Abstract
While the immunodeficient status of NOD.Cg-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) and NSG-related mice provides utility for numerous research models, it also results in increased susceptibility to opportunistic pathogens. Over a 9-week period, a high rate of mortality was reported in a housing room of NSG and NSG-related mice. Diagnostics were performed to determine the underlying etiopathogenesis. Mice submitted for evaluation included those found deceased (n = 2), cage mates of deceased mice with or without diarrhea (n = 17), and moribund mice (n = 8). Grossly, mice exhibited small intestinal and cecal dilation with abundant gas and/or digesta (n = 18), serosal hemorrhage and congestion (n = 6), or were grossly normal (n = 3). Histologically, there was erosive to ulcerative enterocolitis (n = 7) of the distal small and large intestine or widespread individual epithelial cell death with luminal sloughing (n = 13) and varying degrees of submucosal edema and mucosal hyperplasia. Cecal dysbiosis, a reduction in typical filamentous bacteria coupled with overgrowth of bacterial rods, was identified in 18 of 24 (75%) mice. Clostridium spp. and Paeniclostridium sordellii were identified in 13 of 23 (57%) and 7 of 23 (30%) mice, respectively. Clostridium perfringens (7 of 23, 30%) was isolated most frequently. Toxinotyping of C. perfringens positive mice (n = 2) identified C. perfringens type A. Luminal immunoreactivity to several clostridial species was identified within lesioned small intestine by immunohistochemistry. Clinicopathologic findings were thus associated with overgrowth of various clostridial species, though direct causality could not be ascribed. A diet shift preceding the mortality event may have contributed to loss of intestinal homeostasis.
Immunodeficient mice play an invaluable role in biomedical research. Since the first immunocompromised mouse strain, the nude mouse carrying a mutation in the Foxn1 gene, was reported in the 1960s, 8 tremendous progress has been made in the fields of immunology, oncology, autoimmunity, infectious disease, and regenerative medicine.4,24 Many of these advancements have been made possible by the development of a vast array of immunocompromised and immunodeficient mouse strains, each possessing unique immunologic profiles to be utilized by the research community. The NOD.Cg.-Prkdc scid Il2rg tm1Wjl /SzJ (NOD-scid gamma, NSG) mouse is among the most immunodeficient strains, combining a host of deficiencies in innate and adaptive immune function as well as altered cytokine signaling, 25 This strain has gained popularity due to its ability to support engrafted human cells and tissues for the generation of humanized models.24,25 Further modifications have been made to this genetic background, such as expression of the human IL3 gene, granulocyte macrophage colony-stimulating factor, and the stem cell factor, KITLG, in the NOD.Cg.-Prkdc scid Il2rg tm1Wjl Tg (CMV-IL3, CSF2, KITLG) 1Eav/MloySzJ (NSG-SGM3) mouse, which confer additional advantages for biomedical research. 34 While there are numerous valuable applications for NSG mice and related strains as research models, their severely impaired immunity also presents several challenges. Specialized husbandry and hygiene, strict biosecurity, and vigorous disease surveillance are paramount to maintaining the health of these animals and preventing the introduction of primary or opportunistic pathogens that could risk research integrity.22,33
Multiple etiologies associated with morbidity and mortality have been reported in NSG colonies, sometimes in the absence of known breaks in biosecurity or identifiable sources. The outcomes of disease are, not surprisingly, dependent upon the etiologic agent involved. An evaluation of spontaneous disease affecting naïve NSG breeder mice at 3 commercial vendor facilities found that ascending urinary tract infections and/or bacterial nephritis caused by Enterococcus spp. and Klebsiella oxytoca were a major cause of morbidity despite the mice being maintained in maximum-level barrier facilities. 10 Both are typically considered commensal or opportunistic agents in immunocompetent mice. 10 Fungal ascending pyelonephritis associated with Candida albicans was also identified as a cause of acute mortality in a group of NSG, and closely related NOD.Cg-Rag1 tm1Mom Il2rg tm1Wjl / SzJ (NRG) mice, that had been further immunosuppressed using chemotherapeutics prior to a humanization surgery. 35 In this case, the source of contamination was believed to be fomite transmission through the use of a shared restraint device. Another report described an epidemic of antibiotic resistant Klebsiella pneumoniae with gastrointestinal disease leading to systemic infections and sudden death in an NSG breeding colony. 27 NSG and NSG-related mouse colonies have more recently been the target of diarrheal outbreaks in multiple facilities.18,21 In one report, rapid spread of low-mortality diarrheal disease throughout a barrier facility led to its depopulation before fecal microbiome profiling suggested intestinal dysbiosis may have been causal. Although speculative, their results pointed toward a potential role of Candidatus Arthromitus (segmented filamentous bacterial, SFB) and Clostridium celatum in the disease pathogenesis. 21 More recently, Clostridioides difficile was implicated in 2 outbreaks of typhlocolitis; the first affected NSG and related strains following treatment with the antibiotic, amoxicillin, while the second involved an experimentally naïve NOD-scid breeding colony at the same institution. 18 These case reports corroborate existing evidence suggesting enhanced susceptibility of immunodeficient mouse strains to Clostridium spp. relative to their immunocompetent counterparts.1,16
Clostridia are a diverse group of mostly gram-positive, spore-forming anaerobes that are widespread and persistent in the environment, while some species are also considered commensal inhabitants of the gastrointestinal tract. Virulence is dependent upon the production of a vast array of exotoxins. 29 While a multitude of multisystemic disease manifestations can occur depending on the bacterial and host species involved, intestinal diseases, including enterotoxemia, enteritis, and enterocolitis, are well-described in humans and a variety of other animal species.5,13,20,23,26,29–32
Herein, we describe a spontaneous disease outbreak affecting a single housing room of NSG and NSG-related strains, characterized by high mortality with most affected animals being found dead in the absence of premonitory signs. In a minority of cases, sick mice were identified with nonspecific clinical signs and/or evidence of diarrhea. Through a comprehensive diagnostic investigation, we aimed to identify the underlying etiopathogenesis.
Materials and Methods
Mice
All mice involved in the disease outbreak were NOD.Cg.-Prkdc scid Il2rg tm1Wjl /SzJ (NSG, Strain #:005557) or closely related strains, NOD.Cg-Prkdc scid Il2rg tm1Wjl Tg (CMV-IL3, CSF2,KITLG) 1Eav/MloySzJ (NSG-SGM3, Strain #:013062), NOD.Cg-Prkdc scid Il2rg tm1Wjl Tg (CMV-IL3, CSF2,KITLG)1Eav Tg (CSF1)3Sz/J (NSG-Quad, Strain #:028657), or NRG.Cg-Rag1 tm1Mom Il2rg tm1Wjl /SZJ (NRG, Strain #:007799), which were supplied directly by the Jackson Laboratory (Sacramento, CA) or bred in the affected room after arrival. Animals were housed at an AAALAC International-accredited institution and ethically cared for in accordance with The Guide for the Care and Use of Laboratory Animals. 11 Affected mice belonged to 4 different principal investigators in a single housing room on protocols approved by Stanford University’s institutional animal care and use committee.
Husbandry and Health Surveillance
The housing room involved contained exclusively high health status immunodeficient NSG or NSG-related (NSG-SGM3, NSG-Quad, or NRG) mice from approved vendors, with the exception of 2 immunocompetent Crl:CD-1 (ICR) (Charles River, Hollister, CA) sentinel mice for disease surveillance purposes. It was populated in September 2021 and was managed as a direct-from-vendor room; breeding and/or experimental manipulations were permitted in the multi-investigator room, but any animals removed from the room (e.g., for imaging procedures) were not permitted reentry. Such cages were transferred to an adjacent experimental housing room which remained unaffected throughout the disease outbreak.
The affected room, with its dedicated anteroom and procedural space, was located within a small vivarium housing several rodent, avian, and aquatic species (including dedicated gnotobiotic and biohazard suites) with a shared common corridor. While not a barrier facility, multiple barrier-level protocols were implemented for this NSG housing room; these included restricted entry, enhanced personal protective equipment requirements (Tyvek coveralls, sleeve protectors, shoe covers, bonnet, surgical mask, and nitrile gloves), and strict hygiene practices. Where possible, manipulations (e.g., cage changing) were performed within a laminar flow hood, although experimental procedures using the adjoining procedural space were not and containers of shared supplies (e.g., rodent feed, enrichment items) were exposed to room air while in use. The room was entered first by dedicated husbandry and research staff prior to entering any other animal housing areas. Mice were housed in positive pressure individually ventilated cages (Innocage, Innovive, San Diego, CA) with pre-filled, irradiated cellulose bedding (Alpha-dri, Innovive, San Diego, CA) with acidified bottled water (Aquavive, Innovive, San Diego, CA). Animals were fed commercially available irradiated rodent diet, Teklad 2918 (Envigo, Madison, WI), ad libitum from September 2021 until early February 2022, at which point they were switched to irradiated PicoLab Rodent Diet 20 (LabDiet, St Louis, MO) due to supply chain issues.
Quarterly routine processing of immunocompetent Crl:CD:-1 (ICR) sentinel mice in December 2021 and April 2022 was performed to screen for infectious agents by necropsy, serology, parasitology, microbiology (i.e., tracheal aerobic culture), and molecular diagnostics (i.e., polymerase chain reaction [PCR]). The Multiplexed Fluorometric ImmunoAssay serological assay was performed in-house to screen for the Mouse Assessment Plus Profile (Charles River Laboratories, Wilmington, MA), including mouse parvovirus, minute virus of mice, mouse hepatitis virus, murine norovirus, Theiler’s murine encephalomyelitis virus, mouse rotavirus (epizootic diarrhea of infant mice), Sendai virus, pneumonia virus of mice, reovirus 3, lymphocytic choriomeningitis virus, mouse adenovirus, ectromelia virus, K virus, polyoma virus, mouse cytomegalovirus, mouse thymic virus, Prospect Hill virus, lactate dehydrogenase elevating virus, Hanta virus, Encephalitozoon cuniculi, cilia-associated respiratory bacillus, Mycoplasma pulmonis, and Clostridium piliforme. Parasitology consisted of direct examination of pelage and intestinal contents for detection of fur mites and pinworms, respectively, as well as perineal cellophane tape test and direct smear of intestinal contents for endoparasites. Microbiologic culture screened for Rodentibacter spp., Pasteurella multocida, Corynebacterium kutscheri, and Streptococcus pneumoniae. Additional diagnostics performed during the outbreak included collection of fecal pellets from affected mice which were shipped to Charles River Laboratories for PCR Rodent Infectious Agent screening for C. difficile and the Rodent Surveillance Plus panel (Charles River Laboratories, Wilmington, MA) to screen for Hanta virus, K virus, lymphocytic choriomeningitis virus, lactate dehydrogenase elevating virus, mouse adenovirus 1 and 2, mouse cytomegalovirus, mouse hepatitis virus, mouse norovirus, ectromelia virus, mouse parvovirus, minute virus of mice, mouse rotavirus (epizootic diarrhea of infant mice), mouse thymic virus, polyoma virus, pneumonia virus of mice, reovirus 3, Sendai virus, Theiler’s murine encephalomyelitis virus, β-hemolytic Streptococcus (groups A, B, C, and G), Bordetella bronchiseptica, Bordetella pseudohinzii, Campylobacter, cilia-associated respiratory bacillus, Citrobacter rodentium, Clostridium piliforme, Corynebacterium bovis, Corynebacterium kutscheri, Helicobacter sp., Klebsiella oxytoca, Klebsiella pneumoniae, Mycoplasma pulmonis, Rodentibacter heylii, Rodentibacter pneumotropicus, Proteus mirabilis, Pseudomonas aeruginosa, Salmonella spp., Staphylococcus aureus, Streptobacillus moniliformis, Streptococcus pneumoniae, Cryptosporidium, Demodex, Encephalitozoon cuniculi, Entamoeba, Giardia, mites, pinworms, Pneumocystis, Spironucleus muris, and Tritrichomonas.
An additional set of survival serum samples were collected from sentinel mice early in the outbreak investigation (March 2022) for serological evaluation (Mouse Assessment Plus Profile via the Multiplex Fluorometric ImmunoAssay), as described above.
Histopathology
Mice submitted for necropsy (n = 27) were moribund, found deceased, or clinically healthy cage mates of mice found deceased, with or without evidence of diarrhea. Live mice were euthanized by carbon dioxide inhalation in accordance with the American Veterinary Medical Association Guidelines for Euthanasia of Animals. Following gross evaluation, tissues were immersion-fixed in 10% neutral-buffered formalin for 72 hours. Formalin-fixed tissues were processed routinely, embedded in paraffin, sectioned at 5 µm, and stained with hematoxylin and eosin. The following organs were evaluated histologically: heart, lung, liver, kidneys, spleen, brain, adrenal glands, salivary glands, esophagus, trachea, urinary bladder, haired skin, reproductive tracts, and gastrointestinal tracts. Elaborated evaluation of the gastrointestinal tracts included longitudinal sections of stomach (cardia to pylorus), duodenum, jejunum, and ileum. Both halves of longitudinally sectioned cecum were evaluated in addition to the proximal, middle, and distal colon. Select small and large intestinal sections were stained with Gram stain.
Immunohistochemistry
The following immunohistochemical protocols for Clostridium perfringens, Paeniclostridium sordellii, Clostridium novyi, Clostridium septicum, and Clostridium chauvoei were performed on select formalin-fixed, paraffin-embedded 4-µm-thick sections of lesioned small and large intestine. Antigen retrieval was performed using pepsin. Primary antibodies included rabbit polyclonal anti-C. perfringens (GenWay Bio, San Diego, CA; 1:100), goat anti-P. sordellii (VMRD, Pullman, WA; 1:750), goat anti-C. novyi (VMRD, Pullman, WA; 1:750), goat anti-C. septicum (VMRD, Pullman, WA; 1:750), and goat anti-C. chauvoei (VMRD, Pullman, WA; 1:750). Visualization of C. perfringens was obtained using the Dako EnVision Kit (Dako, Carpenteria, CA). Remaining secondary antibodies and detection were achieved with rabbit anti-goat immunoglobulin G (IgG; Vector Laboratories, Burlingame, CA) and the VECTASTAIN ABC HRP Kit (Vector Laboratories, Burlingame, CA). Positive controls included tissues of several animal species in which each of the corresponding clostridia had been detected by culture and/or PCR. Negative controls were sections of the case mice inoculated with rabbit nonimmune serum instead of the anti-clostridial antibodies sera.
Bacteriology
Enteric cultures were performed on 23 mice. Aerobic, anaerobic, Salmonella spp., and Campylobacter spp. cultures of small and large intestinal contents (n = 13 mice) were performed in the microbiology laboratory of the Stanford University Veterinary Service Center Animal Diagnostic Laboratory using prepared media (Remel, Lenexa, KS and Anaerobe Systems Inc., Morgan Hill, CA). All cultures were performed under proper atmospheric conditions (Mitsubishi Gas Chemical, New York, NY and BD GasPak, East Rutherford, NJ). Fecal samples were streaked to blood agar, Columbia CNA agar, and MacConkey agar for aerobic culture to screen for common pathogens, including Salmonella spp. (incubated in 5% CO2 at 36°C ± 1); anaerobic cultures were inoculated to bacteroides bile esculin/Laked blood agar with kanamycin and vancomycin (BBE/LKV) bi-plate, Brucella agar, and phenylethyl alcohol (PEA) agar (anaerobic conditions with an indicator at 36°C ± 1), and Campylobacter CVA agar was streaked to screen for Campylobacter spp (microaerophilic atmosphere at 42°C ± 1). Isolated organisms were evaluated through Gram stain, colony morphology, biochemical analysis (indole, oxidase, catalase, etc), Omnilog identification (Biolog, Hayward, CA), MALDI-TOF analysis (Bruker, Billerica, MA), and RapID ANA (Remel, Lenexa, KS). Additional enteric cultures were performed at Charles River Laboratories (n = 10 mice). Identification of select culture isolates were subsequently confirmed via MALDI-TOF by 3 external laboratories: University of California Davis Veterinary Medical Teaching Hospital Clinical Diagnostic Laboratory (n = 15 isolates), Stanford Health Care’s Microbiology Laboratory (n = 4 isolates), and California Animal Health and Food Safety Laboratory San Bernadino (n = 6 isolates).
Molecular Analysis
Fecal samples from 23 mice were submitted to Charles River Laboratories for the Charles River Laboratories Mouse Surveillance Plus PCR Rodent Infectious Agent (see Husbandry & Health Surveillance section, above, for complete pathogen list) as well as additional PCR assays for astrovirus-1, C. difficile, murine chapparvovirus, and Leptospira spp (Charles River Laboratories, Wilmington, MA).
C. perfringens and C. difficile Enzyme-linked Immunosorbent Assay
Pooled small and large intestinal contents from one mouse exhibiting diarrhea were tested for C. perfringens organisms and toxins (alpha (CPA), beta (CPB), and epsilon (ETX)) via a commercial capture ELISA (enzyme-linked immunosorbent assay) kit (BIO-X, Brussels, Belgium) and for C. difficile organisms and toxins (A and B) using a commercial ELISA kit (Techlab, Blacksburg, VA), all according to the manufacturer’s instructions.
C. perfringens Multiplex PCR Toxinotyping
Culture samples from n = 2 mice from which C. perfringens was isolated were further analyzed for toxin gene expression via PCR. To screen for various toxin genes, 50 µl PCRs were conducted with 100 ng of sample DNA template and toxin-specific gene primers, as previously described (Supplemental Table, S1), along with Multiplex PCR Kit (Qiagen, Catalog # 206143), according to the manufacturer’s guidelines. The following PCR cycle parameters were used: initial denaturation at 95°C for 15 minutes, followed by 35 cycles each of 94°C for 30 seconds, 55°C–62°C for 30 seconds (applying an appropriate annealing temperature for each primer set), and 72°C for 90 seconds with a final 10-minute extension period at 72°C. Upon completion of PCR, 10 µl of products were run on an E-Gel EX 1 % agarose (Invitrogen, Thermo Fisher Scientific).
Control Cohort
A control group of naïve, healthy, NSG (n = 4, 2 males and 2 females) and NSG-SGM3 (n = 4, 2 males and 2 females) were purchased from The Jackson Laboratory (Sacramento, CA) in January 2023. Upon arrival at our facility, they were immediately euthanized via carbon dioxide inhalation and samples were collected, pooled by sex and strain, and subjected to aerobic and anaerobic cultures as well as C. perfringens and C. difficile toxin ELISA to screen for presence of organisms and/or toxins (CPB, ETX, C. difficile toxins A and B), which were performed as described above. Six healthy NSG mice (3 males and 3 females) were also sourced directly from the vendor in August 2021 as part of an unrelated project and routine histology was performed, as described above. In addition, in December 2021, a paraffin block prepared from a naïve, healthy, NSG male mouse (n = 1) was processed for C. perfringens immunohistochemistry (IHC) as part of an unrelated investigation.
Cell Line Pathogenicity and Culture
All investigators housing mice in the affected room were surveyed regarding their experimental activities prior to and during the outbreak. Three of 4 groups indicated that they had performed only breeding in experimentally naïve animals prior to and throughout the disease outbreak. One group had begun performing experimental manipulations in the room in January 2022, prior to the disease outbreak. This study involved the intramuscular inoculation of osteosarcoma cell lines, 143B and MG-63.3, into mice, which were then treated with various forms of laboratory-engineered T cells. The osteosarcoma cell lines were tested previously at our institution and were confirmed negative for murine pathogens. To evaluate the pathogenic potential of the osteosarcoma cell lines, 4 frozen aliquots were obtained from the investigator for testing.
Experimentally naïve female NSG mice (n = 20) were obtained directly from the vendor (Jackson Laboratories, Sacramento, CA) and housed in a dedicated housing room in a separate facility. Mice were ear-tagged and housed in groups of 4 per cage. One cage served as a negative control and was not experimentally manipulated. Each of the remaining cages was randomly assigned to 1 of the 4 cell lines. Three mice per cage were inoculated subcutaneously with 100 µl of cell suspension, and the fourth mouse received 100 µl enterally via oral gavage. All animals were subsequently monitored for 4–6 weeks for morbidity or mortality.
The remaining cell line suspensions were pooled and sent out for aerobic and anaerobic bacterial and fungal cultures (IDEXX, Westbrook, ME).
Results
Epidemiology
In late February 2022, approximately 6 months after populating the room, an increased mortality rate was noted. Over a span of 3 days, 9 mice from 8 different cages (i.e., approximately 4% of cages housed in the room at the time; 8/186) belonging to 2 different investigators were found dead. This 3-day mortality count equaled the cumulative number of mortalities from September 2021 through February 8, 2022. Over the following weeks, disease prevalence and corresponding morbidity and mortality continued to rise, peaking in early April 2022 (Fig. 1). Over the course of the outbreak, 234 mice were found dead by husbandry or veterinary staff and 28 morbidities were reported for veterinary evaluation. These values reflect only what was reported by husbandry and veterinary staff through the electronic medical records system and do not account for animals that were found by research staff and subsequently euthanized or collected without notifying the veterinary staff. The room was ultimately depopulated at the end of April 2022 due to widespread disease to allow for disinfection and repopulation. Throughout the course of the outbreak and prior to depopulation, animals were submitted for diagnostics, as described above, and clinically healthy immunocompetent sentinel mice were processed according to routine disease surveillance schedules, in addition to adjunct diagnostic tests.
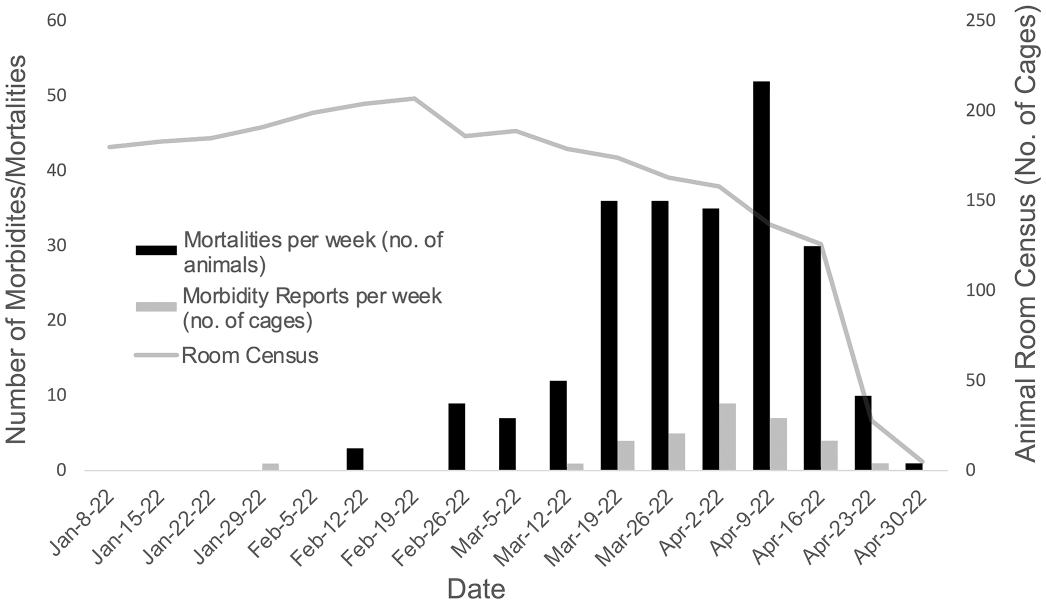
Epidemiologic trend for the affected NOD.Cg-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) and NSG-related housing room from January 1, 2022 through April 30, 2022 based on retrospective analysis of facility census and veterinary medical record data. Mortalities per week represent the number of deaths reported by husbandry or veterinary staff and do not include mice found dead/euthanized by research staff during this period. Morbidities are reported by cage and may represent multiple animals within a cage and unrelated veterinary concerns. During the peak of the outbreak, the high ratio of mortalities to morbidities reported highlights the rapid onset of clinical signs and disease progression in the absence of premonitory signs. Gradual reduction in room census due to increased mortality is depicted prior to depopulation of the room in April 2022.
Health Surveillance
Routine quarterly health surveillance via processing of Crl:CD-1 (ICR) sentinel mice revealed that the colony was negative for all excluded pathogens (see the “Materials and Methods” section for pathogens list) in December 2021 and April 2022. Additional serology results for sentinel mice in March 2022 yielded negative results for all agents included in the Mouse Assessment Plus Profile (Charles River Laboratories, Wilmington, MA).
Gross Findings
Individual clinicopathologic data are presented for all mice in Supplemental Table S2. Mice were submitted for diagnostic evaluation if they were moribund (n = 8), found deceased (n = 2), diarrheic cage mates of a deceased mouse (n = 3), or asymptomatic cage mates of a mouse found deceased (n = 14). All moribund and found dead mice had dark red to black small and large intestinal serosa with loose to absent, dark colonic contents (Fig. 2a) or moderate to severe small intestinal and cecal dilation characterized by abundant gas and/or fluid digesta (Fig. 2b). Cage mates of found dead mice had intestinal gas or fluid distension (13/17, 76%), dark red to black serosa (1/17, 6%), or were grossly normal (3/17, 18%). All grossly normal mice were submitted as cage mates of a deceased mouse.
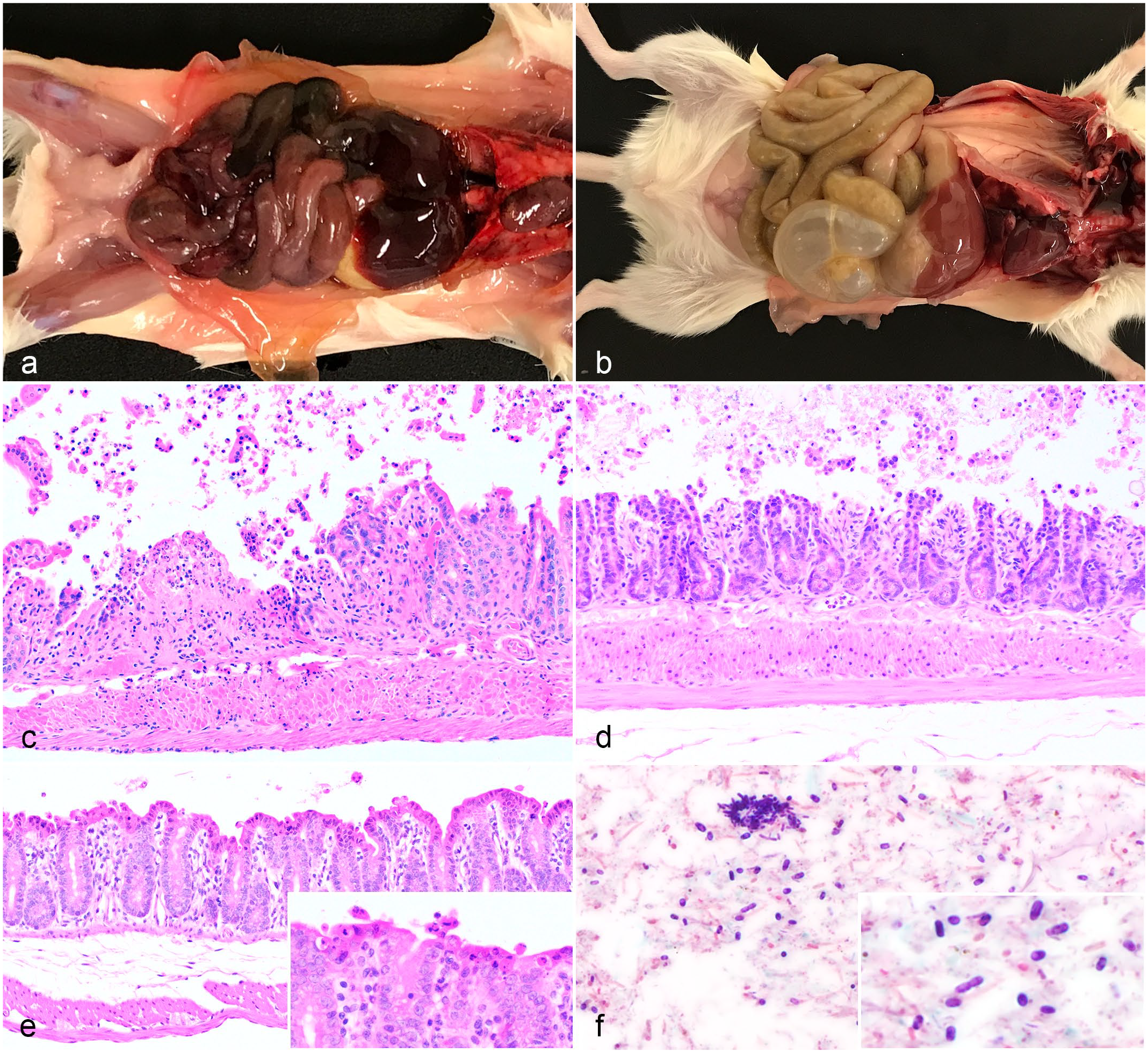
Gross and histopathologic features of enterocolitis in NOD.Cg.-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) or NSG-related adult mice. (a) Moribund mouse with dark red to black small and large intestinal serosal surfaces. (b) Cage mate of moribund mouse with markedly dilated, fluid- and gas-filled small intestine and cecum. (c) Moribund mouse with ulceration of the colonic epithelium, lamina proprial fibrin deposition, and neutrophilic infiltrates. Hematoxylin and eosin (HE). (d) Moribund mouse with multifocal erosion and sloughing of superficial ileal epithelium without inflammation. HE. (e) Cage mate of moribund mouse with widespread superficial epithelial cell death within the cecum. Inset: higher magnification of superficial epithelial cell death and sloughing. HE. (f) Diarrheic cage mate of moribund mouse with high numbers of luminal, gram-positive, approximately 1 µm × 4 µm square-ended bacilli with occasional clear, oval, subterminal endospores. Clostridium perfringens was cultured from fecal material. Gram stain.
Histopathology and Immunohistochemistry Findings
A total of 25 mice from the affected room were evaluated histologically. Overall, histologic lesions were more frequent and severe in moribund and found dead mice. All moribund and found dead mice had erosive to ulcerative enterocolitis (Fig. 2c, d) and/or scattered individual enterocyte cell death with luminal sloughing (Fig. 2e). Erosive to ulcerative enterocolitis was present in the ileum (7/7), colon (6/7), cecum (5/7), and jejunum (5/7). Erosive to ulcerative enterocolitis was not identified in any cage mates of found dead mice. A single moribund mouse exhibited transmural coagulative necrosis of the distal jejunum and ileum with thrombosis of mucosal and submucosal vessels. Scattered individual enterocyte cell death and luminal sloughing was seen in decreasing order of frequency in the cecum, colon, and ileum. Individual cell death and sloughing was seen most frequently within diarrheic cage mates (2/2), found dead mice (1/2), moribund mice (4/8), and less frequently in asymptomatic cage mates (6/14, 43%). Submucosal edema (13/25, 52%) and compensatory mucosal hyperplasia (4/25, 16%) of the ileum, cecum, and/or colon were occasionally noted.
Murine intestinal tissue is typically embedded whole (nonlinearized) in longitudinal segments. This provides the distinct advantage of visualizing ingesta and fecal contents histologically in-situ. Comparative pathologists thus become accustomed to what generally constitutes “normal flora” in a healthy mouse, at least in terms of relative amounts and proportions based on bacterial morphology. This is most evident within the cecum, which is typically filled by abundant filamentous bacteria and partially digested feed (Fig. 3a, b). In 18 of 24 (75%) mice, there was a distinct shift in both the amount of cecal contents and the overall morphology of the luminal bacteria (Fig. 3c, d). Overall, ceca were sparsely populated and contained relatively increased proportions of bacterial rods with a reduction in filamentous bacteria. This shift in the relative abundance and proportion of cecal bacteria will be henceforth referred to as dysbiosis. Dysbiotic mice displayed increased proportions of gram-positive, approximately 1 × 4 µm, square-ended bacilli (Fig. 2f, Fig. 3d) that were free within the lumen and rarely adherent to necrotic intestinal villi (n = 1 mouse). Gram-positive bacilli occasionally exhibited clear, oval, subterminal endospores (Fig. 2f). Bacilli that were positive by IHC for C. perfringens, P. sordellii, C. novyi, C. septicum, and C. chauvoei were identified free within the lumen of lesioned small intestinal tissue (Fig. 3e, f) and adherent to necrotic intestinal villi. No significant histologic lesions were noted in any other evaluated organs.
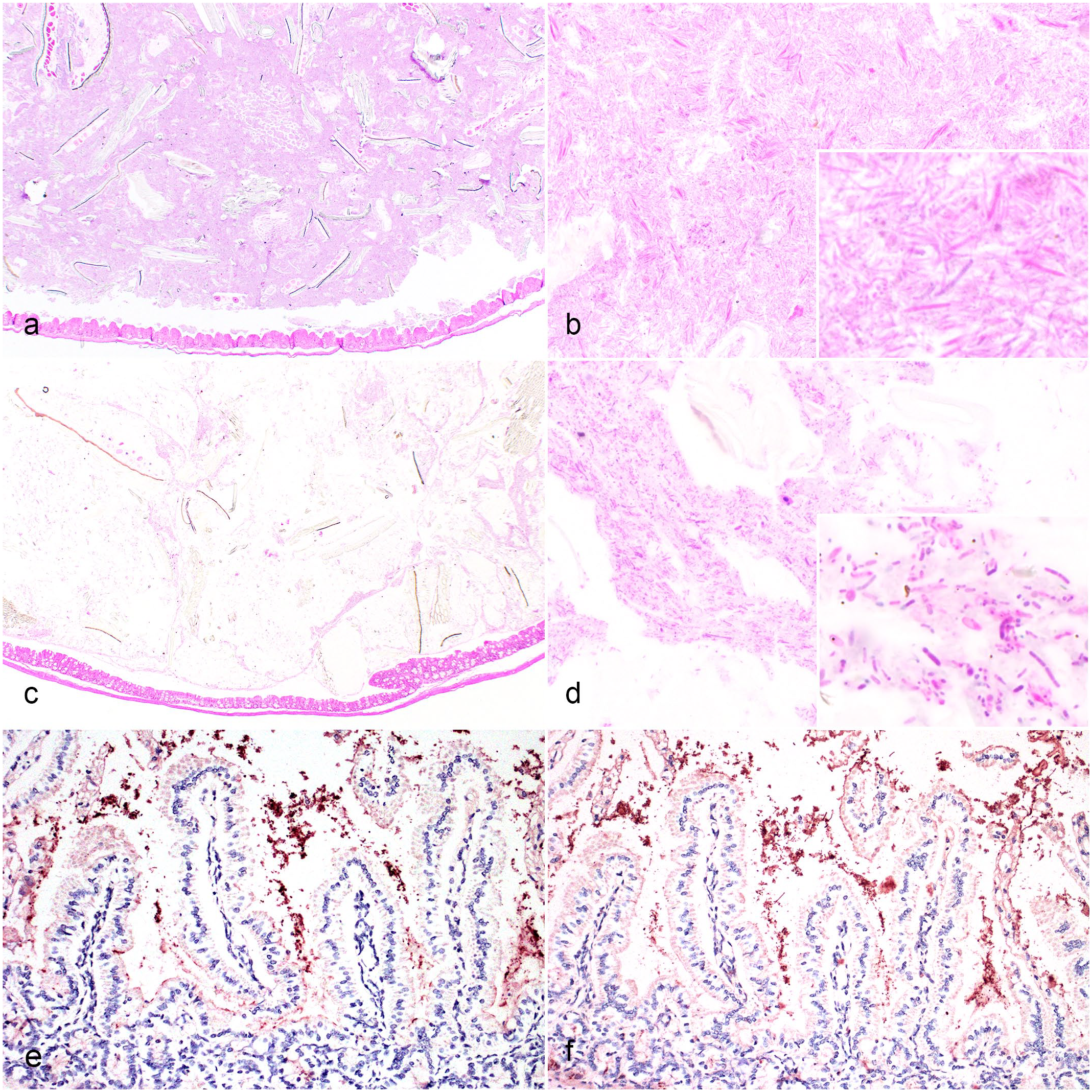
Cecal dysbiosis and mixed small intestinal clostridial immunoreactivity. (a) Typical filamentous bacteria admixed with partially digested feed material fills the cecum of a healthy NOD.Cg.-Prkdc scid Il2rg tm1Wjl /SzJ (NSG) control mouse. Hematoxylin and eosin (HE). (b) Higher magnification of densely packed cecal filamentous bacteria. Inset: higher magnification of filamentous bacteria. HE. (c) Sparsely filled cecum from a cage mate of a moribund mouse. This mouse exhibited small intestinal and cecal gas distension and was Clostridium perfringens positive via microbiological culture. HE. (d) Higher magnification of cecal contents demonstrating a marked reduction in filamentous bacteria and increased number of bacilli. Inset: higher magnification of bacilli in the cecal contents. HE. (e) Small intestinal luminal bacilli with immunoreactivity to Clostridium perfringens in a moribund NSG mouse. Clostridium perfringens immunohistochemistry. (f) A serial section demonstrates immunoreactivity to Paeniclostridium sordellii. Paeniclostridium sordellii immunohistochemistry.
Bacteriology
C. perfringens and P. sordellii were most frequently isolated from the intestine of moribund mice (5/6), diarrheic cage mates (2/2), found dead mice (1/2), and less frequently, asymptomatic cage mates (4/13, 31%). More generally, anaerobic enteric microbiological cultures yielded Clostridium spp. in 13 of 23 (56%) mice and P. sordellii in 7 of 23 (30%) mice. Mixed clostridial species were identified in 4 of 23 (17%) mice and co-isolates of clostridial species and P. sordellii were identified in 3 of 23 (13%) mice. The most commonly isolated clostridial species included C. perfringens (7/23, 30%), Clostridium disporicum (4/23, 17%), Clostridium saudiense (3/23, 13%), Clostridium ramosum (2/23, 9%), and Clostridium butyricum (1/23, 4%). Aerobic enteric microbiological cultures yielded varying levels of Staphylococcus xylosus, Enterococcus faecalis, and/or Lactobacillus johnsonii in all surveyed mice.
Molecular Analysis
All mice were fecal PCR positive for astrovirus. A cohort of asymptomatic, 8-week-old NSG mice (n = 6, 3 males and 3 females) received directly from the vendor and immediately processed for fecal PCR were also PCR positive for astrovirus suggesting that astrovirus is enzootic within the NSG strain and is unrelated to the outbreak. Mice were fecal PCR negative for all other murine pathogens screened for on the Charles River Laboratories Mouse Surveillance Plus PCR Rodent Infectious Agent plus Clostridium PCR panels (see the “Materials and Methods” section for pathogen list).
C. perfringens and C. difficile Toxins ELISA
ELISA for C. perfringens (CPA, CPB, and ETX) and C. difficile (A and B) toxins were negative.
C. perfringens Multiplex PCR Toxinotyping
Toxinotyping was performed on n = 2 culture samples from which C. perfringens was isolated. In both instances, genetic profiling was consistent with C. perfringens type A. Samples were α-toxin (cpa) positive, and β-toxin (cpb), β2-toxin (cpb2), ε-toxin (etx), ι-toxin (iap), and CPE (cpe) negative.
Control Cohort
No anaerobic bacteria were isolated from intestinal content obtained from naïve male and female NSG and NSG-SGM3 control groups. ELISAs in all groups were negative for C. difficile organisms and toxins. All groups were also negative for C. perfringens toxins, although 2 groups (NSG males and NSG-SGM3 females) were positive for C. perfringens organisms. Aerobic microbiological cultures performed on pooled intestinal contents revealed E. faecalis, S. xylosus, and mixed normal flora. Whole-body histology performed on 6 healthy, naïve, NSG mice did not reveal gastrointestinal lesions. Cecal contents were considered normal for NSG mice (i.e., abundant filamentous bacteria, Fig. 3a) and did not exhibit dysbiosis as defined above. IHC results from an unrelated investigation in December 2021 indicated C. perfringens positivity within the lumen of the colon—but not the small intestine—of a healthy control male NSG mouse.
Cell Line Pathogenicity and Culture
Aside from the expected tumor development in subcutaneously inoculated mice, signs of morbidity or mortality were absent, regardless of the cell line or route of administration. Fungal and aerobic and anaerobic bacterial cultures performed at an external diagnostic laboratory (IDEXX BioAnalytics, Columbia, MO) yielded no growth.
Discussion
An approximately 9-week epizootic of high mortality with limited premonitory clinical signs (e.g., diarrhea) resulted in the death of more than 200 adult male and female NSG and NSG-related mice from a single housing room. Histopathologic lesions included erosive to ulcerative enterocolitis in addition to individual epithelial cell death and luminal sloughing. Notable histologic shifts in the amount and proportion of typical cecal bacteria (i.e., dysbiosis) were present in 75% of evaluated mice. Variable numbers of gram-positive, spore-forming bacilli were identified within the small and large intestinal lumen. Enteric microbiological culture and immunohistochemistry identified several different clostridial species within lesioned intestinal tissues and intestinal lumens. Those samples for which toxinotyping was performed identified C. perfringens type A. In the absence of additional toxin gene identification, causality could not be definitively determined. Together, these findings suggest cecal dysbiosis, with associated intestinal clostridial overgrowth, was related to morbidity and mortality. However, the causal relationships between the preceding diet shift, onset of disease, histopathologic lesions, and overgrowth of clostridial organisms is complex. This report highlights the diagnostic challenge of ascribing causality to clostridial organisms, particularly in the context of severely immunodeficient mice.
Over the past several decades, there have been multiple observational studies linking the onset of murine intestinal disease with clostridial organisms, typically C. perfringens or C. difficile.7,14,15,19 Lactating dams of immunocompetent strains, particularly those in their second week of lactation, are most frequently cited as experiencing peracute death. Affected dams exhibit a range of intestinal lesions, including superficial epithelial cell death of the small and large intestine to necrotizing enterocolitis. In published reports, C. perfringens was most frequently isolated following anaerobic culture7,14,19 and C. perfringens alpha-toxin genes have been occasionally identified via PCR. 14 Identification of C. perfringens toxins via ELISA has, to date, not been identified in these mice, presumably due to the availability of assays, the lability of the toxins, or true absence of toxin production by the clostridial organisms. However, because many species of Clostridium are considered commensal and nonpathogenic, demonstrating toxin production is required for a definitive diagnosis.31,33 The mice reported herein share overlapping clinicopathologic features with these lactating dams, namely sudden mortality, gross small intestinal and cecal dilation, erosive to ulcerative enterocolitis or superficial epithelial cell death, and luminal gram-positive bacilli.
Despite the clinicopathologic findings in this report, untangling the causal relationships between the observed enterocolitis, cecal dysbiosis, and clostridial overgrowth is diagnostically challenging. Due to supply chain issues in February 2022 (approximately 3 weeks prior to the initial mortality events), mice in the affected room were shifted from one commercially available irradiated rodent diet to another (see the “Materials and Methods” section for details). According to vendor datasheets (Supplemental Table S3), the percentage of calories from carbohydrates increased from 58% (Teklad 2918, Envigo) to 62.4% (PicoLab, LabDiet). High carbohydrate diets have been associated with C. perfringens overgrowth and disease in domestic ruminants 2 and have been anecdotally implicated in C. perfringens outbreaks in mice.3,6,14 It is plausible that the diet change resulted in room-wide dysbiosis and clostridial overgrowth, as previously described in other species.17,31 It should be noted, however, that NSG mice routinely undergo a change in diet upon arrival at our institution, when they are switched to irradiated diets from autoclaved feed (LabDiet 5K52, LabDiet) provided at the vendor. Despite changes in nutrient composition and decontamination method, there is typically no evidence of intestinal perturbation during this transition. Because of the feed shift in February 2022, it is diagnostically difficult to discern if the diet change resulted in a loss of intestinal homeostasis allowing for clostridial overgrowth and subsequent enterocolitis or if enterocolitis due to an undocumented cause led to dysbiosis and nonpathogenic clostridial overgrowth.
The diagnostic conclusions are further complicated by the variety of clostridial organisms isolated via microbiological culture, as well as an inability to identify C. perfringens toxin genes known to be pathogenic. Though diarrheic, moribund, and found deceased mice had a higher incidence of C. perfringens via culture than cage mates of found deceased mice (Supplemental Table S2), a breadth of clostridial organisms were isolated from both symptomatic and asymptomatic mice. Furthermore, toxinotyping of select C. perfringens cultures were identified as C. perfringens Type A. Extensive reviews of clostridial infections in animals and humans, as well as their associated toxinotypes, have been published elsewhere and are beyond the scope of this discussion. 30 In short, controversy exists around the ability of C. perfringens type A to induce enteric disease in animals. It remains unclear, however, if the immunodeficient status of NSG mice renders them uniquely susceptible to otherwise traditionally nonpathogenic clostridial species. Similar to previous reports in mice, we were unable to definitively identify toxin production via ELISAs for both C. perfringens and C. difficile. It should be noted that due to sampling and logistical limitations, the singular animal for which toxin screening was performed was not moribund, though it did exhibit perianal fecal staining and small intestinal/cecal dilation.
During the epizootic, and prior to obtaining ancillary diagnostic results, investigation into potential sources of pathogen introduction was thoroughly evaluated. Pathogen entry via fomites, experimental biologics, or breaches in biosecurity practices was explored. Introduction of pathogenic agents via contaminated feed or feed packaging has been documented in the literature. 12 A recent report from the Quality Assurance Laboratory at the National Institute of Environmental Health Sciences identified several Clostridium species (including C. perfringens) within 23 lots of nonsterile laboratory animal diets screened for via anaerobic culture. 12 Based on PCR for toxin genes, most of these C. perfringens strains were toxinotype A, although 2 strains possessed the beta-toxin gene, which is reported to be highly fatal in mice.12,28 Gamma irradiation of rodent diets is a common and effective means of pathogen reduction or elimination depending on the irradiation dose, but sterility is not guaranteed and doses of irradiation required to kill bacterial spores (greater than 30 kGy) may exceed doses typically used by feed manufacturers (20–25 kGy). 9 Autoclaving of rodent diets is typically performed on-site and can achieve adequate sterilization (including elimination of spores) providing the autoclave process is appropriately validated. 9 Neither direct examination and culture of experimental biologics used in the affected colony (i.e. osteosarcoma cell lines), nor infectivity studies with said products yielded identification of known pathogens or murine morbidity/mortality, respectively, rendering them an unlikely source of contamination. Personnel-associated breaches in biosecurity were considered unlikely, as room entry procedures were designed to mimic those of our high-health status barrier facility.
Unlike previous reports in immunologically competent murine strains, the impacted population of mice described herein consisted of over 200, adult, male and female severely immunodeficient mice. The spontaneous and explosive nature of this disease outbreak highlights inherent challenges of maintaining severely immunodeficient mouse colonies within the confines of a conventional level facility. Others have suggested that the successful maintenance of NSG colonies requires specialized training for individuals working with them, comprehensive disease surveillance, and standard operating procedures designed to minimize the risk of opportunistic infections.10,33 Despite risk mitigation through the implementation of such procedures and allowing only clean, vendor-supplied animals to enter the room, observations from this outbreak investigation indicate that some threats to the health of severely immunodeficient mice remain difficult to avoid. Notably, immunocompetent soiled-bedding CD-1 sentinel mice housed within the same room under identical husbandry conditions remained clinically healthy throughout the epizootic event, despite being directly exposed to large quantities of presumably contaminated fecal matter from the NSG colony. These observations suggest that NSG and NSG-related mice may be exquisitely sensitive to intestinal perturbations.
In summary, high mortality within a single housing room of adult NSG and NSG-related mice was associated with enterocolitis, cecal dysbiosis, and clostridial overgrowth. The onset of clinical disease was temporally related to a shift in diet due to supply chain disruptions, which may have impacted intestinal homeostasis. Following depopulation of the remaining mice in April 2022 and subsequent decontamination, the room was repopulated, and the colony switched to an autoclaved diet. To date, no recurrent morbidity or mortality has been identified.
Supplemental Material
sj-pdf-1-vet-10.1177_03009858231217197 – Supplemental material for Epizootic of enterocolitis and clostridial overgrowth in NSG and NSG-related mouse strains
Supplemental material, sj-pdf-1-vet-10.1177_03009858231217197 for Epizootic of enterocolitis and clostridial overgrowth in NSG and NSG-related mouse strains by Justin D. Arthur, Jeannie L. Mullen, Francisco A. Uzal, Claude M. Nagamine and Kerriann M. Casey in Veterinary Pathology
Footnotes
Acknowledgements
We thank Shellie Hyde, Dave Schumacher, Amber Durnal, and My Hoang for assistance gathering epidemiological and husbandry data. We wish to recognize the technical contributions of Elias Godoy, Greg Nelson, and Sergio Koba of Stanford University’s Veterinary Service Center, as well as the efforts of Roberta Moorhead, Rina Barouch-Bentov, Brittany Accetturo, and John Duque from the Stanford University Animal Diagnostic Laboratory. Histology processing was completed using the Stanford University Animal Histology Services.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded in part by the Stanford University Department of Comparative Medicine.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
