Abstract
c-Kit mutations have been reported in 15% to 40% of certain human melanoma subtypes, including those histologically similar to canine oral malignant melanomas. Therapeutic response to tyrosine kinase inhibitors has been demonstrated in those human patients. As canine oral malignant melanomas tend to have a poor prognosis despite aggressive surgical removal, evaluation of KIT expression and identification of c-Kit mutations in canine oral melanocytic neoplasms was performed to determine if there is any indication that tyrosine kinase inhibitor drugs might effectively treat any of these cases. This study evaluated 27 canine oral malignant melanomas and 12 canine histologically well-differentiated oral melanocytic neoplasms for activating c-Kit mutations, determined differences in immunohistochemical expression of KIT and c-Kit mutation status, and determined if KIT expression could predict c-Kit mutation status. Among samples that contained intraepithelial nests of neoplastic melanocytes in the KIT-labeled sections, KIT was expressed within cells in these nests in 22/23 (96%) malignant melanomas and 5/7 histologically well-differentiated neoplasms. KIT was expressed in 10% to 30% of neoplastic melanocytes in the lamina propria in 3/24 (13%) malignant melanomas, but 0/9 (0%) histologically well-differentiated neoplasms. Next-generation sequencing identified 85 variants in c-Kit, including 9 nonsynonymous mutations that resulted in amino acid changes predicted to affect protein function. c-Kit mutations with predicted deleterious protein effects were more common in malignant melanomas (8/27 [30%] vs 1/12 [8%]). There was no apparent relationship between detected c-Kit mutations and KIT expression. These results do not support the use of therapies that target c-Kit.
Keywords
Malignant melanoma (MM) is the most common form of canine oral malignancy across all breeds. Canine oral MMs are locally invasive with a high degree of metastatic potential and a poor prognosis for long-term survival, similar to human malignant melanomas. They also have similar morphologic features as human mucosal and acral lentiginous malignant melanomas, the most notable being prominent lentiginous spread. Other common features include ulceration, an extensive vertical growth phase, and frequent lymph node metastasis. 8,9,44,47,53,55 Historically, it had been thought that all canine oral melanocytic neoplasms were locally invasive with a high degree of metastatic potential and a poor prognosis for long-term survival. However, recently, a subset of canine oral melanocytic neoplasms has been identified that exhibit more benign biological behavior with a low rate of recurrence and metastasis and a favorable overall prognosis for survival. 7,17,56 These are referred to as histologically well-differentiated oral/lip melanocytic neoplasms (HWDMs), 17 or canine oral melanocytic neoplasms of low malignant potential, 53 and they resemble blue nevi in humans. 11,53 HWDMs are generally <2 cm in diameter, heavily pigmented, lack cellular atypia, have rare mitoses, and are rarely ulcerated. 17,55
While morphologic and immunohistochemical (IHC) classification of canine oral melanocytic neoplasms has identified specific phenotypes, there are very few studies that have examined genetic features of these neoplasms as a means of classification. A recent trend in human medicine is to integrate both morphologic and genetic features in order to improve melanoma classification. In several studies, epidemiological, clinical, histopathological, genetic, and biological aspects of human melanocytic neoplasms have been shown to support distinct subtypes, causal pathways, and cells of origin. 4,10,14,60,62 This improved classification is useful for determining appropriate targeted therapies for patients and may assist in the design and interpretation of clinical trials. 10
The oncogene c-KIT codes for KIT, which is a transmembrane tyrosine kinase receptor found in hematopoietic cells, mast cells, germ cells, interstitial cells of Cajal, and melanocytes. 1 In melanocytes, c-KIT is involved in regulating pigmentation, migration, differentiation, survival, and proliferation. 1,22,23 Tyrosine kinase inhibitors (TKIs) target protein kinases, such as c-KIT, by blocking their ATP-binding site. The regulation of the KIT pathway is highly intricate and dependent on several factors. 1,22,23 Different patterns of expression of KIT have been reported for different subtypes of melanocytic neoplasms in humans, 45 and it has been noted that overexpression of KIT, or mutations within the gene, are more common in mucosal than cutaneous melanomas. 51 Certain subtypes of human melanocytic neoplasms with more intense immunolabeling for KIT have also been shown to be more likely to have a mutation in the c-KIT gene, generally in exon 11. 2,51 Other studies have shown that overexpression of KIT is more common in benign or less aggressive forms of human melanocytic neoplasms than in aggressive forms. 38 c-KIT mutations have been identified in 15% to 40% of 3 subtypes of human melanomas, these being mucosal, acral, and chronic sun-induced melanomas, and a response to TKI therapy has been demonstrated in a significant proportion of these patients. 2,6,15,25 –28,30,49,52,58 However, the role of c-KIT in human melanocytic neoplasms appears to be highly variable depending on the subtype of melanocytic neoplasm, and different types of c-KIT mutations appear to have opposite effects on melanocyte proliferation and response to TKI therapy. In fact, it is possible that TKI therapy may be detrimental in certain types of human melanocytic neoplasms. 5,54 Some KIT inhibitors may have inhibitory effects on RAF kinases, which could lead to activation of the MAPK pathway and thereby accelerate the growth of RAS-driven tumors. 5 Furthermore, it has been shown that KIT also has tumor-suppressive functions and inhibition of KIT expression may lead to accelerated progression of early-stage cutaneous melanomas. 54
In dogs, gain-of-function c-Kit mutations have been identified in mast cell tumors, leukemias, and gastrointestinal stromal tumors. 21,31,34,59,61 A few studies have evaluated canine melanocytic neoplasms for c-Kit mutations in a limited number of cases, 13,20,39 but none have performed c-Kit exome sequencing. KIT immunolabeling has also been reported in a proportion of canine melanocytic neoplasms. 13,19,20,39,40,53 However, very little is yet known about the role of c-Kit in canine melanocytic neoplasms and no significant association between KIT labeling and survival time has been demonstrated.
Data collected from our institution suggest that patients with a high Ki67 index have a very guarded prognosis regardless of adjuvant therapies. 43 Despite the current lack of evidence for c-Kit mutations in canine melanomas, recently, veterinary oncologists have begun using TKIs to treat dogs with oral MM. Use of such drugs may be more for their antiangiogenic abilities rather than for c-Kit inhibition. However, to our knowledge, no published studies have evaluated their effectiveness for these tumors. Masitinib 16 (Kinavet, AB Science) and toceranib (Palladia, Zoetis) are TKIs that have been approved for use in dogs, have few clinical side effects, and are generally tolerated well. They have been shown to be effective in treating canine mast cell tumors with a c-Kit mutation, as well as for treating a proportion of mast cell tumors without a c-Kit mutation. 25,33,35,48 Masitinib, however, is not available in the United States. Because canine and human mucosal/acral MMs have similar morphology and clinical progression, and do not respond well to chemotherapy and radiation treatment, 8,9,44,47,53 further evaluation of KIT expression, c-Kit mutation status, and response to TKI therapy in canine oral MM patients is warranted to determine the likelihood of therapeutic response to c-Kit inhibitors.
The aims of this study were to screen canine oral melanocytic neoplasms for activating c-Kit mutations using next-generation sequencing, to confirm any c-Kit mutations by Sanger sequencing, to determine differences in KIT expression and c-Kit mutation status between 2 subtypes of canine oral melanocytic neoplasms, and to determine if KIT expression can be used to predict c-Kit mutation status.
Materials and Methods
Case Material
Thirty-nine canine oral melanocytic neoplasms that had been submitted as formalin-fixed tissue for routine biopsy were selected from the Michigan State University Veterinary Diagnostic Laboratory clinical database. Cases were selected if tissue blocks were available and if it was deemed there was enough tissue in the block to perform all necessary tests. Cases for which immunohistochemical labeling for Ki67 had already been performed and for which slides were still available were preferentially selected. Archived case material consisted of neoplasms that had been fixed in 10% neutral buffered formalin, embedded in paraffin wax, and previously diagnosed as melanocytic neoplasms. Signalment and location of the neoplasm were recorded from archived reports. Survival and disease-free interval data were not collected for this study, as this was not the aim. Due to the retrospective nature of the study, and how cases were selected, complete and useful survival data would not have been possible to obtain, especially since treatment protocols likely varied greatly between patients.
Histological Assessment
Five-micrometer sections were cut from paraffin blocks containing sections of each neoplasm and stained with hematoxylin and eosin (HE) for confirmation of the diagnosis and histological assessment. If needed for better discernment of nuclei, sections were bleached to remove melanin pigment prior to staining with HE. Microscopic characteristics of each neoplasm were described by a board-certified pathologist (RCS) and included cellular morphology (round or epithelioid, spindloid, or mixed); ulceration (present or absent); intraepithelial nests (present or absent); junctional activity (present or absent); lentiginous spread (present or absent); depth of invasion (superficial lamina propria, deep lamina propria, skeletal muscle, bone invasion); lymphatic invasion (present or absent); degree of pigmentation (less than 50% or greater than 50% of tumor cells contain melanin); nuclear atypia (defined below); mitoses in 10 high-power (40×) fields (hpf; 2.37 mm2); necrosis (present or absent); and inflammation (present or absent). Nuclear atypia was defined by Spangler and Kass, 56 and each neoplasm was scored as either having low (<30% of cells with poorly differentiated nuclei) or high (≥30% of cells with poorly differentiated nuclei) nuclear atypia according to Bergin et al. 7
IHC for Ki67
Each neoplasm was immunohistochemically labeled for Ki67, as previously described by Bergin et al. 7 Ki67 immunolabeling was performed on the Benchmark Automated Staining system (Ventana Medical Systems, Inc) following heat-induced epitope retrieval. Briefly, a mouse monoclonal anti-Ki67 antibody (MIB-1, Dako Cytomation) at dilution 1:50 was applied and detected using the Enhanced V-Red detection system (Ventana), which utilized alkaline phosphatase and the chromagen AEC. Positive controls included canine lymph nodes and mast cell tumors. Evaluation of Ki67 immunolabeling was performed by a board-certified pathologist (RCS) without knowledge of the case outcome and following a previously described method. 7 A neoplasm was diagnosed as an oral MM if 2 of the 3 following criteria were met: a high Ki67 index (≥19.5 positive nuclei per 1 cm2 grid reticle at 40× 6,7 ); high nuclear atypia (≥30% atypical nuclei 7,55,56 ); and a high mitotic count (≥4/10 hpf 7,55 ); otherwise, it was diagnosed as a HWDM.
IHC for KIT
Each neoplasm was immunohistochemically labeled for KIT protein. KIT immunolabeling was performed using a Dako Autostainer (Dako). Antigen retrieval was performed with HIER using a high pH cc1 for 30 minutes. Sections were incubated with rabbit polyclonal anti-KIT antibodies (Dako Cytomation) at a 1:300 dilution for 32 minutes. An enhanced V-red kit detection system with hematoxylin counterstain was used. Canine mast cell tumors were used as positive controls.
Cytoplasmic labeling of both intraepithelial and lamina proprial neoplastic cells was evaluated separately. Labeling of intraepithelial nests of neoplastic cells was recorded as positive or negative. The percentage of positive lamina proprial cells was recorded as 0 (negative), <10% of cells labeling, 10% to 30% cells labeling, 30% to 75% cells labeling, and >75% cells labeling. The lamina proprial portion of the neoplasm was considered positive for KIT if greater than 10% of the neoplastic cells in the lamina propria showed cytoplasmic labeling.
Sequencing of the c-KIT Gene
Sequencing was performed for each neoplasm and for normal tissue adjacent to 13 of the MMs for which enough normal tissue was present. Areas containing viable neoplasms (39 cases) and areas containing normal adjacent tissue (13 cases; cases 15–27) were marked on the HE-stained slides and then matched to the paraffin-embedded blocks.
Tumor area and nontumor area (for 13 of the MMs) on the paraffin-embedded block were macrodissected with a scalpel, deparaffinized with 1 ml FisherBrand CitriSolv Clearing Agent (Fisher Scientific), and washed twice in 1 ml of 100% ethanol prior to genomic DNA extraction using the RecoverAll Kit (Thermo Fisher Scientific, Inc) following the manufacturer’s protocol. The quantity of the extracted DNA was determined using a Qubit 2.0 fluorometer (Thermo Fisher Scientific, Inc). PCR amplifiability of DNA samples extracted from formalin-fixed, paraffin-embedded tissue was determined using a primer set for beta-2 microglobulin housekeeping gene with 221 bp amplicon.
Sequencing of all exons of the c-KIT gene was performed using custom Ampliseq Ion Torrent next-generation sequencing (Thermo Fisher Scientific, Inc). AmpliSeq library samples were prepared using the Ion AmpliSeq Library Kit 2.0 according to the manufacturer’s protocol (Thermo Fisher Scientific, Inc), with some modifications. The preparation was started with 100 to 1000 ng of genomic DNA. The prepared library was quantified using a Qubit 2.0 Fluorometer (Thermo Fisher Scientific, Inc) and a Bioanalyzer high-sensitivity DNA chip (Agilent Technologies Inc). Emulsion PCR (polymerase chain reaction) was carried out to clonally amplify the library on ion sphere particles (ISPs) using the Ion OneTouch 200 system and Template Kit v2.0 (Thermo Fisher Scientific, Inc), according to the manufacturer’s protocols. ISPs were enriched for template-positive ISPs using Dynabeads MyOne Streptavidin C1 beads and Ion OneTouch Enrichment System Instrument (Thermo Fisher Scientific, Inc). Sequencing primer and polymerase were added to the final enriched ISPs prior to loading onto an Ion 318 chip v2. Sequencing was performed on the PGM (Thermo Fisher Scientific, Inc). To sequence an authentic variant, an ideal average coverage for each amplicon of 100× was used in this study.
Data Analysis and Experimental Validation
Sequence data were processed using the Torrent Suite software v5.0.4 (Thermo Fisher Scientific, Inc) for alignment with the canine reference genome (canFam3), base calling, trimming of barcoded adapter sequences, and filtering of poor signal reads. Torrent Variant Caller plugin software v4.0 was used for coverage analysis and variant calling. The variant caller parameter setting was somatic PGM low stringency. Neutral or deleterious variants were determined after analysis using PROVEAN (Protein Variation Effect Analyzer) software tool (http://provean.jcvi.org/index.php). 12
Sanger sequencing was used to confirm the deleterious variants identified by the PGM platform. Primer pairs for the c-Kit gene were designed using Primer3 software. Standard PCR reactions were performed using GoTaq DNA polymerase (Promega). PCR products were visualized on The QIAxcel Capillary Electrophoresis System (Qiagen) to confirm the presence of single product amplification. PCR products were purified using the QIAquick PCR Purification Kit (Qiagen) and further submitted for sequencing using capillary electrophoresis on an ABI PRISM 3130 Genetic Analyzer (Thermo Fisher Scientific, Inc). Sanger sequencing results were aligned to reference normal gene sequence in order to confirm the variants.
Statistical Analysis
Fisher’s exact tests with 2-sided alternative hypotheses were used to assess evidence of a relationship between KIT expression and presence of c-Kit mutations, as well as for any association of these variables with the categorical histologic features. To assess evidence of an association between KIT expression and presence of c-Kit mutations with quantitative histologic features (age, Ki67 index, and mitotic count), Wilcoxon rank sum tests were used. Observations for which at least one of the results for KIT expression, c-Kit mutation presence, or value/status of the histologic parameter was unknown were removed from the respective statistical analysis. A Bonferroni adjustment to an overall error rate of 5% for each group of comparisons was made, such that individual pairwise comparisons were assessed at a 0.05/n level of significance (n = 13, 15, and 17 for KIT intraepithelial pairwise comparisons, KIT submucosal pairwise comparisons, and c-Kit mutation pairwise comparisons, respectively). Association tests were conducted using the combined data from MM and HWDM samples. The data analyzed in this study are available as Supplemental Materials or by request to the authors.
Results
Patient Demographics
Complete demographic data are included in Supplemental Table S1. Dogs with MM ranged in age from 3 to 15 years (mean 11.5 years). Eleven castrated males, 13 spayed females, 1 intact male, and 2 of unknown sex were included. Breeds are specified in Supplemental Table S1. The locations of the MMs were the gingiva (9), the mucosal surface of the lip (8), buccal mucosa (3), soft palate (2), oral cavity not otherwise specified (2), sublingual (1), base of tongue (1), and “lower side of jaw” (1).
Dogs with HWDM ranged in age from 5 to 17 years (mean 9.5 years). There was 1 intact male, 6 castrated males, and 5 spayed females. Breeds are specified in Supplemental Table S1. The most common location of these HWDMs was the gingiva (5 dogs). Other reported locations included lip (3 dogs); buccal mucosa (2 dogs); sublingual (1 dog); and oral cavity not otherwise specified (1 dog).
Histological and Ki67 Index Assessment
Twenty-seven neoplasms (cases 1–27) were classified as MMs based on meeting at least 2 of the 3 following criteria: ≥30% nuclear atypia; ≥4 mitoses in 10 hpf (2.37 mm2); and ≥19.5 Ki67-positive nuclei per 1 cm2 grid reticle at 400×. Twelve neoplasms (cases 28–39) that lacked these features, and were highly pigmented, were classified as HWDMs. MMs were generally poorly pigmented, ulcerated, contained intraepithelial nests of neoplastic cells, and exhibited junctional activity and lentiginous spread (Fig. 1A). Of 27 MMs, 25 (93%) had less than 50% pigmentation, and 19 (70%) exhibited lentiginous spread. HWDMs were often raised, heavily pigmented, nonulcerated, and had abundant collagenous stroma (Fig. 1B). Only 3 out of 12 (25%) HWDMs exhibited lentiginous spread and in 2 of these it was very mild. Complete histology and Ki67 data are included in Supplemental Table S1.
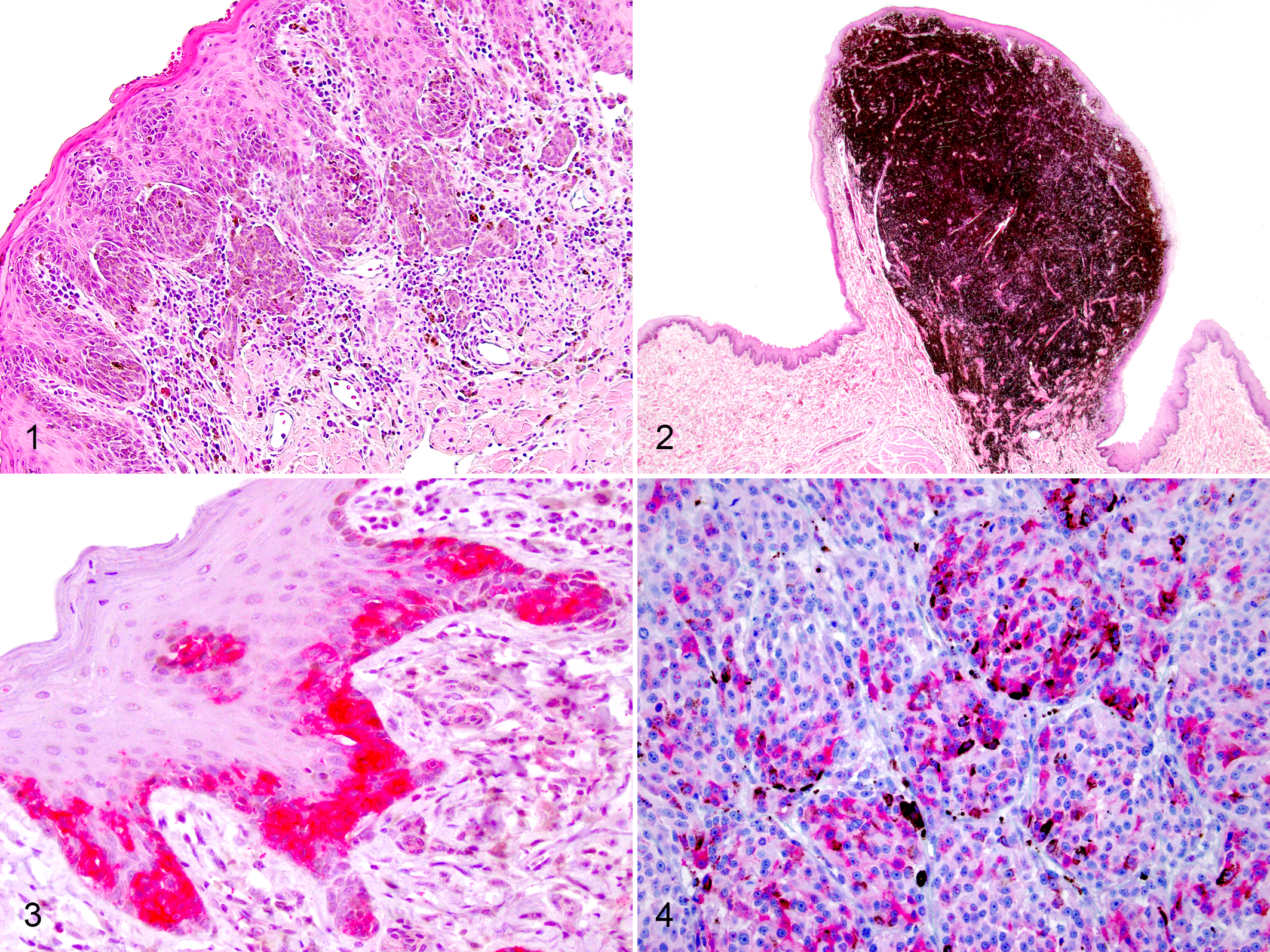
Melanocytic neoplasms, oral tissues, dog. (A) Case 3. An oral malignant melanoma that is poorly pigmented, contains intraepithelial nests of neoplastic cells, and exhibits junctional activity and lentiginous spread. Hematoxylin and eosin (HE). (B) Case 30. A histologically well-differentiated melanocytic neoplasm that is highly pigmented, raised, nonulcerated, <2 cm diameter, and has abundant collagenous stroma. HE. (C, D) Case 13. An oral malignant melanoma with membranous to cytoplasmic immunolabeling for KIT protein that is most frequent in neoplastic melanocytes within the overlying epithelium (C) but occasionally within aggregates of melanocytes in the lamina propria (D). Anti-KIT immunohistochemistry, red chromogen label, hematoxylin counter stain.
Assessment of IHC for KIT
Sections from 23 of the MMs and 7 of the HWDMs contained overlying epithelium and had intraepithelial nests of neoplastic melanocytes within the KIT-labeled sections. KIT protein expression was noted within intraepithelial nests of neoplastic melanocytes in 22 of these 23 (96%) MMs and 5 of these 7 (71%) HWDMs (Fig. 1C). KIT protein expression was noted in 10% to 30% of the submucosal neoplastic melanocytes in 3 of 24 (13%) MMs (Fig. 1D), but none of the 9 HWDMs for which enough tissue was available for analysis. KIT labeling was membranous to cytoplasmic. Complete KIT IHC data are included in Supplemental Table S1.
Next-Generation Sequencing
Using samples from all 39 neoplasms (27 MMs and 12 HWDMs), next-generation sequencing identified 85 variants in c-Kit: 40 were within introns; 21 were synonymous mutations; and 24 were nonsynonymous mutations. Ten of the 24 nonsynonymous mutations (1 each in 8 MMs, 2 in 1 HWDM) resulted in amino acid changes that were predicted to affect protein function by PROVEAN analysis. Thus, 8/27 (30%) of the MMs and 1/12 (8%) HWDMs had c-Kit mutations with predicted deleterious effects on protein function. One of the 8 MMs and one of the HWDMs had the same mutation, K345T (exon 6). The second mutation in this HWDM was G194S (exon 3). Other mutations in the MMs included the following: G865E (exon 19); P168S (exon 3); S400F (exon 7); D287N (exon 5); G663E (exon 14); K785E (exon 16); and M637I (exon 13) (Table 1).
c-KIT Mutations With Deleterious Protein Effect Identified by Next-Generation Sequencing and Correlation With KIT Immunolabeling.
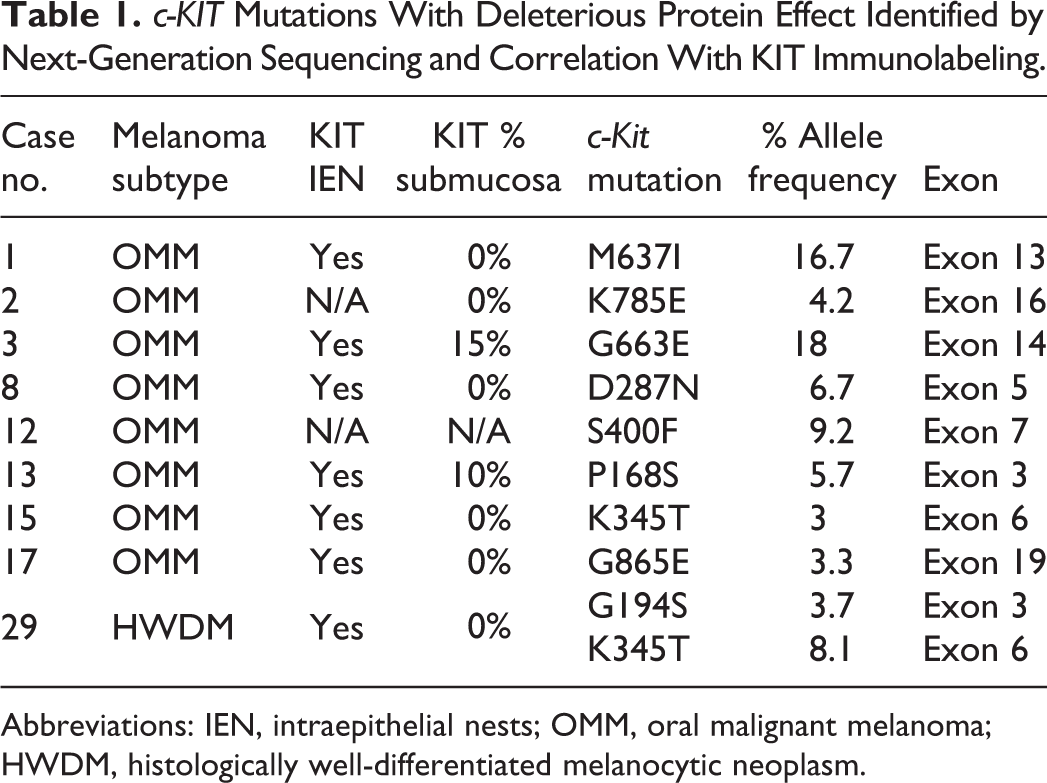
Abbreviations: IEN, intraepithelial nests; OMM, oral malignant melanoma; HWDM, histologically well-differentiated melanocytic neoplasm.
In addition, next-generation sequencing was performed using normal tissue collected from areas adjacent to 13 of the 27 MMs (cases 15–27; Table 2). This identified 1 nonsynonymous c-Kit mutation in each of 2 MMs that were not present in the adjacent normal tissue, and 1 nonsynonymous mutation that was present in the normal tissue but not the adjacent neoplasm.
Number of c-Kit Sequence Variants Found in 13 Canine Oral Malignant Melanomas and Adjacent Normal Tissue.
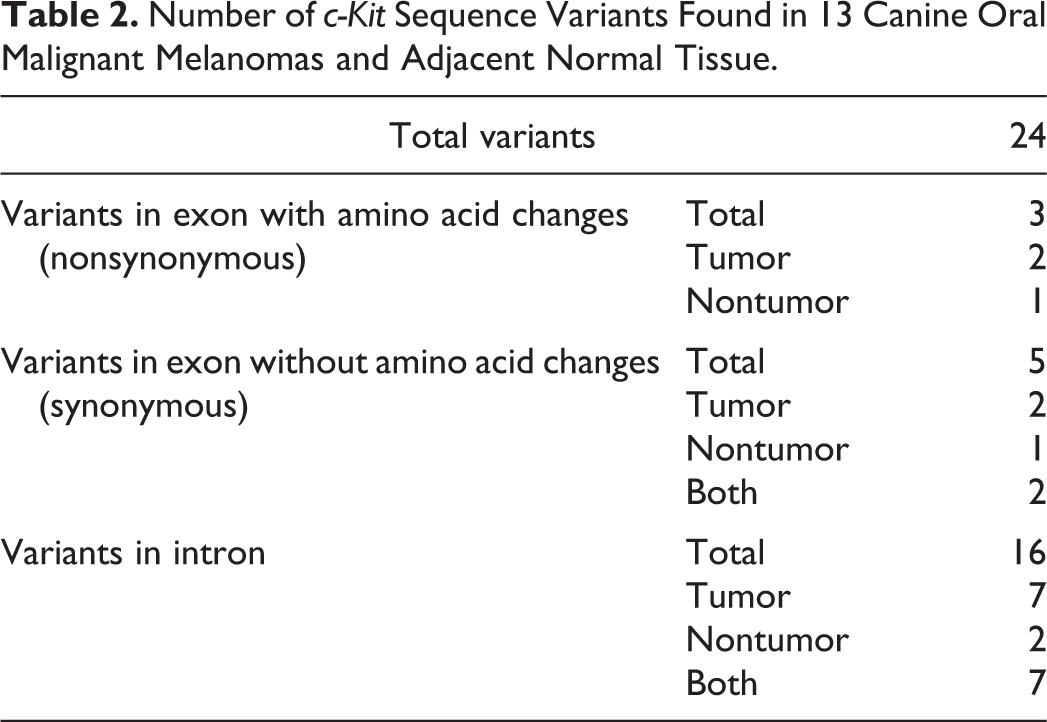

Sanger Sequencing
Sanger sequencing was performed for the 9 neoplasms that were shown to have deleterious mutations. Wild-type c-Kit was amplified in all specimens, and none of the mutations could be confirmed by Sanger sequencing.
Statistical Analysis
Two of the 3 MMs that had KIT labeling in at least 10% of the lamina proprial melanocytes had a deleterious mutation in c-Kit (Table 1), but there was no apparent relationship between c-Kit mutation status and KIT expression or any particular histologic feature based on the results of Fisher’s exact tests (categorical variable comparison) and Wilcoxon rank sum tests (quantitative variable comparison), with a Bonferroni-adjusted level of significance.
Discussion
The current study used whole exome sequencing of c-Kit in canine melanocytic neoplasms. Very few studies have investigated the role of c-Kit in canine melanocytic neoplasms. One study sequenced c-Kit using mRNA in 2 canine melanomas, 20 and a few studies have used PCR to attempt to identify mutations in exon 11. 13,39 Most reported mutations are synonymous single nucleotide polymorphisms that are of no significance to tumorigenesis. Only 2 missense, or nonsynonymous, mutations (L579P in exon 11; G442D in exon 8) have been identified in 2 separate studies, each in a single melanoma. 13,20 These 2 mutations were also identified by Sanger sequencing, indicating that the mutations were present in a significant number of neoplastic cells. One of these mutations (L579P) 13 was an activating mutation that is equivalent to human KIT L576P, the most common activating KIT mutation found in human melanomas. 2,6,32,63
In our study, 9 predicted deleterious c-Kit mutations were identified. A K345T (exon 6) mutation was present in 1 MM and 1 HWDM. A second mutation in the same HWDM neoplasm was a G194S (exon 3) mutation. One mutation each was found in 7 other MMs: G865E (exon 19); P168S (exon 3); S400F (exon 7); D287N (exon 5); G663E (exon 14); K785E (exon 16); and M637I (exon 13). To the authors’ knowledge, none of these mutations have been identified previously in canine or human melanocytic neoplasms. In human melanocytic neoplasms, the most common site of c-KIT mutations is in exon 11, but mutations in other exons, such as exons 13, 17, and 18, have also been reported. 3,6 In the current study, no mutations were found in exons 11, 17, or 18, and only one mutation was found in exon 13.
In addition, next-generation sequencing was performed on both the neoplasm and adjacent histologically normal tissue for 13 of the 27 MMs in order to determine if mutations within the neoplasms were spontaneous or not. Two nonsynonymous mutations (K345T and G865E) were identified in 2 of these MMs and these mutations were not present in the adjacent tissue, indicating that these mutations were likely spontaneous mutations that occurred within the neoplasms. One nonsynonymous mutation was only identified in the normal tissue of one dog, but this mutation was a neutral mutation that did not result in a change in predicted protein function and likely was a spontaneous mutation. Sanger sequencing failed to confirm the identified deleterious mutations in any of the neoplasms in this study, which indicates that there was a low frequency of these mutations within the neoplasms. Thus, most likely, these mutations are not driver mutations. However, it is still possible that these mutations play some role in the differentiation or progression of these neoplasms and could be responsible for acquiring a more aggressive phenotype.
Based on the current genetic data, there is no indication for generalized use of drugs that specifically target c-Kit mutations in canine melanocytic neoplasms. TKI drugs that target multiple tyrosine kinase receptors may still provide some benefit if other receptors are present in canine melanocytic neoplasms. However, at this time, these other receptors have not been investigated in these neoplasms. Future studies to evaluate for the expression of other tyrosine kinase receptors, such as platelet-derived growth factor receptor, vascular endothelial growth factor receptor, and so on, may be helpful.
Meaningful comparative analysis of KIT expression patterns and mutational status requires accurate classification of morphologically distinct subtypes of melanocytic neoplasms. We used histologic features in combination with the Ki67 index to classify the neoplasms in the current study, as this approach has been shown to most accurately predict the behavior of canine oral/lip melanocytic neoplasms by distinguishing those neoplasms with a low malignant potential from those with a high malignant potential. 7 In the current study, we identified 27 oral MMs and 12 HWDMs by using this method. Only one previous study compared KIT expression in cutaneous MMs to various histopathologic features (such as cell type, junctional activity, etc) and did not find any association except between labeling intensity and degree of tumor pigmentation. 19 Similarly, in the current study, there was no apparent association between KIT expression and any particular histopathologic feature. The c-Kit mutation status was also not associated with any histologic feature, except that 27% of oral MMs compared to 8% of HWDMs had a predicted deleterious c-Kit mutation.
KIT labeling of intraepithelial nests of neoplastic cells in the current study was similar to that seen in human acral lentiginous and mucosal melanomas where the vast majority of the in situ component had consistent diffuse and strongly positive immunolabeling for KIT regardless of c-KIT mutation status. 58 Thus, KIT labeling of intraepithelial nests of neoplastic cells does not appear to be correlated with the c-Kit mutation status in dogs. Similarly, there was no apparent association between KIT labeling of the neoplastic melanocytes in the lamina propria and the c-Kit mutation status in the current study. One study of human acral lentiginous/mucosal melanomas identified KIT expression in less than 10% of neoplastic melanocytes in the invasive component as a strong negative predictor of c-Kit mutation status, 58 whereas in another study, the c-KIT mutation status did not correlate with KIT expression in various human melanoma subtypes including acral and mucosal melanomas. 6 While abnormal KIT expression has been used in addition to c-Kit mutation status to predict response to TKIs in canine mast cell tumors and certain subtypes of human melanomas, 2,3,5,25,27,30,33,35,46,48,49 the rationale of this approach is based on the hypothesis that abnormal KIT expression may indicate an underlying mutation in an exon that had not been tested for mutations. Our data do not support this hypothesis for canine melanocytic neoplasms. However, one of the TKIs that is licensed for use in dogs, toceranib (Palladia, Zoetis), targets other tyrosine kinase genes besides c-Kit. Thus, additional studies are needed to evaluate the potential effectiveness of TKIs in treating melanomas with expression of KIT or other tyrosine kinases, as there may be upstream effects and other overexpressed tyrosine kinase receptors that could be targeted. Furthermore, increased expression of KIT may be secondary to an upstream activation of stromal cell-derived factor-1α (SDF-1α), the main ligand for KIT, instead of a genetic alteration in c-Kit. Histone deacetylase inhibitors (HDACi) have been shown to decrease c-Kit mRNA levels, total KIT protein, and cell surface KIT expression in human mastocytosis, indicating that epigenetic changes, including DNA methylation and histone modifications, of SDF-1α may have a major impact on KIT expression. 36 Clinical trials using HDACi in KIT expressing canine melanomas should be considered for future studies.
The potential use of KIT expression as a prognostic marker for human and canine melanocytic tumors is still uncertain and may be dependent on the subtype of melanocytic tumor. In human malignant melanomas, both increased expression 2,18,37,50,58 and decreased expression 38,41,45,57 of KIT have been reported and may be related to stage of tumor progression. The location of the labeling (epidermal vs dermal) is also an important factor to consider. Several studies reported a decrease or loss of KIT labeling in the dermal vertical growth phase of human MMs and in metastatic melanomas. 24,29,38,41,42,57 One proposed reason for this is that KIT signaling plays a role in cell differentiation and loss of its expression may be an important step in tumor progression. 2 Six studies 13,19,20,39,40,53 have examined KIT expression in canine melanocytic neoplasms in a limited number of cases, but no significant associations with mutation status or survival have been reported. One study reported that KIT was expressed in 37 of 44 (85%) canine melanomas, which consisted mostly of oral/lip MMs; however, the percentage of positive labeling cells was very low in 27 of the 37 positive cases. 53 In that same study, oral HWDMs had virtual absence of KIT expression. Another study reported that 20/39 (51%) of canine oral MMs had labeling for KIT, but 6 of these neoplasms had labeling in less than 10% of the neoplastic cells, and the location of the positive neoplastic cells was not indicated as intraepithelial or lamina proprial. 39 Chu et al 13 reported that 36/36 (100%) canine MMs, which included 28 oral MMs and 8 MMs from various other sites, had weak or strong KIT labeling, but the location of the positive neoplastic cells was not described. Thus, it is not possible to compare the results of our study with these latter 2 studies. Newman et al 40 reported KIT expression in neoplastic melanocytes in the subepithelial portion of the neoplasms in 6/61 (10%) of malignant mucosal melanocytic neoplasms that consisted of 58 oral MMs and 3 conjunctival MMs. In our study, 3/24 (13%) of the oral MMs had KIT labeling of neoplastic melanocytes in the lamina propria, which is similar to the percentage reported by Newman et al. 40 However, 22/23 (96%) oral MMs in our study had KIT labeling in the in situ component, which is higher than that reported by Newman et al, 40 which described 44/61 (72%) mucosal MMs with KIT labeling in either basilar mucosal melanocytes or junctional neoplastic melanocytes. Gomes et al 19 only examined cutaneous melanocytic neoplasms and found a greater percentage and intensity of KIT expression in dermal melanocytomas compared to cutaneous MMs. Newman et al 40 reported strong and diffuse cytoplasmic labeling for KIT in 14 of 14 dermal melanocytomas and showed that dermal melanocytomas were more likely to label for KIT than were malignant mucosal melanomas; cutaneous MMs and oral HWDM were not evaluated in that study. Thus, it appears that KIT expression in canine melanocytic neoplasms varies considerably.
In summary, this study further demonstrates that spontaneously occurring canine oral melanocytic neoplasms share histopathologic and biological features with human acral lentiginous and mucosal melanomas. We identified 9 deleterious novel c-Kit mutations in a set of canine oral melanocytic neoplasms, none of which were in exon 11. While inability to confirm these mutations with Sanger sequencing indicates that these mutations occurred at a low frequency and that these mutations most likely are not driver mutations, they may be acquired mutations that could be associated with tumor progression or an increasingly aggressive phenotype. There was also no apparent association between KIT labeling and c-Kit mutation status, or any histologic parameter. KIT labeling of intraepithelial nests in canine oral melanocytic neoplasms was similar to human acral lentiginous and mucosal melanomas where >50% of the in situ component was consistently diffusely and strongly positive for KIT, regardless of the c-Kit mutation status and, therefore, not useful for predicting a mutation in c-Kit. Furthermore, 27% of oral MMs compared to 8% of HWDMs had a predicted deleterious c-Kit mutation. While there is no current indication for the generalized use of TKIs to specifically target c-Kit mutations in canine oral MMs, integrating genetic features of canine oral melanocytic neoplasms with morphologic phenotyping may lead to more accurate diagnosis and selection of targeted therapy. More research, ideally in the form of well-designed prospective survival studies, is needed to determine the potential role of other tyrosine kinase receptors as prognostic indicators as well as measures for determining response to TKI therapy for various subtypes of canine melanocytic neoplasms.
Supplemental Material
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211009784 - Correlation Between KIT Expression and c-Kit Mutations in 2 Subtypes of Canine Oral Melanocytic Neoplasms
Supplemental Material, sj-pdf-1-vet-10.1177_03009858211009784 for Correlation Between KIT Expression and c-Kit Mutations in 2 Subtypes of Canine Oral Melanocytic Neoplasms by Rebecca C. Smedley, Tuddow Thaiwong, Lorna E. Deeth and Matti Kiupel in Veterinary Pathology
Footnotes
Acknowledgements
We would like to thank the Michigan State University Veterinary Diagnostic Laboratory’s Histopathology Department for processing the histologic samples and for performing immunohistochemical labeling.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
