Abstract
Several studies based on histopathology or molecular investigations suggest a causal relation between Felis catus papillomavirus (FcaPV-2) infection and bowenoid in situ carcinoma (BISC) in cats. Nevertheless, data on distribution of viral DNA for different F. catus papillomavirus types (FcaPV-1, 2, 3, 4, 5) in precancerous skin lesions are lacking. In this study, incisional and excisional skin biopsies from 18 cats with BISC were investigated for the presence of FcaPV DNA by quantitative polymerase chain reaction (qPCR) and chromogenic in situ hybridization (CISH) using specific probes to detect each of the FcaPVs that have been identified so far. By qPCR analysis, 15 of 18 samples were positive for FcaPV-2, 2 were positive for FcaPV-4, and 1 sample was negative for all FcaPVs studied. Two cases were positive for FcaPV-5 by qPCR only. FcaPV-1 and FcaPV-3 were not detected by either method. CISH positivity for FcaPV-2 and FcaPV-4 was 100% concordant with qPCR. FcaPV-2 CISH signal was observed as nuclear dots within grouped neoplastic keratinocytes in 12 BISCs and in the perilesional skin of 9 biopsies. In 3 of these 9 cases, the signal was not observed within the BISC. FcaPV-4 CISH positivity was detected only within BISCs in 2 cases. The overall rate of concordance for FcaPV detection between PCR and CISH was 97.8%. This study suggests that CISH is a reliable method to detect FcaPV-2 and FcaPV-4 infection in cats and provides useful information on the type, rate, and localization of infected cells.
Feline bowenoid in situ carcinoma (BISC) is an uncommon skin disease frequently associated with species-specific papillomaviruses (PVs). 12,22 Lesions may occur at any cutaneous site, even in pigmented and densely haired skin, as well as in areas with or without sun exposure. 12 Among Felis catus papillomaviruses described in cats (FcaPV-1 to FcaPV-5), the most studied for its association with BISC is FcaPV-2. 1,5,14,19,20,28 –30,32 FcaPV-1, the first feline virus to be fully sequenced (and initially named FdPV-1), was detected in a plaque and associated with oral papillomas but never with BISCs. 16,29 FcaPV-3, which induces histopathological features different from those shown by other FcaPVs, has been associated with multicentric BISC in a Sphynx cat and recently in 4 other cases. 15,21 FcaPV-4 DNA has been identified by real-time polymerase chain reaction (PCR) in oral tissue of a cat with stomatitis and in 2 cats with BISCs. 5,11 FcaPV-5 is a very recently identified papillomavirus that was associated with a feline viral plaque. 14
Therefore, most BISCs associated with PV infection are related to FcaPV-2 and, to a lesser extent, to FcaPV-3 and FcaPV-4. 11,22 In the past decade, the causal relation between PVs (in particular FcaPV-2) and BISC has also been suggested by several authors based on the presence of histopathological indicators of viral infection, immunohistochemistry for p16 accumulation as a result of impaired cell cycle, and identifying enhanced E6/E7 gene expression in precancerous lesions. 13,18,32 Immunohistochemistry has also been used to detect the major viral capsid protein (L1). However, since L1 is produced only late in differentiated keratinocytes, the presence of this viral antigen is restricted to the superficial layers of the epidermis, thus reducing the overall sensitivity of this method. 10,32 Moreover, the lack of antibodies specific to FcaPV types and the use of cross-reacting L1 antibodies influence both sensitivity and specificity of this approach.
Molecular assays, such as PCR, have been largely used to investigate the viral etiology of BISC. 9,15,18 –20,23 Nevertheless, the interpretation of a positive PCR result could be ambiguous, since positive results have also been reported for FcaPV-2 in normal skin of cats. 31 In a previous study, cytobrush samples collected from healthy cats were found to be positive for FcaPV-2 by PCR, raising the question of whether this technical approach is diagnostically useful. Thus, a positive result by conventional PCR might be due to the presence of virions on the skin surface and does not demonstrate the virus as the causative agent of the lesion. 6 Recently, a specific FcaPV-1-2-3-4 quantitative PCR assay has been proposed to quantify FcaPV DNA. 11 This approach has proven useful to show that lesions with high copy numbers of FcaPV DNA may be more likely associated with viral pathogenesis. Very recently, RNA in situ hybridization (ISH) proved to be a reliable method to demonstrate active FcaPV-2 E6 and E7 gene transcription within the neoplastic cells in feline squamous cell carcinoma (SCC), and fluorescent ISH detected FcaPV-2 in 37.5% of the examined BISCs. 2,8
The primary aims of this study were (1) to investigate the feasibility and reliability of chromogenic in situ hybridization (CISH) to study papillomavirus infection in cats, (2) to compare quantitative PCR (qPCR) and CISH methods in terms of sensitivity and specificity for diagnostic purposes, and (3) to investigate the type and localization of infected cells in skin samples.
Materials and Methods
Sample Collection and Histopathology
Thirty-four archived, formalin-fixed, paraffin-embedded specimens from incisional and excisional feline skin biopsies with a diagnosis of BISC were retrieved from the Department of Veterinary Sciences, University of Pisa (Italy), Laboratory “LaVallonea,” Passirana di Rho (MI, Italy) and the Histopathology Department, Istituto Zooprofilattico Sperimentale delle Venezie, Legnaro (PD, Italy), from 2016 to 2017. As negative control cases, 9 biopsy specimens were selected, including normal skin (n = 3) and skin from cats with allergic (n = 3), neutrophilic (n = 1) and herpetic dermatitis (n = 2). Three veterinary pathologists (M.V., C.Z., F.A.) independently reviewed the slides, and when diagnostic agreement was achieved, cases with complete signalment, history, and clinical data were selected and further used for qPCR and CISH.
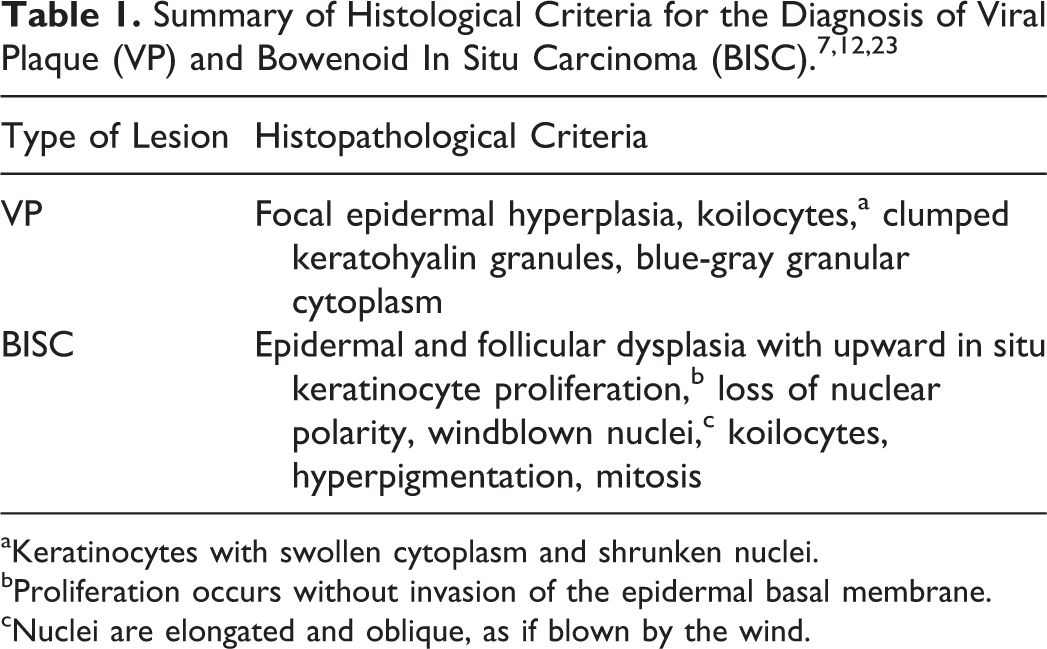
From all cases, 5- to 10-μm-thick tissue sections were serially cut: 1 section was routinely stained with hematoxylin and eosin (HE), while the next 10- and 5-μm-thick sections were used for qPCR and CISH analysis, respectively. To prevent carryover of contaminating DNA, the microtome overlay was covered with a new piece of adhesive tape and a new blade was used for each sample. Slices for CISH were mounted onto adhesive glass slides (Roche, Monza, Italy). Perilesional skin from excisional biopsies of BISC cases was evaluated for alterations with features of viral plaques. Histological lesions were investigated and classified based on parameters previously reported (Table 1). 7,32
aKeratinocytes with swollen cytoplasm and shrunken nuclei.
bProliferation occurs without invasion of the epidermal basal membrane.
cNuclei are elongated and oblique, as if blown by the wind.
qPCR
DNA extraction was performed on formalin-fixed, paraffin-embedded (FFPE) sections, using the DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany) following the manufacturer’s instructions and applying a preliminary removal of paraffin by extraction with xylene. DNA was eluted in 100 μl and stored at –20°C until analyses were performed.
The extracted DNA was amplified using 4 specific sets of primers amplifying a portion of FcaPVs’ L1 gene. 11 To detect the recently identified FcaPV-5, primers FcaPV-5-F (5′-ACTGCAGTACCCCCTAAGGA-3′) and FcaPV-5-R (5′-CtttAAAAACTTCCGCCCCAGTG-3′) were designed based on the available reference sequence (accession number: NC_035479.1). Assays were performed on a Rotorgene thermocycler (Corbett Research, Sydney, Australia) using SSCO SybrGreen Master Mix (Bio-Rad, Hercules, CA) and 5 μl of extracted DNA. A plasmid generated by inserting FcaPV-1, 2, 3, and 4 gene segments into a pMA-T vector using GeneArt technology (ThermoFisher Scientific, Waltham, MA) was used to determine if nonspecific products were amplified during the reaction by melt curve analysis and to evaluate qPCR efficiencies. Serial dilutions of pFcaPV plasmid ranging from 106 to 102 copies/reaction in DNase-RNase free water were prepared (prep A). The same plasmid dilution series was spiked in feline genomic DNA (prep B) obtained from a negative control sample. The efficiency for each FcaPV qPCR assay was calculated and compared among all FcaPV types and between prep A and prep B for each FcaPV-specific assay.
All samples, including controls, were tested in duplicate and the results were analyzed by 2–ΔΔCq quantitative method as described by Mazzei et al 11 using albumin as the reference gene. In detail, the number of copies of the target viral gene, measured as Cq value, generates a ΔCq value when compared to the corresponding Cq value of the reference gene. Moreover, a ΔΔCq value was calculated comparing the ΔCq value of the sample of interest to ΔCq values obtained from the negative control group. The ΔΔCq value represents a normalized measure of DNA viral quantity and has been calculated using REST software. 24 All samples with a positive 2–ΔΔCq value, indicating the fold change of target viral gene copies compared to the reference gene, were identified as positive and submitted for sequence analysis (BMR Genomics, Padova, Italy). qPCR for albumin was also performed to confirm the adequacy of extracted DNA.
CISH
Probes for CISH analysis were generated by PCR using a DNA labeling kit with digoxigenin-dUTP (DIG DNA Labeling Mix; Roche). Briefly, DIG-labeled DNA probes were synthetized using HotstartTaq DNA Polymerase mix (Qiagen) with addition of 200 μM DIG DNA Labeling Mix. The PCR protocol consisted of 30 amplification cycles using FcaPV-specific primers followed by 10 amplification cycles after the addition of 2 units of @Taq polymerase (Euroclone, Milano, Italy).
For FcaPV-1, 2, 3, and 4, labeled DNA was synthesized using the pFcaPV plasmid that was generated by inserting the FcaPVs’ gene segments into the pMA-T vector using GeneArt technology (Thermo Fisher Scientific) as previously described. 11 The FcaPV-5 probe was generated using a qPCR-positive sample as a DNA template. PCR products were purified by the QIAquick Nucleotide removal kit (Qiagen). The purity and concentration of the probes were quantified by running an aliquot on a 2% agarose gel with a quantitative 100-bp DNA marker (SharpMass; Euroclone).
CISH for FcaPV-1, 2, 3, 4, and 5 DNA was performed in automation on the Ventana BenchMarck ULTRA platform (Ventana Medical System; Roche). FFPE tissue sections were deparaffinized, rehydrated, and permeabilized as described below. Target site unmasking was performed by incubating the slides 3 times in cell conditioning buffer (ULTRA CC2; Roche) at 86°C, followed by ISH Protease 3 treatment, at 36°C for 16 minutes. Denaturation was obtained at 80°C for 20 minutes. Sections were incubated with specific digoxigenin-dUTP labeled probes at a 1.5-μg/ml concentration and underwent hybridization at 42°C for 9 hours. After being washed 3 times with buffer for 8 minutes, the detection substrate UltraView Red ISH DIG Detection Kit (Roche) was applied. Finally, sections were counterstained with hematoxylin. Positive signals were detected as red-purple deposits within the nucleus of epithelial cells. Negative control sections, consisting of non-BISC lesions and normal skin, were included in each run. No positive controls were available for FcaPV-1, 3, or 5.
Results
Sample Collection and Histopathology
Eighteen cases with clinical and histopathological diagnoses of BISC and complete signalment data were selected. Data on signalment of all cats (with BISC, non-BISC, and healthy) are provided in Supplemental Table S1. In all cases, morphological alteration typical of papillomavirus infection was seen with different combinations (2 or more of the histological findings mentioned in Table 1). Perilesional skin was included in 11 of 18 cases, and 9 of 11 cases had viral plaques (Table 2); in some cases with perilesional skin, focal thickening and hypergranulosis of the perilesional epidermis were seen, which led to doubtful interpretation (hyperplastic vs viral plaques). Negative control cases, which included normal skin and allergic, neutrophilic, and herpetic dermatitis, did not have lesions consistent with viral plaques or BISC.
Detection of Results of FcaPV-1 to FcaPV-5 by qPCR and CISH.
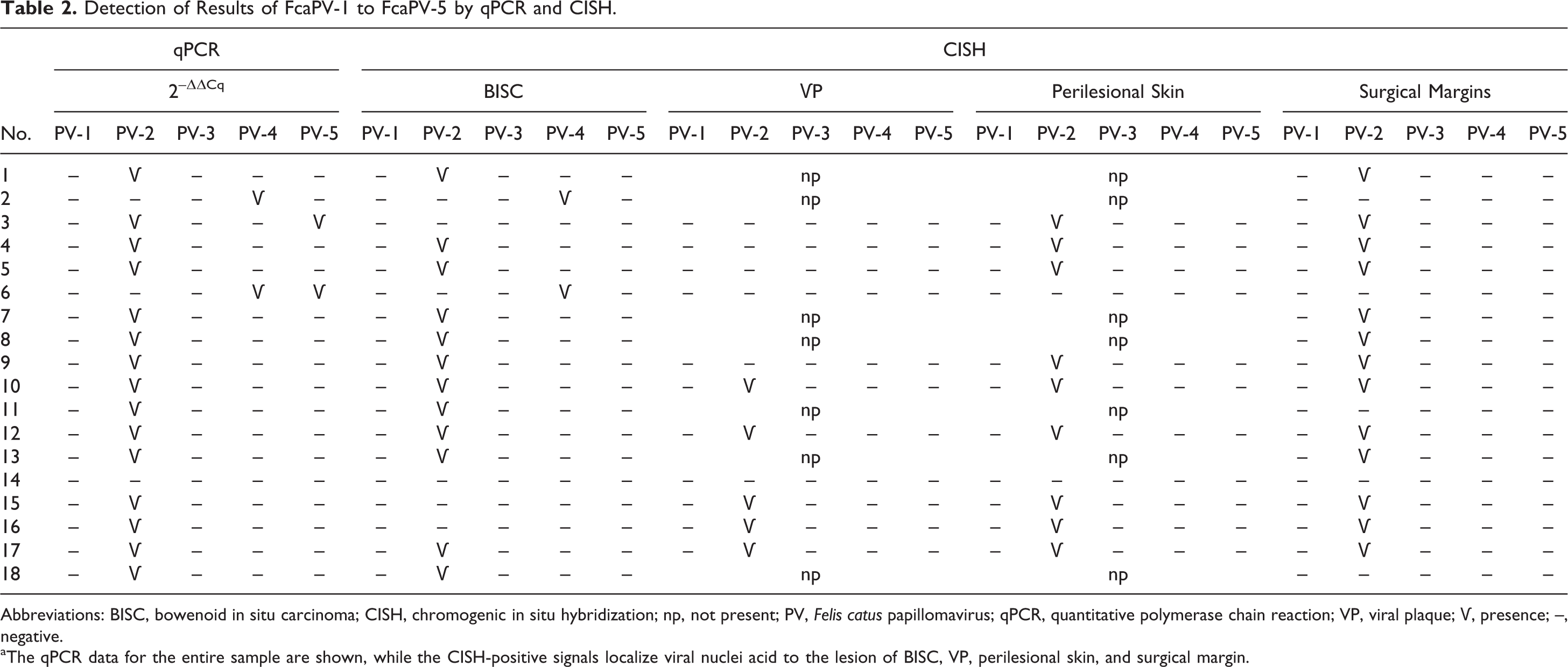
Abbreviations: BISC, bowenoid in situ carcinoma; CISH, chromogenic in situ hybridization; np, not present; PV, Felis catus papillomavirus; qPCR, quantitative polymerase chain reaction; VP, viral plaque; Ѵ, presence; –, negative.
aThe qPCR data for the entire sample are shown, while the CISH-positive signals localize viral nuclei acid to the lesion of BISC, VP, perilesional skin, and surgical margin.
qPCR
To determine the efficiency of each qPCR assay, preparations of pFcaPV plasmid in DNase-RNase free water (prep A) were compared to the same preparations spiked with genomic feline DNA (prep B). The efficiency of each qPCR assay was comparable for both prep A and prep B (prep A efficiency: 94.5% ± 2.4%; prep B efficiency: 93.0% ± 4.2%; mean ± SD). Efficiency values between prep A vs prep B were also similar (Table 3).
Quantitative Polymerase Chain Reaction Efficiency.a
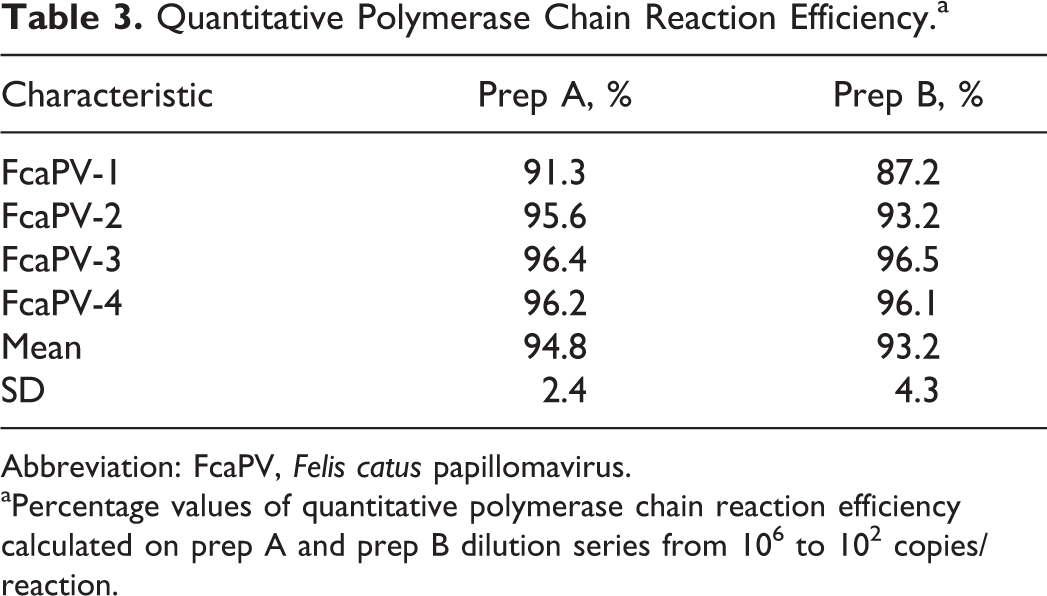
Abbreviation: FcaPV, Felis catus papillomavirus.
aPercentage values of quantitative polymerase chain reaction efficiency calculated on prep A and prep B dilution series from 106 to 102 copies/reaction.
The reference gene, albumin, was amplified in all samples. Overall, 2–ΔΔCq analysis was positive in 17 of the 18 samples from cases of BISC. Specifically, 15 of 18 (83%) BISC cases were positive for FcaPV-2, 1 case was positive only for FcaPV-4, 1 case was positive for both FcaPV-2 and FcaPV-5, 1 case was positive for both FcaPV-4 and FcaPV-5, and 1 case was negative for all of the PV types tested (Table 2). FcaPV types assigned by qPCR were confirmed by sequence analysis. No PV was detected by 2–ΔΔCq analysis in control cases, including both non-BISC inflammatory lesions and normal skin.
CISH
Overall, considering BISC and perilesional skin, CISH identified 17 of 18 samples positive for either FcaPV-2 or FcaPV-4. FcaPV-2 signal was detected in BISCs in 12 of 18 cases. Labeling was exclusively localized within nuclei of grouped neoplastic keratinocytes and was detected exclusively in the suprabasal layers, from stratum spinosum to the stratum corneum (Figs. 1, 3). The nuclear signal intensity ranged from very mild (with small red-purple spots) to strong (with large and dense red-purple patches), and the distribution varied from focal to diffuse. Focal positivity was represented by 1 to 2 groups of a few positive nuclei within the entire BISC lesion. The signal was strong and diffuse in 5 cases, strong and focal in 2 cases, and mild and focal in 5 cases (Figs. 1, 3, 4). Within perilesional skin, FcaPV-2 was detected in the nuclei of suprabasal epidermal keratinocytes (stratum granulosum to stratum corneum) of 9 of 11 cases, in viral plaques, and in the hyperplastic epidermis (Figs. 4–6). In 3 of these 9 cases, the labeling was present in the perilesional skin but was not observed within the associated BISC. CISH positivity was detected at the surgical margins of both incisional and excisional biopsies in 13 of 18 cases (Figs. 7, 8).
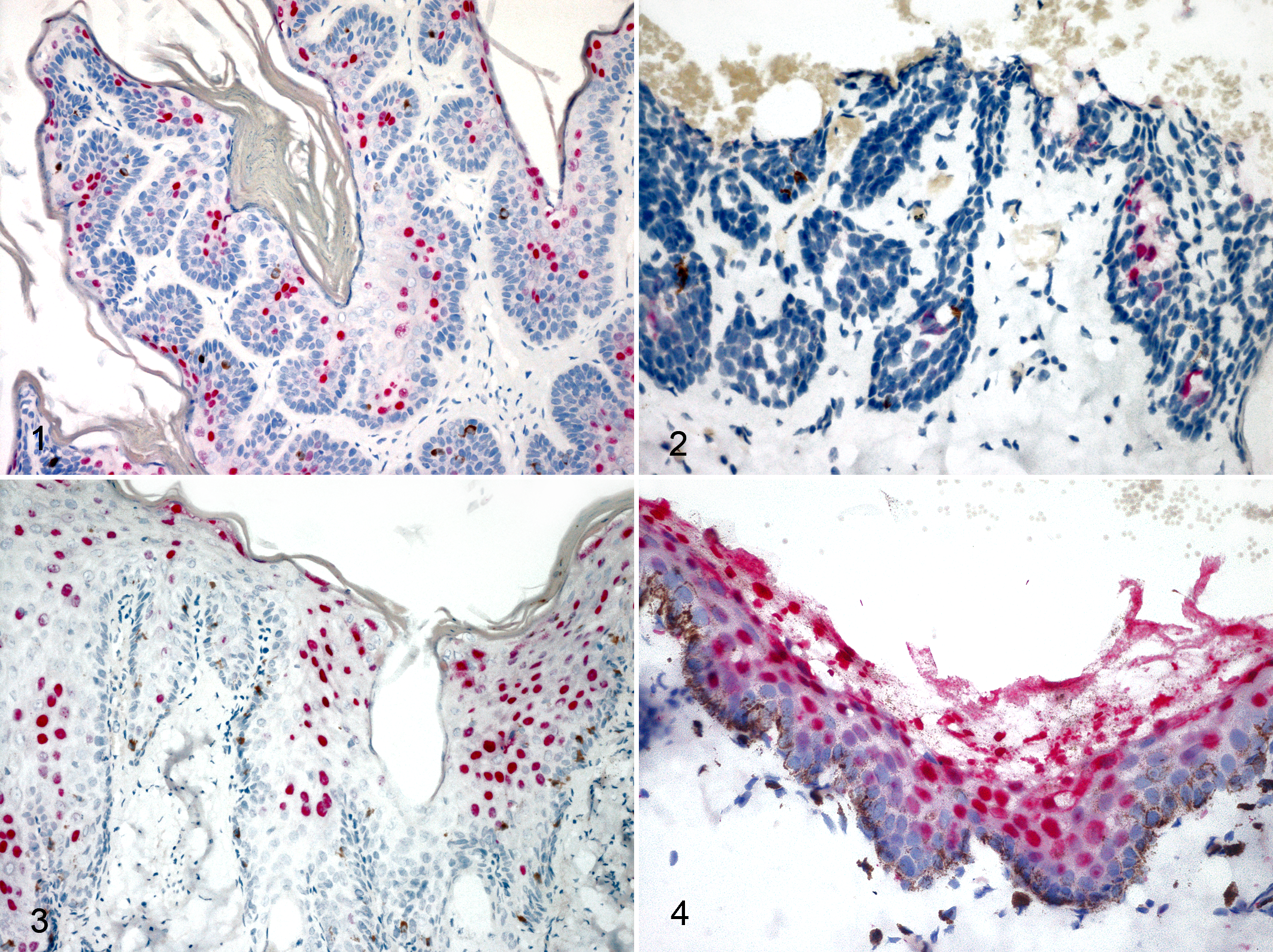
Felis catus papillomavirus (FcaPV) infection, skin, cats. Chromogenic in situ hybridization. Figures 1 and 3 (case Nos. 1 and 7): the red nuclear labeling for FcaPV-2 is strong and diffuse within bowenoid in situ carcinoma (BISC). Figure 2 (case No. 2): the nuclear labeling for FcaPV-4 is strong and focal within a BISC. Figure 4 (case No. 15): strong and diffuse red nuclear labeling in a viral plaque.
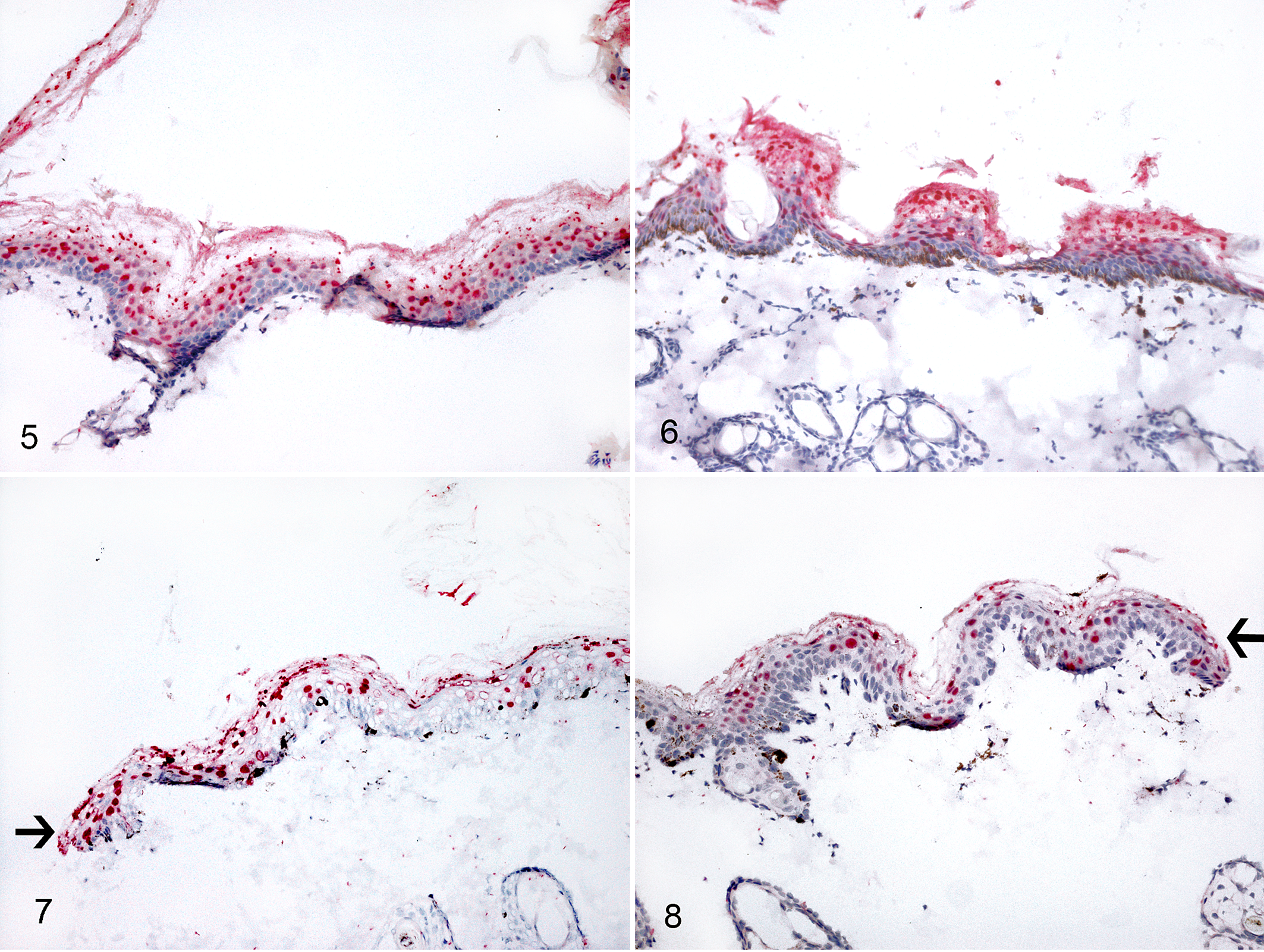
Felis catus papillomavirus (FcaPV) infection, skin, cats. Chromogenic in situ hybridization. Within the perilesional skin, strong and diffuse red labeling is present in the nuclei of suprabasal keratinocytes in regularly (Fig. 5; case Nos. 17 and 16) or focally (Fig. 6; case No. 15) hyperplastic epidermis and at the surgical margins (arrows) of 2 cases in which the perilesional skin was sampled together with the bowenoid in situ carcinoma (Figs. 7 and 8; case Nos. 10 and 17).
Regarding FcaPV-4, the signal was detected in 2 cases as a focal and strong positive nuclear signal of suprabasal keratinocytes within BISC (Fig. 2), while no labeling was detected in the perilesional skin. No FcaPV-1, 3, and 5 labeling was detected in any of the examined samples. One case was negative for all of the tested PV types. No CISH signal was detectable in control samples, which included normal skin and skin affected by dermatitis. The overall rate of concordance for FcaPV-1, 2, 3, 4, and 5 detection between PCR and CISH was 97.8%; CISH demonstrated a specificity and sensitivity of 100% for FcaPV-2 and FcaPV-4 (Table 2).
Discussion
The frequent identification of FcaPV-2 in cutaneous lesions by PCR has implicated this virus in the development of feline BISC. 1,19,20 However, FcaPV-2 DNA has also been detected in a proportion of asymptomatic cats. 6,31 Many PCR-based studies could not discriminate active infection marked by a high number of intracellular viral DNA from positivity due to the presence of virions on the skin surface. This issue was partially solved by the development of a reverse transcription (RT) qPCR method detecting enhanced E6 and E7 transcription activity in BISCs and by fluorescent ISH to detect FcaPV-2 DNA. 2,32
This study detected and localized each FcaPV DNA (FcaPV-1 to FcaPV-5) within the lesions of feline BISC by a combination of qPCR and CISH. The goal of any FcaPV detection strategy would be not only to ascertain the presence of FcaPV but to also characterize its role in oncogenesis. It is known that a positive result of conventional PCR alone is prone to misinterpretation, while qPCR provides a quantitative measure of viral load with useful diagnostic implications. 11 In our study, PV DNA was detected by CISH in 94.4% of the examined samples (17/18), within the nuclei in the BISC and/or perilesional skin. There was 100% concordance of CISH and qPCR for FcaPV-2 and FcaPV-4. Recently, Demos et al 2 demonstrated a lower prevalence of intralesional FcaPV-2 DNA (35.7%) using FISH, within a series of BISCs that were positive for FcaPV-2 by PCR. This lower prevalence might be due to false-positive results obtained by conventional PCR, as well as to low sensitivity of the fluorescent in situ hybridization (FISH) method. CISH has the advantage of preserving the morphological context much better than FISH in FFPE samples. In addition, the availability of many commercial immunostainers that allow automation, as well as time and cost sparing, makes this method more feasible for diagnostic purposes.
It is hypothesized that when PV comes into contact with a mucocutaneous epithelium, microabrasions allow infection of basal cells, resulting in the production of small numbers of circular PV DNA copies with low replication activity. 26 However, the viral life cycle is completed as infected keratinocytes differentiate, 3,27 and the expression of PV E6 and E7 proteins promotes replication of the normally postmitotic suprabasal cell and allows greater amplification of the viral genome. 4 Thus, we consider that the presence of the CISH signal in nuclei of the suprabasal epithelium suggests a role in tumorigenesis and tumor maintenance by the FcaPV in most of the examined BISCs. Moreover, qPCR and CISH were both negative in all the negative controls (ie, samples from animals that did not have BISC), suggesting that these 2 methodologies are specific and useful tools to diagnose active PV infection in cats. qPCR has been previously shown to be helpful in discriminating between infection and the mere presence of the virus on the skin surface, 11 while CISH adds useful information on the type and localization of infected cells. PCR quantitation is more diagnostically feasible than CISH; however, the information obtained by PCR is different from that derived from CISH and does not provide insights into the distribution of the viral DNA within the skin.
Feline viral plaques and BISCs have been traditionally classified as separate disease entities. However, since both are typically caused by FcaPV-2 and since transitional lesions between viral plaques and BISCs have been reported, they probably represent different severities of the same disease process. 33 Histologically, feline viral plaques comprise mild epidermal hyperplasia that often contains prominent PV-induced cell changes. Nonetheless, the histological diagnosis may be difficult if cytopathic effects are not evident. In these cases, CISH could be a useful method to demonstrate the viral origin of these lesions. In fact, in some of the examined cases in this study, CISH positivity was observed in perilesional skin within viral plaques and hyperplastic epidermis.
In 3 cases, CISH labeling for FcaPV-2 was absent within the BISC lesion but detectable in the perilesional skin. It is already known that neoplastic transformation can lead to loss of the virus or to minimal replication in advanced cases. 2 Our results reinforce prior observations suggesting that PV productive infection is maintained in precancerous lesions of human cervical cancer. 4,25 Moreover, it has been demonstrated that areas of hyperplastic epithelium surrounding feline SCC consistently contained intense ISH nuclear signals for FcaPV-2. 8 The detection of CISH-positive cells in the perilesional skin, but not within the BISC, also suggests that incisional biopsies may not be reliable to confirm a viral cause. Signal visible in perilesional skin suggests cats with BISCs may have more widespread infections that predispose to new lesions. In this view, CISH may be helpful to investigate the infective status of the epidermis.
FcaPV-4 DNA was demonstrated in cells within BISCs in 2 cases, supporting the possible role of this PV type in BISC development. FcaPV-4 has been previously detected by real-time PCR from a cat with stomatitis and from 2 cats with BISC. 5,11 In the positive cases herein described, FcaPV-4 DNA was restricted to the BISC, with a focal pattern.
In our study, no positivity for FcaPV-1 and FcaPV-3 was detected by qPCR or by CISH. Although CISH did not detect FcaPV-1, 3, or 5, the sensitivity of this method is not known because no positive controls were available. Nonetheless, our findings are consistent with the very low prevalence of these PVs reported in feline skin lesions. 14,15,17,29,30 Sequence analysis for FcaPV-1, 3, and 5 supported the specificity of this method. FcaPV-5 was detected by qPCR in 2 cases, while CISH failed to detect virus in both of these samples. Since these 2 cases were positive by CISH for FcaPV-2 and FcaPV-4, respectively, the lack of FcaPV-5 signal should not be ascribed to fixation or processing interference. A possible explanation for the lack of detection could be either the loss of infected cells when cutting sections or ineffectiveness of probe annealing for FcaPV-5. Further studies are needed to better understand the role of FcaPV-5 in BISC development.
Conclusion
In conclusion, this study documents the presence of nuclear FcaPV-2 and FcaPV-4 DNA in feline BISCs and in perilesional skin, both within viral plaques and in apparently hyperplastic epidermis, supporting the pathogenic role of FcaPV-2 and FcaPV-4 in the development of BISCs. Our results suggest that CISH is a reliable method to detect PV infection and provides additional useful information on the type, rate, and localization of infected cells. Thus, CISH may be proposed as useful method, in both diagnostic and research fields, to detect FcaPV-2 to FcaPV-4 infection in cats.
Supplemental Material
Supplemental Material, DS1_VET_10.1177_0300985819859874 - Felis catus Papillomavirus Types 1, 2, 3, 4, and 5 in Feline Bowenoid in Situ Carcinoma: An In Situ Hybridization Study
Supplemental Material, DS1_VET_10.1177_0300985819859874 for Felis catus Papillomavirus Types 1, 2, 3, 4, and 5 in Feline Bowenoid in Situ Carcinoma: An In Situ Hybridization Study by Marta Vascellari, Maurizio Mazzei, Claudia Zanardello, Erica Melchiotti, Francesco Albanese, Mario Forzan, Marta Francesca Croce, Alberto Alberti and Francesca Abramo in Veterinary Pathology
Footnotes
Acknowledgements
We thank all the colleagues who referred the cases.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financially supported by Fondi di Ateneo University of Pisa.
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

