Abstract
Newcastle disease is an important disease of poultry caused by virulent strains of Newcastle disease virus (NDV). During the 1998 to 2002 outbreaks of Newcastle disease in Australia, it was observed that the mild clinical signs seen in some chickens infected with NDV did not correlate with the viruses’ virulent fusion protein cleavage site motifs or standard pathogenicity indices. The pathogenicity of 2 Australian NDV isolates was evaluated in experimentally challenged chickens based on clinical evaluation, histopathology, immunohistochemistry, and molecular techniques. One of these virus isolates, Meredith/02, was shown to induce only very mild clinical signs with no mortalities in an experimental setting, in contrast to the velogenic Herts 33/56 and Texas GB isolates. This minimal pathogenicity was associated with decreased virus replication and antigen distribution in tissues. This demonstrates that the Australian Meredith/02 NDV, despite possessing a virulent fusion protein cleavage site, did not display a velogenic phenotype.
Keywords
Newcastle disease is a viral disease of birds, which is endemic in many countries and can have severe impacts on domestic poultry and in some cases wild birds. It is caused by virulent strains of Newcastle disease virus (NDV), which is classified within the genus Avulavirus in the family Paramyxoviridae and is synonymous with avian paramyxovirus serotype 1 (APMV-1). 3 NDV is a negative-sense RNA virus, containing 6 genes (3′-NP-P-M-F-HN-L-5′) that encode 7 proteins: the nucleocapsid protein (NP), the phosphoprotein (P), the matrix protein (M), the fusion protein (F), the hemagglutinin-neuraminidase protein (HN), the large polymerase protein (L), and the V protein that is produced by RNA editing of the P gene. 26 There is wide variability in pathogenicity between isolates of NDV, which has led to grouping of the various viruses into 5 pathotypes (from most pathogenic to least pathogenic): viscerotropic velogenic, neurotropic velogenic, mesogenic, lentogenic, and asymptomatic. 3,32 Viscerotropic velogenic viruses are highly pathogenic with lesions associated with the gastrointestinal tract. Neurotropic velogenic viruses are also highly pathogenic, but respiratory and neurological signs predominate. Mesogenic viruses vary in their pathogenicity but often present with respiratory signs in younger birds and low morbidity and mortality. Lentogenic viruses are associated with mild respiratory disease, and birds infected with asymptomatic viruses show little to no clinical signs at all. 32 The intracerebral pathogenicity index (ICPI) is commonly used to classify these viruses and is conducted by intracerebral inoculation of 1-day-old chicks. In addition, sequencing of the F protein cleavage site has also been found to correlate with pathogenicity. 6 According to the World Organisation for Animal Health (OIE) guidelines, virulent, notifiable strains are those that contain multiple basic amino acids (lysine or arginine) at the F protein cleavage site with a phenylalanine at residue 117 or have an ICPI of 0.7 or greater. 32 Therefore, both mesogenic and velogenic viruses are all classified as virulent despite some mesogenic viruses causing minimal disease. 9,10
From 1998, Australia experienced multiple outbreaks of Newcastle disease in commercial poultry, including broiler and layer chickens and quail, before it was subsequently eradicated in 2002. 18,20 While outbreaks of Newcastle disease have not occurred in Australia since 2002, Newcastle disease viruses continue to circulate in wild birds, and outbreaks of pigeon paramyxovirus (APMV-1 genotype VIb) have occurred in pigeons and wild birds from 2011, with the potential to cause disease in poultry. 7 Currently circulating avirulent viruses in wild birds in Australia are from both class I and class II genotype I, and while it was speculated that the 1998 to 2002 outbreak viruses may have originated in wild birds, testing of wild birds at the time of the outbreak was not able to definitively prove this. 14,15,24
The Newcastle disease outbreaks in 1998 were predominantly located within the Mangrove Mountain region of New South Wales (NSW) and were immediately preceded by the isolation of a Newcastle disease virus in chickens from Peats Ridge, NSW (PR/98). PR/98 was detected in association with respiratory disease in broiler chickens and was shown to have an avirulent F protein cleavage site. 14 It was found that the sequence of the PR/98 virus cleavage site varied by only 2 nucleotides from the sequence found in virulent viruses. These 2 nucleotide changes induced 2 amino acid changes from 112RRQGRL117 (PR/98) to 112RRQRRF117 (virulent). 14 The PR/98 virus was thought to be the precursor to the virulent viruses detected during this time, as it was isolated on the majority of properties that went on to have outbreaks of Newcastle disease.
While most of the NDVs isolated from the 1998 to 2002 disease outbreaks were classified as virulent on account of their F protein cleavage site motifs, it was noted that the disease syndrome often differed from what was expected of a velogenic virus. 18,30 There was slower spread throughout the flock, less severe clinical signs, and a lower case fatality rate than would be observed with typical velogenic NDVs. Of particular note was an outbreak in 2002 on a layer farm in Meredith, Victoria, in which egg production had decreased by 40%, production of soft-shelled eggs had increased, and there was a very slight rise in mortalities per month from 0.4% to 0.8%. 30 Despite not having the clinical appearance of a velogenic virus, the ICPI of the isolated NDV was determined to be 1.61, and the F protein cleavage site motif was 112RRQRRF117. 17 As a result, a stamping-out policy was instituted and the birds were culled.
Given the significant trade and economic implications of Newcastle disease outbreaks, it is important to determine whether the current OIE definition of virulent, notifiable strains of NDV based on the F protein cleavage site motif is justified. A number of studies have shown that while the F protein is critical in predicting the behavior of an NDV, pathogenicity is multifactorial and proteins other than the F protein may play key roles. 11,22,28
There have been limited in vivo experimental studies using Australian NDV isolates. Previous experimental studies using an Australian virus with a virulent F protein cleavage site of 112RRQRRF117, isolated from Glenorie, NSW (9809-19-1107), found that affected birds showed mild to severe depression, respiratory disease, and neurological signs with no mortalities and recovery after 10 days. 12,27
Although the field data indicate that some Australian viruses isolated from 1998 to 2002 with virulent cleavage sites are not highly pathogenic, it is not understood why there is this apparent contradiction. Given the paucity of information on the disease from these outbreaks, one of our objectives was to confirm the field observations that chickens infected with these viruses develop only mild clinical disease. A second objective was to determine the pathogenesis of the viral infections in chickens evaluated experimentally.
In this study, we characterized the infection caused by a virulent Australian NDV (Meredith/02) and compared it with an avirulent Australian virus (PR/98). Two highly virulent reference viruses, the viscerotropic Herts 33/56 and neurotropic Texas GB, were included for comparison. The results of this work will be important in understanding the risk framework of Australian NDVs, the appropriate outbreak response to their detection in poultry flocks, and whether the current OIE definition of Newcastle disease is applicable in the Australian situation.
Materials and Methods
All of the animal experiments included in this article were conducted in accordance with the Australian National Health and Medical Research Council’s Australian Code of Practice for the Care and Use of Animals for Scientific purposes. The design of the experiments and care of the animals were approved by the CSIRO Australian Animal Health Laboratory’s Animal Ethics Committee.
All experimental work involving the use of live virus was conducted at biosafety level 3 (BSL-3).
Animals and Handling
Fifty 8-week old specific-pathogen-free (SPF), White Leghorn chickens were used in the study. The birds were randomly assigned to 1 of 5 groups of 10 birds (including 1 control group). Each group of chickens was housed in a separate room at BSL-3 comprising bare floor and a retreat area with artificial perches and sawdust. The male to female ratios of the birds in each group varied from 1:1 to 2:3. The birds were allowed 4 days to acclimatize to the rooms before challenge and had free access to feed and water at all times. Animals were monitored twice daily when clinically healthy and up to every 2 hours during the day when clinical signs were evident. Prior to euthanasia, birds were anesthetized with a combination of xylazine at 5 mg/kg (Xylazil-20; Ilium, Smithfield, Australia) and ketamine at 53 mg/kg (Metamil; Ilium) delivered via the intramuscular route.
Virus Isolates
Four Newcastle disease virus isolates were used in this study: 2 Australian viruses and 2 virulent viruses exotic to Australia (Table 1). The Meredith/02 virus was originally isolated from a cloacal swab collected during an outbreak of Newcastle disease at Meredith, Victoria, in 2002 and was classified as virulent because it contained a polybasic F protein cleavage site and an ICPI of 1.61. 17 The PR/98 virus was isolated at Peats Ridge, NSW, in 1998 and is known as the avirulent precursor virus to the subsequent virulent NDV outbreaks. The PR/98 virus has an ICPI of 0.6. 17 Both of these Australian viruses had undergone 1 passage in eggs prior to their use in this study. The 2 exotic viruses were Herts 33/56, which is a viscerotropic velogenic virus (class II, genotype IV), and Texas GB (class II, genotype II), which is a neurotropic velogenic virus with ICPIs of 1.9 and 1.75, respectively. 4 Both viruses were imported from the Central Veterinary Laboratory, Weybridge, England, in 1992 with an unknown passage history.
Newcastle Disease Virus Isolates Used in the Study.
HN, hemagglutinin-neuraminidase.
aBasic amino acids are indicated in bold.
Prior to use, each virus isolate was grown in 9- to 11-day-old embryonated SPF chicken eggs (Charles River Laboratories, Frenchs Forest, Australia) via inoculation of 0.2 ml into the allantoic cavity and incubation at 37°C for 4 days.
Experimental Design
Each group of 10 birds was challenged with 1 of the 4 virus isolates using allantoic fluid diluted in phosphate-buffered saline (PBS) to give a titer of 105 50% embryo infectious doses (EID50) in 200 μl per bird. The inoculum was equally divided between the ocular, nasal, and oral routes and administered into the conjunctival sac, both nares, and caudal pharynx, respectively. The control birds were inoculated similarly with 200 μl PBS. At 2 and 4 days postchallenge (dpc), 2 birds from each group were euthanized to assess viral replication in tissues. The remaining 6 birds were then observed over 14 days. Birds were euthanized at the point at which they developed moderate clinical signs or at the end of the trial period on day 14. Moderate clinical signs were defined as inactivity, infrequent eating, huddling, and not reverting to normal behavior on stimulation.
Prior to challenge and at 7 dpc, 1 to 2 ml of blood was collected from the ulnar vein and placed into serum separator tubes. At euthanasia, blood was collected via cardiac puncture after anesthesia. The blood was then allowed to clot at room temperature for 30 minutes before being centrifuged at 1300 g for 10 minutes. The serum was removed and stored frozen at –20°C until testing. Immediately prior to testing, sera were heat inactivated at 56°C for 30 minutes.
At euthanasia, a range of tissue samples was collected from all birds. Samples for histology and immunohistochemistry were fixed in 10% neutral buffered formalin for 24 to 48 hours before processing. Tissues for histology included trachea, lung, air sac, heart, spleen, eyelid, thymus, cecal tonsils, bursa, pancreas, proventriculus, gizzard, duodenum, skin, skeletal muscle, and brain. For molecular analysis and virus isolation, approximately 0.5-cm3 samples of fresh brain, spleen, lung, kidney, and small intestine were collected into tubes containing 970 μl viral transport media (PBS, penicillin, streptomycin and gentamicin, and 1% bovine serum albumin) with 2 mm aluminum silicate beads (Biospec Products, Bartlesville, OK). These samples were stored frozen at –80°C until required. Tissues from control birds were only tested by histopathology and immunohistochemistry.
Serology
The titer of antibodies was measured by hemagglutination inhibition (HI) assay using standard methods. 32 The only variations from the OIE test method were the use of 8 hemagglutinating units (HAUs) of inactivated homologous antigen and the addition of 0.5% chicken red blood cells. Antigen was produced for all 4 viruses by propagation of each virus in the allantoic cavity of 10-day-old embronated eggs for 4 days, clarification of allantoic fluid by centrifugation, and inactivation by gamma-irradiation at 5 Mrads (50 kGray).
Histopathology and Immunohistochemistry
After fixation, tissues were routinely processed and 4-μm sections were stained with hematoxylin and eosin. For immunohistochemistry, antigen retrieval was performed using the DAKO PT LINK (Dako, Glostrup, Denmark) by heating the slides to 97°C for 30 minutes and then cooling to 70°C in the EnVision FLEX Target (Dako) high pH retrieval solution and washing for 5 minutes in Tris buffer. After this, endogenous peroxidases were quenched by the addition of a 3% H2O2 solution. The tissues were then incubated with a mouse monoclonal antibody (Q91-6, produced at The Australian Animal Health Laboratory) against the NDV nucleoprotein at a 1/800 dilution. The EnVision FLEX+ Mouse Linker (Dako) was used to amplify the antibody signal prior to the addition of the EnVision Flex/horseradish peroxidase (HRP) conjugate and aminoethylcarbazone (AEC) chromogen (Dako). Slides were then counterstained with Lillie-Mayer hematoxylin using 0.5% hematoxylin Certistain (Australian Biostain, Traralgon, Australia) and Scott’s tap water before mounting.
RNA Isolation and Quantitative Reverse Transcriptase Polymerase Chain Reaction
Tissue samples were homogenized using a bead beater for 30 seconds (FastPrep-24; M. P. Biomedicals, Irvine, CA) and then centrifuged at 13 000 rpm for 1 minute using a benchtop microcentrifuge. Subsequently, 100 μl of the tissue supernatant was combined with 260 μl MagMAX Lysis/Binding solution (Ambion, Life Technologies, Mulgrave, Australia). RNA was then extracted using the MagMAX-96 Viral RNA Isolation kit (Ambion) according to the manufacturer’s instructions. After isolation, RNA was quantified via NanoDrop (ND-1000; NanoDrop Technologies, Wilmington, DE) and used immediately with the remainder stored at –80°C. Quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) in triplicate was conducted using the AgPath-ID One-Step RT-PCR kit (Applied Biosystems, Victoria, Australia). Previously published NDV M gene primers and probe were multiplexed with eukaryotic 18S primers and probe to allow for quantitation of NDV RNA copy numbers (Table 2). 31 Reactions in a 20-μl volume were run using a 200-nM final concentration of each of the NDV M gene forward and reverse primers and 100 nM of the probe. The 18S forward and reverse primers were used at 50 nM, along with 200 nM of the probe. qPCR was performed with an ABI 7500 real-time machine (Applied Biosystems) using cycling conditions of 45°C for 10 minutes and 95°C for 10 minutes, followed by 45 cycles of 95°C for 15 seconds and 60°C for 45 seconds. Positive samples were defined as those with a cycle threshold (CT) less than 40. Relative quantification was conducted via standard curves generated by cloning the NDV M gene and chicken 18S segments into the pGEM-T easy vector (Promega, Madison, WI) using standard methods. 25 NDV M gene copy numbers were then expressed relative to 10 8 copies of 18S RNA.
Quantitative Polymerase Chain Reaction Primer and Probe Sequences.
NDV, Newcastle disease virus.
Virus Isolation
Virus isolation was only conducted on PCR-positive tissue samples. An aliquot (200 μl) of the tissue homogenate supernatant was inoculated into the allantoic sac of 10-day-old SPF embryonated chicken eggs in duplicate, which were then incubated at 37°C for 7 days. Allantoic fluid was then tested for hemagglutination activity (HA).
Sequencing
The genome of each of the virus isolates was sequenced via the MiSeq (Illumina, San Diego, CA) platform using a variation of a published method. 19 Briefly, viruses were grown in 9- to 11-day-old SPF eggs and then purified using a 20% and 50% discontinuous sucrose gradient and ultracentrifugation. RNA was then extracted from the purified allantoic fluid using the MagMAX-96 Viral RNA Isolation kit (Ambion) according to the manufacturer’s instructions. Reverse transcription was performed using a 17-mer primer combined with a random octamer (5′-GTT TCC CAG TAG GTC TCN NNN NNN N-3′) and Superscript III Reverse Transcriptase (Invitrogen, Carlsbad, CA). The complementary DNA (cDNA) was then amplified via PCR using the Expand High Fidelity kit (Roche, Indianapolis, IN) and products purified with the Wizard SV Gel and PCR Clean-Up System (Promega) before sequencing. Read mapping and de novo assembly were then used to assemble the genomes of the viruses (CLC Genomics Workbench 6.0.5; CLC, Inc, Aarhus, Denmark). Read mapping was performed for the Meredith/02, PR/98, and Texas GB viruses using previously published sequences AY935490, AY935497, and GU978777, whereas the alignment for the Herts 33/56 virus consisted of 2 independent read mappings and 2 de novo assemblies. For the purposes of this study, the ends of the sequences were not required and therefore not determined.
At the conclusion of the trial, partial resequencing of the virus isolates was performed on RNA extracted from tissue samples to ensure that the F protein cleavage site and the HN extension had not mutated during in vivo replication. The sequencing was performed by Sanger sequencing using published primers, 5,17 the Big Dye Terminator Reaction Kit v. 3.1 (Applied Biosystems), and an ABI 3130xl Genetic Analyzer (Applied Biosystems).
Statistical Analysis
Kaplan-Meier survival curves were generated using GraphPad Prism 5.02 for Windows (GraphPad Software, San Diego, CA). Survival curves were analyzed using the Mantel-Cox log-rank test with pairs of survival curves compared at a time. GraphPad Prism was also used to analyze serology and PCR data using Student t tests and the Mann-Whitney U test, respectively. Statistical significance was set at P < .05 for all analyses.
Results
Clinical Signs
All birds challenged with the Meredith/02 virus survived until the end of the trial (apart from the clinically healthy birds euthanized at 2 and 4 dpc according to the study design). However, at 3 dpc, the 5 birds were seen to be slightly less active in the morning, with 2 birds displaying very mild labored respiration. These clinical signs had resolved within 6 hours.
No clinical signs were observed in any of the birds inoculated with the PR/98 virus.
All birds inoculated with the exotic Herts 33/56 and Texas GB viruses showed moderate clinical signs as defined by a decreased response to stimulation, decreased feed intake, and huddling before the end of the trial period and were euthanized at that point. Eight birds in the Herts 33/56 group showed clinical signs at 2 dpc, including moderate depression, head tucking, and an unwillingness to move away when approached. All birds in this group were then euthanized at 2 dpc, including 2 healthy birds as per the study design. Two birds inoculated with the Texas GB isolate showed moderate depression, head tucking, and mild ataxia at 4 dpc and were euthanized at that time. At 5 dpc, the remainder of these birds (n = 4) displayed ataxia, with 3 birds showing an obvious head twitch. These birds were then euthanized at 5 dpc. No abnormalities were detected clinically in any of the control birds.
A survival curve comparing the euthanasia time points for birds infected with each of the different viruses is given in Figure 1, and further detail is provided in Supplemental Table S1. The survival times for the birds infected with the Australian viruses were significantly different from those infected with the Herts 33/56 (P = .0009) and Texas GB viruses (P = .0012).
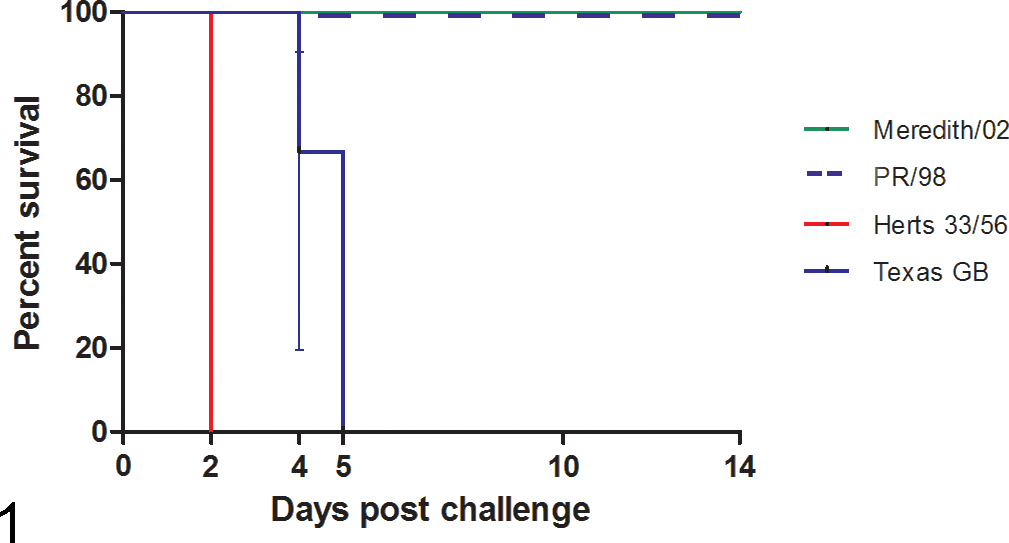
Kaplan-Meier survival curve of the birds infected with the 4 viruses over 14 days. Birds were euthanized at the point at which they showed moderate inactivity, infrequent eating, and an inability to revert to normal behavior on stimulation. The plot does not include the 4 birds in each group that were euthanized on 2 and 4 days postchallenge (dpc) according to the study design. Error bars indicate 95% confidence intervals.
Serology
All birds were negative for serum antibodies prior to challenge. All birds in the Meredith/02 and PR/98 groups had seroconverted by 7 dpc with significantly different (P = .01) geometric mean titers of 28.2 and 26.3, respectively. By 14 dpc, mean titers had reached 210.2 in the Meredith/02 group and 27.2 in the PR/98 group, again showing significant differences (P = .0004) between the virus strains. None of the birds inoculated with the Herts 33/56 virus had seroconverted due to the early euthanasia time point at 2 dpc. Low levels of antibodies (22 and 24) were detected at 5 dpc in 2 birds inoculated with the Texas GB virus, with the rest of the birds being euthanized without detectable antibodies.
Gross Pathology
Gross pathological findings were restricted to the birds infected with the exotic isolates of NDV. In birds infected with the Herts 33/56 isolate, the most consistent findings were hemorrhagic cecal tonsils (10/10), hemorrhage at the junction of the proventriculus and esophagus (4/10), and mild splenic enlargement with pale mottling (4/10). Birds inoculated with the Texas GB isolate displayed moderate cloudiness of the air sacs (2 birds on 4 dpc and 2 birds on 5 dpc), along with 1 bird with mild splenomegaly on 5 dpc. There were no gross lesions detected in birds inoculated with the Australian NDVs or in control birds.
Histopathology
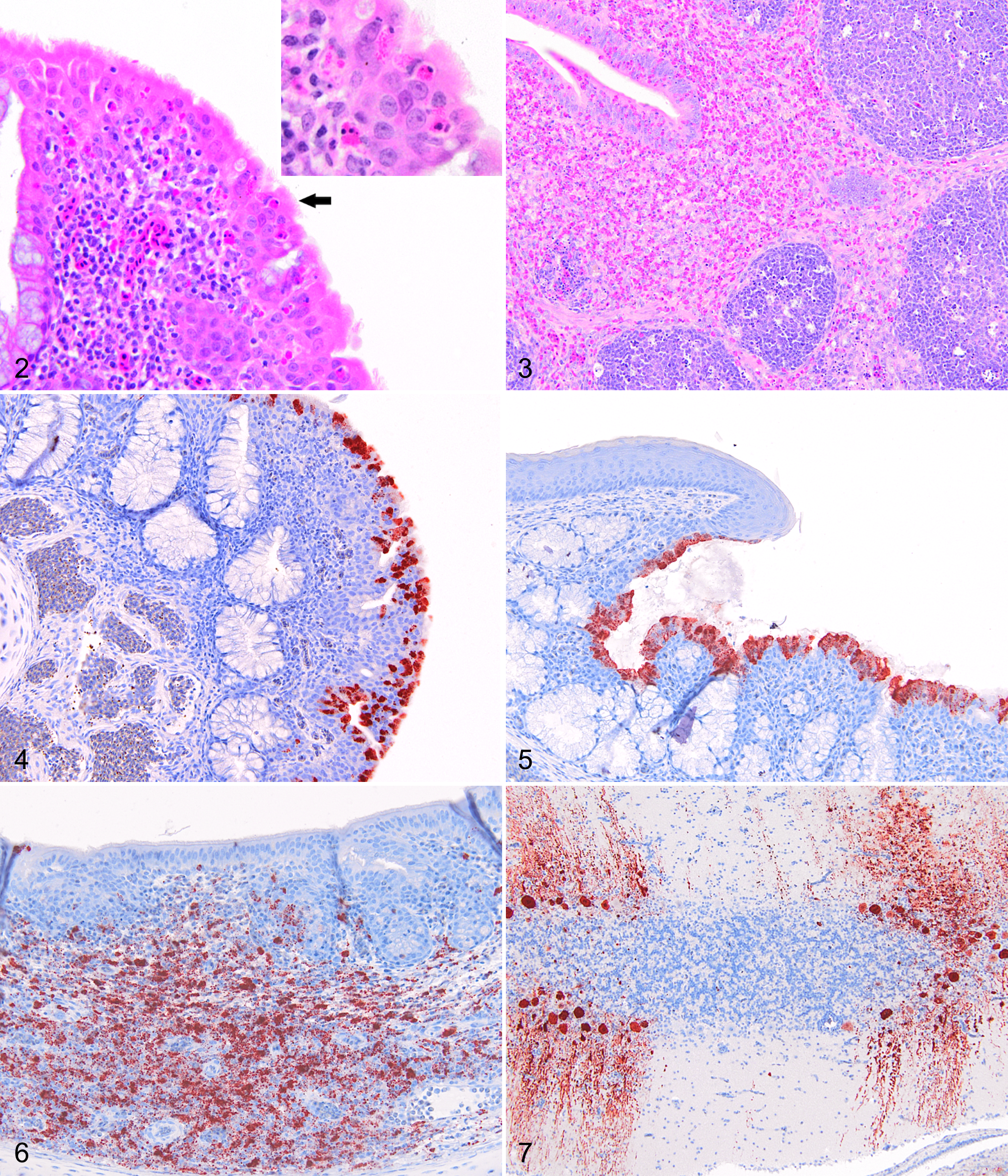
The histological lesions noted in birds infected with the Meredith/02 and PR/98 isolates of NDV were minimal. The lesions involved mild infiltrates of heterophils in the epithelium and submucosa around sites of inoculation, that is, the nasal turbinates (Fig. 2), trachea, and conjunctiva. These features were observed in the Meredith/02 group within all birds euthanized on 2 and 4 dpc and in 2 birds euthanized on 14 dpc. Only the 2 birds euthanized on 2 dpc in the PR/98 groups showed mild heterophilic infiltrates in the nasal turbinates and conjunctiva.
Eight-week-old White Leghorn chickens infected with Newcastle disease virus (NDV) isolates Meredith/02 (virulent), PR/98 (avirulent), Herts 33/56 (virulent), and Texas GB (virulent).
In birds infected with the Herts 33/56 virus, histological lesions were predominantly seen in sites of lymphoid tissue accumulation such as the thymus (10/10), conjunctiva (10/10), spleen (9/9), proventriculus (at the esophageal and gizzard junctions, 10/10), cecal tonsils (10/10), bursa (9/10), and nasal turbinates (8/10). In these regions, there were moderate to marked necrosis and apoptosis of lymphocytes and macrophages, with edema, fibrin deposition, and heterophil infiltration (Fig. 3).
In birds inoculated with the Texas GB virus, there was moderate loss of cilia throughout the tracheas, with occasional necrosis of epithelial cells and moderate numbers of inflammatory cells, predominantly heterophils (8/10). The other significant lesions were seen within the central nervous system and comprised perivascular lymphocyte cuffing and glial nodule formation throughout the cerebrum, cerebellum, and brainstem (8/10). Occasional neuronal necrosis was also seen, most commonly in the cerebellum or brainstem. These lesions were seen in all birds except those euthanized on 2 dpc. There were no histological lesions detected in any of the control birds.
Immunohistochemistry
No immunohistochemical staining was detected in any tissues from control birds. Antigen was detected in birds infected with all 4 viruses, but the distribution and quantity of the staining varied between the different virus groups (Table 3). Antigen staining was present in both the cytoplasm and the nucleus of cells as fine- to moderate-sized granules. At euthanasia of the Herts 33/56 birds, antigen was detected in the brain (9/10), nasal turbinates (10/10), eyelids (10/10), larynx/trachea (4/8), lung (10/10), heart (10/10), spleen (9/9), kidney (7/10), and cecal tonsils (10/10). In the Texas GB birds, antigen was also detected in the brain (8/10), nasal turbinates (10/10), eyelids (8/9), larynx/trachea (9/10), lung (8/10), heart (8/10), spleen (10/10), kidney (8/10), and cecal tonsils (8/10). The quantity of antigen was scored on a 3-point scale (Table 3). In contrast, birds inoculated with the Meredith/02 virus showed antigen predominantly restricted to sites of inoculation, with staining seen in the nasal turbinates (Fig. 4; 7/10), eyelid (4/10), larynx/trachea (3/8), spleen (4/4), and cecal tonsils (3/9). Birds inoculated with the PR/98 virus also showed antigen staining in the nasal turbinates (Fig. 5; 4/10), eyelid (2/10), and larynx/trachea (2/10) only on 2 and 4 dpc. There was also antigen staining in the spleen of 1 bird euthanized at 14 dpc. The antigen detected in the spleen and cecal tonsils at 2 and 4 dpc in the Meredith/02 birds indicates a greater degree of systemic replication than with the PR/98 virus.
Immunohistochemical Findings in Chickens Following Challenge With Difference Isolates of Newcastle Disease Virus.
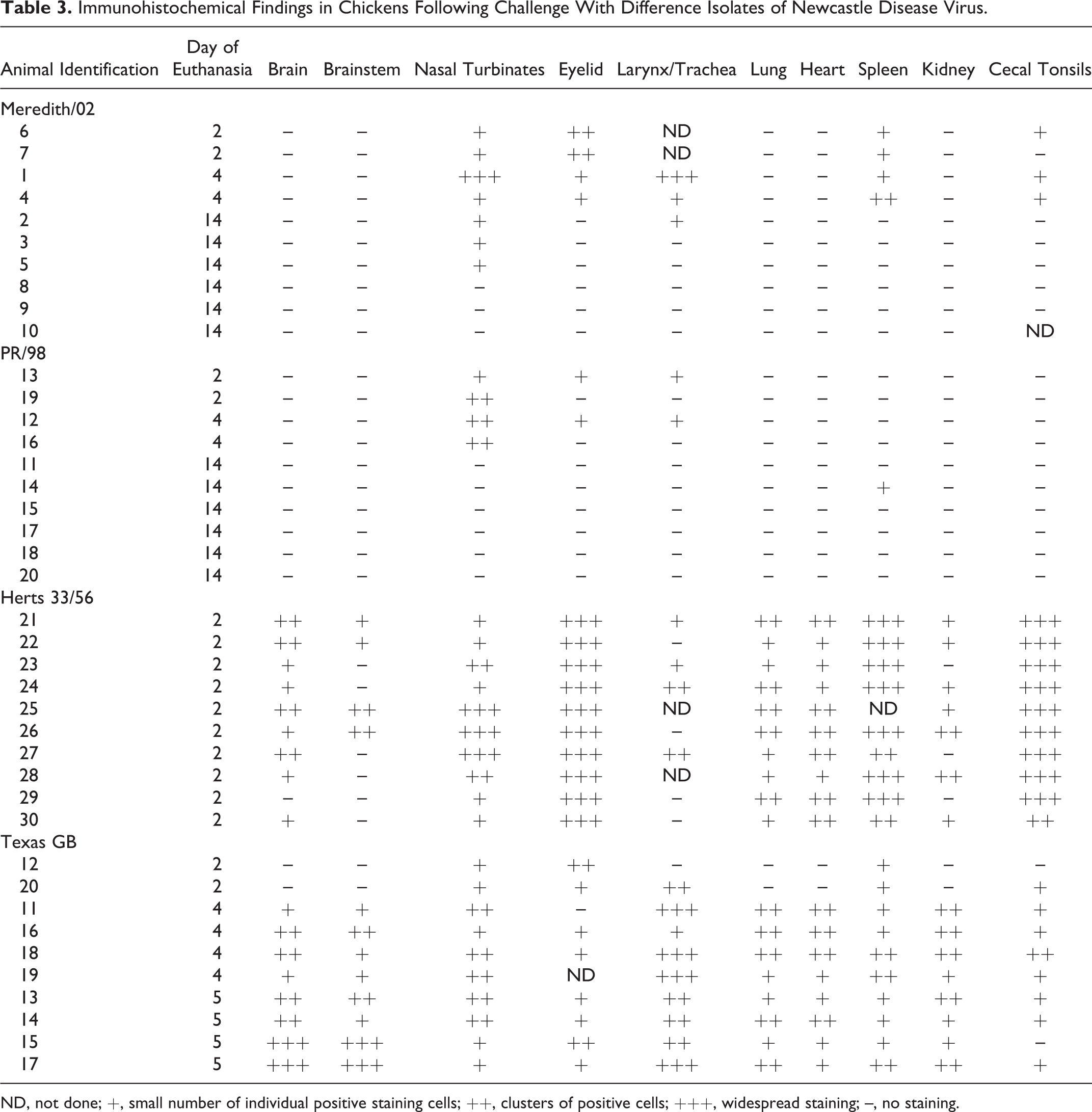
ND, not done; +, small number of individual positive staining cells; ++, clusters of positive cells; +++, widespread staining; –, no staining.
The cellular tropism also varied between viruses, with birds infected with the Herts 33/56 viruses showing a stronger lymphoid tissue tropism than both the Australian viruses and the Texas GB virus. The birds infected with the Herts 33/56 virus showed a predominance of staining in lymphoid tissue within mononuclear cells in both lymphocytes and macrophages (Fig. 6). The positive staining lymphoid tissue was found throughout the upper respiratory tract and gastrointestinal tract. Antigen was also detected in the brain in both neurons and glial cells. The most notable difference between the Texas GB virus and the other viruses was the large amount of staining in the central nervous system, in which cell bodies and processes of large neurons were stained, particularly Purkinje cells in the cerebellum (Fig. 7) and brainstem nuclei. These birds also displayed greater staining of respiratory epithelial cells in the upper respiratory tract compared with the Herts 33/56–infected birds (Table 4).
Cellular Tropism of Newcastle Disease Virus Within the Nasal Turbinates by Immunohistochemistry.

+, small number of individual positive staining cells; ++, clusters of positive cells; +++, widespread staining; –, no staining.
Polymerase Chain Reaction
Five tissue samples (brain, spleen, lung, kidney, small intestine) from each of the birds were tested for NDV RNA via qRT-PCR (Table 5). All 5 tissue samples from all 10 birds infected with the Herts 33/56 virus showed detectable levels of NDV RNA at the time of euthanasia (2 dpc). Most tissue samples collected from birds infected with the Texas GB virus at 2, 4, and 5 dpc were positive for NDV RNA (42/50 samples). Initially, only the spleen samples from the 2 birds euthanized on 2 dpc were positive, but all tissues from the remaining birds were positive on both 4 dpc and 5 dpc. A total of 15 samples were NDV RNA positive from the 10 birds inoculated with the Meredith/02 virus, including all 10 spleen samples with 1 positive kidney sample (4 dpc), 3 positive lung samples (2 and 4 dpc), and 1 positive duodenum sample (4 dpc) . Only 5 tissues were viral RNA positive from the birds infected with the PR/98 virus, consisting of 2 spleen samples from 2 and 4 dpc, 2 duodenum samples from 2 and 14 dpc, and 1 cerebrum sample from 14 dpc.
Distribution of Newcastle Disease Virus in Tissues of Chickens, Based on PCR and Virus Isolation.a
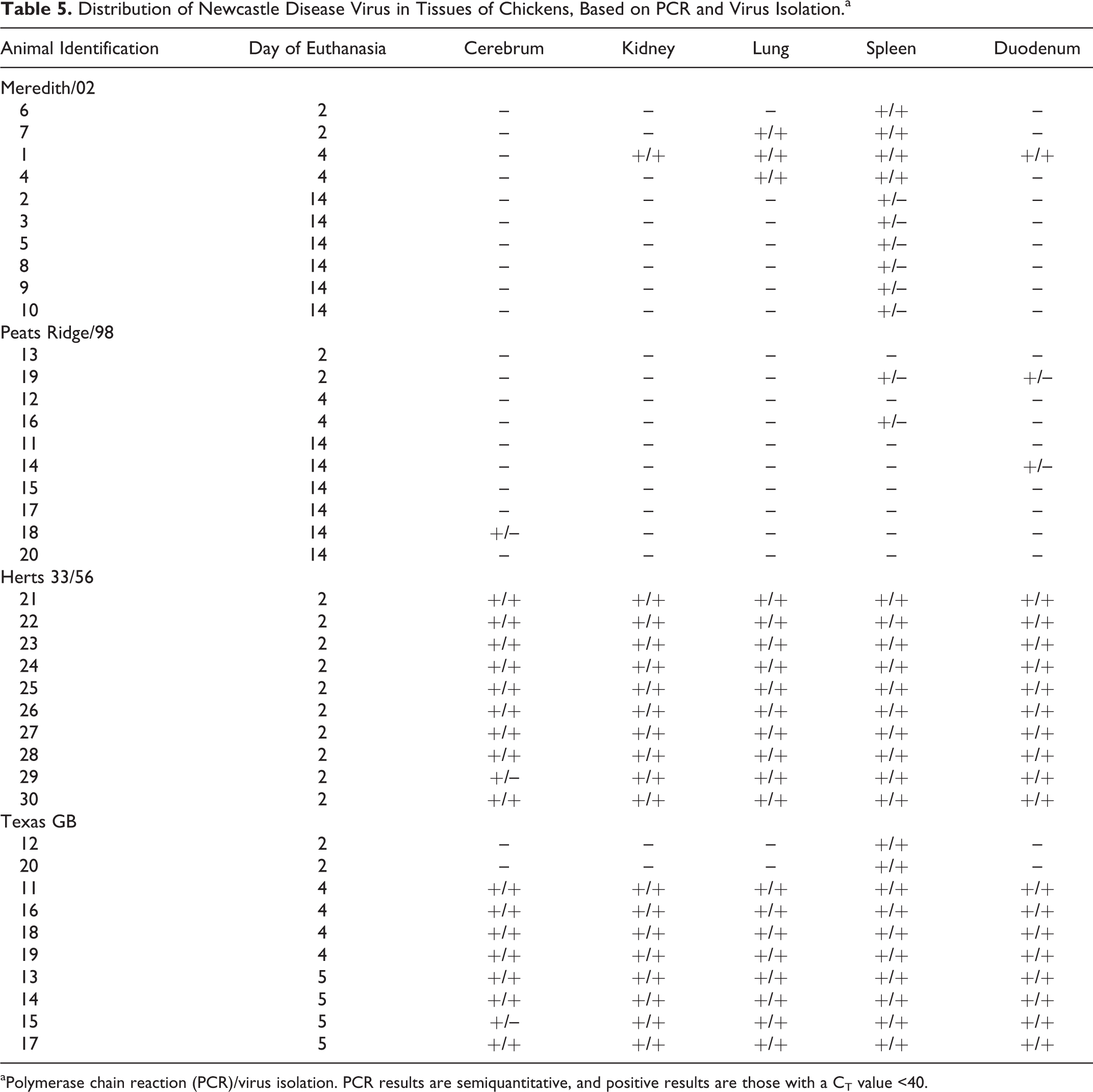
aPolymerase chain reaction (PCR)/virus isolation. PCR results are semiquantitative, and positive results are those with a CT value <40.
When comparing viral RNA loads between the viruses, 2 dpc was chosen as the comparative time point because at the point of euthanasia at 14 dpc, all of the birds infected with the exotic velogenic viruses had already been euthanized. Comparisons of RNA loads in the spleen using the Mann-Whitney U test showed that compared with each other, all 4 viruses had significantly different copy numbers at 2 dpc when normalized to 18S (P < .05; Fig. 8).
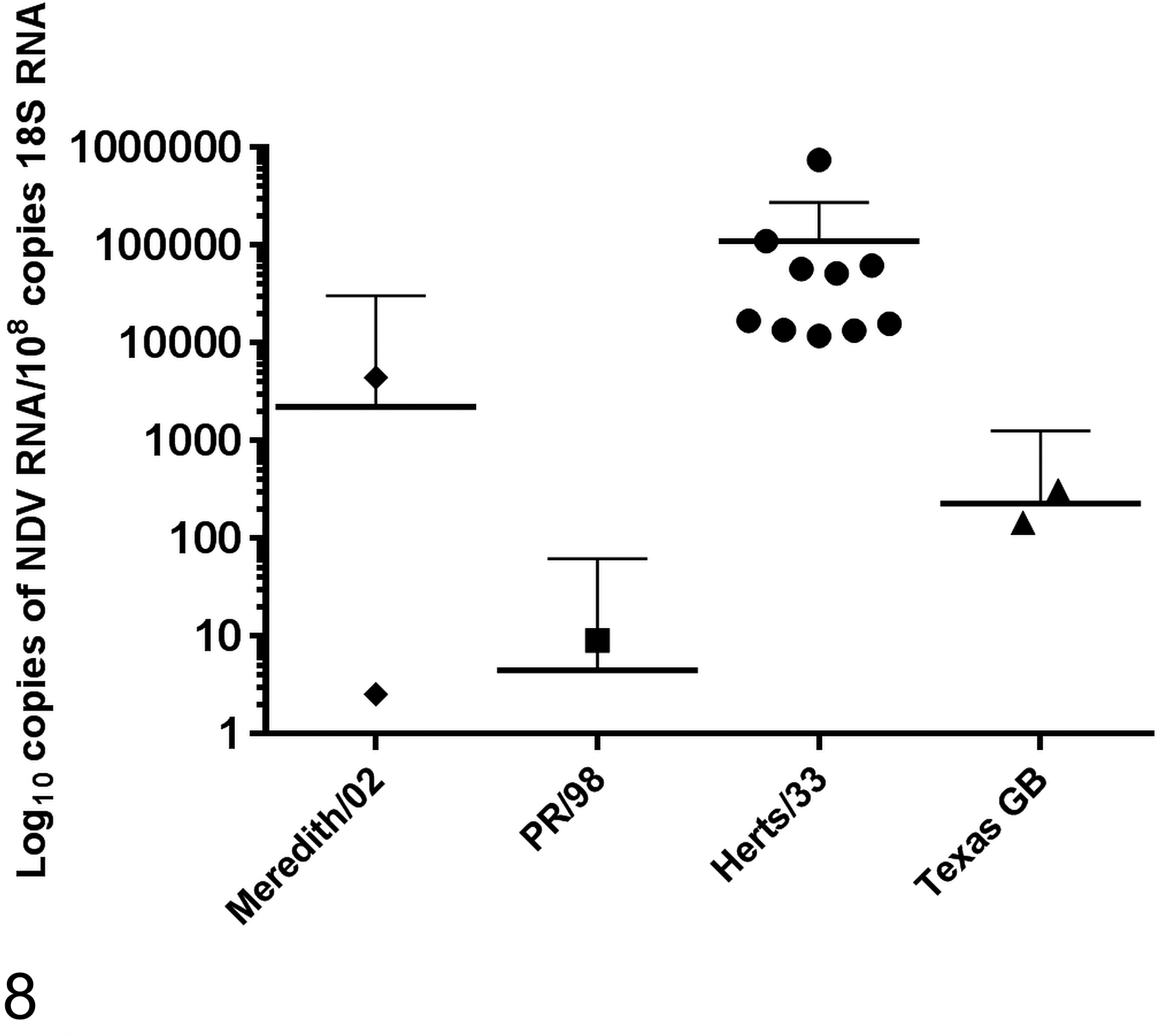
Newcastle disease virus (NDV) RNA copy numbers in spleen samples at day 2 after challenge with different isolates of NDV. The graph shows the means with 95% confidence intervals. All comparisons between viruses were significantly different (P < .05, Mann-Whitney U test).
Virus Isolation
Virus isolation was attempted from 5 different tissues (brain, kidney, lung, spleen, and small intestine) when PCR results were positive (Table 5). Virus was reisolated from all but 1 PCR-positive tissue in 1 bird in both the Herts 33/56 and Texas GB virus groups. In both groups, the PCR-positive, virus isolation–negative sample was cerebrum.
In birds exposed to the Meredith/02 NDV, virus was only recovered from birds euthanized at 2 and 4 dpc and not from those euthanized at 14 dpc. No virus was isolated from birds inoculated with the PR/98 virus at any time point.
Sequencing
Whole-genome sequencing of the virus isolates showed that the viruses had the same F cleavage site sequences as previously reported (Table 1). 6,17,23
Partial sequencing of the F protein and HN extension from samples collected from the Meredith/02, Herts 33/56, and Texas GB inoculated birds after the infection trial were consistent with the sequence results of the virus inoculum, indicating that these regions had not mutated at these sites during replication in the birds. Due to the low virus titers in the birds infected with the PR/98 virus, sequencing was not successful on samples from these birds.
Discussion
This study investigated the pathogenicity of the atypical Australian NDV Meredith/02 by comparison with an avirulent Australian virus and 2 exotic velogenic viruses. The atypical Australian virus (Meredith/02) possessed a virulent F protein cleavage site sequence according to the OIE definition but did not induce severe clinical signs, which is consistent with the field observations at the time of its isolation. 30 This is unusual given that NDVs with multiple basic amino acids at the F protein cleavage site and a phenylalanine at position 117 are usually associated with a virulent phenotype. 6
While birds exposed to the Meredith/02 virus appeared very mildly depressed for a short period of time on 3 dpc, they all completely recovered within hours. The PR/98 virus was typical for most avirulent NDVs, in that no clinical signs were observed in inoculated birds. As expected, the velogenic viruses Herts 33/56 and Texas GB induced severe clinical signs, leading to the euthanasia of all birds by 2 and 5 dpc, respectively. The clinical signs seen in the birds inoculated with Herts 33/56 and Texas GB viruses were typical for viscerotropic and neurotropic viruses and were similar to those previously reported. 2,12
This work has confirmed the observation made by Susta et al 27 that some Australian isolates of NDV obtained from 1998 to 2002 are less pathogenic in an experimental setting than other viruses with similar virulent cleavage site motifs. This is despite the Meredith/02 virus and the APMV-1/chicken/Australia/9809-19-1107/1998 viruses having the same virulent F protein cleavage site of 12 RRQRRF117 and high ICPIs of 1.61 and 1.88, respectively. The Meredith/02 virus in particular appears even less pathogenic than the APMV-1/chicken/Australia/9809-19-1107/1998 virus. However, previously, the Australian NDV isolates responsible for the 1998 to 2002 outbreaks, including the Meredith/02 virus, had been described as velogens, based on F protein cleavage site sequence data and ICPI values. 18,27 Clearly, the cleavage site sequence and ICPI data alone are not sufficient to classify pathotype, although currently these are the best measures available to predict the behavior of an NDV.
Phylogenetic analysis of Australian NDVs detected during the 1998 to 2002 outbreaks placed them within class II, genotype I, of which the majority of isolates are avirulent and found in wild birds. 8 The presence of a 9–amino acid extension to the HN protein identifies these viruses as uniquely Australian. 17,29 The Meredith/02 isolate was shown to cluster with the other 1998 to 2002 isolates, although it was slightly more divergent from others in the group. 17 This slight increase in phylogenetic distance may account for the difference in pathogenicity between the 9809-19-1107 virus reported by Susta et al 27 and the Meredith/02 virus reported here. In addition, Susta et al 27 used slightly younger 4-week-old chickens, which may have led to increased susceptibility to disease.
The presence of a virulent F protein cleavage site enables the NDV to be cleaved by proteases that are widespread throughout numerous cell types. This allows the virus to enter and replicate in a greater range of cells and organ systems than those with an avirulent cleavage site motif. 13,21 Therefore, in accordance with its cleavage site motif, it would be expected that the Meredith/02 virus would also have a wide antigen distribution. The distribution and concentration of virus as measured by immunohistochemistry, PCR, and virus isolation indicated that while the virus was able to replicate systemically in the spleen of all birds and in the kidney, lung, and duodenum of 3 of 10 birds, live virus was cleared from all tissues by 14 dpc. Compared with the virulent exotic viruses, the virus load of the Meredith/02 virus was markedly reduced, corresponding with fewer and less severe histopathological lesions, as well as minimal clinical signs in the inoculated birds. The antigen was primarily restricted to the mucosa at the sites of inoculation. In comparison, the viscerotropic Herts 33/56 virus preferentially targeted lymphoid tissue early in the course of infection. This indicates that the presence of a virulent F protein cleavage site in the Meredith/02 virus, while allowing for some systemic viral replication, was not associated with the high viral loads in tissues associated with typical velogenic viruses.
In general, the pathogenicity of a virus is determined by a number of factors, including the route of exposure to the virus, the immune status and immune response of the birds, and the ability of the virus to replicate and disseminate throughout the host tissues. 3 Increased virus replication and a greater inflammatory response within lymphoid tissues have been associated with increased pathogenicity of velogenic NDVs. 16 The results of our study indicate that the low pathogenicity of the Meredith/02 virus must be attributed to regions of the viral genome other than the F protein cleavage site, given that the cleavage site contains a virulent motif. Studies using reverse genetics have also indicated that regions of the NDV genome other than the F protein cleavage site may also contribute significantly to pathogenicity. 9,11,22 For example, the viral replication complex has been associated with the decreased pathogenicity of the pigeon paramyxovirus for chickens, despite its virulent cleavage site motif. 11
Overall, this experimental work has confirmed the field observations that, despite containing a virulent F protein cleavage site as defined by the OIE, the Meredith/02 isolate of NDV is not highly pathogenic for chickens. Clinical signs, pathological lesions, and antigen distribution in the experimentally infected birds were instead consistent with a lentogenic or mesogenic virus.
The risk posed by these Australian viruses is unclear. This evolution of an avirulent to virulent virus is unusual and has only been described once before in Ireland. 1 While the PR/98 virus has been shown to only require 2 nucleotide changes to become pathogenic, it is uncertain whether the Meredith/02 virus is also a significant risk to the poultry industry. While the Meredith/02 virus displayed minimal pathogenicity in the field and very limited pathogenicity in an experimental setting, there is still a possibility that this virus may increase in pathogenicity given appropriate circumstances. It is possible that the Meredith/02 was not yet well adapted to chickens. High pathogenicity may evolve with intensive production systems, such as most commercial chicken operations in Australia, due to the large numbers of birds held in such facilities allowing numerous passages of the virus. In addition, concurrent infection with agents such infectious bronchitis virus or infectious bursal disease virus may induce immunosuppression and allow for virus evolution. Immune pressure induced by vaccination may also contribute to an increased NDV mutation rate and result in increased pathogenicity. However, the genetic changes necessary for this transformation are unknown. Conversely, this virus may be a stable genetic type that may not continue evolving greater pathogenicity and therefore have a low risk for commercial poultry. Currently, we are not able to assess this on the genetic information that we have, but knowledge of the impacts of potential mutations would be important for risk assessment and outbreak response. At the time of its detection, a conservative approach was taken and the affected flocks were depopulated. However, this approach may not be necessary if it can be demonstrated that such viruses have little risk of further evolution toward higher pathogenicity.
The Australian NDV, Meredith/02, while containing a virulent F protein cleavage site (RRQRRF) consistent with a virulent virus, was only mildly pathogenic in chickens in an experimental setting. Viral replication primarily occurred in sites of inoculation, and there were no mortalities associated with the viral infection. The decreased pathogenicity of the virus may therefore be associated with regions of the genome other than the F protein cleavage site.
Footnotes
Acknowledgements
We are very grateful for the assistance of the animal studies team in housing and caring for the birds used in the study. We also acknowledge the help of Ina Smith, Adam Foord, Kelly Davies, Tyrone McDonald, and Vicky Stevens with molecular work, including sequencing. Thank you also to Paul Selleck for valuable conversations about Australian Newcastle disease throughout the course of this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Australian Poultry Cooperative Research Council.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
