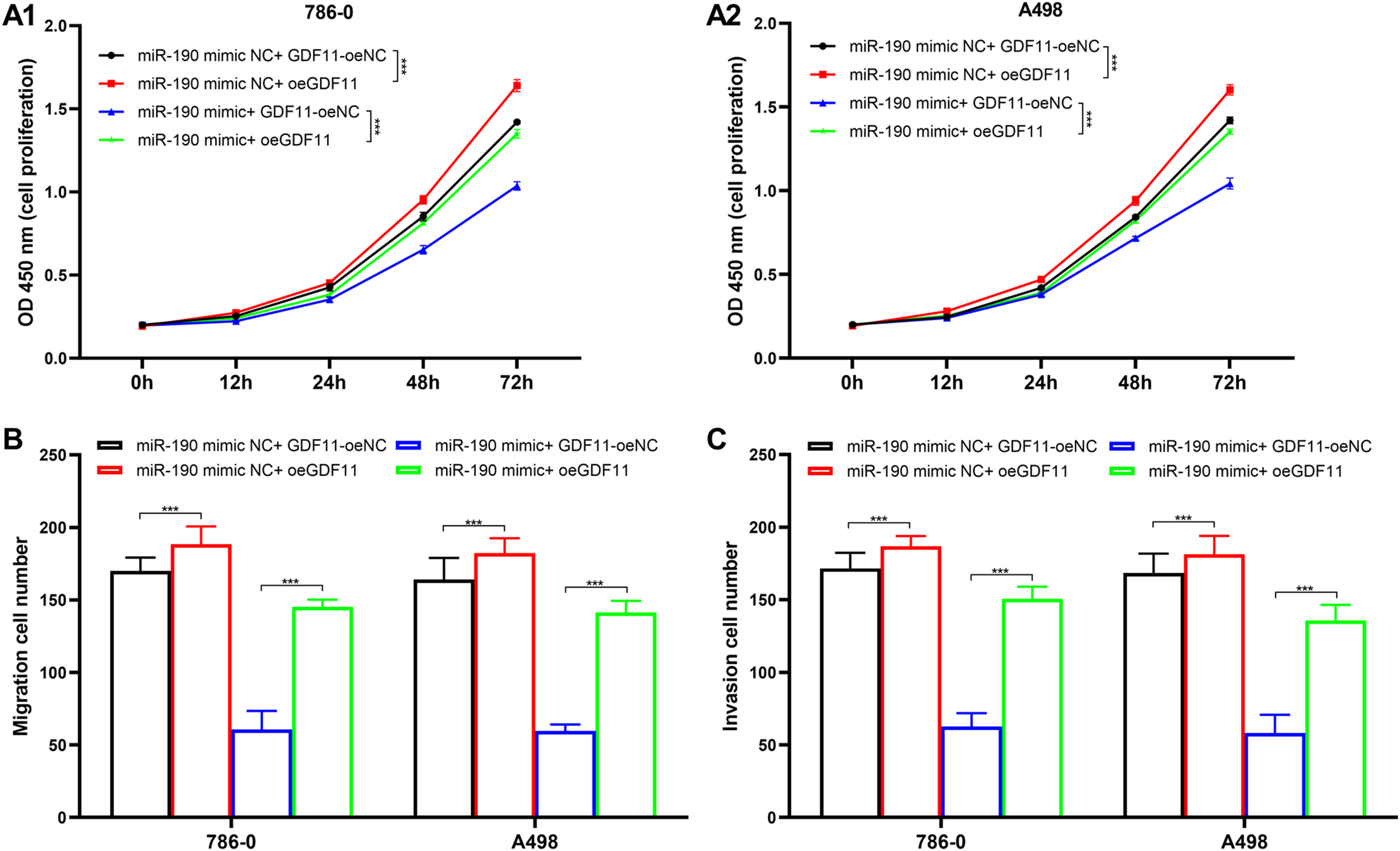
Effects of GDF11 for cell viability. (A1) Effects of GDF11 for cell proliferation in 786–0 cells. (A2) Effects of GDF11 for cell proliferation in A498 cells. (B) Effects of GDF11 for migration of 786–0 and A498 cell lines. (C) Effects of GDF11 for invasion of 786–0 and A498 cell lines. ***
Discussion
Since RCC is a common tumor in the urinary system, many patients have tumor metastasis and recurrence after surgery.21,22 Studies have confirmed that abnormal expression of miRNA has been found in several tumors, including RCC, and miRNA plays a crucial role in the occurrence and progression of RCC.23,24 Also, it shows good research potential in tumor development, providing a new way for diagnosis and therapy of RCC. Current research discovered that miR-190a-5p was decreased in RCC tissues compared to adjacent tissues. Based on the mean miR-190a-5p level in tumor tissues, RCC cases were divided into low and high groups. Patients in the low group had more lymph node metastasis, high TNM stages, and large tumor size. Previous studies suggested that high miR-190 levels were closely related to better outcomes of renal cancer. 19
In this study, the 5-year overall survival frequency of RCC cases was 68.77%; RCC patients with high miR-190a-5p levels had a longer survival time than patients in the low-levels group. Cox regression analysis demonstrated that low miR-190a-5p level and high TNM stage were independent risk factors for worse RCC prognosis. miR-190a-5p mimics were transfected into 786–0 and A498 cells; overexpressed miR-190a-5p could suppress the proliferation, removal capability, and invasiveness. Similar results were revealed in previous research. Yu et al. 25 suggested that miR-190 was reduced in estrogen receptor (ER)-BC cells than ER + cells. Elevated miR-190 could suppress the proliferation and decrease the percentage of the S phase of ER-BC cells. Reduced miR-190 could promote the EMT and metastasis of BC via SMAD2. 26 However, miR-190 was up-regulated in gastric cancer tissues. 12 This difference might be induced by the type of tumors.
We selected target genes from TargetScanHuman, StarBase, and miRDB databases, and found five intersection genes. We selected GDF11 for the following function analysis. GDF11 is upregulated in mice with renal diseases and promotes the proliferation and EMT of epithelial cells via SMAD3. 27 Therefore, we speculated that GDF11 might contribute to RCC development. Elevated GDF11 levels were discovered in 786–0 and A498 cells compared with HKb-20 cells. GDF11 was directly targeting to miR-190a-5p. Overexpressed GDF11 was transfected into miR-190a-5p mimic cells. Proliferation, removal capability, and invasiveness of 786–0 and A498 cells were significantly facilitated by miR-190a-5p mimic + oeGDF11. Increased GDF11 could promote the proliferation, migration, and invasion of 786–0 and A498 cells, which is induced by overexpressed miR-190a-5p. All of these results suggested that oeGDF11 could invert the inhibition of proliferation, removal capability, and invasiveness of 786–0 and A498 cells caused by miR-190a-5p. Current results were confirmed with previous studies. Pons et al. 27 suggested that GDF11 was elevated in kidney tissues with acute injury. High GDF11 could induce EMT and kidney injury in mice models. Gu et al. 28 suggested that GDF11 level was positively with the differentiation and proliferation of esophageal cancer cells.
Shortcomings of the current study should be mentioned. First, only one population and two cell lines were used in this study; other populations and more cell lines should be gathered for prognostic and functional analyses. Second, subgroup analysis based on different RCC subtypes was not performed due to the sample size not being large enough. Third, the influences of miR-190a-5p for the downstream signaling pathway of GDF11, apoptosis, inflammatory responses, and other responses were not analyzed in this study. Also, the influence of miR-190a-5p for RCC development was not certified in animal models. Therefore, further studies focused on the function analysis of miR-190a-5p should be certified in other cell lines and animal models in the future.
In a word, reduced miR-190a-5p in RCC tissues predicted worse outcomes for RCC patients. Overexpressed miR-190a-5p could restrain the proliferation, removal capability, and invasiveness of RCC cells via suppressing GDF11.
Supplemental Material
sj-docx-1-jbm-10.1177_03936155241290251 - Supplemental material for Prognostic value of miR-190a-5p in renal cell cancer and its regulatory effect on tumor progression
Supplemental material, sj-docx-1-jbm-10.1177_03936155241290251 for Prognostic value of miR-190a-5p in renal cell cancer and its regulatory effect on tumor progression by Jun Liu, Lili Zhang, Zhancheng Wang, Hu Li, Bo Wang and Xiaoqiang Liu in The International Journal of Biological Markers
Supplemental Material
sj-docx-2-jbm-10.1177_03936155241290251 - Supplemental material for Prognostic value of miR-190a-5p in renal cell cancer and its regulatory effect on tumor progression
Supplemental material, sj-docx-2-jbm-10.1177_03936155241290251 for Prognostic value of miR-190a-5p in renal cell cancer and its regulatory effect on tumor progression by Jun Liu, Lili Zhang, Zhancheng Wang, Hu Li, Bo Wang and Xiaoqiang Liu in The International Journal of Biological Markers
Footnotes
Acknowledgments
Not Applicable.
Ethical considerations
Ratification was offered by the Institutional Review Board of Tianjin Medical University General Hospital.
Consent to participate
Signed informed consent was provided by all patients before enrolling in research.
Consent for publication
Not applicable.
Data availability
All data generated or analyzed during this study are included in this article and its supplementary material files. Further enquiries can be directed to the corresponding author.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was funded by National Natural Science Funds of China (82171594).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
