Abstract
The intestinal microbiota is believed to play a role in the pathogenesis of inflammatory bowel disease in humans and chronic inflammatory enteropathy (CIE) in dogs. While most previous studies have described the gut microbiota using sequencing methods, it is fundamental to assess the spatial distribution of the bacteria for a better understanding of their relationship with the host. The microbiota in the colonic mucosa of 22 dogs with CIE and 11 control dogs was investigated using fluorescence in situ hybridization (FISH) with a universal eubacterial probe (EUB338) and specific probes for select bacterial groups. The number of total bacteria labeled with EUB338 probe was lower within the colonic crypts of dogs with CIE compared to controls. Helicobacter spp. and Akkermansia spp. were decreased on the colonic surface and in the crypts of dogs with CIE. Dogs with CIE had increased number of Escherichia coli/Shigella spp. on the colonic surface and within the crypts compared to control dogs. In conclusion, the bacterial microbiota in the colonic mucosa differed between dogs with and without CIE, with depletion of the crypt bacteria in dogs with CIE. The crypt bacterial species that was intimately associated with the host mucosa in control dogs was composed mainly of Helicobacter spp.
Keywords
The gut microbiota in humans and animals is host-specific but diverse, with variations along the longitudinal axis of the intestine. 17,23,50 The luminal microbiota is associated with the digesta and encompasses the majority of the bacteria in the gut. 54 In the small intestine, the microbiota is predominated by facultative anaerobes 19 that can tolerate the acidic environment that contains oxygen and antimicrobial molecules such as bile acids. The bacterial density increases in the cecum and colon with the most diverse and dense bacterial communities within the body, especially composed of strict anaerobes. 19,30 This higher abundance of bacteria in the lumen of the large intestine is explained by the slower transit, presence of fermentable polysaccharides, and decreased concentration of antimicrobial substances. 18,54
Besides the variation in distribution of facultative anaerobes and strict anaerobes along the intestinal segments, the intestinal compartments (eg, lumen, mucus layer and crypts) also show a distinct bacterial composition, considered as niches. 16 The bacteria associated with the mucus on the mucosal surface and crypts can be defined as the mucosal microbiota. The spatial organization of microorganisms is dependent on the metabolism of the bacteria and the host, the folding of the mucosa in the large intestine, and the organization of the mucus. 7,33 Some organisms are able to penetrate the mucus and colonize the colonic crypts, maintaining direct contact with the host epithelium. 12,37 The environment closer to the epithelium has a higher oxygen level that selects for facultative aerobic taxa such as Proteobacteria and Actinobacteria. 1 A clear differentiation between luminal and mucosa-associated communities has been reported in several species, including humans, 17,62 mice, 57 pigs, 8 cattle, 28 and primates. 61
Humans with inflammatory bowel disease and dogs with chronic inflammatory enteropathy (CIE) have been shown to have both fecal and intestinal dysbiosis. 20,31,55 Similar to inflammatory bowel disease in humans, the cause of canine CIE is likely a combination of genetic predisposition, environmental factors and alterations in the intestinal microbiota that together induce an aberrant immune response in the host. 59 In dogs with CIE, the fecal microbial changes comprise increased abundance of the Proteobacteria, especially Enterobacteriaceae, and decreased representation of the Firmicutes, including Faecalibacterium, Blautia, Turicibacter, and Clostridium cluster XIV species. 31 Metagenomic sequencing methods are widely used to characterize the intestinal microbiota in dogs. 31,51 These tools generate information about the relative abundance of bacterial taxa present in the feces and intestinal lumen. In order to study the bacterial populations in situ, it is crucial to utilize techniques such as fluorescence in situ hybridization (FISH) that preserve the mucosal biofilm, thereby providing an opportunity for spatial analysis of bacterial load and diversity. 5 Few studies have investigated the in situ microbiota in dogs with CIE. 9,60 Cassmann et al described the mucosal bacteria in the ileum and colon of dogs with CIE, reporting increases in bacteria belonging to the Clostridium coccoides/Eubacterium rectale group, Bacteroides spp., Enterobacteriaceae, and E. coli compared to control dogs. 9 The objective of this study was to characterize and describe the bacteria in the colonic mucosa of control dogs and dogs with CIE using FISH to target bacterial groups that have been demonstrated to be altered in dogs with CIE and human patients with inflammatory bowel disease. Additionally, we investigated a possible correlation between the localization and number of bacteria with the canine inflammatory bowel disease activity index (CIBDAI) and severity of colonic histological lesions.
Material and Methods
Ethics Statement
The collection and analysis of samples from dogs with CIE was approved by the Texas A&M University Institutional Animal Care and Use Committee (Animal use protocols 2012-083 and 2015-0069) or by the Finnish National Animal Experiment Board (ESAVI/6973/04.10.03/2011 and ESAVI/10384/04.10.07/2014), for dogs enrolled at Texas A&M University or the University of Helsinki, respectively. Written consent was obtained from all owners of dogs. Dogs in the control group were euthanized for reasons unrelated to this study and the bodies were donated for teaching and research purposes. The post-euthanasia collection of samples was exempted by the Texas A&M University Institutional Animal Care and Use Committee.
Animal Population and Samples
The control group comprised 11 adult dogs without a history of gastrointestinal disease. The dogs presented to the Texas A&M Veterinary Teaching Hospital with traumatic injuries and were euthanized at the owner’s request. None of the dogs had received antibiotics or immunomodulatory therapy in the preceding 6 months. Full-thickness samples from the gastrointestinal tract, including the ascending, transverse, and distal colon, were collected within 20 minutes of euthanasia, fixed in 10% formalin, routinely processed for histopathology, and stained with hematoxylin and eosin. None of the control dogs had significant gross or microscopic lesions involving the gastrointestinal tract.
The group of dogs with CIE included 22 client-owned dogs with samples retrieved from the archives of the Gastrointestinal Laboratory at Texas A&M University and the Faculty of Veterinary Medicine at University of Helsinki. The inclusion criteria included histological findings of inflammation in colonic biopsy specimens, persistent (>3 weeks’ duration) gastrointestinal signs, and failure to document other causes for gastroenteritis (eg, infectious, parasitic, neoplastic) by thorough diagnostic testing. The diagnostic evaluation in all dogs with CIE consisted of a clinical examination and analysis of clinical history, hematological and serum biochemistry analysis, urinalysis, fecal examination, diagnostic imaging, and histopathological examination of mucosal biopsy specimens. The CIBDAI was determined at presentation by the attending veterinarian, according to the dog’s attitude/activity, appetite, vomiting, fecal consistency, defecation frequency, and weight loss; with the cumulative score ranging from 0 to 18. 25 Most of the dogs with CIE (18/22) had not received antibiotics in the 4 months before sample collection. Four dogs had received antibiotics 3–4 weeks prior sample collection. None of the dogs were treated with immunosuppressive drugs before sampling. Colonic samples used for FISH were collected via colonoscopy. In most dogs, the preparation for colonoscopy included feeding a low-residue diet (boiled cod fish or duck) for 4 days, fasting for 24 hours before the procedure, and an enema with 20 mL/kg of warm water immediately before the procedure. Endoscopically obtained samples of colonic mucosa were fixed in 10% formalin, routinely processed for histopathology, and stained with hematoxylin and eosin. Histological changes in the colon were classified using a grading system for histopathologic criteria (0 = normal, 1 = mild lesions, 2 = moderate lesions, and 3 = severe lesions), which was developed by the World Small Animal Veterinary Association Gastrointestinal Standardization Group. 2,13 Individual scores were assigned for each parameter (ie, surface epithelial injury, crypt dilation, fibrosis, goblet cell count, and lamina propria infiltrate with lymphocytes/plasma cells, eosinophils, neutrophils, and/or macrophages, as individual cellular infiltrates).
FISH and Bacterial Quantification
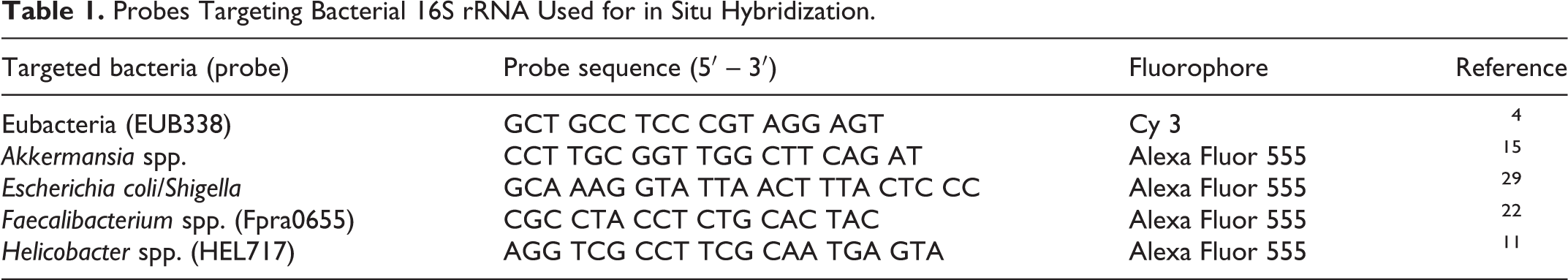
The colonic samples were routinely processed for histology and 4 µm formalin-fixed paraffin-embedded sections were used for FISH. After deparaffinization in xylene and hydration in ethanol, serial sections were individually hybridized with oligo probes diluted to 5 ng/µL in hybridization buffer. The probes (Table 1) targeting the 16 S rRNA gene of bacteria included the universal bacterial probe EUB338 for total bacterial counts, and the specific probes for Helicobacter spp., Escherichia coli/Shigella spp., Faecalibacterium spp., and Akkermansia spp. Each intestinal section was also separately hybridized with a negative control, non-sense probe (non-EUB-Alexa Fluor 488) to evaluate possible bacterial autofluorescence. 56 After hybridization, the slides were extensively washed in buffer, mounted with antifade solution containing 4′,6-diamidino-2-phenylindole (DAPI), and analyzed using an epifluorescence microscope (Olympus BX43). For each case and each probe, 10 random fields with labeled bacteria on the surface above the epithelium and in 10 crypts were captured with a 40× and 60× objective, respectively. A minimum of 3 different endoscopic biopsies were evaluated for mucosal bacterial content. Each field was captured with the DAPI filter for identification of host cell nuclei, fluorescein isothiocyanate (FITC) filter for background autofluorescence from the host epithelium, and tetramethylrhodamine-isothiocyanate (TRITC) filter for identification of probe-labeled bacteria. The images from the 3 filters were combined before quantification. Due to the presence of areas with dense numbers of bacteria that often overlapped, bacterial quantification was performed using ImageJ software. Quantification was only performed when the hybridization signals were strong and orange to red with the combined filters. A positive hybridization signal was not observed using similar technical conditions with the non-EUB338 probe. The individual size of bacteria was established by measuring 20 individualized bacteria (with a 60× objective for the crypts and 40× for the surface) in each bacterial group and calculating the mean size. The number of pixels representing positive labeling was recorded for each of the captured fields using ImageJ software and the median number of bacteria per field was calculated.
Probes Targeting Bacterial 16S rRNA Used for in Situ Hybridization.

Statistical Analyses
The datasets were tested for normality and equality of variances using the Shapiro-Wilk test and the Brown-Forsythe test, respectively. The mean number of labeled bacteria per field for each probe and compartment in dogs with CIE and controls was compared using a Mann-Whitney U test or Student’s t test where appropriate. A Spearman’s rank correlation coefficient was calculated to test for possible correlations between the number and location of labeled bacteria, CIBDAI scores, individual and cumulative histopathological scores in dogs with CIE. Tests were performed on JMP software (JMP 13, SAS software Inc.), with significance set as P ≤ .05.
Results
Study Population
The CIE group comprised 22 dogs (12 males and 10 females; median age: 3.2 years; age range: 0.5–13 years). The most commonly represented breeds included German shepherd (n = 4), soft coated wheaten terrier (n = 2), and mixed breed dogs (n = 3). The median body weight was 23.9 kg (range: 5.5–43.4 kg). All dogs with CIE presented for variable and chronic gastrointestinal signs indicative of enterocolitis. The median CIBDAI score was 5 out of 18 (range: 2–9). Based on the CIBDAI scores, 13 dogs with CIE had insignificant or mild disease activity (score: 2–5), 8 dogs had moderate disease activity (score: 6–8), and 1 dog had severe disease activity (score ≥ 9). When the histopathological scores for each morphologic parameter in the colon of dogs with CIE were summed up, the median cumulative score was 2 (range: 1–6).
The control group was composed of 11 dogs (5 males and 6 females) and the median age was 4 years (age range: 1–13 years). The median body weight was 23.6 kg (range: 2.7–32 kg). There was no history of gastrointestinal signs or abnormalities of the gross or histopathological analysis of the gastrointestinal tract (histopathological score = 0). The age (P = .44) and body weight (P = .95) of control dogs were not statistically different from dogs with CIE.
Mucosal Microbiota
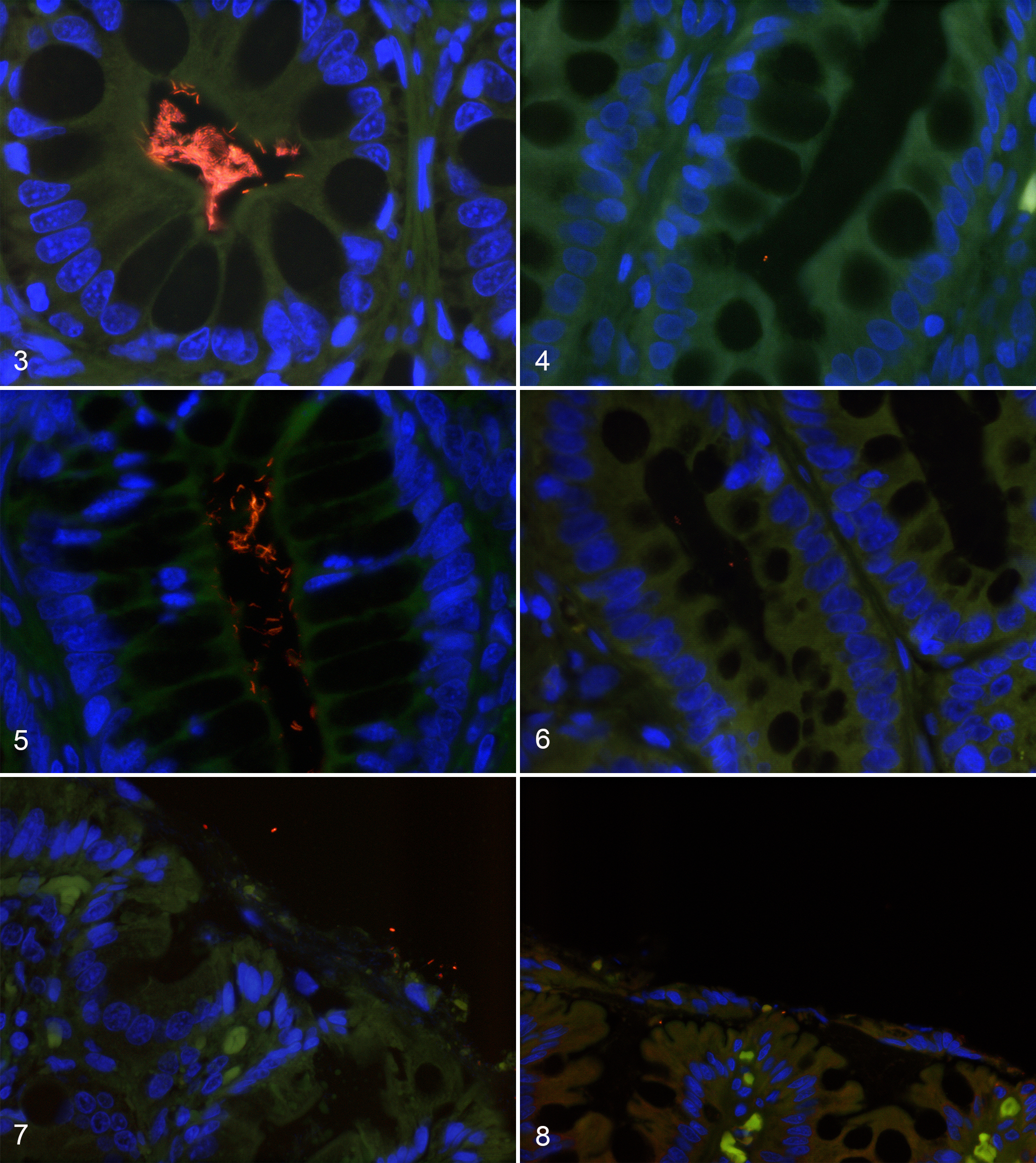
All the bacterial groups investigated with FISH were located on the colonic surface and within the crypts in both control dogs and dogs with CIE (Figs. 1, 2 and Table 2, Table S1). The number of total bacteria labeled with EUB338 probe (P = .002) was decreased within the crypts of dogs with CIE (Figs. 3, 4) compared to control dogs. Dogs with CIE had higher numbers of Escherichia coli/Shigella on the colonic surface (P = .009) and within the crypts (P = .015) as compared to control dogs. Both Helicobacter spp. (Figs. 5, 6) and Akkermansia spp. (Figs. 7, 8) were decreased on the colonic surface (P = .03; P = .012, respectively) and in the crypts (P = .008; P = .0003, respectively) of dogs with CIE. No differences were detected in the number of Faecalibacterium spp. or the total number of bacteria on the colonic surface between control dogs and dogs with CIE. There were no statistical differences between the data of the 4 dogs with CIE that received antibiotics and the dogs with CIE that had not been treated with antibiotics.
Number of Bacteria on the Colonic Surface and Within Crypts of Dogs With Chronic Inflammatory Enteropathy (CIE) and Control Dogs.
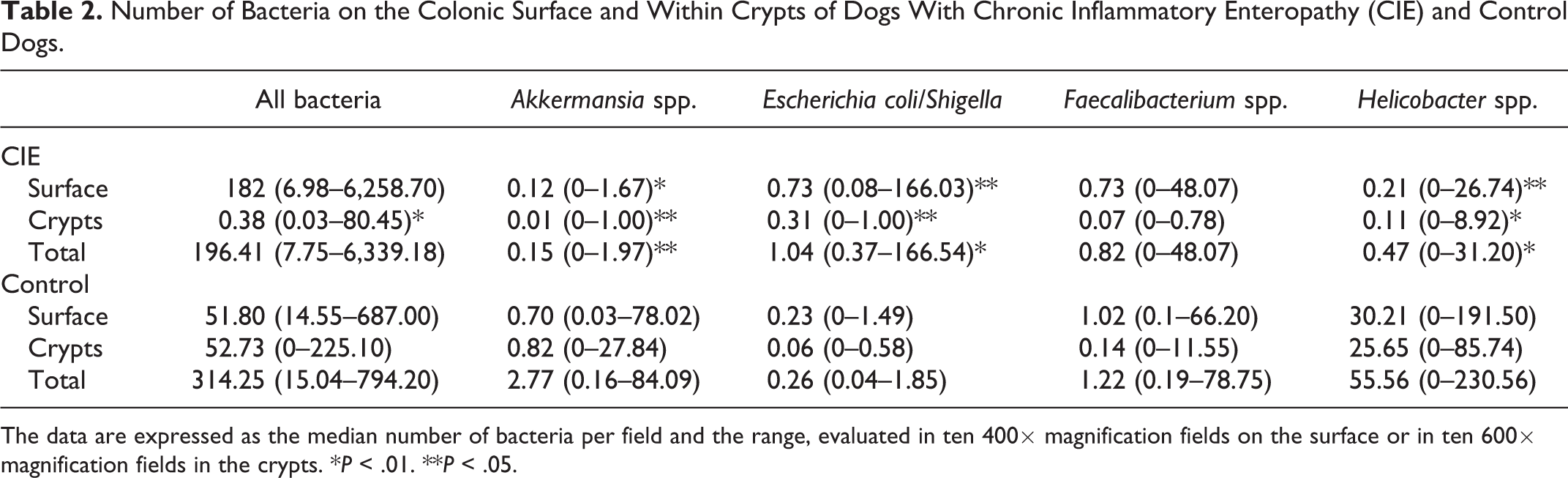
The data are expressed as the median number of bacteria per field and the range, evaluated in ten 400× magnification fields on the surface or in ten 600× magnification fields in the crypts. *P < .01. **P < .05.
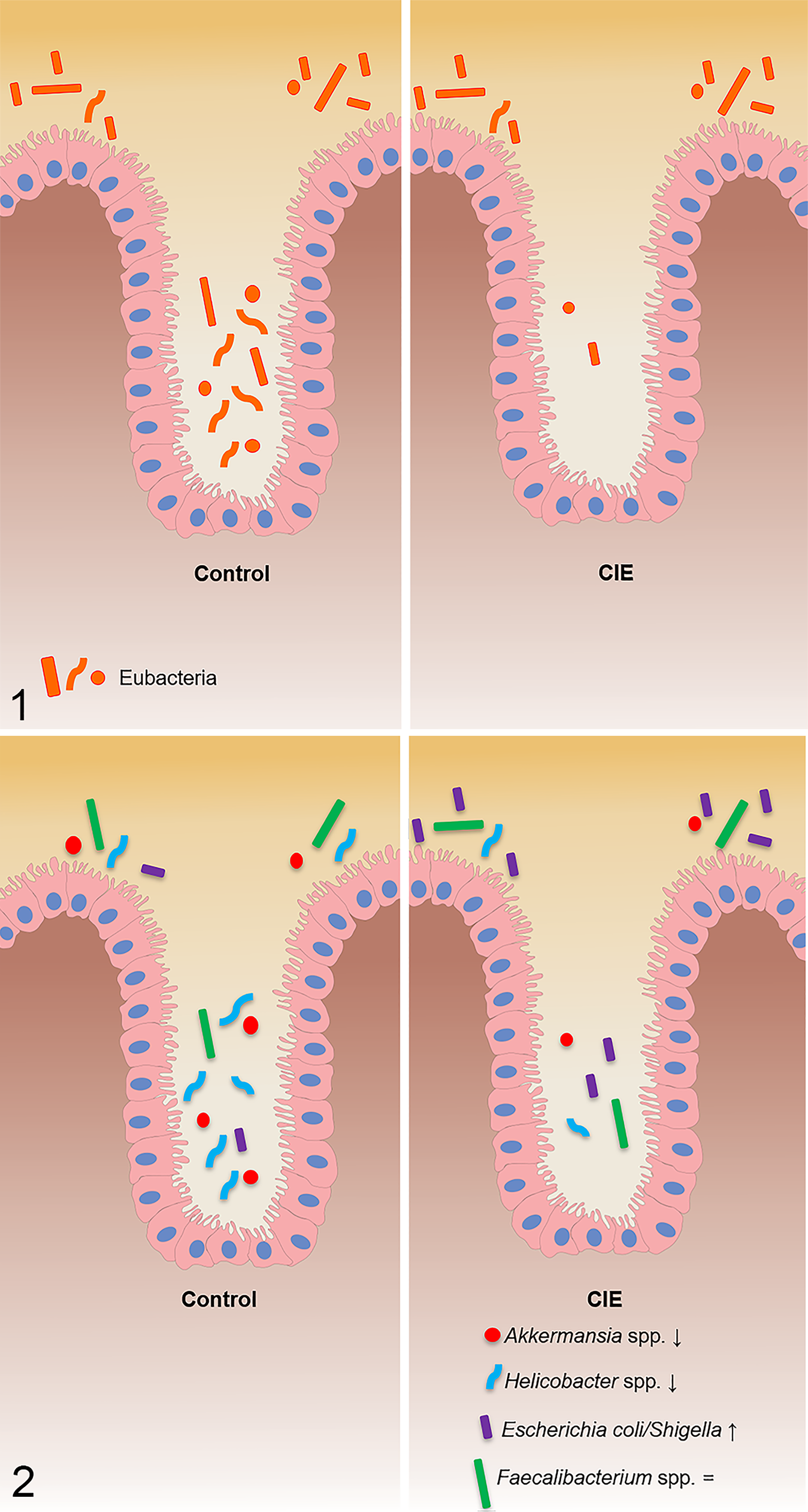
Correlations
Spearman’s rank correlation coefficient showed no significant correlation between mucosal bacterial populations in the colon and the CIBDAI or histopathological scores (individual or cumulative) in dogs with CIE.
Discussion
In this study we demonstrated reduced numbers of total cryptal bacteria in the colon of dogs with CIE. Similar to our findings, a reduction in the cryptal bacteria has previously been described in the colon of humans with acute ulcerative colitis, although the bacterial species were not characterized. 44 The cryptal microbiota is thought to play a role in mucosal homeostasis, promoting immunomodulation and serving as a reservoir for bacteria, allowing recolonization of the gut after disruption by antibiotics or infection. 16 The predominance of a bacterial species in a specific niche is not an uncommon concept. In mice, the segmented filamentous bacterium dominates at the epithelial surface of the gut associated lymphoid tissue and Alcaligenes spp. is an almost-exclusive inhabitant of murine Peyer’s patches. 35 In mice, the colonic crypts are known to be colonized by Acinetobacter spp., 37 Helicobacter spp., 58 and Bacteroides spp. 43
Helicobacter spp. was the predominant bacterial group colonizing the colonic crypts in the control dogs of our study. Helicobacter spp. was previously demonstrated to colonize the stomach and small intestine via adherence to epithelial surface glycans. 27 Helicobacter spp. can be classified as enterohepatic or gastric. 46 Gastric strains are larger “corkscrew” spiral-shaped bacteria while enterohepatic strains are slightly curved to spindle-shaped with tapered ends. 41 Helicobacter pylori is an important pathogen associated with the development of peptic and gastric ulceration as well as gastric cancer in humans. 36 However, natural H. pylori infection has not been described in the dog, which harbors H. bizzozeronii and H. felis in the gastric mucosa. 41 Enterohepatic species of Helicobacter have previously been reported to be the main bacterial group colonizing the colonic crypts of healthy dogs. 41 Helicobacter spp. DNA has previously been amplified from the colonic mucosa in all 6 of 6 healthy dogs tested. 41 These cryptal bacteria have been described to belong to H. bilis/flexispira taxon 8, H. cinaedi, and H. canis, with different species colonizing the same individual. 41 Helicobacter spp. can be cultured from the feces of both healthy dogs and dogs with gastrointestinal disease with a similar prevalence. 42 The significance of enterohepatic species of Helicobacter that colonize the intestinal tract and the liver of humans and animals is controversial. 46 Because enterohepatic strains are highly prevalent in healthy humans and animals, they have been considered as members of the commensal bacterial microbiota. 46 However, some studies have associated these bacteria with gastroenteritis, hepatitis, and other diseases. 46 We considered Helicobacter spp. a commensal bacterium in control dogs, contrasting a previous study 10 that described a positive association between mucosal fibrosis/atrophy and the abundance of Helicobacter spp. in the colonic crypts of dogs with gastrointestinal disease. The study by Castiglioni et al 10 analyzed samples from dogs with either acute or chronic gastrointestinal disease using immunohistochemistry, which is considered to be less specific than FISH. Their study did not include a control group and did not identify a positive correlation between colonization by Helicobacter spp. and intestinal inflammation. Our results support a previous study that reported decreased numbers of Helicobacter spp. in the colonic mucosal surface of dogs with CIE. 9
The colonic mucus layer of humans is enriched in Akkermansia muciniphila, a mucin-utilizing, anaerobic, gram-negative bacterial species. 7,14 A depletion of the mucosal population of Akkermansia muciniphila is a common finding in humans with inflammatory bowel disease, 39,40 suggesting that it may have potential anti-inflammatory properties. Akkermansia muciniphila beneficially affects glucose metabolism, lipid metabolism, and intestinal immunity. 18,38,48 Akkermansia spp., within the Verrucomicrobia phyla, is rarely identified in feces of dogs through next generation sequencing, 51 probably because this bacterial species is present in low abundance and closely associated with the colonic mucus layer and crypts in dogs. Further studies are warranted to investigate the role of Akkermansia spp. in gastrointestinal health and disease in dogs.
Human patients with inflammatory bowel disease have an increased abundance of bacteria at the inflamed colonic mucosal surface, 52 including Bacteroides spp. and E. coli. 6,32 Although the abundance of total bacteria on the colonic surface did not differ between control and CIE dogs, our findings support a previous study that described increased E. coli and Enterobacteriaceae bacteria within the colonic and ileal mucosa of dogs with CIE. 9 Previous studies utilizing 16 S rRNA sequencing and qPCR found that Faecalibacterium spp. bacteria were decreased in the feces of dogs with CIE, 3,20 while we were unable to detect significant differences between control dogs and dogs with CIE in the mucosa. The lack of statistical correlation between mucosal bacterial numbers and the CIBDAI or histopathology scores could be related to the marked individual variability in bacterial counts in dogs with CIE or due to the predominance of dogs with mild clinical disease. Although alterations in intestinal microbial composition have long been associated with chronic intestinal inflammation; it has been difficult to prove whether dysbiosis is a cause or effect of disease. 34
In this study, we used archived formalin-fixed samples from clinical cases to characterize the mucosal microbiota. Although Carnoy’s fixative can be superior to formalin in preserving the mucus layer, 53 several studies have utilized formalin-fixed samples for quantification of bacteria in humans 6,49,52 and animals 9,24,26 and demonstrated a largely intact mucus layer. In control dogs, full-thickness samples were collected immediately after the dogs were euthanized for reasons unrelated to this study, to avoid the unnecessary use of animals for experiments. On the other hand, the colonic samples in dogs with CIE were collected endoscopically, with routine colonic cleansing prior to collection. It is possible that the cleansing might affect the mucosal biofilm. However, it seems unlikely that the cleansing would affect the deeper microbiota within the colonic crypts or alter the composition of specific bacterial groups. Additionally, the age, sex, breed, and diet can influence the gut microbiota. While very high protein diets (∼45 g/100 g of dry matter) may affect the intestinal microbiota of dogs, such high levels of protein are not commonly found in commercial diets (∼27 g/100 g of dry matter). 21,45,47
Conclusions
This study showed that dogs with CIE had alterations in the composition of their colonic mucosal microbiota. Interestingly, the cryptal bacteria that are intimately associated with the host mucosa and composed mainly of Helicobacter spp., were depleted in dogs with CIE. Helicobacter alone, or associated with other bacterial species described here in lower abundance could play a role in excluding colonization by pathogenic bacteria or in regulating inflammation in the canine colon. Future immunologic and metagenomic studies are warranted to better understand the host–microbiome interaction in canine CIE.
Supplemental Material
Supplemental_Table_S1 - Bacterial Biogeography of the Colon in Dogs With Chronic Inflammatory Enteropathy
Supplemental_Table_S1 for Bacterial Biogeography of the Colon in Dogs With Chronic Inflammatory Enteropathy by Paula R. Giaretta, Jan S. Suchodolski, Albert E. Jergens, Jörg M. Steiner, Jonathan A. Lidbury, Audrey K. Cook, Mohsen Hanifeh, Thomas Spillmann, Susanne Kilpinen, Pernilla Syrjä and Raquel R. Rech in Veterinary Pathology
Footnotes
Acknowledgements
The authors are thankful to the Finnish Center for Laboratory Animal Pathology (FCLAP) for providing unstained histologic slides. The technical assistance of So Young Park, Kathrin Burke, and Laura Parikka is gratefully acknowledged.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Paula R. Giaretta is supported by a fellowship from the Brazilian National Council for Scientific and Technological Development (CNPq).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
