Abstract
A juvenile, male tiger shark (Galeocerdo cuvier) developed illness after capture in Florida waters and was euthanized. Gross lesions included mild skin abrasions, hepatic atrophy, and coelomic fluid. Histologically, gills contained multifocal lamellar epithelial cell necrosis and thromboses. Scattered gill and esophageal epithelial cells had large, basophilic, intracytoplasmic, and intranuclear inclusions. Ultrastructurally, lamellar epithelial cells contained arrays of intracytoplasmic viral particles and scattered intranuclear nucleocapsids. Capsulated virions were 148 ± 11 nm with an 84 ± 8 nm icosahedral nucleocapsid and an electron-dense core. Next-generation sequencing, quantitative polymerase chain reaction, and in situ hybridization performed on formalin-fixed tissue confirmed a herpes-like viral infection. The viral polymerase shared 24% to 31% protein homology with other alloherpesviruses of fish, indicating a divergent virus. This report documents the pathologic findings associated with a molecularly confirmed novel herpes-like virus in an elasmobranch.
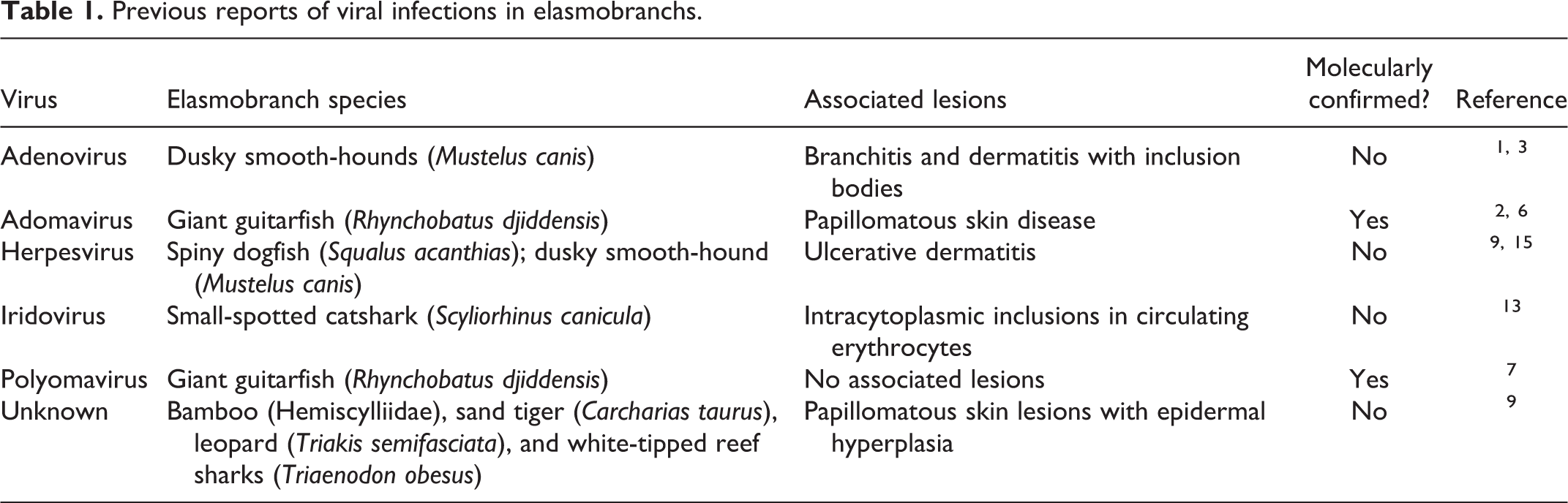
Elasmobranchs (sharks, skates, rays, and sawfish) are popular display animals and have essential roles in global ecosystems. Infectious diseases, particularly viral diseases (Table 1), afflicting elasmobranch species are relatively unknown. 1,2,6,7,9,13,15 One retrospective study included suspected herpesviral and adenoviral particles in skin and gill lesions, respectively, although presumed viral diseases comprised less than 1% of 1546 cases. 9
Previous reports of viral infections in elasmobranchs.

Herpesviruses are a diverse group of enveloped, double-stranded DNA viruses, with large complex genomes, and occur in animals from bivalves to humans. Collectively, they share biological and structural features including similar replicative strategies and icosahedral nucleocapsids of approximately 100 nm surrounded by a proteinaceous tegument and a lipid envelope. 16,17 However, sequence similarities between the 3 herpesviral families are limited to several core genes, particularly the terminase gene involved in viral packaging. 3 –5,11 The family Alloherpesviridae, composed of 2 monophyletic clades, contains host-specific pathogens that cause significant disease in cyprinid, anguillid, ictalurid, acipenserid, salmonid, gadid, clupeid, percid, and cichlid fish. 8,10,11 Notable examples include ictalurid herpesvirus 1 (channel catfish virus) and cyprinid herpesvirus 3 (koi herpesvirus). 10 Like mammalian herpesviruses, the alloherpesviruses of fish produce variable disease signs and losses, some severe, and develop latency and recrudescence. 11 Typical lesions include skin and gill epithelial proliferation, hemorrhages, ascites, and renal and hepatic necrosis. 10
Tiger sharks (Galeocerdo cuvier) are iconic top predators that play a vital role in ecosystem function and ecotourism. No viruses and only a few infectious diseases have been reported in tiger sharks, including mycotic dermatitis and branchitis. 9 This report describes gross, histopathologic, transmission electron microscopic (TEM), in situ hybridization (ISH), and initial molecular findings associated with necrotizing branchitis in a tiger shark caused by a novel herpes-like virus.
A wild-caught, 5.8 kg, 120 cm long, juvenile, male tiger shark was collected off the Florida Keys, Florida, in December 2018. The shark was transferred within 4 hours of capture to a 40,000-gallon holding tank (17.4 × 6.7 × 1.5 m), which contained 2 captive-born sandbar sharks (Carcharhinus plumbeus). The diet offered consisted of a variety of proteins, including salmon, mackerel, tuna, beef steak, and duck, via broadcast feeding. Approximately 9 days after collection, the shark had progressively decreased activity and appetite, accompanied by rostral abrasions and clinical signs of respiratory distress including spyhopping and gill flaring. Due to poor quality of life, euthanasia was elected 6 days after the onset of signs, using 2-phenoxyethanol via immersion in a 300-gallon holding tank. The specimen was maintained overnight on ice, and a necropsy was performed the following morning. Gross lesions were limited to a few skin abrasions on the ventral mandible and pectoral girdle, minimal serosanguinous coelomic fluid, and a small dark-brown liver indicative of atrophy. 9
Tissues were received by the Aquatic Pathology Service, University of Georgia College of Veterinary Medicine, and fixed in 10% neutral-buffered formalin. Representative sections of the brain, epigonal organ, esophagus, gill, heart, intestine, kidney, Leydig organ, liver, pancreas, rectal gland, skeletal muscle, skin, spleen, stomach, and testicle were routinely processed; sectioned at 5 µm; and stained with hematoxylin and eosin. Select sections were stained with Fuelgen stain to highlight DNA within viral inclusions and Fraser-Lendrum and phosphotungstic acid hematoxylin (PTAH) stains to highlight fibrin.
When compared with normal tiger shark gill structure (Fig. 1), histologic examination revealed an acute, multifocal, necrotizing branchitis, affecting up to 25% of lamellae, accompanied by scattered lamellar thrombi. Individual and small clusters of lamellar epithelial cells were moderately to severely enlarged. Cells with increased cytoplasmic volume occasionally had cytoplasmic inclusions markedly distorting their morphology and peripheralizing nuclei. Enlarged nuclei frequently contained central inclusions surrounded by less intensely stained marginated chromatin (Figs. 2, 3). The amphophilic to hyperbasophilic, finely granular, cytoplasmic, and nuclear inclusions both stained Fuelgen-positive, indicating DNA presence (Fig. 3). Potentially confounded by mild autolysis, affected epithelial cells often sloughed from lamellae. Scattered fibrin thrombi segmentally occluded and expanded lamellar capillaries (Fig. 2). Additional lamellae were segmentally necrotic, with loss of tinctorial properties and morphologic detail, nuclear pyknosis, and cellular fragmentation. Foci with thrombosis and necrosis were frequently associated with an absence of lamellar epithelium. Fine eosinophilic granulocytes often circulated in lamellar capillary lumens, but no additional inflammatory changes were present. Scattered superficial esophageal mucosal cells contained large nuclear and cytoplasmic inclusions with morphologic and staining properties as described in the gills (Fig. 4). Additional microscopic findings were limited to the liver and stomach. Hepatocytes contained less cytoplasmic lipid than healthy elasmobranch livers, consistent with mild hepatocellular atrophy and gross findings, suggesting the animal may not have been feeding well post-capture. Incidentally, the fundic stomach contained fragments of a cestode within the submucosa.
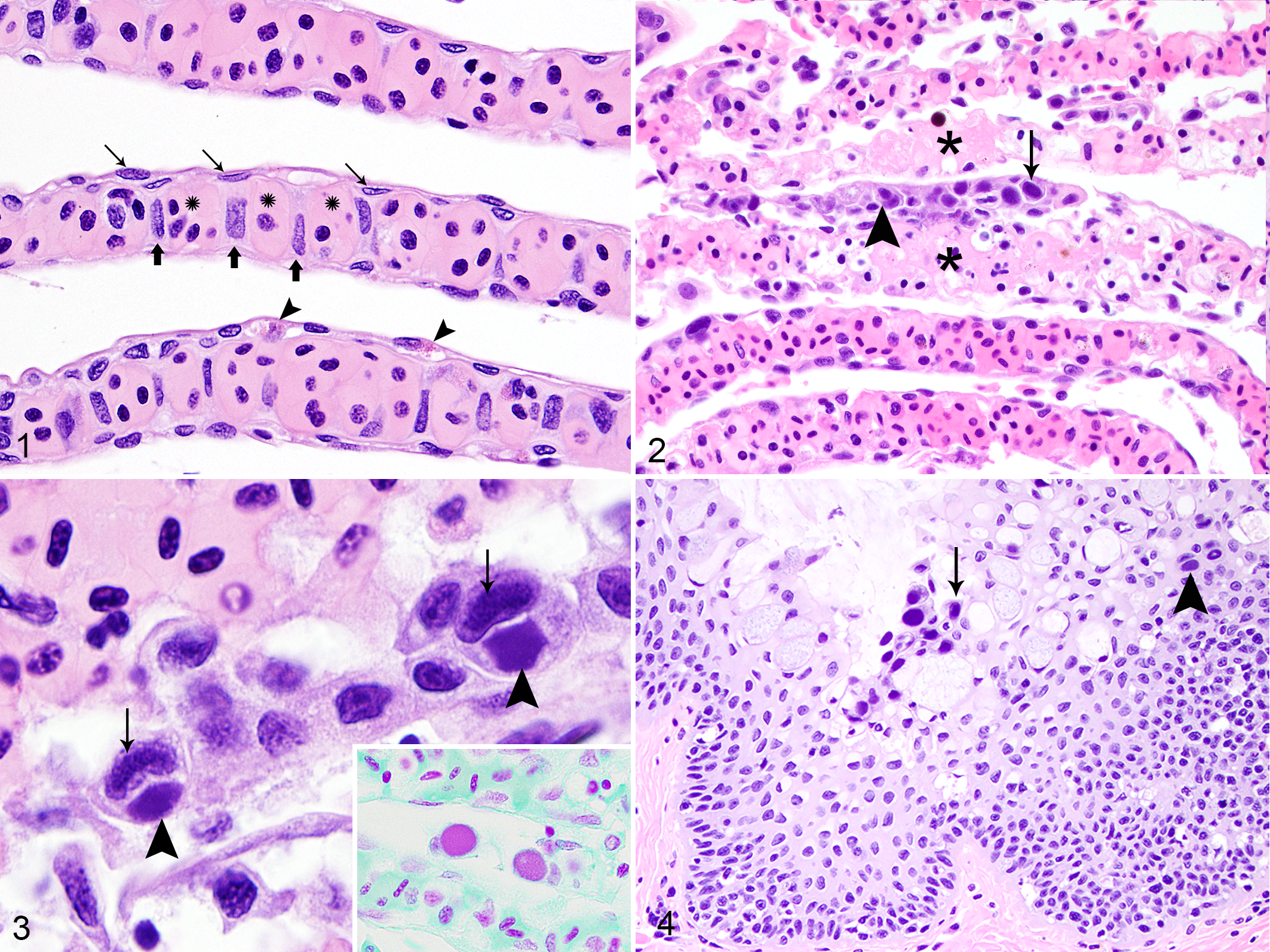
Gill, tiger shark.
Formalin-fixed gill lamellae were routinely processed for TEM after transferring overnight to a solution of 2% glutaraldehyde and 2% paraformaldehyde in phosphate buffer. Ultrathin sections, stained with lead citrate and examined on a transmission electron microscope, revealed lamellar epithelial cells with small numbers of intranuclear, 84 ± 8 nm (mean ± standard deviation; n = 20) icosahedral nucleocapsids with an electron-dense core. Large arrays of complete, 148 ± 11 nm (n = 50), enveloped viral particles were present within membrane-bound cytoplasmic vesicles (Figs. 5, 6).
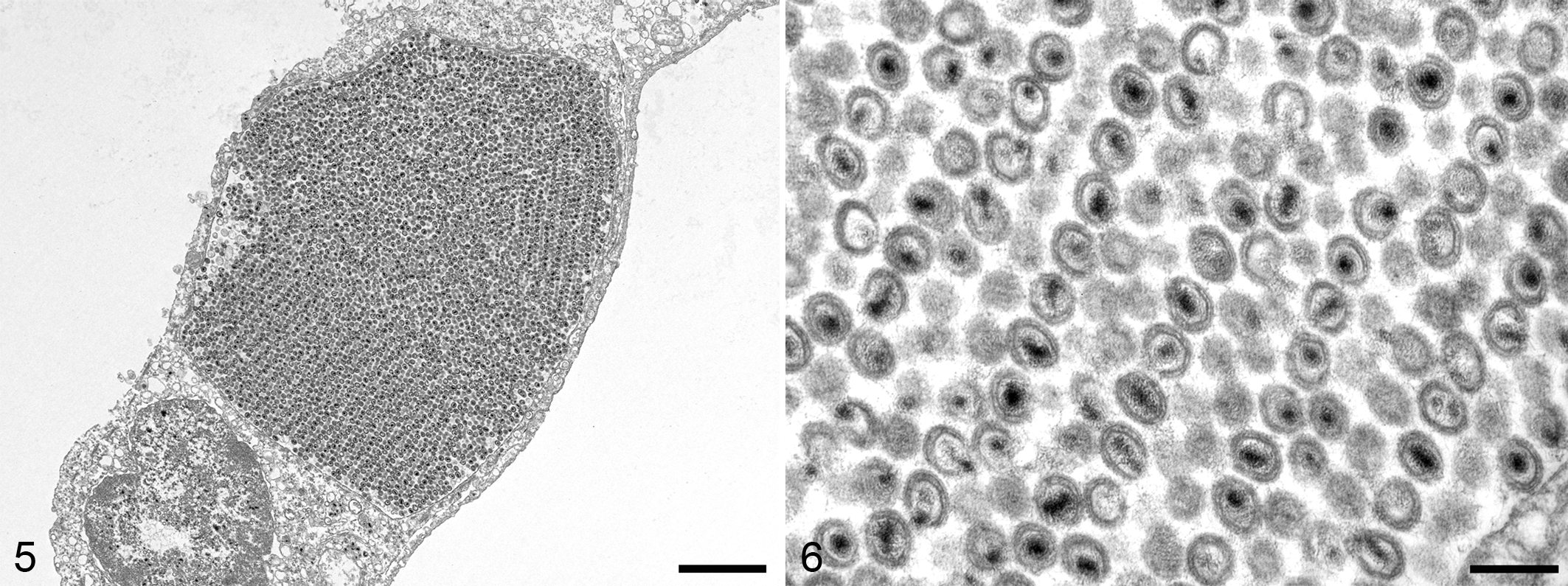
Herpesvirus infection, gill, tiger shark.
Total DNA was isolated from formalin-fixed gills using a QIAamp DNA Mini Kit (QIAGEN). Based on particle size and morphology, a herpesvirus was suspected. Conventional polymerase chain reaction (PCR) protocols to identify the viral polymerase gene were attempted but produced no successful amplification. 12,20 A next-generation sequencing (NGS) library was then generated using a Nextera XT DNA Library Preparation Kit (Illumina Inc) and sequenced using Illumina HiSeq with 2 × 150-bp paired-end sequencing reagents (Genewiz). Over 0.8 billion reads were assembled into contigs using SPAdes 18 and CLC genomics workbench (QIAGEN) followed by a protein homology search using GHOSTX 19 with an in-house database curated from all proteins in the GenBank non-redundant database. A partial genome of ∼240,000 bp was assembled, which contained core herpesvirus genes including the DNA polymerase, major capsid protein, and terminase. Sequences for these core genes are available in Genbank under accession numbers MZ381455, MZ381456, and MZ381454, respectively. The polymerase shared 24% to 31% predicted protein homology with other known fish alloherpesviruses (BLASTp, Query coverage >50%, e-value <10−50), indicating a divergent herpes-like virus. The draft genome sequence was used for quantitative PCR (qPCR) and ISH assay development. Further genome assembly is ongoing.
A Taqman hydrolysis probe-based qPCR assay specific to the divergent virus was designed to quantify viral loads in formalin-fixed gills using PCR primers directed against a region of the polymerase gene (SHV3668F: 5′-AAAATACGCGTCGCATCACG-3′; SHV3772R: 5′-TTTCTGTTTCTGGTTGCGCG-3′) and a hydrolysis probe (5′-/56-FAM/TGCCACTAC/ZEN/GACAAAAACGAAACGC/3IABkFQ/-3′). Using endpoint PCR, the reaction generated a 105-bp amplicon identical to that produced by the earlier metagenomic sequencing. The amplicon was gel-purified, quantified using a Qubit fluorometer (Thermofischer Scientific Inc), and Sanger-sequenced to verify sequence identity. Serial 10-fold dilutions from 1:10 to 1:100,000,000 were prepared for standard curve generation. All samples and standards were run in triplicate. The qPCR assay quantified 8.8 ± 0.5 × 102 viral copies/mg of tissue in the DNA originally submitted for NGS. An additional DNA preparation used in qPCR assay development had similar viral DNA levels (1.18 ± 0.05 × 103 copies/mg of tissue).
An anti-sense ISH probe (V-SHV-MCP-POL, Catalog No. 887731, RNAscope 2.5, Advanced Cell Diagnostics Inc) targeted the viral mRNA (sense strand) and DNA. The probes were designed against the polymerase (20 ZZ combinations, nucleotides 1700–2626) and major capsid protein (20 ZZ combinations, nucleotides 2114–3062) gene sequences, avoiding binding to known related herpesviruses identified by BLASTn search of the non-redundant nucleotide database in NCBI GenBank. Stock probe solution was diluted by half concentration using RNAscope probe diluent (Catalog No. 300041, Advanced Cell Diagnostics Inc). The chromogenic assay (RNAscope 2.5 HD Detection Kit [RED] User Manual, Part 2, Catalog No. 322360-USM, Advanced Cell Diagnostics Inc) was performed according to the manufacturer’s protocols on infected gill and esophagus, along with spleen, stomach, intestine, and pancreas, with a target retrieval time of 15 minutes. Tissues from previous tiger shark submissions were used as host negative controls. Adipose fin of channel catfish infected with Henneguya adiposa and specific RNAscope ISH probe (H.adiposa-18S, Catalog No. 566921, Advanced Cell Diagnostics Inc) were used as a positive control for assay performance. The ISH positively labeled intranuclear and intracytoplasmic viral inclusions in lamellar and esophageal epithelial cells (Figs. 7, 8). There was no positive labeling in the spleen, stomach, intestine, or pancreas, and no cross-reactivity with host tissues.
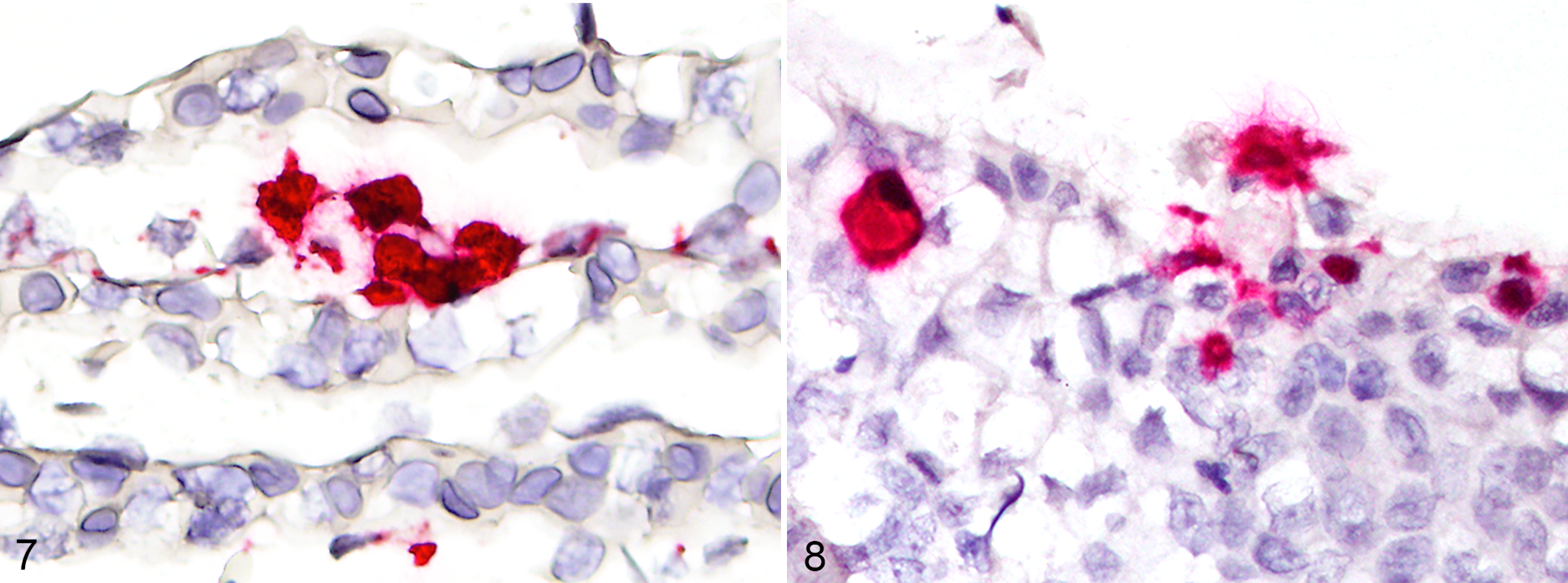
Herpesvirus infection, gill, tiger shark. Multiple lamellar epithelial cells positively label with the novel herpesvirus-like polymerase and major capsid protein probe. In situ hybridization (ISH), red chromogen, hematoxylin counterstain.
Consistent with the epitheliotropic nature of many herpesviruses, lesions in this case and the suspected herpesviral dermatitis of dogfish involved epithelial surfaces. 9,15 In the dogfish, skin lesions consisted of basal epidermal cell degeneration, necrosis, vesiculation, and ulceration, accompanied by eosinophilic intranuclear and intracytoplasmic inclusion bodies. 15 However, no molecular analysis was performed on the dogfish virus, and the diagnoses were based on ultrastructural particle morphology. Lesions were observed over 2.5 years in both newly captured and aquarium-housed sharks, although incidence appeared to increase in captive specimens over time. That report provided no discussion of lesion resolution or impacts on mortality. 15
In contrast to the dogfish study, the tiger shark in the current report had moderate and minimal lesions of gill respiratory epithelium and esophageal mucosa, respectively. Antemortem observation of respiratory distress supports the significance of histologic changes, suggesting that viral branchial lesions likely contributed to the shark’s decline. The viral source, timing of infection, and transmission route are unknown. Similarly, it is unclear whether the lesions represent a newly acquired infection or reactivation of a latent infection. With either possibility, immune compromise induced by various stressors could have influenced the disease onset. In bony fish, primary herpesviral infections often result in mild disease and dissemination in a population, while stressful conditions can promote severe disease and high mortalities. 10 Although the significance to wild tiger shark populations is unknown, infections introduced from latently infected sharks could pose a risk to susceptible naïve specimens under human care. In this case, the co-housed sandbar sharks remained unaffected.
Reports of viral disease in cartilaginous fish are rare, although they are expected to increase as interest in elasmobranch conservation, research, and public display grows and environmental pressures on wild populations increase. Due to a lack of elasmobranch cell lines and fresh tissue for use in virus isolation, recognition and characterization in this case relied upon metagenomic sequencing following histologic and ultrastructural evaluation. Even with metagenomic sequencing, detection of divergent viruses is difficult, as sequence recognition and alignments are dependent on existing data in public databases. 14 This is the first report of a molecularly identified herpes-like virus in an elasmobranch species, the tiger shark. The diagnosis was supported by histologic and TEM findings as well as qPCR and ISH assays based on NGS findings from formalin-fixed tissue.
Footnotes
Acknowledgements
The authors thank Mary Ard for tissue preparation for transmission electron microscopy, Dr. Natalie Stilwell for assistance with RNAscope, and the Athens Veterinary Diagnostic Laboratory Serology and Virology Department for providing the generic herpesviral qPCR primers.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
