Abstract
Canine cutaneous mast cell tumors (ccMCTs) are frequent neoplasms with variable biological behaviors. Internal tandem duplication mutations in c-KIT exon 11 (c-KIT-11-ITD) are associated with poor prognosis but predict therapeutic response to tyrosine kinase inhibitors. In a previous work, deep learning algorithms managed to predict the presence of c-KIT-11-ITD on digitalized hematoxylin and eosin-stained histological slides (whole-slide images, WSIs) in up to 87% of cases, suggesting the existence of morphological features characterizing ccMCTs carrying c-KIT-11-ITD. This 3-stage blinded study aimed to identify morphological features indicative of c-KIT-11-ITD and to evaluate the ability of human observers to learn this task. 17 untrained pathologists first classified 8 WSIs and 200 image patches (highly relevant for algorithmic classification) of ccMCTs as either positive or negative for c-KIT-11-ITD. Second, they self-trained to recognize c-KIT-11-ITD by looking at the same WSIs and patches correctly sorted. Third, pathologists classified 15 new WSIs and 200 new patches according to c-KIT-11-ITD status. In addition, participants reported microscopic features they considered relevant for their decision. Without training, participants correctly classified the c-KIT-11-ITD status of 63%–88% of WSIs and 43%–55% of patches. With self-training, 25%–38% of WSIs and 55%–56% of patches were correctly classified. High cellular pleomorphism, anisokaryosis, and sparse cytoplasmic granulation were commonly suggested as features associated with c-KIT-11-ITD-positive ccMCTs, none of which showed reliable predictivity in a follow-up study. The results indicate that transfer of algorithmic skills to the human observer is difficult. A c-KIT-11-ITD-specific morphological feature remains to be extracted from the artificial intelligence model.
Keywords
The development of artificial intelligence (AI)-based tools for diagnostic assistance purposes in the medical field has advanced rapidly in recent years. Among other applications, deep learning models (DLMs) have shown their efficacy and reliability in human medicine for detecting diabetic retinopathies, 11 pulmonary nodules, 22 heart diseases, 6 or even in identifying primary tumors from cancers of unknown primary origin 29 based on digital histologic samples or diagnostic imaging. In veterinary medicine, several DLMs have been developed, for example, for the histologic classification of skin neoplasms,16,43 automated mitotic count,3,4 or automated nuclear pleomorphism measurement. 21 One of the key elements often discussed when transferring DLMs to a clinical context is the interpretability of the AI-generated results.20,40 In some cases, such as mitotic detection, verification of the result using heat map-highlighted areas is relatively easy to interpret and understand by the human user. In contrast, for tasks where the AI models are trained under weakly supervised conditions to identify unknown patterns (eg., molecular expression prediction tasks), it is not necessarily clear how the AI model decides.
To diagnose canine cutaneous mast cell tumors (ccMCTs), a plethora of prognostic morphological factors in hematoxylin and eosin (HE)-stained slides, as well as immunohistochemical factors, have been identified over the years.17,18,41,42 The presence of internal tandem duplication in the exon 11 of the c-KIT gene (c-KIT-ITD-11) has been associated with higher malignancy and better response to therapy with tyrosine kinase inhibitors. 9 The European Medicines Agency requires polymerase chain reaction (PCR) confirmation of this mutation to start treatment with mastinib. 15
In a previous work, DLMs were trained under weak supervision to predict the presence of c-KIT-ITD-11 in HE-stained digitalized histological slides (whole-slide images, WSIs). 37 In this context, weakly supervised training means that the only information provided for each WSI was whether it was c-KIT-ITD-11 PCR-positive or PCR-negative. The DLMs achieved an average accuracy of 79% (75%–87%) in predicting c-KIT-11-ITD status, suggesting the existence of an as-yet unidentified morphological feature distinguishing c-KIT-ITD-11-positive from -negative ccMCTs. Describing this feature would make the DLMs’ predictions more interpretable and could potentially be integrated into histologic prognostic assessment.
This study aimed at identifying one or more specific morphological features of c-KIT-ITD-11-positive ccMCTs and to evaluate the ability of pathologists to learn this prediction task from AI. In a follow-up study, 4 pathologists assessed the predictability of 4 potential morphological features suggested in the first experiment: cellular pleomorphism, anisokaryosis, cytoplasmic granularity, and extracellular meshwork.
Materials and Methods
Participants
The target group for this study was veterinary pathologists of varying experience levels. Seventeen participants registered via an online form between March and September 2024, through which information about the participants’ work experiences in veterinary diagnostic pathology, yearly case load, frequency of ccMCT diagnosis, and frequency of requested PCR-based analysis of c-KIT-11-ITD were collected. A blank registration form is available for reference in the supplementary materials (Supplemental Form S1). The participants were divided into 2 groups (9 in group A and 8 in group B), aiming to mirror the spectrum of experience levels in both.
Digital Slides and Patches
All WSIs and patches used in this study were generated in a previous study by this group. To briefly summarize, 368 HE-histological slides of distinct ccMCTs, produced by the Veterinary Diagnostic Laboratory of Michigan State University in the USA (stain A) between 2018 and 2022 as part of routine diagnostics (including a PCR evaluation of c-KIT-11-ITD) were digitalized at 40x magnification with 3 different slide scanners. The same slides were de-stained and re-stained in HE at the Institute for Veterinary Pathology of Free University Berlin in Germany (stain B), as described in a previous work, 37 and digitalized with the same 3 scanners, resulting in 6 distinct data sets. To train and compare 6 distinct DLMs, all data sets were identically divided into training, validation, and test splits. To process the large WSIs, the DLMs segmented the background from the tissue area and divided the latter in multiple 256 × 256-pixels patches (little snippet of the original image, also referred to as image “tiles”), which were analyzed individually. A label (positive or negative) was attributed to each WSI, in line with the weakly supervised nature of training (ie, the c-KIT-11-ITD status is known only at the WSI level, not at the patch level). During training, the model assigned a relevance score (from 0 for nonrelevant to 1 for highly relevant) to each patch, reflecting their estimated weight in the global WSI label. Note that these scores were not supervised directly; they were learned internally by the model, using an attention-based mechanism implicitly highlighting patches most informative for the WSI label. 24 Supplemental Figure S1 schematically depicts this approach. The relevance scores can be visualized as heat maps, with bluish colors indicating low relevance and reddish tones high relevance. 37
The image subsets selected for this study were extracted from the 2 data sets scanned with an Aperio AT2 scanner (best image resolution) and were labeled A and B. Group A worked with subset A and group B with subset B.
WSI selection
Eight and 15 WSIs were randomly selected (of 368 WSIs) for phase 1A and the challenge (see main study structure below), respectively. Attention was paid to ensure parity between high- and low-grade MCTs as well as between c-KIT-ITD-11-positive and c-KIT-ITD-11-negative cases. WSIs exhibiting at least one of the following criteria were excluded: very faint staining, blurry areas, artifacts, very few neoplastic cells, extensive necrotic areas. Subsets A and B both contained 23 WSIs of the same tumor section, each in their respective staining.
Patch selection
A total of 400 highly relevant patches were selected: 200 for phase 1B and 200 for the challenge (see study structure below). The 50 patches with the highest relevance scores of WSIs correctly labeled by the DLMs (true positive and true negative) were extracted out of the training and validation splits of the data sets for phase 1B and from the test split for the challenge. Patches showing at least one of the following exclusion criteria were eliminated: less than 5 neoplastic cells (mostly tumor margins or areas with hemorrhage), blurriness, presence of artifacts, more eosinophils than neoplastic cells, necrotic areas. Up to 3 patches per WSI were randomly selected out of the remaining pool. Again, parity between high- and low-grade ccMCTs as well as between c-KIT-ITD-11-positive and c-KIT-ITD-11-negative cases was maintained. Note that the patches in the 2 subsets were not identical and did not necessarily originate from the same slides, as they were segmented and analyzed by 2 distinct DLMs with differing performance levels. Subset A was composed of 217 different WSIs, while subset B contained 211 WSIs. Twenty-six WSIs appeared only in subset A and 20 WSIs appeared only in subset B.
Patch selection for the follow-up study
Forty patches were selected from the phase 1B subsets in both stains A and B (the first 20 c-KIT-11-ITD-positive and first 20 c-KIT-11-ITD-negative cases, based on patch number). Those patches originated from 37 WSIs from subset A, and from 38 WSIs from subset B; 24 and 25 WSIs appeared only in the A and B cuts, respectively.
Study Structure
The study consisted of 4 phases (Fig. 1), which participants were required to complete within 3 weeks.
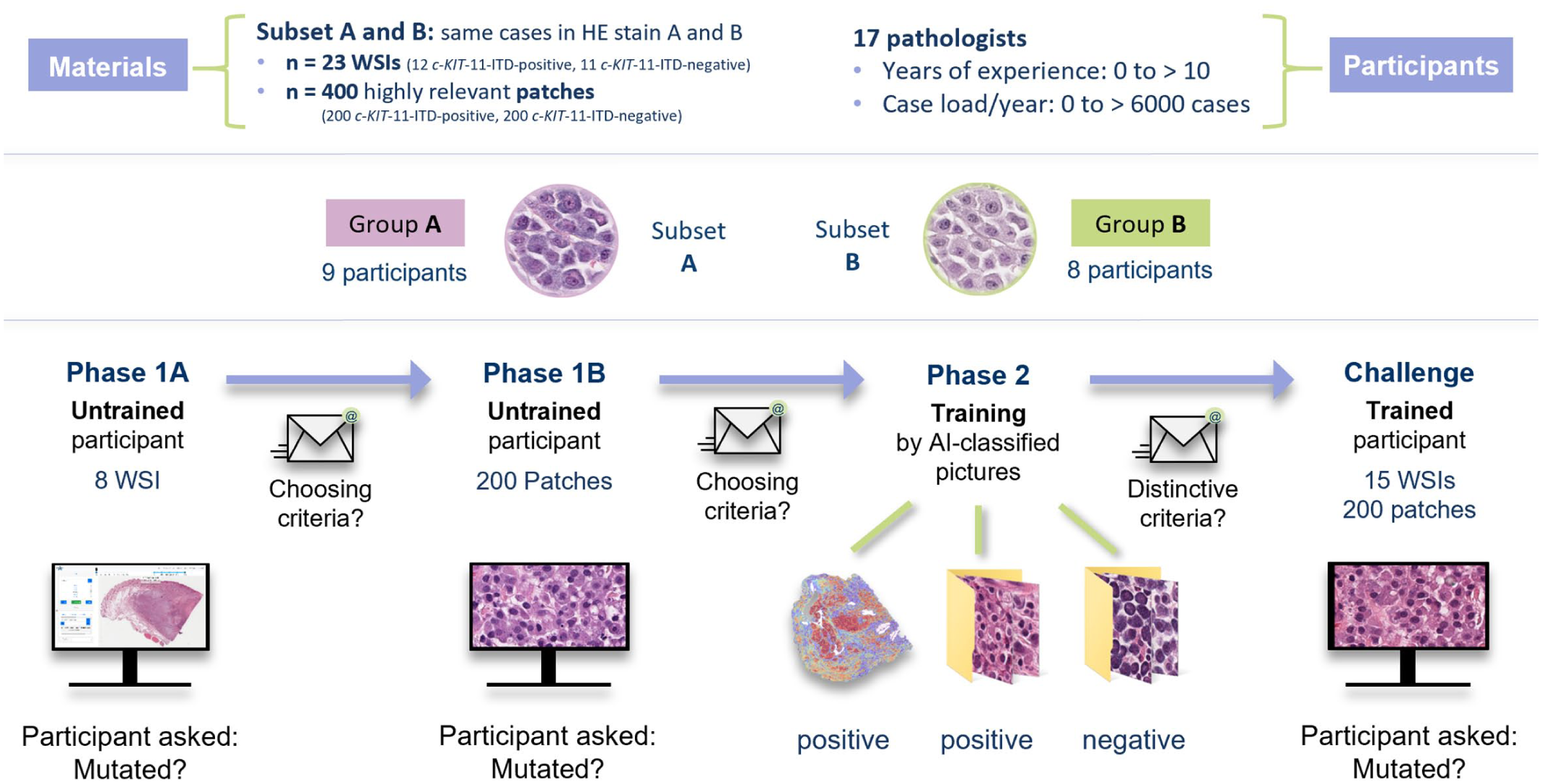
Study scheme. Two groups of participants (groups A and B) worked with digitalized histologic images of canine cutaneous mast cell tumors from subsets A and B, respectively. The subsets differed in their hematoxylin and eosin (HE) stains. Stain A corresponded to the original slide staining from the Veterinary Diagnostic Laboratory of Michigan State University (USA), while stain B was produced by destaining and restaining these slides at the Institute for Veterinary Pathology, Free University Berlin (Germany). Both groups followed the same study scheme. In phase 1, participants labeled 8 unknown whole-slide images (WSIs, 1A) and 200 highly relevant patches (1B) as either positive or negative for ITD in exon 11 of c-KIT (c-KIT-11-ITD). After each step, they submitted a form listing the morphological features their decisions were based on. In phase 2, the same WSIs (with heat maps highlighting relevant areas) and patches were displayed along with their correct labels, allowing participants to learn and identify morphological features, which they reported in a third form. The challenge consisted of labeling 15 new WSIs and 200 new patches. AI, artificial intelligence; ITD, internal tandem duplication.
Phase 1A (WSI baseline)
To assess the participants’ baseline performance, they were asked to review 8 WSIs and label them as c-KIT-11-ITD “positive” or “negative” according to their intuition.
Phase 1B (Patch baseline)
In this second baseline phase, participants were asked to intuitively sort 200 patches into “positive” or “negative.”
Phase 2 (Training)
Participants revised the same WSIs and patches from phases 1A and 1B; however, this time they were labeled and sorted by the DLMs. A heat map overlay indicating the relevance of the patches was provided, with a color scale ranging from blue (low relevance) to red (high relevance).
Challenge
Participants were now considered “trained” by the DLMs. They were tasked with labeling 15 new WSIs and 200 new highly relevant patches according to their c-KIT-11-ITD status.
Data Availability
The WSIs were made available on the open-source online platform EXACT (EXpert Algorithm Collaboration Tool), 30 a digital microscopy and labeling tool allowing the participants to navigate and zoom freely through the WSIs. The patches were uploaded to a Microsoft OneDrive folder, where participants could sort them directly into a “positive” and “negative” subfolder. Alternatively, the folder could be downloaded, sorted into the respective subfolders offline, and returned sorted to the study organizer as a .zip file.
Collection of Morphological Features
At the end of phases 1A, 1B, and 2, participants were asked to fill in a Microsoft Form (Supplemental Form S2) with 3 open-ended questions concerning the morphological features they identified as suggestive for c-KIT-11-ITD-positive and c-KIT-11-ITD-negative cases, as well as the time spent on the study phase. For phases 1A and 1B, the questions referred to the participants’ personal opinions. For phase 2, the questions referred to what seemed to be the decision features of the DLM.
A list of morphological features was compiled from the open answers for each phase, replacing descriptive or colloquial terms with corresponding technical terms. Opposite terms or semi-quantitative statements were grouped into summarizing features (eg, “monomorphism” and “polymorphism” into “shape”). A detailed description of the methodology can be found in Supplemental Material S1.
Follow-Up Study—Testing of Morphological Features
A follow-up study (Fig. 2) was conducted to determine the discriminative ability of the following morphological features: cell shape (monomorphism or pleomorphism), nucleus size (isokaryosis or anisokaryosis), and cytoplasmic granules (visible or not visible) as the top 3 global features of the main study. The prevalence of these 3 morphological features was consecutively assessed by 3 participants (1 from group A and 2 from group B) in each 40 stain A and B patches (randomly selected of subsets A and B, 20 c-KIT-11-ITD-positive and 20 c-KIT-11-ITD-negative cases). The patches are available in the Supplemental File “Patches.zip.”
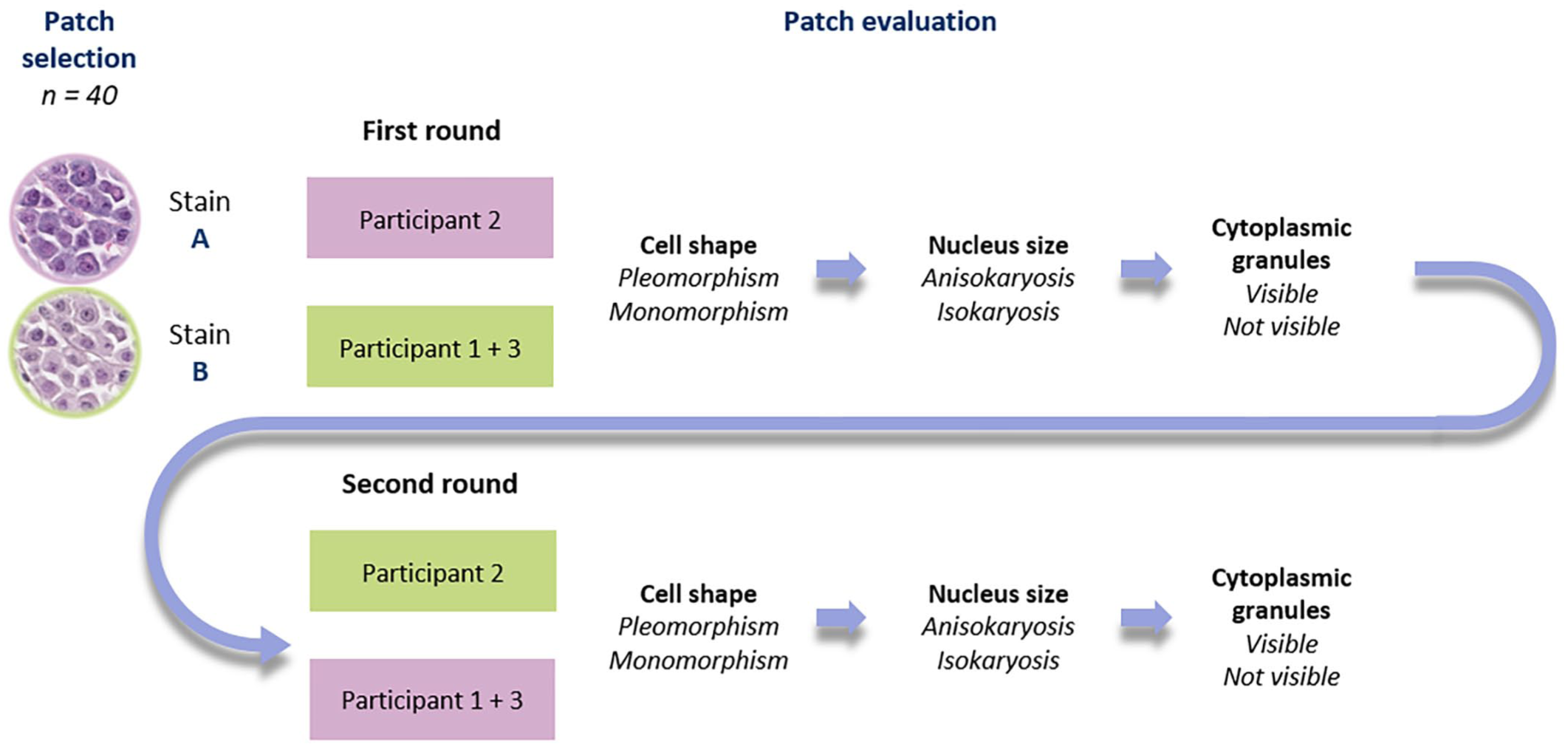
Follow-up study scheme. Forty patches were randomly selected of subsets A and B (parity of c-KIT-11-ITD-positive and c-KIT-11-ITD-negative cases). Three participants (1–3) were divided into 2 groups. In the first round, the stain A patches were given to participants 2, while participants 1 and 3 received the stain B patches. They consecutively assessed 4 morphological features: cell shape (pleomorphism or monomorphism), nucleus size (anisokaryosis or isokaryosis), and cytoplasmic granules (visible or not visible). In the second round, the participants repeated the consecutive assessment looking at 40 patches in the other stain (stain A for participants 1 and 3 and stain B for participant 2).
Statistical Analysis
In this article, the percentage of correct classifications will be referred to as classification performance (CP, sum of true positives and true negatives divided by the sum of true and false positives and negatives, multiplied by 100). The sensitivity (Sen) measures the true-positive detection rate (number of true positives divided by the sum of true positives and false negatives), whereas the specificity (Spe) measures the true negative detection rate (number of true negatives divided by the sum of true negatives and falls positives). CP, Sen, and Spe are reported as averages for the 2 different groups. These performance metrics were calculated using Microsoft Excel 2021. For all these metrics, a score of 70% and above is considered good. Regarding the follow-up study, an interrater reliability analysis was performed in R (version 4.4.2) using Cohen’s kappa and weighted kappa statistics. 19 A kappa score greater than 0.6 indicated acceptable interrater reliability, with values closer to 1 reflecting higher prediction consistency.
Results
Participants
All 17 participants were veterinarians. Two participants did not currently work as diagnostic pathologists and 1 participant never did. Fourteen participants were currently working in diagnostic pathology, with experience ranging from first-year residents examining 10–500 histological cases per year, to certified veterinary pathologists with over 10 years of experience and a case load of over 3000 cases per year. Six participants reported seeing canine MCTs only rarely, 1 participant monthly, 8 participants weekly, and 1 participant daily. Three participants only requested PCR-based analysis on c-KIT-ITD-11 for high-grade MCTs, while the other 14 never did. The anonymized participant information is available in Supplemental Table 1. The 2 groups ended up slightly unequal in size (9 participants in group A and 8 participants in group B). Nevertheless, the overall experience level remained balanced between the groups (Supplemental Figure S2).
CP, Sensitivity, and Specificity
Figure 4 displays a box plot representation of the average CP of all participants across the different study phases compared with the in-domain average CPs of the DLMs. In phase 1A (pretraining labeling of WSIs), participants reached an average CP of 74.3% (50.0%–100.0%), which dropped in the challenge (posttraining labeling) on WSIs to an average CP of 60.4% (40.0%–73.3%). Concerning the patches, the average CP of 53.7% in phase 1B (35.0%–67.0%) improved to 63.7% (48.0%–75.0%) in the challenge.
The groups were compared to examine the potential influence of the staining on the CPs. The average results for the 2 groups are reported in Table 1, while the detailed results are available in Supplemental Table S2.
Group comparison of the average classification performances, sensitivity, and specificity pre- and posttraining.
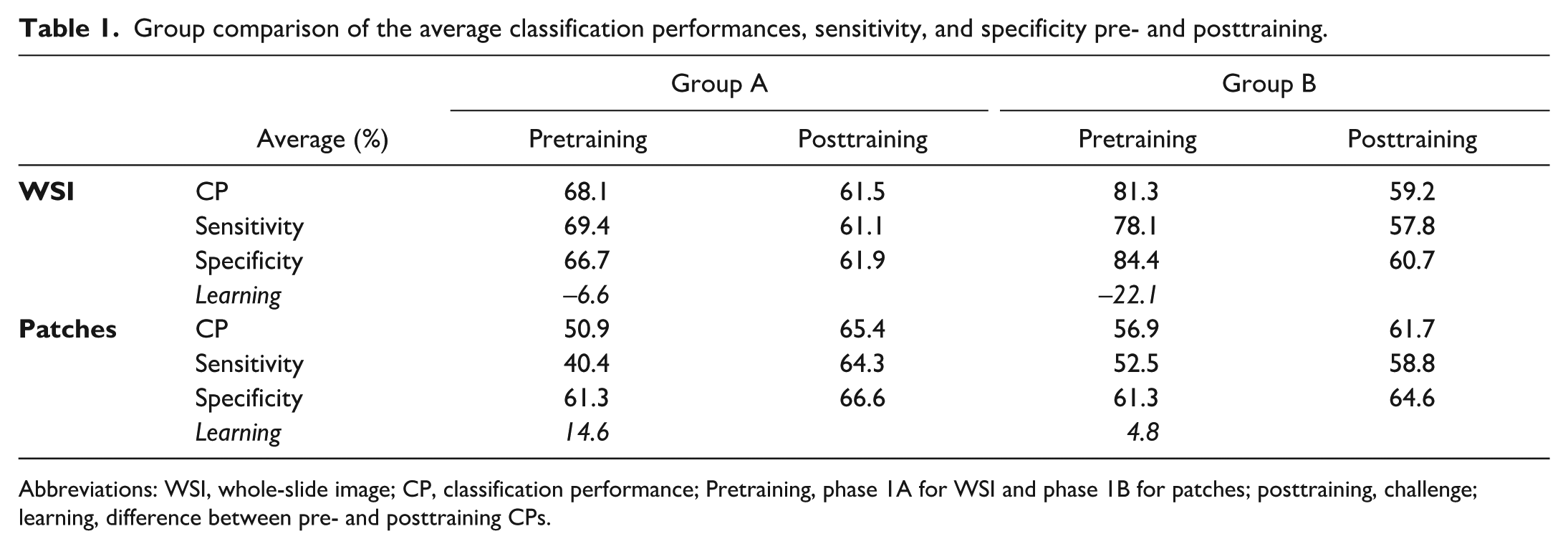
Abbreviations: WSI, whole-slide image; CP, classification performance; Pretraining, phase 1A for WSI and phase 1B for patches; posttraining, challenge; learning, difference between pre- and posttraining CPs.
In phase 1A (intuitive WSI prediction), group A showed an average CP of 68.1% with an average Sen of 69.4% and an average Spe of 66.7%. Group B surprisingly showed an average CP of 81.3% with an average Sen of 78.1%, and an average Spe of 84.4%. Regarding the WSI challenge, after training, group A dropped to an average CP of 61.5%, average Sen of 61.1%, and average Spe of 61.9%. Group B showed a stronger performance drop with an average CP of 59.2%, average Sen of 57.8, and average Spe of 60.7%. In total, 4 participants (numbers 7, 15, and 16 from group A and number 1 from group B) reached a CP over 70% (all 4 at 73.3%).
In phase 1B (intuitive patch prediction), the difference between the 2 groups was less prominent. Group A showed an average CP of 50.9% with an average Sen of 40.4%, and average Spe of 61.3%. Group B showed an average CP of 56.9% with an average Sen of 52.5%, and average Spe of 61.3%. In the patch challenge, after training, group A rose up to an average CP of 65.4% with an average Sen of 64.3% and an average Spe of 66.6%, while group B progressed to an average CP of 61.7% (54.5-71.0%), average Sen of 58.8%, and average Spe of 64.6%. In total, 4 participants (numbers 7, 12, and 16 from group A and number 9 from group B) reached a CP over 70% (71.0%–75.0%). Participants 1, 7, 9, 12, 15, and 16 were considered “high performers,” as they achieved CPs greater than 70.0% in a posttraining task.
The global pre- and posttraining Sen and Spe were calculated for each participant (Supplemental Table S2). Only 3 participants (numbers 7, 8, and 15 from group A) showed an improvement in both Sen and Spe posttraining, but only 1 of them (number 7) achieved both posttraining Sen and Spe over 70.0% (Sen = 75.0% and Spe = 73.2%).
Learning Effect
Globally, the learning effect, calculated as the difference between the pre- and posttraining CPs, turned out negative regarding the WSIs (global average learning effect of –13.9%). Group B showed a stronger negative average learning effect of –22.1% than group A with -6.6%. Only participant 7 from group A managed to score a positive learning effect of 23.3%, progressing from a pretraining CP of 50.0% (Sen and Spe = 50.0%) to a posttraining CP of 73.3% (Sen = 75.0%, Spe = 71,4%) (Fig. 3).
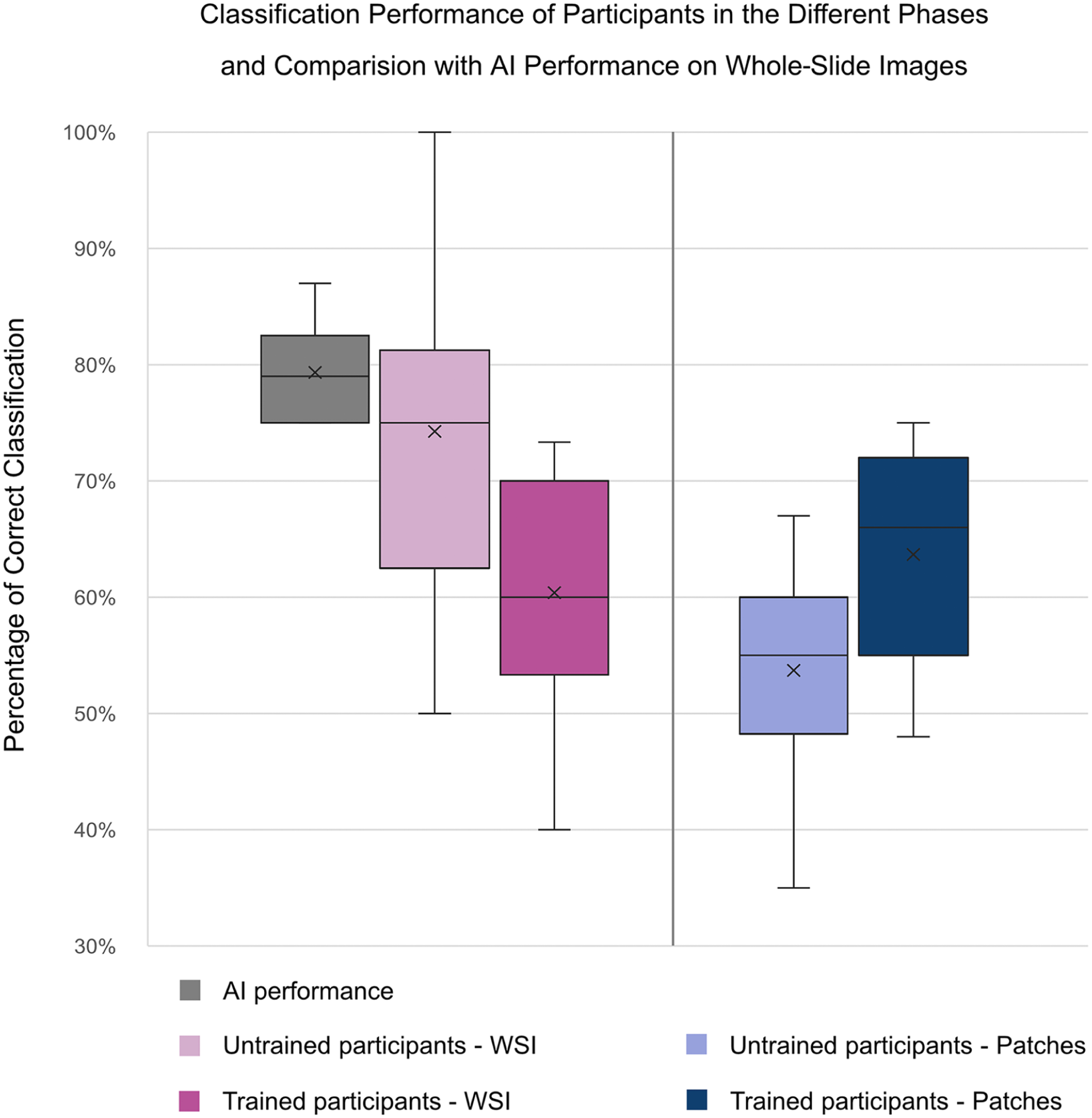
Box plot visualization of the classification performances of all participants across the study phases, compared with the average case-level classification accuracy of 6 independently trained deep learning models. The reduced posttraining global average classification performance on whole-slide images (WSIs) indicates a negative learning effect, while the augmented posttraining average classification performance on patches is indicative of a positive learning effect. Note that the upper interval of the trained participants on patches reaches the lower interval of the artificial intelligence (AI) performance. However, it must be taken into account that AI performance reflects WSI-level classification based on weakly supervised training using slide-level PCR labels, with no available ground truth at the patch level. The patches shown to participants were selected based on high relevance scores from cases correctly classified by the AI. Therefore, direct comparison between human patch-level performance and global AI performance should be interpreted with caution.
Concerning the patches, the global average learning effect was +10.0%. Group B showed less progression than Group A with an average learning effect of 4.8% against 14.6%, respectively. Only 2 participants (number 5 from group A and number 10 from group B) scored a negative learning effect (–3.5% and –1.5%, respectively). Participant 7 scored the largest positive learning effect (+34.5%).
Morphological Features
A total of 55 morphological features for c-KIT-11-ITD-positive and 43 features for c-KIT-11-ITD-negative MCTs were collected and compiled into 21 summarized morphological features. A list of summarized morphological features identified by the 6 “high performers” was also composed. Detailed counts can be found in Supplemental Table S3. Figure 4 displays the top 15 summarized morphological features identified by the “high performers” on the left side and by all the participants on the right side. For the “high performers,” cell size, cytoplasmic granules, and nuclear size were the most relevant morphological features differentiating c-KIT-11-ITD-positive from c-KIT-11-ITD-negative WSIs and patches. As for the rest of the participants, the cell shape was the most relevant distinctive morphological feature, followed by cytoplasmic granules and nucleus size, while the cell size was only listed in the 12th position.
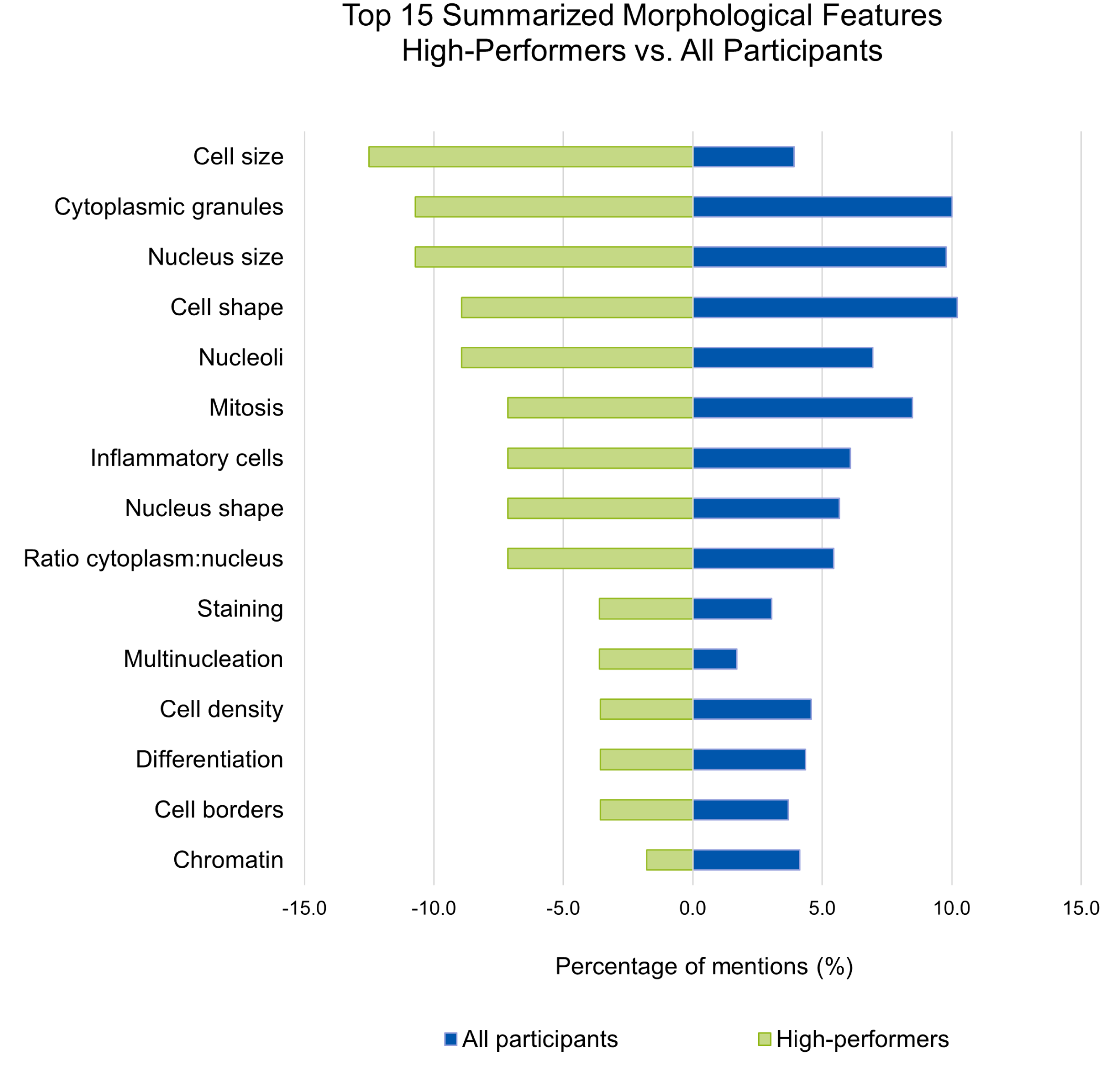
Comparison of the top 15 summarized features for all participants and for “high performers.” Global features summarize opposite terms or semi-quantitative estimations. For the “high performers,” cell size, cytoplasmic granules, and nuclear size were the most relevant features differentiating c-KIT-11-ITD-positive (internal tandem duplication in c-KIT exon 11) from c-KIT-11-ITD-negative whole-slide images and patches. For the rest of the participants, the cell shape was the most relevant distinctive feature, followed by cytoplasmic granules and nucleus size. Note that the x-axis reports percentages of mentions, as 2 groups of different sizes were compared.
Follow-Up Study—Testing of Morphological Features
Of the 3 binarily assessed morphological features, the cytoplasmic granules had the highest predictivity for all participants (average Sen = 73.1%; average Spe = 63.1%). Cell shape and nucleus size were not consequently predictive, with both average Sen and average Spe below 70% (Fig. 5). The detailed morphological assessment is available in Supplemental Tables S4 and S5. All average kappa scores were below 0.6 (0.10–0.38), indicating poor interrater reliability (Supplemental Table S6).
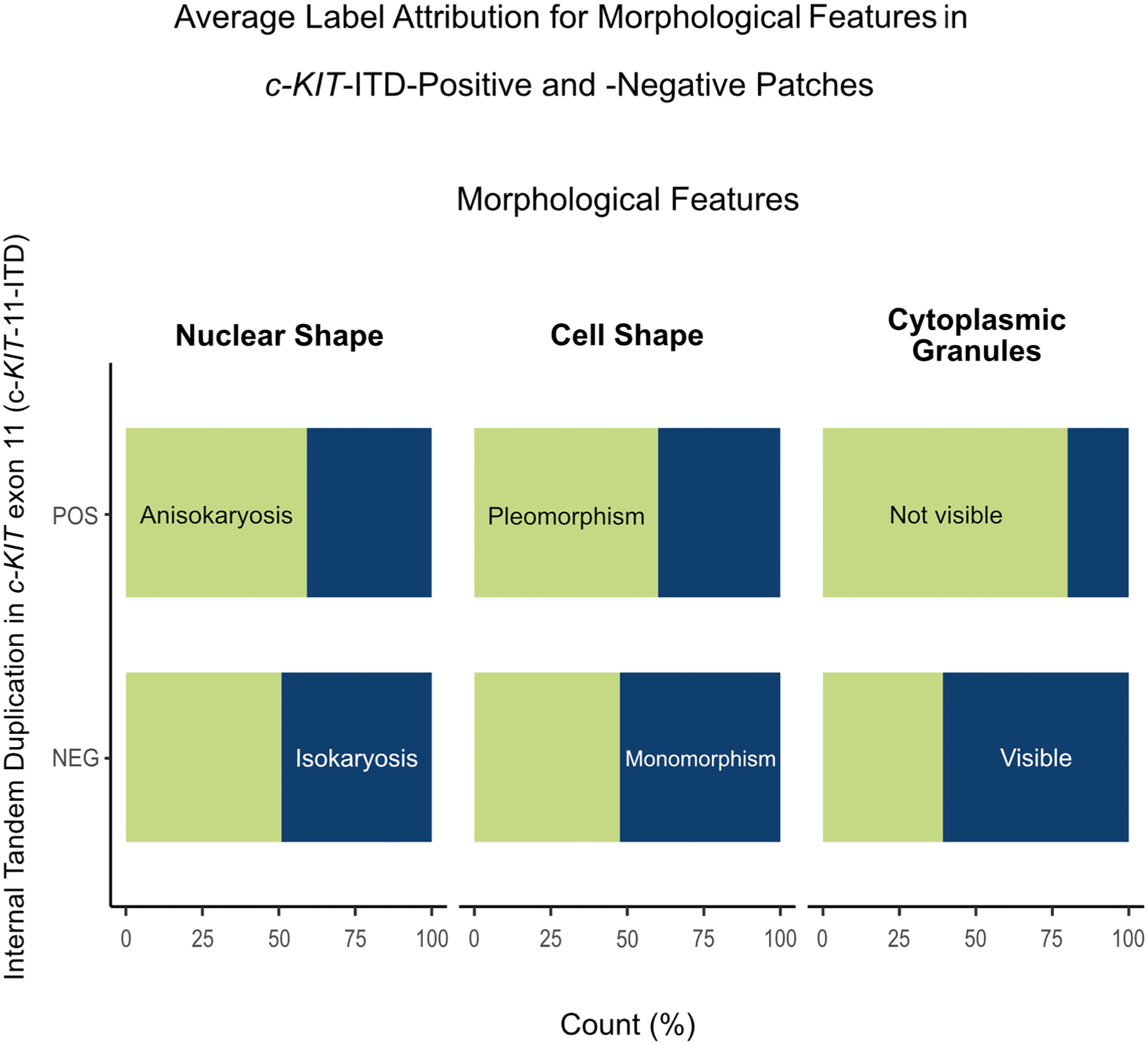
Follow-up study—Average label attribution for morphological features in c-KIT exon 11 internal tandem duplication (c-KIT-11-ITD)-positive and -negative patches of canine cutaneous mast cell tumors. The upper bar (POS) of the stalked diagram can be read as the average sensitivity of the label, while the lower bar (NEG) reflects the average specificity of this same label. “Not visible cytoplasmic granules” show the highest predictivity, being present in over 70% of the positive cases and absent in over 60% of the negative cases. However, these scores are not sufficient to consider these labels as specific for c-KIT-11-ITD-positive patches.
Discussion
The development of AI-assisted medical diagnostic tools is a rapidly evolving field, already being integrated into clinical workflows. 14 A very promising application is the prediction of molecular markers from standard histologic samples.1,8,10,13,39 This work builds upon a previous publication by our group, 37 and is the first in veterinary medicine aiming at extracting morphological features linked to a mutation by interpreting AI-generated results. To this date, no specific morphological feature has been associated with internal tandem duplication in exon 11 of the c-KIT gene (c-KIT-11-ITD) in ccMCTs.
To the best of our knowledge, so far, only one group published a similar study in the field of human medicine. Here, weakly supervised DLMs were trained to predict the p16 status, as a marker for human papilloma virus mediated carcinogenesis, in oropharyngeal squamous cell carcinoma. 1 Morphological features were extracted through histologic examination of 140 highly relevant patches by 8 pathologists. Three morphological features were identified as significant differentiators between p16-positive and p16-negative samples, which were then confirmed by cycle-consistent adversarial network (CycleGAN) image translation (the AI generates exemplary patches for the labels). In this study, a group of 17 participants reviewed 23 WSIs and 400 patches with 2 different HE stains, aiming to extract specific morphological features linked to the c-KIT-11-ITD status of ccMCTs and assess the learnability of predicting this mutation.
In contrast to the human study, we show that participants were unable to predict c-KIT-11-ITD after AI-based self-training. The surprisingly good average CPs in phase 1A (untrained participants) were attributed to a statistical hazard due to a small sample number and chance. This hypothesis was supported by the performance drop on patches in phase 1B and the further decline in the challenge on WSIs, indicating a negative learning effect on WSIs. In addition, the morphological features identified after phase 1A, largely corresponding to broad malignancy criteria, were repeated in phase 2 (morphological features after AI-based self-training) without improving performance.
Even though a positive average learning effect could be observed in the challenge on patches, the overall performance remained below the 70% threshold (considered here as decently better than chance) and inferior to the AI’s performance. Four participants achieved CPs over 70% on patches, approaching the CP of the least accurate DLM. It is important to note here that human patch-level performance is being compared with WSI-level performances of DLMs, as the ground truth labels used to train the models (PCR results) represent blended signals from entire tissue samples without spatial resolution. Nevertheless, the patches assessed in this study were all highly relevant patches, selected from cases correctly classified by the DLMs, and thus likely contained key morphological features associated with the WSI-level label.
Several factors may explain the weak learning effect and the overall failure to identify specific c-KIT-11-ITD morphological features in this study. Customary pathology training is based on direct learning from expert knowledge, displayed as an image with arrows pointing at specific features (visual information) precisely described in a legend (verbal information), a combination proven to be highly effective for learning. 31 In this study, the AI acted as an “expert,” providing only large image groups without verbal description.
Another potential limiting factor was the unsupervised training method without direct feedback. Studies based on Ebbinghaus’s work demonstrate that repetition over time is critical to retain information; spaced repetitions being more effective than block learning.12,25,28,32 This study comprised a single training phase. In addition, the learning environment was not standardized, as participants were allowed to complete the study flexibly within 3 weeks, leading to varying exposure durations. Furthermore, cognitive learning theories suggest that the brain connects new information with preexisting knowledge.5,12,26 Participants may have struggled to fully distance themselves from prior associations between malignancy features and mutation probability, potentially complicating the identification of new patterns.
Overall, a very low interobserver agreement was measured, indicating an important variability in feature interpretation. This variability might be due to the subjective nature of the task, since thresholds for defining object sizes and shapes are often imprecise. A study measured the limitations of categorical assessment of nuclear features, even by experimented pathologists, reporting kappa values below 0.6 as well. 21 In the present follow-up study, the relatively limited experience of some participants in recognizing specific structures may have further contributed to this variability. Repeated training sessions could potentially standardize interpretation and improve consistency. Furthermore, given the small number of participants, it is likely that a larger and more diverse group of pathologists would yield slightly higher kappa, potentially converging toward a more robust consensus.
Most of the morphological features mentioned by the participants are well known, broad criteria of malignancy (eg, cellular and nuclear pleomorphism, high mitotic count, etc.) included in different grading systems for mast cell tumors,27,34 as well as for other neoplasms. 2 The top morphological features, cellular pleomorphism followed by nuclear size and cytoplasmic granularity, were poorly predictive.
Another observation was the influence of the staining protocol on the learning effect. Group A showed a better learning effect working on a more intense stain that particularly highlighted basophilic cytoplasmic granules, while group B worked with a less saturated staining showing a lower detail level. The only morphological feature consistently mentioned by group A and not by group B was the presence of cytoplasmic basophilic granules (indicative of c-KIT-11-ITD-negative ccMCTs). This suggests that morphological features found in one database are not necessarily transferable to another database. Interestingly, DLMs performed better with stain B (less saturated stain), while pathologists performed better with stain A (intense stain). This suggests that group A might have identified a visually prominent feature, cytoplasmic granules, either not detected or not prioritized by the DLMs. The stain A saturation might have modified feature visibility, aiding human perception while introducing variability or noise that was irrelevant, or even detrimental, for DLM performance.
This underlines that machine learning models and human brains process images differently, 33 and that the AI might be able to detect features invisible to the human eye (such as pixel-level details). To assess whether human-recognizable features are actually diagnostically relevant or misleading for DLMs, future work could focus on systematic feature ablation. This consists of testing the relevance of specific features or regions by selectively hiding (eg, blurring) or modifying (eg, color channel removal) them and observing the effects on model performances.7,23 While our study does not directly implement uncertainty quantification techniques, such as Bayesian neural networks,35,36 or generative tools, such as CycleGANs or unpaired image-to-image translation, 44 these approaches represent promising directions for future work to better assess model confidence in ambiguous cases and to independently verify if the morphological features suggested in the present study hold up against automated feature synthesis.
It is important to note, however, that even with advanced techniques from the field of explainable AI, a definitive identification of single, human-interpretable histologic features may not necessarily be achievable. A likely outcome might be the generation of idealized, synthetic examples representing clear-cut c-KIT-11-ITD-positive and c-KIT-11-ITD-negative cases, while ambiguity and difficulty will persist in borderline cases, just as for human experts.
The prediction of mutations based on standard digitalized histologic slides holds an interesting potential for future diagnostic workflows. Should the underlying decision criteria of the AI remain unknown, such tools could still serve as rapid and cost-efficient screening tests. In human medicine, the question of whether AI-based prediction should be excluded from routine diagnostics solely due to a lack of full interpretability remains actively debated. The American Food and Drug Administration values clearly demonstrated performance, reproducibility, and clinical safety over full interpretability, while the European Coordinating Research and Evidence for Medical Devices consortium takes valid clinical association as well as technical (including interpretability) and clinical performances in account to rate medical devices incorporating AI tools. 38 Circling back to the present study, another line of future work should concentrate efforts on improving the DLMs performances as well as clinically testing and validating the DLMs.
To conclude, although Adachi et al 1 showed that extraction of specific morphological features with broad visual information without verbal information is possible, it did not work particularly well in the present study. A specific or interpretable feature for c-KIT-11-ITD, possibly very subtle or a combination of features, remains to be identified. Only a limited positive learning effect was observed after an AI-based training with WSIs or cropped highly relevant small tumor areas (patches). Even “high performers” achieving a moderate predictivity remained inferior to AI performances. While AI as a diagnostic tool holds an interesting potential as a screening test for mutational status in HE slides, interpretability and translation for human cognition may be challenging.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858251380284 – Supplemental material for Artificial intelligence predicts c-KIT exon 11 genotype by phenotype in canine cutaneous mast cell tumors: Can human observers learn it?
Supplemental material, sj-pdf-1-vet-10.1177_03009858251380284 for Artificial intelligence predicts c-KIT exon 11 genotype by phenotype in canine cutaneous mast cell tumors: Can human observers learn it? by Chloé Puget, Jonathan Ganz, Christof A. Bertram, Thomas Conrad, Malte Baeblich, Anne Voss, Katharina Landmann, Alexander F. H. Haake, Andreas Spree, Svenja Hartung, Leonore Aeschlimann, Sara Soto, Simone de Brot, Martina Dettwiler, Heike Aupperle-Lellbach, Pompei Bolfa, Alexander Bartel, Matti Kiupel, Katharina Breininger, Marc Aubreville and Robert Klopfleisch in Veterinary Pathology
Footnotes
Acknowledgements
ChatGPT-4 (December 2024–August 2025), the latest free language model by OpenAI, was used to proofread spelling and grammar as well as to improve language.
Author Contributions
CP designed and conducted the studies, participated in the follow-up study, and wrote the manuscript with contributions of RK and the other authors; RK, MA, MK, CAB, KL, and AV contributed to the study design; MK provided the original histological slides; JG, MA, and KB provided the segmented image sets for the studies and managed the EXACT digital annotation tool; CP and AB performed the statistical analysis and created the figures; RK, TC, CAB, AV, KL, AFHH, AS, MB, SH, LA, SS, MD, SdB, HAL, and PB were study participants; all authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Author Christof Bertram is a member of the Editorial Board of Veterinary Pathology and has no further conflicts to declare. The author did not take part in the peer review or decision-making process for this submission.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
ORCID iDs
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
