Abstract
Ranaviruses have been detected in over 12 families of reptiles including many genera of turtles, tortoises, and terrapins, but the pathogenesis of these infections is still poorly understood. Krefft’s river turtle hatchlings (N = 36; Emydura macquarii krefftii) were inoculated intramuscularly with Bohle iridovirus (BIV, Ranavirus, isolate) or saline, and euthanized at 9 timepoints (3 infected and 1 control per timepoint) over a 24-day period. Samples of lung, liver, kidney, and spleen were collected for quantitative polymerase chain reaction (PCR); internal organs, skin, and oral cavity samples were fixed for histopathological examination. The earliest lesions, at 8 days postinoculation (dpi), were lymphocytic inflammation of the skin and fibrinoid necrosis of regional vessels at the site of inoculation, and mild ulcerative necrosis with lymphocytic and heterophilic inflammation in the oral, nasal, and tongue mucosae. Fibrinonecrotic foci with heterophilic inflammation were detected in spleen and gonads at 16 dpi. Multifocal hepatic necrosis, heterophilic inflammation, and occasional basophilic intracytoplasmic inclusion bodies were observed at 20 dpi, along with ulcerative lymphocytic and heterophilic tracheitis and bronchitis. Tracheitis, bronchitis, and rare bone marrow necrosis were present at 24 dpi. Of the viscera tested for ranaviral DNA by PCR, the liver and spleen had the highest viral loads throughout infection, and thus appeared to be major targets of viral replication. Testing of whole blood by qPCR was the most-effective ante-mortem method for detecting ranaviral infection compared with oral swabs. This study represents the first time-dependent pathogenesis study of a ranaviral infection in turtles.
Ranaviruses are globally distributed pathogens responsible for disease in many species of ectothermic vertebrates, including fish, amphibians, and reptiles. 13 Ranaviruses are emerging pathogens: they were only discovered in the 1960s and major epizootics (in wild and captive populations) were not recorded until the 1980s. 13 At least 175 species of ectothermic vertebrates are susceptible to ranaviral disease, but due to underreporting, there are likely many more. 10 Ranaviruses cause disease in wild and captive reptiles, often producing high levels of mortality. 13 Yet, when compared with other ectothermic vertebrates, we know little about the factors that influence ranaviral pathogenesis in reptiles as, for the most part, research on ranaviral pathogenesis has focused on fish and amphibians. 41 A significant amount of research has been performed in some reptilian species, such as the eastern box turtle (Terrapene carolina carolina), but most of the studies have looked at infection at a single timepoint of infection.1,6,18
Australia has a large diversity of reptiles, including several species reported to be susceptible to ranaviral infection.4,20,27 Globally, freshwater turtles are the group of reptiles most commonly reported with ranaviral infections. Australia is home to many freshwater turtle species, primarily side-necked turtles from the family Chelidae. 37 More than 40 % of the freshwater turtle species found in Australia have a conservation status of threatened or worse (at state or federal levels, or according to the IUCN, or some combination of these). 35 Because they have small geographic ranges, extirpation of Australian freshwater turtle populations can result in significant loss to the species as a whole. In 2015, a nidovirus disease outbreak killed almost all adult Bellinger River snapping turtles (Myuchelys georgesi) which were restricted to a small coastal catchment in Australia, the Bellinger River.33,42 As a result of this single disease outbreak, the Bellinger River snapping turtle is now listed as critically endangered and is one of the top 25 most threatened freshwater turtle species in the world, exemplifying the need to have a better understanding of viral diseases in Australian freshwater turtles. 34
Ranaviruses are present in Australian frogs in environments where freshwater turtles are also found. 32 Ante-mortem serological surveys of Australian reptiles (turtles, crocodiles, and snakes) suggest that reptiles are exposed to ranaviruses, and that, particularly amongst turtles and crocodiles, older adults might serve as subclinical carriers. 3 We know some reptile species, such as Krefft’s river turtle (Emydura macquarii krefftii) hatchlings, can develop severe disease when experimentally challenged with ranaviruses. 4 However, to date no ranaviral epizootics have been identified in wild Australian turtles. This is potentially a result of insufficient monitoring of the remote areas where these turtles are found, and a lack of effective testing strategies. The dose-dependent clinical signs and postmortem lesions of ranaviral infection in Krefft’s river turtles have been described; however, we do not have a detailed understanding of the development of the disease. 40
While there have been studies of temporal ranaviral pathogenesis in amphibians and one reptile species,11,26 all published studies in turtles have looked at an endpoint infection. End-stage ranavirosis in turtles often presents as respiratory signs, including nasal and oral discharge and edema of the neck and eyes resulting from rhinitis, stomatitis, and conjunctivitis. 41 However, we do not know how the duration of ranaviral infection influences the underlying histopathological changes for any turtle species. Understanding the time-dependent pathogenesis of ranaviral infection in turtles will help to develop better conceptualize models of ranaviral infection and the disease process, thus allowing recommendations about diagnostic protocols (e.g., when and where to sample). We present the results of an experimental infection with time-staggered sampling designed to determine the pathogenesis of ranaviral infection in Australian Krefft’s river turtle hatchlings.
Materials and Methods
All experiments were carried out under a JCU Animal Ethics permit (A2344) and a Department of Environment and Science Scientific Research Permit (WISP13270413).
Source of Virus
The Bohle iridovirus (BIV) isolate used in this study was kindly provided by Alicia Maclaine and Narges Mashkour following the methods described. 27 This isolate was the original 1992 isolate obtained from ornate burrowing frogs (Platyplectrum ornatum) and has been sequenced.19,32 The viral isolate was propagated at 25°C in fathead minnow cells in Dulbecco’s modified eagle medium (DMEM; Thermo Fisher Scientific, NY, USA), supplemented with 100× antibiotic–antimycotic (Thermo Fisher Scientific, NY, USA) and 10 % fetal bovine serum. The inoculum was titrated to an approximate LD50 dose for Krefft’s river turtle hatchlings, 104.33 TCID50 mL−1. 38
Source of Host
Cathedral traps were used to obtain five gravid Krefft’s river turtles (1.4–1.8 kg) during November 2017. Although vertical transmission has not been documented in turtles, all gravid turtles were tested and found to be negative for the presence of ranaviruses in blood through a qPCR assay (described below). Turtles were temporarily housed in the JCU Veterinary School quarantine area, before being induced to lay their eggs. Oviposition was induced with an injection of 1 IU 100 g−1 of syntocinon (synthetic oxytocin 10 IU mL−1, Troy Laboratories). Eggs were incubated in autoclaved vermiculite for approximately 50 days (µ = 28.9°C, SD = 0.27). Newly emerged hatchlings were housed in the JCU Turtle Health Research Facility.
Experimental Design
Two weeks prior to commencing the experiment, 36 three-month-old hatchlings were randomly selected and assigned to the infection (n = 27) or control (n = 9) groups. Animals were randomly assigned in groups of 4 (3 infected, 1 control) to a predetermined endpoint for euthanasia: 0.33, 1, 2, 4, 8, 12, 16, 20, and 24 days postinoculation (dpi). Infected and control hatchlings were kept in separate rooms to prevent contamination. Hatchlings were housed in slanted 1-l plastic containers with a dry area for basking and 500 ml of water that was changed daily after feeding. The hatchlings were fed ad libitum with commercial turtle pellets (Exo Terra, aquatic turtle food) for 10 to 15 minutes per day. Room temperature was maintained at 28°C via air conditioning, water temperature was an average of 25.5°C (SD = 0.57) throughout the experiment. On day 0 of the experiment, hatchlings were intramuscularly (upper left thigh) inoculated with a 1 × 104.33 TCID50 dose of BIV, or a placebo control (phosphate buffered saline). Morphometric measurements (mass, straight carapace length, and straight carapace width) were recorded at the start of the trial and dietary intake was monitored throughout.
Clinical signs
Prior to, and during, the infection trial, animals were monitored for the development of clinical signs of ranaviral infection in Krefft’s river turtles as previously described. 38 These clinical signs included: inappetence (at least 2 consecutive days of eating less than 25% of provided food), lethargy, skin lesions (single or multifocal raised white to yellow plaques), increased basking behavior, and edema of the neck and limbs.
Endpoint
Turtles were euthanized by MS222 overdose at their predetermined endpoint or when they developed one of more of the severe clinical signs previously reported in similarly infected hatchlings: large epidermal plaques, extensive edema, complete anorexia, or extreme lethargy. 40 Each turtle received an intracoelomic injection of 200 µL of 5% neutral-buffered MS222. Following loss of response to stimulus, a second 300 µL 50% overdose of MS222 was administered via intracoelomic injection. 8
Gross pathology and sample collection
On the day of euthanasia, final morphometric data were recorded and a 1-ml unfiltered water sample was taken from turtle holding containers (after feeding, before water change). A separate tray and instrumentation were used for each animal. Tools were sterilized in a 4% hypochlorite solution for at least 5 minutes before each use and washed with surgical soap after each use. Gloves were changed between animals and surfaces were decontaminated with a hypochlorite solution (4%). Prior to euthanasia, photos were taken of any external lesions. Oral swabs were collected by rolling a cotton swab back and forth on the tongue 10 times and stored in 500 µl of DMEM. A 27½-gauge needle was used to collect between 100 and 200 µl of blood from the external jugular vein. The blood sample was left to clot and then centrifuge-separated after an overnight incubation at 4°C. Following euthanasia, a necropsy was performed, and gross pathological changes were noted. Kidney, liver, and lung samples from all animals in a group, and spleen from one animal per group, were collected and stored at −80°C until DNA extraction and qPCR analysis. The remaining viscera and hind legs were fixed in 10 % neutral-buffered formalin for histological examination.
Real-Time Quantitative PCR (qPCR)
DNA was extracted from the thawed samples (blood clot, swab, water, kidney, liver, lung, and spleen) using an ISOLATE II Genomic DNA Kit (Bioline) following the manufacturer’s protocol for DNA extraction from the appropriate sample type. The quantitative PCR assays were produced and run according to the methods described by Leung et al. 25 In short, a section of the BIV major capsid protein gene (MCP) and EBF3N (a single copy gene conserved in vertebrates) were cloned into pGEM-easy vectors following the manufacturer’s protocol (Promega). Plasmids were linearized using a PstI restriction digest (New England Biolabs) and quantified using a QuantiFluor dye system (QuantiFluor, Promega). Absolute standard curves were constructed in triplicate from 108 to 101 copies µl−1 and 107 to 101 copies µl−1 for the MCP and EBF3N plasmids, respectively. The qPCR was performed on a MIC real-time thermocycler (Applied Biosystems) with the conditions described by Leung et al. The assays were run as a duplex with the MCP probe on the green channel (FAM probe) and ENF3N on the red channel (Cy5 probe). Viral loads were determined by dividing the number of MCP copies µl−1 multiplied by two by the number EBF3N copies µl−1 and are referred to as copies per cell throughout this manuscript.
Histopathology
Formalin-fixed tissues were processed for histological examination as per routine standard operating procedures. 5 Samples selected for histological processing included: head (right half), tongue, leg (injection site), liver (right lobe), heart (cross section of atria), lung (caudal half of right lobe), intestine (at the level of the yolk sac remnant, or Meckel’s diverticulum), stomach (pylorus), spleen, a kidney, and a gonad. Sections that included bone were decalcified in Gooding and Stewart’s decalcifying fluid before embedding. Immunohistochemistry (IHC) and in situ hybridization (ISH) were used to visualize ranaviral protein and ranaviral RNA in infected tissue, respectively. IHC and ISH was performed on 32 (27 infected and 5 controls) and 9 (8 infected and 1 control) hatchings, respectively. The methods for IHC and ISH followed those described. 26
Results
Animals
The median starting mass of hatchlings used in this study was 7.2 g (range, 5.4–8.9 g). Two hatchlings were euthanized prior to their scheduled endpoint because they developed severe clinical signs; specifically, skin lesions (single or multifocal raised white to yellow plaques), extensive edema, lethargy, and loss of appetite. Hatchling A13 was euthanized 6 days ahead of schedule on day 18, and hatchling E3 was euthanized 4 days ahead of schedule, on day 16. For analysis, A13 was grouped with those euthanized on day 20, and E3 was grouped with those euthanized on day 16, unless otherwise stated. There were no mortalities or unscheduled euthanasia in the control animals.
Clinical Signs
Clinical signs observed in this study, in order of first appearance, included: slight swelling at the inoculation site (leg) (4 dpi), increased basking behavior (7 dpi), development of raised white to yellow epidermal plaques at or near the site of injection (8 dpi), swelling around the neck (14 dpi), and inappetence (15 dpi). No clinical signs were observed in any control animals.
PCR
To determine the ranaviral tissue tropisms in addition to viral spread in Krefft’s river turtle hatchlings, viral loads were determined using qPCR at selected time points after infection. Ranaviral DNA was detectable in tissue samples (kidney, liver, lung, and spleen) from some individuals at every time point, but there was large variation in the replicates. The highest viral loads were detected in the 2 animals that were euthanized early due to severe clinical signs. Viral loads in the kidney, lung, and liver tissues increased exponentially (ie, linear increase on a log scale). Viral loads in the liver and lung increased at each time point from 2 to 20 dpi, while kidney increased from 4 dpi and began to plateau at 16 dpi. There was an approximately 1000-fold increase in mean viral load in most tissue samples from the first time point (0.33 dpi) to the peak viral load (20 dpi) (Fig. 1).
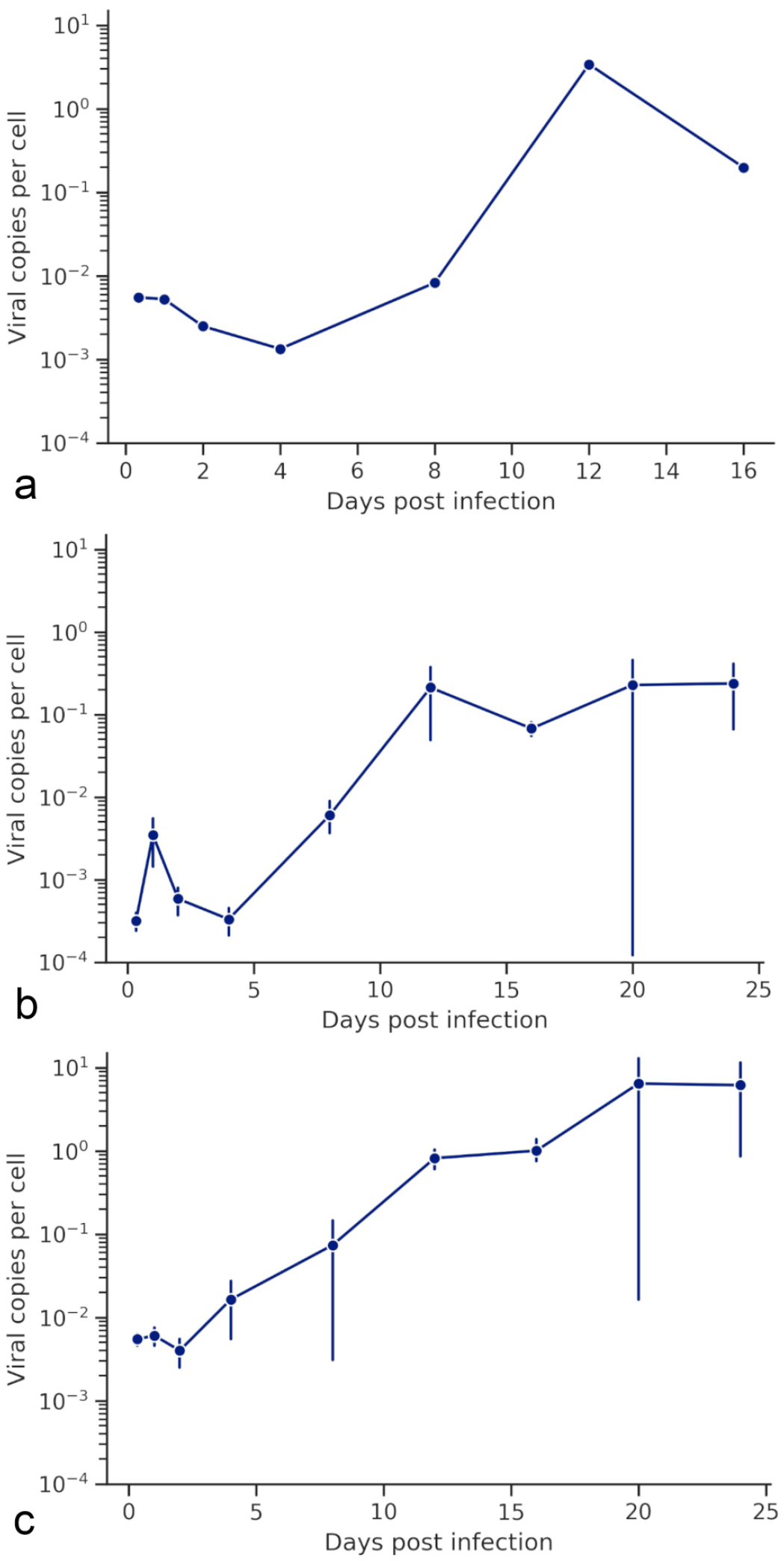
Viral load in spleen, kidney, and liver at various points during infection. These data are expressed as the log10 of the single animal sampled per time point (a, spleen) or as the log10 of the mean viral load +/− standard error of the mean of all animals sampled (b, c; kidney and liver, respectively).
The mean proportion of total viral load was calculated to determine tissue tropisms of this viral infection (ie, which tissues had the highest viral load throughout the course of infection). The highest mean proportion of total viral load was recorded in the liver, which was consistently high and ranged from 50% to 90% of total viral load (Fig. 2). There was large variation in the proportions of total viral load until 12 dpi, after which the liver consistently had the highest load. The spleen had the highest proportion of total viral loads at 0.33 dpi and decreased until 8 dpi and then peaked again in the individual sampled at 12 dpi (Table 1).
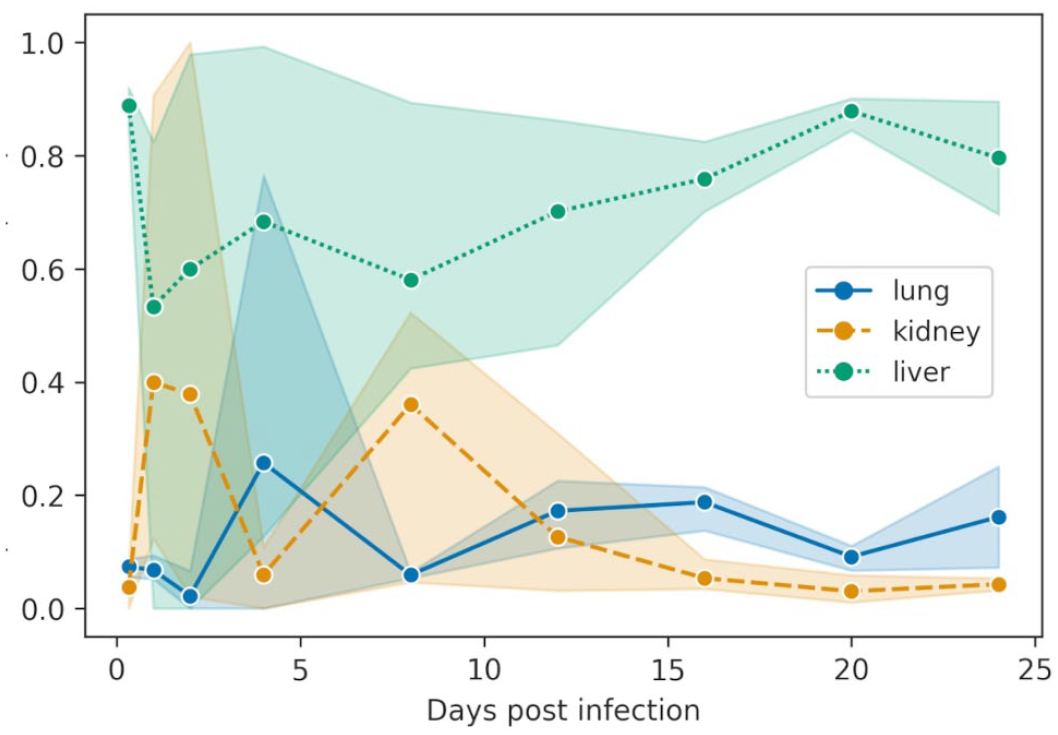
Tropism of Bohle iridovirus for lung, kidney, and liver after intramuscular inoculation of Krefft’s river turtle hatchlings with an LD50 dose (104.33 TCID50) of Bohle iridovirus. The plot shows the mean proportion of total viral load for each sample type (lung, kidney, and liver) at each time point. The shaded area represents the 95 % confidence interval. Day 16 and 20 include data from the animals euthanized early due to severe clinical signs.
Tissue tropisms of BIV in Krefft’s river turtle hatchlings inoculated intramuscularly with an LD50 dose (104.33 TCID50) of Bohle iridovirus.
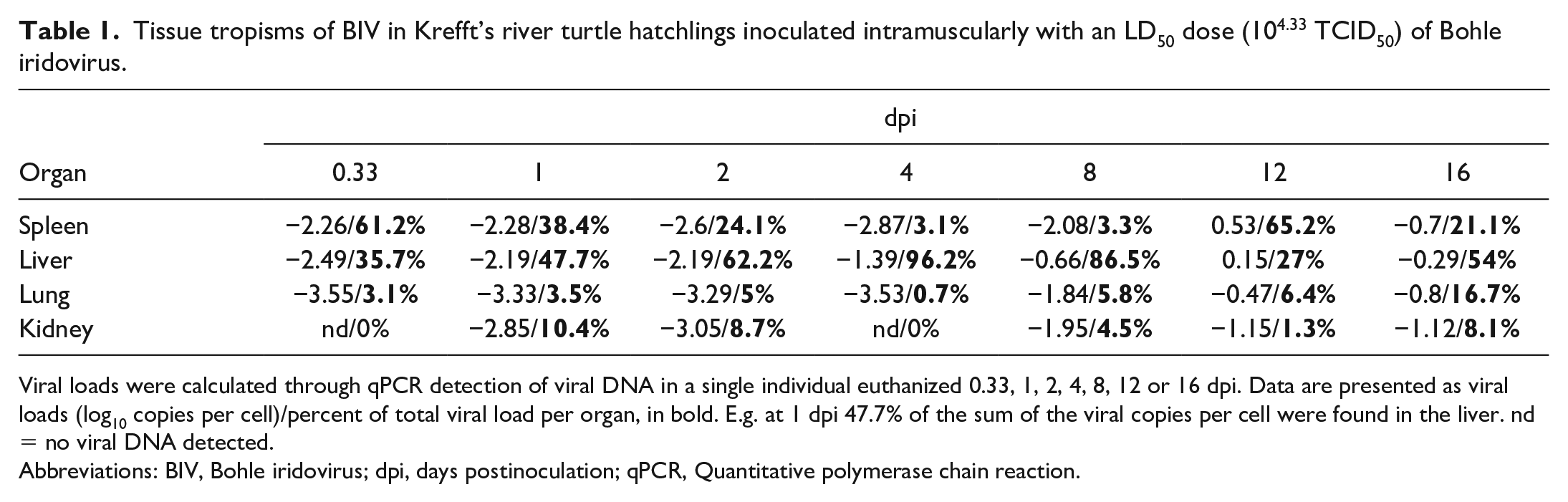
Viral loads were calculated through qPCR detection of viral DNA in a single individual euthanized 0.33, 1, 2, 4, 8, 12 or 16 dpi. Data are presented as viral loads (log10 copies per cell)/percent of total viral load per organ, in bold. E.g. at 1 dpi 47.7% of the sum of the viral copies per cell were found in the liver. nd = no viral DNA detected.
Abbreviations: BIV, Bohle iridovirus; dpi, days postinoculation; qPCR, Quantitative polymerase chain reaction.
To compare the viability of different ante-mortem sample types for ranaviral infection detection, the viral loads of environmental water, blood clots, and oral swabs were also quantified. Viral DNA was detected in the water from infected hatchling environments at 0.33 dpi and decreased to 8 dpi but was not detected in the water at 2 dpi (Fig. 3). No swabs reacted in the ranaviral qPCR assay until 16 dpi. The mean viral copies per cell increased from 16 dpi until the last endpoint (24 dpi); however, only one individual was positive at each time point. Oral swabs from both the individuals that were euthanized early due to severe clinical signs (A13 on day 18 and E1 on day 16) reacted in the qPCR assay. Viral DNA was detected in blood samples from individual turtles from every endpoint, but blood samples from all three infected animals from an endpoint were not positive until 16 dpi and 1 individual was still negative at 20 dpi (Fig. 3). All samples (tissues and ante-mortem) from all control animals were consistently negative at all time points.
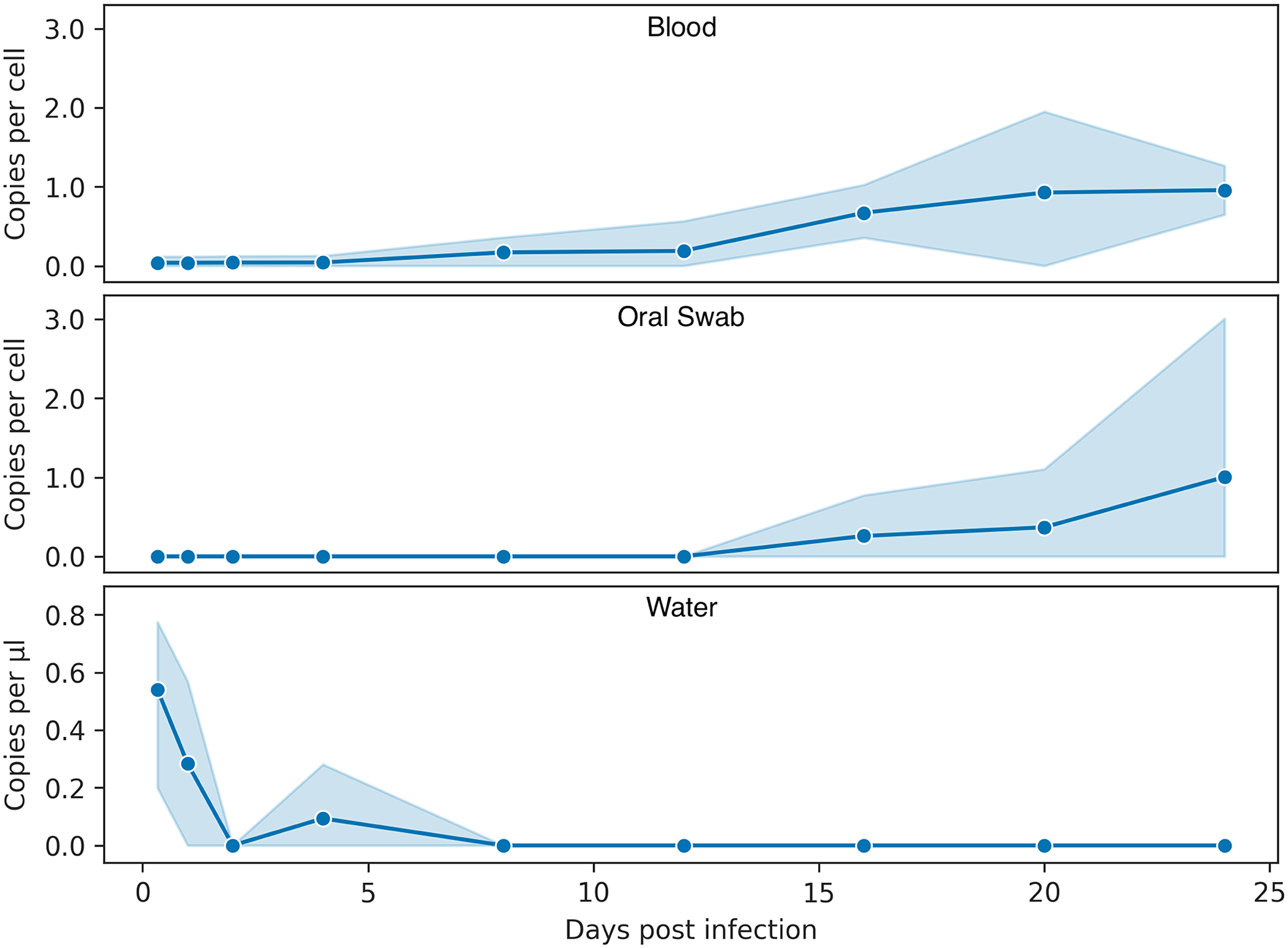
Viral copies in different ranaviral sample types over time. All data were cube-root transformed to reduce right skew and improve visualization. The data can be converted back to standard units by cubing (y 3 ) each value. The data (means and 95% confidence interval) show the viral load (copies per cell) in blood clots (top) and oral swabs (middle), and viral copies per µl of water from the environment of infected turtles (bottom). Days 16 and 20 include data from the animals euthanized early due to severe clinical signs.
Gross Pathology
Gross lesions, only observed at necropsy of infected individuals after 12 dpi, included raised oral plaques surrounded by erythema and petechial hemorrhages, soft tissue edema, splenomegaly, splenic congestion with multifocal to coalescing white discoloration (suggesting multifocal necrosis and coinciding with the highest viral load in the organ) (Fig. 4), air in the digestive track (tympanism), congested intestinal blood vessels, and multifocal discoloration or mottling of the liver suggestive of hepatic necrosis.
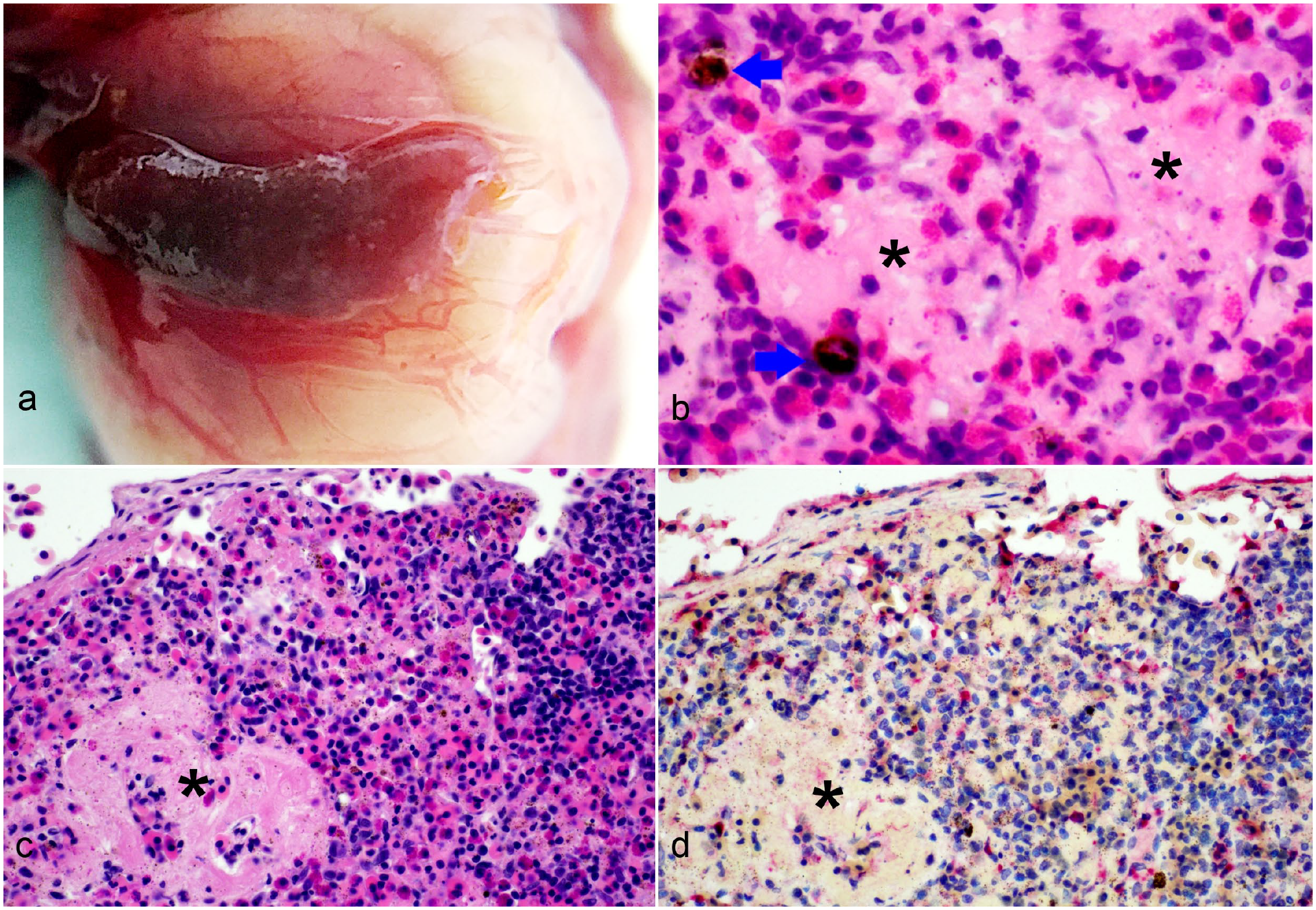
Bohle iridovirus infection, spleen, Krefft’s river turtle hatchling, 18 to 20 days postinfection. (a) The spleen is enlarged and discolored, with congestion of surrounding blood vessels. (b, c, d) Necrosis of the spleen, with amorphous to fibrillar eosinophilic debris (asterisks), degranulated and intact heterophils and occasional pigment-laden macrophages (melanomacrophages or hemosiderophages, arrows) (hematoxylin and eosin, b–c). (d) Immunohistochemical immunolabeling for viral antigen is present in the necrotic foci as well as scattered immune and stromal cells.
Histopathology
Systemic histological lesions were observed in infected individuals only (Table 2). Histopathological lesions, detected first at 8 dpi, consisted of dermal lymphocytic inflammation and vascular fibrinoid necrosis with lymphocytic infiltration of affected vascular walls in the dermis and skeletal muscle near the inoculation site. Mild necrosis of the oral, nasal, or tongue mucosae along with lymphocytic and heterophilic inflammation were also detected at 8 dpi. Multifocal to coalescing necrosis with heterophilic inflammation in the spleen (Fig. 4) and gonads (Fig. 5), and fibrin deposition particularly in the spleen, were present from 16 dpi onwards. In the liver, multifocal necrosis and associated heterophilic inflammation were detected at 20 dpi (Fig. 6). Lymphocytic to heterophilic inflammation and ulcerative necrosis of the trachea and bronchi were observed at 20 dpi, and were more severe at 24 dpi. Occasional necrosis of bone marrow cells, with changes suggestive of intracytoplasmic inclusion bodies, was observed in one individual at 24 dpi. Mild lymphohistiocytic interstitial inflammation was found in a single individual euthanized at 2 dpi, and one euthanized at 24 dpi.
Histopathological lesions in Krefft’s river turtle hatchlings euthanized at different dpi after intramuscular inoculation with an LD50 dose (104.33 TCID50) of Bohle iridovirus.
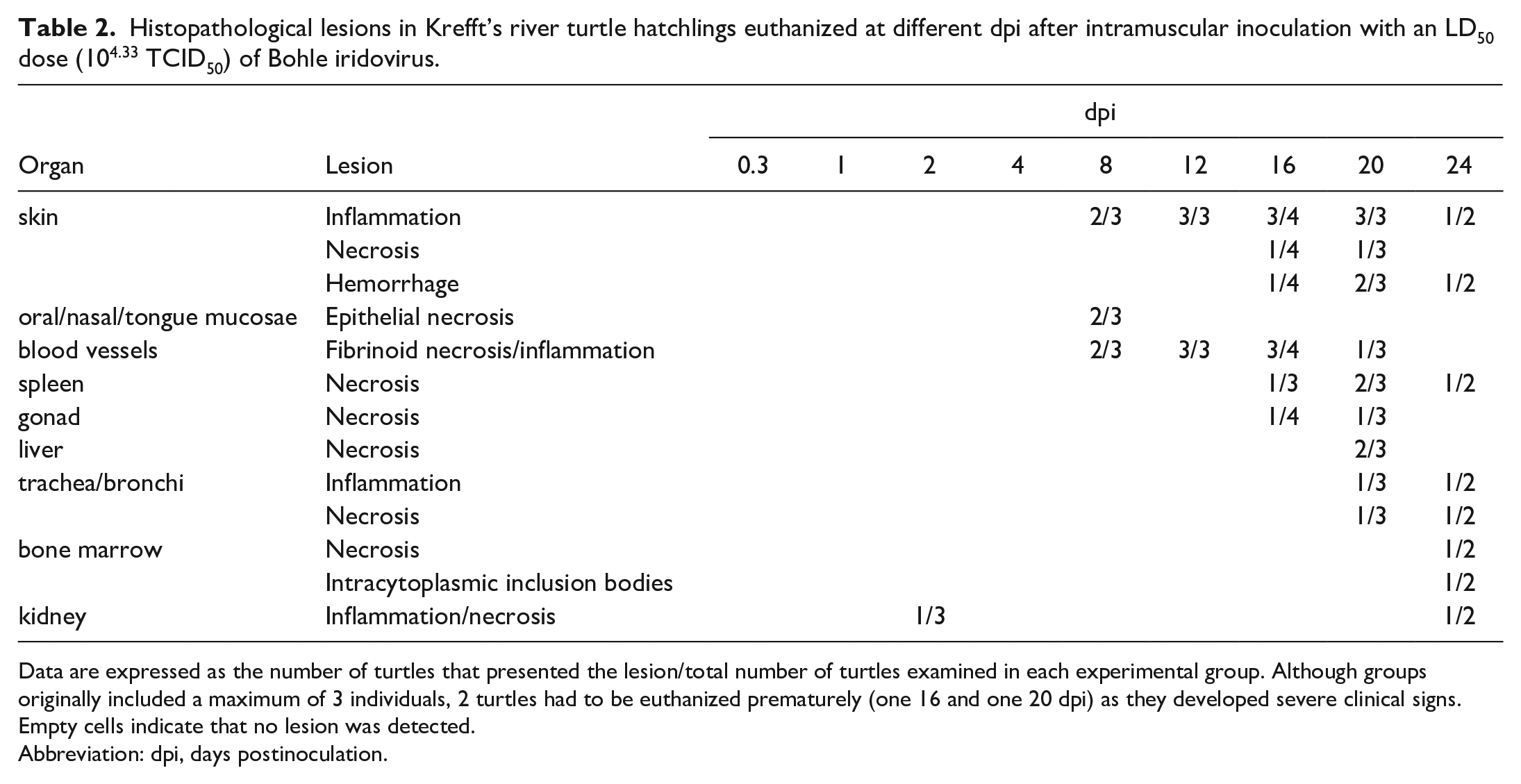

Data are expressed as the number of turtles that presented the lesion/total number of turtles examined in each experimental group. Although groups originally included a maximum of 3 individuals, 2 turtles had to be euthanized prematurely (one 16 and one 20 dpi) as they developed severe clinical signs. Empty cells indicate that no lesion was detected.
Abbreviation: dpi, days postinoculation.
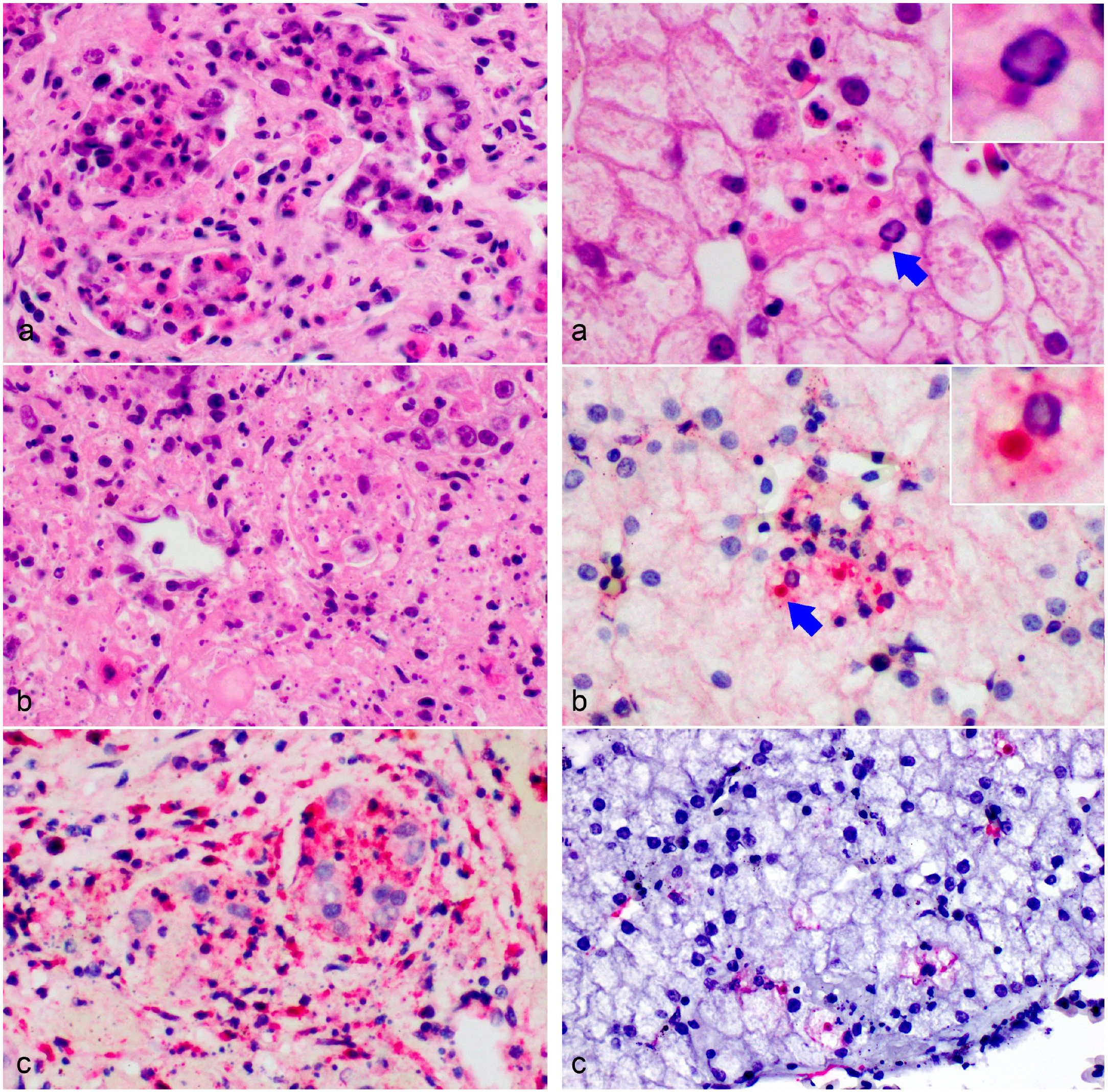
Bohle iridovirus infection, gonads, Krefft’s river turtle hatchling,18 days postinfection. Multifocal to coalescing tubular epithelial degeneration and necrosis (a, b), sometimes resulting in complete obliteration of the tubular architecture (b) associated with interstitial necrosis, and fibrinoheterophilic inflammation (hematoxylin and eosin). (c) Immunohistochemical immunolabeling for viral antigen is present in the necrotic tubules as well as scattered immune and stromal cells.
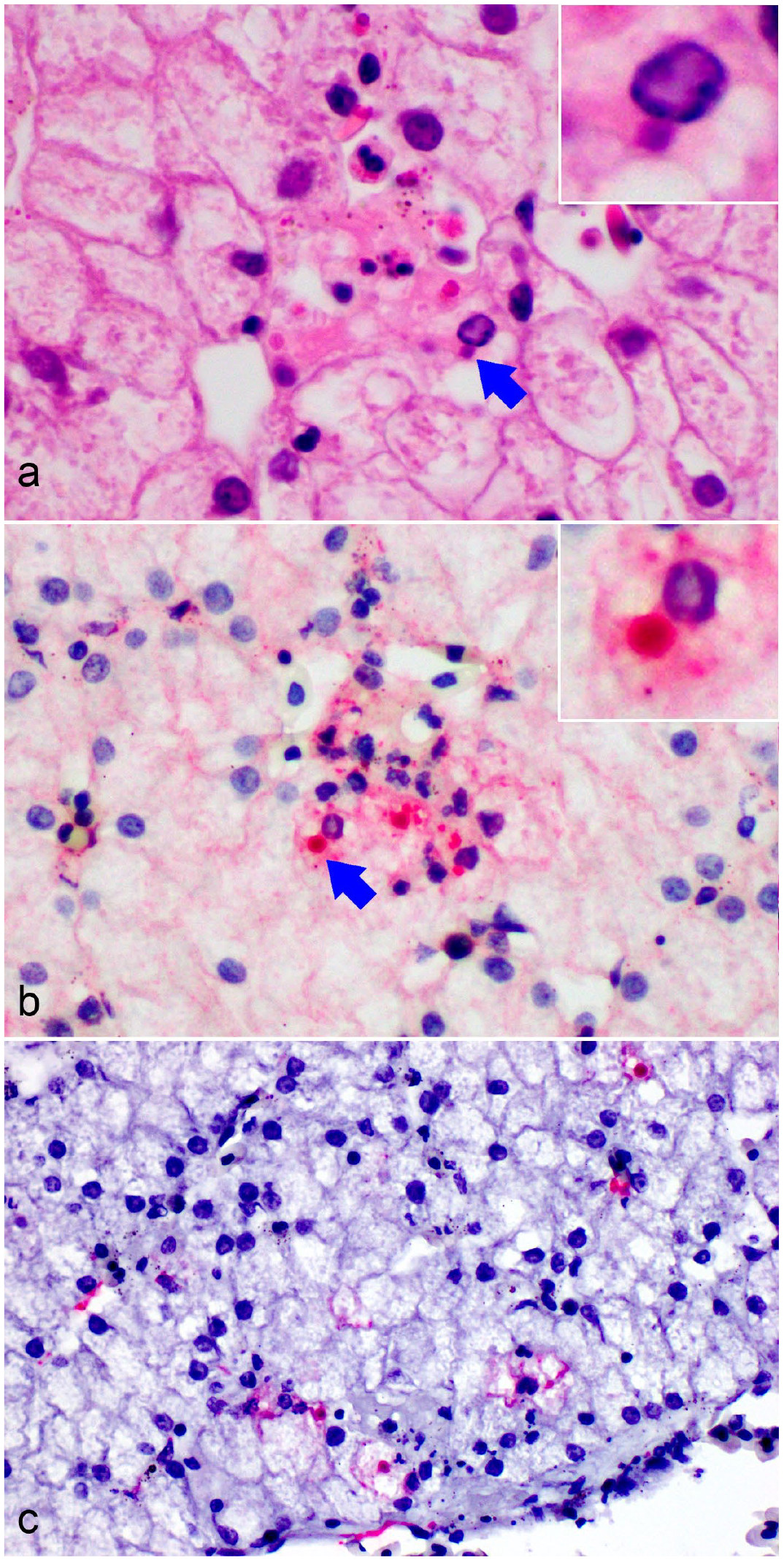
Bohle iridovirus infection, liver, Krefft’s river turtle hatchling,18 days postinfection. (a) A small focus of necrosis, with a small amount of fibrin deposition and few heterophils is associated with hepatocytes that appear to contain intracytoplasmic inclusion bodies (arrow and insert, hematoxylin and eosin). (b) Intense immunohistochemical immunolabeling for viral antigen is present in the focus of necrosis, highlighting the viral inclusions (arrow and insert). (c) Some hepatocytes and sinusoidal endothelial cells are positive for ranavirus major capsid protein gene RNA via in situ RNA hybridization.
No immunolabeling (IHC, Table 3) for ranavirus was present in any tissue prior to 4 dpi, and detection of ranavirus RNA (ISH, Table 4) was not evident until 16 dpi. Ranaviral RNA and protein labeling by ISH and IHC, respectively, were associated with histopathological changes. In the early stages of infection, it was difficult to differentiate true IHC positivity from nonspecific background staining. There was some weak immunolabeling in the bone marrow hematopoietic cells near the injection site at 4 dpi. Skin was positive by IHC on day 12 pi. More generalized immunolabeling was observed at 16 dpi, at which point labeling of viral antigen in spleen, testis, and oral mucosa tissues could be confidently identified (Table 3). Splenic tissues had consistent immunolabeling (two or more individuals) from 16 to 24 dpi (Table 3). Only one liver sample (18 dpi) was clearly positive by IHC (Table 3), this sample was from an animal that was euthanized due to clinical signs (A13). Positively labeled intracytoplasmic inclusion bodies were observed in the liver of A13 at 18 dpi (Fig. 6b). Trachea/bronchi and esophageal immunolabeling was observed at 18 dpi, and the trachea and bronchi were positive in one animal at 24 dpi.
Immunohistochemical labeling of tissues from Krefft’s river turtle hatchlings euthanized at different dpi after intramuscular inoculation with an LD50 dose (104.33 TCID50) of Bohle iridovirus.
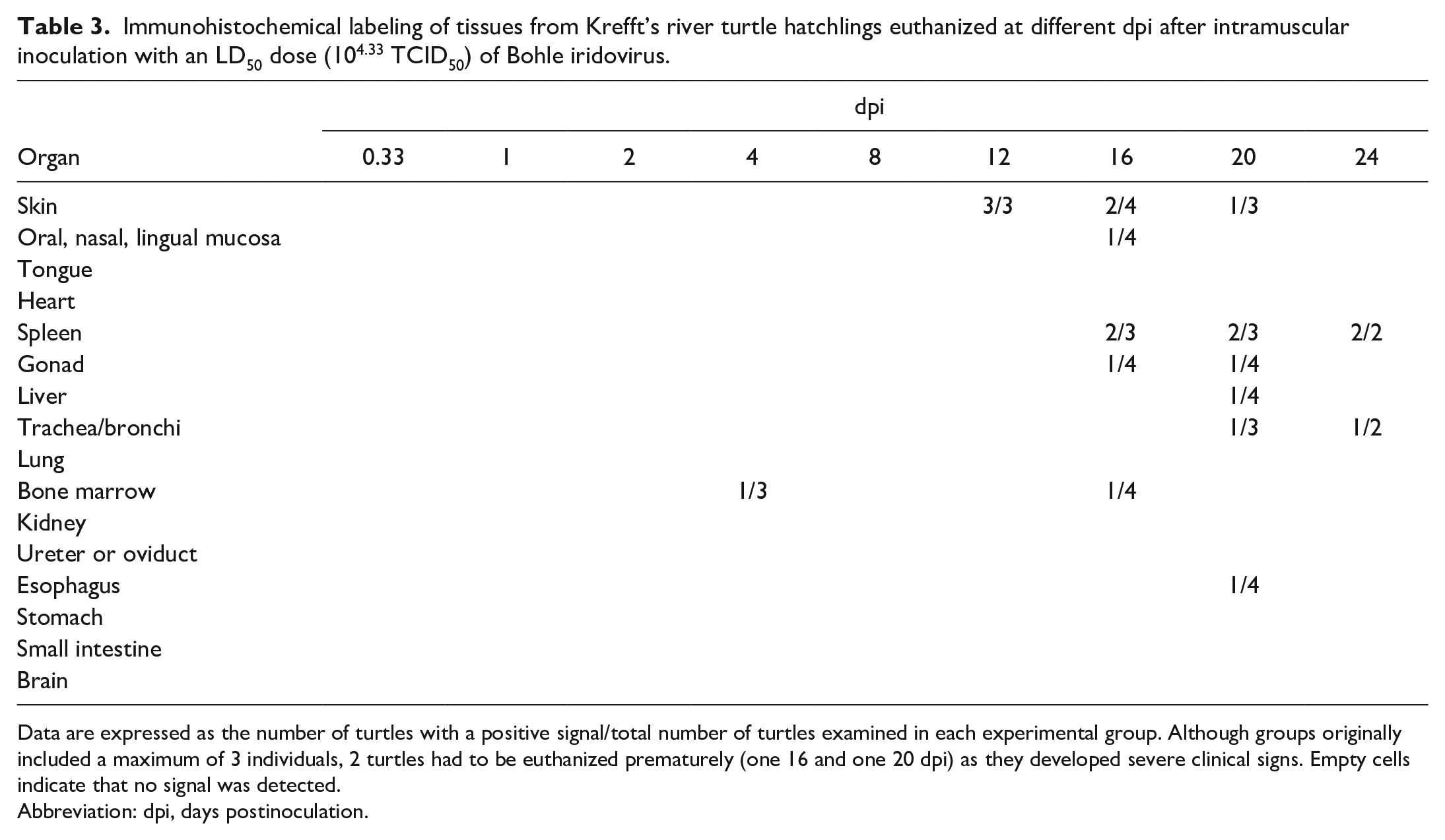
Data are expressed as the number of turtles with a positive signal/total number of turtles examined in each experimental group. Although groups originally included a maximum of 3 individuals, 2 turtles had to be euthanized prematurely (one 16 and one 20 dpi) as they developed severe clinical signs. Empty cells indicate that no signal was detected.
Abbreviation: dpi, days postinoculation.
In situ hybridization labeling of a subset of tissues from one to three Krefft’s river turtle hatchlings euthanized at different dpi after intramuscular inoculation with an LD50 dose (104.33 TCID50) of Bohle iridovirus.
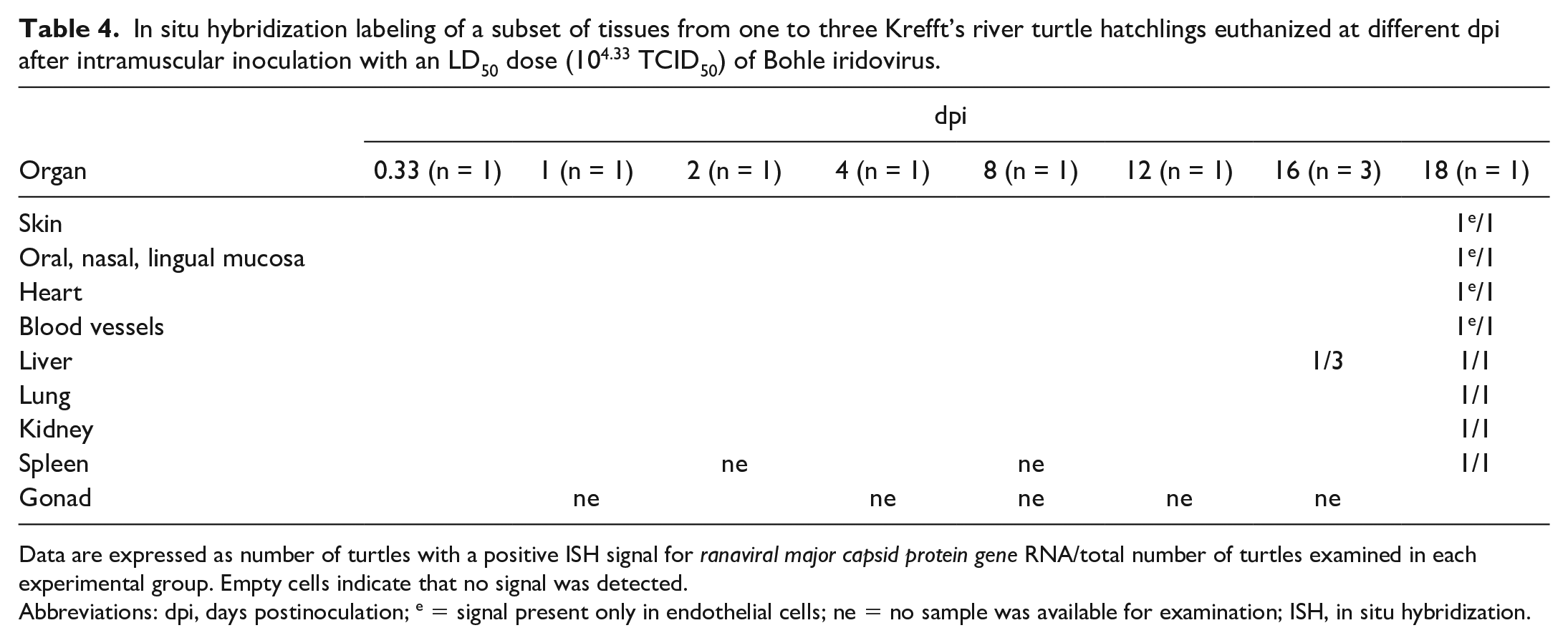

Data are expressed as number of turtles with a positive ISH signal for ranaviral major capsid protein gene RNA/total number of turtles examined in each experimental group. Empty cells indicate that no signal was detected.
Abbreviations: dpi, days postinoculation; e = signal present only in endothelial cells; ne = no sample was available for examination; ISH, in situ hybridization.
By ISH, clear labeling of ranaviral RNA was not observed in the liver until after 16 dpi. Wide-spread labeling of ranaviral RNA was observed in the individuals that were euthanized early (days 16 and 18) (Table 4). This widespread labeling was predominantly observed in endothelial cells rather than parenchymal cells, although clear labeling was observed in the kidney, lung, spleen, and liver. No labeling was observed in any of the control animals.
Discussion
We report the first time-dependent pathogenesis study of a ranaviral infection in turtles, with implications for best sampling protocols, although further study will be required to determine the clinical value of these findings.
The quantitative PCR assay used in this study detected ranaviral DNA in all sample types at some point during infection. The most-effective antemortem method for detecting ranaviral DNA in these turtles was qPCR on blood. At least one turtle blood sample was positive at every time point from 8 hours to 28 dpi. Another study on ranaviral infection in cryptodirid turtles found that oral swabs and whole blood had comparable sensitivities in turtles that were challenged with a ranaviral isolate via intramuscular injection. 2 We found that blood samples were consistently more sensitive for ranaviral detection compared with oral swabs from intramuscularly inoculated pleurodirid turtles. Oral swabs reacted in the ranaviral assay only on the last 3 time points, 16 dpi (1/4), 20 dpi (1/3), 24 dpi (1/2), and two of these reactors were samples from the animals that were euthanized due to severe clinical signs. Therefore, swabs may only be effective at identifying late-stage infections in Australian freshwater turtles. Similarly, oral swabs were also effective only in the late stages of ranaviral infection in experimentally inoculated wood frogs. 11 Studies of wild turtles have also demonstrated inconsistency in detection rate between sample types from the same turtles.1,12,23 While ranaviral loads in blood samples from eastern box turtles (Cryptodira spp) peak first, oral swabs contained viral DNA for a longer period of time. 18 It is possible that as infection continues (ie, past the 24 days of this study) more oral swabs from Krefft’s river turtles would have become positive. It is also unclear what the influence of reinfection would be on optimal sample choice.
In a resource-constrained setting, collecting blood samples from Australian freshwater turtles with potential ranaviral infection should be prioritized over other nondestructive sample types. While blood samples were positive at all sampling points throughout infection, and therefore, potentially a good sample for detecting ranaviral DNA at any time during the infection, this may be artificial and a result of the inoculation route (ie, intramuscular injection) used in the study. Intramuscular injection is an effective method for producing ranaviral disease in turtles; however, we do not know the natural infection route for turtles, and we know clinical signs can change with inoculation route which may have implications for best sampling methods. 40 While oral swabs were not a sensitive diagnostic sampling method until late in infection, they may be more effective in studies of wild animals if ranaviral infection occurs orally. Future studies should examine this possibility by comparing the adequacy of different sample types in wild infections.
Environmental DNA from water has been used to detect ranaviruses in amphibian environments.16,17,29,36 Although ranaviral infection in Krefft’s river turtles has a long incubation period, they do not appear to shed virus during most this time. Ranaviral DNA was only detected in water from the environment of infected hatchlings at the start of infection. It is possible that this early detection (0.33, 1, and 4 dpi) is the result of leakage from the injection site and not viral replication. These results suggest that environmental DNA may not be an effective method for ranaviral detection in Krefft’s river turtles under the current experimental setting. This study was scheduled for termination at 24 dpi and although acute clinical signs appeared well within this time period, potential survivors and carriers may initiate shedding of virus at a later stage. Krefft’s river turtles may not be effective viral spreaders (in a waterborne epidemic); however, more extensive studies will be required to investigate the full extent of viral shedding in this species. Although viral shedding was not observed in this study, as previously suggested, infected turtle hatchlings could still spread the infection to susceptible predacious species.31,40
Ranaviral transmission via vectors has been suggested, and ranaviral DNA has been detected in mosquitoes associated with a ranaviral epizootics in turtles. 24 Freshwater leeches, a common parasite of aquatic turtles (and often found on Krefft’s river turtles), could also serve as possible ranaviral vectors.28,41 The work presented here adds to the evidence for possible vector-borne transmission. We show that ranaviral DNA was detectable in the blood of infected turtles throughout infection. Although DNA presence is not evidence of infectious virus, it is plausible that a blood-feeding vector (mosquito, leech, etc.) could ingest the virus when consuming the blood meal. Ranaviruses share a common ancestor with iridoviruses that infect invertebrates, 21 so it is also conceivable that these invertebrate vectors still play a role in their biology, although transmission studies will be required to elucidate the potential of vector transmission in ranaviral disease.
A study on ranavirus-inoculated red eared sliders (suborder Cryptodira) found that kidney had the highest viral load in postmortem samples (except for tissues at the inoculation site). 2 In this study, liver had a constantly higher mean viral load when compared with lung and kidney samples. If samples are collected at post-mortem from turtles of the Pleurodira ranaviral screening, it is advisable to collect liver or spleen samples. Both sample types were positive throughout infection, although only a few spleens were obtained for qPCR. These samples also had the highest viral loads during infection, which would improve detection rates when using a less sensitive assay. Bone marrow has been identified as an effective sample type for ranaviral detection in turtles (suborder Cryptodira), especially in cases where other samples have degraded. 6 While we did not test bone marrow with qPCR, we did see immunolabeling at 4 and 16 dpi, and mild necrosis 24 dpi. This suggests that bone marrow samples may also be suitable for Krefft’s river turtles, although a direct study would be useful to determine the suitability of this sample type in Australian turtles.
Clinical signs observed in this study were consistent with those reported in ranaviral infection of reptiles.4,40 Clinical signs included edema of the neck and legs, increased basking behavior, single or multifocal raised white to yellow plaques at or near the site of injection, and inappetence. Ranaviral infection in Krefft’s river turtle hatchlings was a slow process: clinical signs appeared only after 16 dpi. Based on previous work on this species, 40 an oral inoculation, which would arguably be more similar to a natural infection, would have probably resulted in an even later onset of clinical signs. In comparison, wood frogs (Rana sylvatica, Lithobates sylvaticus) infected with frog virus 3 and Eastern water dragons (Intellagama lesueurii lesueurii) infected with BIV developed mild lesions at 2 and 3 dpi, respectively, and were euthanized (or had died) by dpi 14.11,26 The long incubation period in Krefft’s turtles means that infected yet asymptotic animals may go unnoticed, especially in areas where populations are not routinely monitored. The lack of specificity of clinical signs further complicates detection of ranaviral infection outside of experimental settings.
Histological labeling methods like IHC and ISH can be used to show the association of histologic lesions with viral protein and nucleic acid; however, these methods were only useful in the late stage of infection. Using a combination of these labeling methods it was possible to determine that viral replication was occurring via multiple lines of evidence. The spleen had the most severe histopathological changes (common for ranaviral infection). Surprisingly, although ISH is generally considered a more sensitive detection method, IHC was more effective in detecting viral presence at earlier stages of infection. Budgetary constraints resulted in only a small subset of animals being tested by ISH, so it is possible that our lack of detection reflected individual variability or poor RNA quality in our samples. As RNA quality was not tested in the tissues used for histopathology (no other probe was available to use as an internal control), we cannot discount the possibility that low RNA quality may have resulted in a lack of ISH labeling. RNA quality seems an unlikely explanation, however, since all tissues were processed in exactly the same manner, we used a technique specifically applicable to formalin-fixed paraffin-embedded tissues, 26 and strong labeling was indeed present in turtles at late stages of infection. Alternatively, the difference may have to do with the techniques themselves. The ISH method used in this study is based on detection of viral RNA, whereas the IHC antibodies detect capsid proteins. Perhaps a paucity of cells in active RNA transcription at the time of sampling could explain the low detection sensitivity of the ISH in our study.
Histopathological changes followed from the site of inoculation to the surrounding tissues, to the spleen and gonads, and finally to the liver and upper respiratory tract. This was similar to the progression of disease in orally infected Eastern water dragons in which splenic necrosis precedes hepatic necrosis. 26 Immunolabeling and qPCR results also suggested that the spleen is a primary site of replication in Krefft’s river turtles. Immunolabeling was consistently observed in the spleen from 16 dpi. In orally infected wood frogs (Rana sylvatica), splenic and hepatic necrosis occurred concurrently. 11 Difference in pathogeneses between these studies may be a result of different study designs (e.g., viral dose and inoculation route) or it may reflect taxonomic differences between hosts.30,41 Although mild mononuclear interstitial nephritis was noted in two individuals, one early and one late in the course of infection, no IHC or ISH labeling confirmed the presence of virus in the tissue, so that it was not possible to determine whether the change was an incidental finding or truly due to BIV infection.
It is possible that ranaviral pathogenesis could change with differing inoculation routes. As histopathological changes were first observed at the site of inoculation, it is likely that these changes are site-dependent and may not be reproduced with different inoculation routes. Indeed, an orally inoculated Krefft’s turtle developed oral lesions, not skin lesions. 40 The extent of these differences in pathogenesis (ie, in the viscera) will have to be examined with more detailed studies.
Ranaviral damage to the host is believed to be driven, at least partly, by the inflammatory response. 14 Inhibition of leukotriene (lipid mediators of inflammation and immune regulation) synthesis can dramatically reduce hepatic damage in rodent models exposed to ranaviruses. 15 While necrosis is observed late in infection, the extent of ranaviral RNA and protein labeling (via ISH and IHC) was limited. Two possible explanations for this immunolabeling pattern include: the necrosis is the direct result of viral replication, but the virus has since dissipated (thus no labeling); or the pattern of immunolabeling is a result of inflammation-driven pathogenesis due to an overactive immune response, as seen in other animals. Indeed, other studies of ranaviral infection in Krefft’s river turtles have also reported an influx of immune cells at the sites of necrosis, although it is difficult to unravel the order of events.4,38
Necrosis in the gonads of turtles inoculated with BIV via intramuscular injection has been previously reported. 38 Here, we showed there was necrosis in the gonads after 16 dpi and that viral antigens were associated with this necrosis. Tropisms for reproductive organs have not been identified in other ranaviral reports and may have implications for species conservation, if this necrosis causes a loss of reproductive potential. Although proving vertical transmission or ranaviruses is beyond the scope of this study, and considering that pathogens may target gonads without necessarily being transmitted to the progeny, our findings suggest the possibility of sexual or vertical transmission in this species.
Descriptions of ranaviral infection in wild turtles often include lesions in the respiratory tract. 41 In our study, histopathological changes in the respiratory tract did not occur until late in infection. It is possible that naturally infected turtles presenting with respiratory clinical signs were already in the later stages of infection. There are several reports of failed treatment for ranaviral infection in turtles,9,22 and it is possible that once these animals appear sick enough to receive veterinary intervention (ie, once they have respiratory distress), the disease is too far advanced to benefit from such treatment. 41 A time-dependent study similar to the one presented here, assessing the effectiveness of treatments (antiviral drugs, fluids, temperature therapy) at different stages of infection (during the incubation, prodromal, and ranavirosis phases) would be useful for veterinarians trying to decide if continued treatment is worth the cost and potentially prolonged suffering of the affected animal. Oral-respiratory disease is also not pathognomonic for ranaviral infection and may be the result of mycoplasmosis, herpesvirus, or adenovirus infection in turtles. Ranaviral infection should not be ruled out if there are no respiratory signs, as the animal might be in the prodromal stage of infection, before respiratory signs have developed.
Reported studies of ranaviral infection and disease have focused on cryptodirid turtles. Turtles from the suborder Pleurodira diverged from turtles in Cryptodira around 157 million years ago. 7 This split represents a deep evolutionary divide between the suborders, and thus, it is important to conduct research on both clades to get a better understanding of ranaviral disease in turtles as a whole.38,41 Pleurodirid turtles, such as Krefft’s freshwater turtle, inhabit Australia, South America, and Africa, as opposed to the cryptodirid turtles that inhabit Europe, Asia, and North America. While members of the suborder Pleurodira are susceptible to ranaviral infection,4,41 no wild or captive epizootics have been documented, although there is evidence of wild exposure from a serological survey. 3 There are many possible causes for the perceived differences between these groups, for example, pleurodirid turtles may be less susceptible than cryptodirid turtles to ranaviral disease, and thus there are fewer pleurodirid epizootics. 41 Ranaviruses themselves may be less abundant in areas inhabited by pleurodirid turtles, as there are also fewer reports of ranaviral outbreaks in other susceptible groups (ie, fish and amphibians) in these areas. 10 It is also possible that there is a difference in research effort on the two suborders: Northern hemisphere (United States and China, in particular) species likely benefit from the higher ranaviral researcher presence and output in these areas. 39 Increased surveys of wild pleurodirid turtles will help clarify the causes of some of these apparent differences.
This research describes the time-dependent histopathological changes in turtles with ranaviral infection and has implications for diagnosis and sampling. Using Krefft’s river turtles as a representative model for the suborder Pleurodira, we have shown that differences between the Pleurodira and Cryptodira (the other extant turtle suborder) extend to ranaviral pathogenesis. Studies of pathogenesis, like the work presented here, provide researchers with a baseline level of knowledge that allows them to assess the factors influencing wild disease, design better experiments, and begin to untangle the complexity of infection.
Footnotes
Acknowledgements
The authors acknowledge Anna Tzamouzaki for her help with husbandry, data collection, and sample processing. The authors thank Robin T. Pasiak and Melissa Fadden, Cornell University, for their assistance with the immunohistochemistry (IHC) and in situ hybridization (ISH) staining, respectively.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
