Abstract
Newcastle disease is a severe threat to the poultry industry and is caused by Newcastle disease virus, a member of the genus Avulavirus, family Paramyxoviridae. The virus is rapidly evolving, and several new genotypes have been discovered in the past few years. Characterization of these strains is important to evaluate field changes, anticipate new outbreaks, and develop adequate control measures. Three Newcastle disease isolates (APMV-1/duck/Vietnam, Long Bien/78/2002, APMV-1/chicken/Australia/9809-19-1107/1998, and APMV-1/double-crested cormorant/USA, Nevada/19529-04/2005) from recent outbreaks were investigated via clinicopathological assessment, immunohistochemistry (IHC), in situ hybridization, virus isolation, and serology in experimentally infected 4-week-old chickens. Phylogenetic studies showed that Australia isolate belongs to class II genotype I, Long Bien to class II genotype VIId, and Nevada cormorant to class II genotype V. Even though all 3 viruses had a virulent fusion protein cleavage site and ICPI values greater than 1.5, they all differed in their ability to cause clinical signs, in their lesions, and in their viral distribution in body tissues. The Long Bien isolate showed the most severe clinicopathological picture and the most widespread viral distribution. The Australia and Nevada cormorant isolates had a milder pathological phenotype, with viral replication restricted to only a few organs. The variability in clinicopathological characteristics despite the similarity in ICPI suggests that full clinicopathological assessment is necessary to fully characterize new isolates and that there are differences in pathogenesis among viruses of different genotypes.
Keywords
Newcastle disease virus (NDV), synonymous with avian paramyxovirus-1 (APMV-1), is the causative agent of Newcastle disease (ND), an infection that threatens poultry worldwide. 1 –3 The poultry industry loses millions of dollars annually from morbidity, mortality, and trade losses due to ND. 3 NDV belongs to the Mononegavirales order, Paramyxoviridae family, Paramyxovirinae subfamily, and Avulavirus genus. The genome is negative sense RNA of approximately 15,200 bases, and it contains 6 genes that encode an equal number of structural proteins, which are from 3′ to 5′: nucleoprotein (NP), phosphoprotein (P), matrix (M), fusion (F), hemagglutinin–neuraminidase (HN), and polymerase (L). Although all ND viruses belong to a single serotype, APMV-1 serotype, isolates differ in virulence in poultry, which translates into a wide range of clinical signs, often making it challenging to recognize NDV as the cause of the clinical problem in the field. 1 Historically, strains were classified according to the time to death after inoculation of embryonating eggs, with those strains that caused death the fastest (and therefore more virulent) termed velogens and those in which the embryos survived for much longer periods of time termed lentogens. Those strains falling in between were called mesogens. 1,31 Today, classification of the different strains is carried out by 2 internationally recognized methods. The intracerebral pathogenicity index (ICPI) involves inoculating 1-day-old chicks intracerebrally and scoring birds as normal, sick, or dead over a period of 8 days, to result in a score from 0.0 to 2.0. Those strains scoring greater than 0.7 are considered “notifiable” to the international community through the World Organization for Animal Health, and any strain with a score greater than 1.5 is considered a velogen. 2,31 The second internationally recognized test is to sequence the F protein cleavage site to evaluate the presence of multiple basic amino acids and phenylalanine, which is considered an indication of virulence and therefore notifiable. 11,29
Outbreaks of ND are reported frequently from many parts of the world, with the most severe disease being caused by virulent strains. The USA is considered free of virulent NDV strains, but occasionally pathogenic strains enter through wild birds or informal (illegal) trade. The last outbreak of ND occurred in California in 2002–2003 and resulted in depopulation of more than 3 million birds and containment costs exceeding US$160 million. 37
There is some correlation between viral virulence and phylogenetic analysis of virus evolution. According to phylogenetic distances, often carried out on the sequence of the F gene, NDV isolates are divided in 2 classes (I and II), and class II is further divided into 9 genotypes. 26,27 Comparison of phylogenetic analysis with biological characteristics of the NDV isolates demonstrates that viruses of class I are mainly composed of avirulent strains, usually circulating among wild birds, whereas viruses of class II are more frequently virulent 8,26 and mainly isolated from poultry. Within class II, most outbreaks worldwide are caused by viruses of genotypes V, VI, and VII, even though occasional outbreaks caused by viruses from other genotypes do occur. Particularly important are viruses of genotype VIId, which lately have caused several outbreaks in China, Vietnam, Indonesia, Pakistan, and Africa. 17,27,28
NDV is a continuously evolving virus with evidence of accelerated evolution of the virulent strains, 26 and there are reports of increased pathogenicity, outbreaks in vaccinated animals, and increased host range. 15,25,26,28,30,38 Therefore, it is important to conduct clinicopathological characterization of new strains that are isolated during the course of outbreaks worldwide to determine how the viruses are changing. Full characterization of the clinical disease and pathogenesis in poultry could also affect the diagnostic aspects of the disease and implementation of control measures. Characterization of the genomic sequences, genotyping of the new isolates, and identification of their nearest ancestors could help identify the source of new outbreaks.
In this study, 3 different strains of NDV from recent outbreaks were characterized through sequencing, phylogenetic analysis, ICPI, and clinicopathological assessment. The latter was performed by inoculating 4-week-old chickens via a natural route (eye drop instillation). The resulting disease was characterized through virus isolation (VI), serology, and clinicopathological observations, including histology, immunohistochemistry (IHC), and in situ hybridization in multiple body tissues.
The first virus, APMV-1/duck/Vietnam, Long Bien/78/2002, in this article referred as Long Bien, was initially isolated from ducks in a Vietnam market during 2002. The second virus, APMV-1/chicken/Australia/9809-19-1107/1998, here referred to as Australia, was isolated during a large outbreak in Australia that occurred from 1998 to 2000. This virus and other isolates from this same outbreak were unusual in that they are the only known virulent representatives of class II, genotype I. There is evidence 12,17,39 that the origin of these viruses can be traced back to low-virulence NDV circulating in waterfowl just prior to the outbreak. The third virus, APMV-1/double-crested cormorant/USA, Nevada/19529-04/2005, and here referred as Nevada cormorant, is an isolate responsible for significant morbidity and mortality in cormorants in the state of Nevada in 2005 and in other US states in 2008. 36
Methods
Viruses
APMV-1/duck/Vietnam, Long Bien /78/2002 (Long Bien), APMV-1/chicken/Australia/9809-19-1107/1998 (Australia), and APMV-1/double-crested cormorant/USA, Nevada/19529-04/2005 (Nevada cormorant) were obtained from the Southeast Poultry Research Laboratory (SEPRL) NDV repository. Each virus was propagated by 1 passage in 9- to 10-day-old embryonated chicken eggs inoculated by the chorioallantoic route.
Eggs and Chickens
The source of embryonating chicken eggs and chickens was the SEPRL SPF White Leghorn flock. Birds were housed in negative pressure isolators under biosafety level-3–enhanced conditions at SEPRL and were provided food and water ad libitum.
Intracerebral Pathogenicity Index Test
To characterize the viruses, ICPI was performed using standard protocols. 2 Briefly, chickens were inoculated at 1 day of age with 0.1 ml of a 1:10 dilution of infective allantoic fluid. Chicks were monitored daily and scored as normal, sick or paralyzed, and dead to compile a score for the 8-day observation period. 2
Clinicopathological Assessment in Chickens
Three groups of ten 4-week-old SPF White Leghorn chickens were inoculated bilaterally in the conjunctival sac with 0.1 ml of viral inoculum. Phosphate-buffered saline was used for the noninfected control birds. The target dose of inoculum was 105.0 50% embryo infectious doses (EID50). The actual infectious doses as determined by back titration in embryonating chicken eggs were 105.5 EID50 for Long Bien, 104.5 EID50 for Australia, and 105.3 EID50 for Nevada cormorant. The birds were monitored clinically every day, and 2 birds of each group were euthanatized after taking oropharyngeal and cloacal swabs at 2, 5, 10, and 14 days post inoculation (dpi). Birds whose condition became critical were euthanatized regardless of the scheduled sampling day.
Blood samples for serology were collected at 10 and 14 dpi. Tissues (eyelid, spleen, bursa of Fabricius, thymus, Harderian gland, proventriculus, small intestine, cecal tonsils, large intestine, air sac, trachea, lung, heart, esophagus, pharynx, crop, brain, liver, pancreas, kidney, comb, head of left femur including bone marrow, and nasal turbinate) were collected and fixed by immersion in 10% neutral buffered formalin for approximately 52 hours. The sections of femur and turbinate were decalcified in 5% formic acid for 3–4 hours. All sampled tissues were routinely processed into paraffin, and 3-μm sections were cut for hematoxylin and eosin staining, IHC, and in situ hybridization (ISH).
Immunohistochemistry
All sampled tissues were examined by IHC to detect viral nucleoprotein. Briefly, after deparaffinization, tissue sections were subjected to antigen retrieval by microwaving for 20 minutes at minimum power in Vector antigen unmasking solution (Vector Laboratories, Burlingame, CA) followed by application of universal blocking reagent (Biogenex, San Ramon, CA) as recommended by the manufacturer. The primary antibody, made in rabbit, was raised against a synthetic nucleoprotein peptide (TAYETADESETRRIC) and used at 1:8000 dilution. The detection system was an alkaline phosphatase–labeled polymer specific for the Fc portion of rabbit immunoglobulin (LabVision, Fremont, CA). Chromogen was a naphthol-based dye (Fast Red, Dako, Carpinteria, CA). Sections were counterstained lightly with hematoxylin and cover-slipped with Permount for a permanent record.
In Situ Hybridization
Selected tissue sections from the NDV-infected birds were probed with a negative-sense digoxigenin-labeled 850 base riboprobe representing the 5′ end of the matrix gene of the NDV Fontana (CA1083) as previously described. 7 The matrix gene from the Fontana strain was cloned into pCRII transcription vector (Invitrogen, Carlsbad, CA), and anti-sense riboprobes were generated using RNA polymerase in the presence of labeled nucleotides. This probe was used for hybridization with all the 3 strains in this study, because sequence information indicated a very high grade of identity (Long Bien 92%; Nevada cormorant 89%; Australia 88%) with the probed strains (NCBI accession no. AY562988).
For hybridization, sections from selected blocks were deparaffinized, rehydrated, digested with 100 μg/ml proteinase K for 15 minutes, and hybridized overnight at 42°C with approximately 20 ng of probe in standard sodium citrate, 50% formamide, 5% blocking reagent (Boehringer Mannheim, Indianapolis, IN), 1% N-lauroylsarcosine, and 0.02% sodium dodecyl sulfate. Following stringent washes, bound probes were visualized by the addition of anti-digoxigenin alkaline phosphatase and the chromogen/substrate nitroblue tetrazolium and 5-bromo-4-chloro-3-indolylphosphate.
Virus Isolation and Titration of Swabs
For those birds used in the clinicopathological assessment experiment, oral and cloacal swabs were obtained from each bird immediately prior to euthanasia and placed in separate tubes containing 1.5 ml of brain–heart infusion broth (BHI) with antibiotics (2000 U of penicillin G/ml, 200 μg of gentamicin sulfate/ml, and 4 μg of amphotericin B/ml; Sigma Chemical Co., St. Louis, MO). Swab sample tubes were centrifuged at 1000 × g for 20 minutes, and the supernatant was removed for virus isolation and titration. Virus infectivity titers were calculated from the result of inoculation of 9- to 10-day old SPF embryonating chicken eggs with serial 10-fold dilutions in BHI containing antibiotics (100 U of penicillin G/ml and 50 μg of gentamicin sulfate/ml). NDV in infected dead or surviving embryos was identified by hemagglutination (HA) activity in amnio-allantoic fluid harvested from chilled eggs. NDV was confirmed in HA-positive samples by hemagglutination inhibition (HI) test with NDV-specific antiserum.
Serology
The HA and HI tests were conducted by conventional microtiter methods with serum separated from the blood samples taken at day 10 and 14 dpi. Four HA units per 25 μ1 of β-propiolactone-inactivated NDV LaSota was used as test antigen in completing the HI test.
RNA Isolation, Polymerase Chain Reaction Amplification, and Sequencing
RNA was extracted from allantoic fluids using Trizol LS (Invitrogen, Carlsbad, CA) according to manufacturer’s instructions. Briefly, 750 μl of Trizol LS reagent was added to 250 μl of allantoic fluid, vortexed, and incubated at room temperature for 7 minutes. The RNA was separated into the aqueous phase with the addition of 200 μl of chloroform, precipitated with isopropanol, and then centrifuged to pellet the RNA. After 1 wash with 70% ethanol, RNA was dried and resuspended in RNase-free water. Polymerase chain reaction (PCR) amplification of the RNA was performed using the Qiagen One Step reverse transcription (RT)–PCR kit (Qiagen, Valencia, CA). Amplified products were separated on a 1% agarose gel, the bands excised and eluted using the QIAquick Gel extraction kit (Qiagen), and the samples quantified using a standard spectrophotometer. All sequencing reactions were performed with fluorescent dideoxynucleotide terminators in an automated sequencer (ABI 3700 automated sequencer; Applied Biosystems Inc., Foster City, CA). Nucleotide sequence editing and analyses were conducted with the LaserGene sequence analysis software package (LaserGene, version 5.07; DNAStar, Inc., Madison, WI) using the full-length genome positions from the NDV LaSota vaccine strain complete genome. The regions sequenced were a 374 base pair partial F gene (positions 4554–4917), and the complete coding region for the F gene (positions 4544–6205), as previously described. 18 All RNA and DNA procedures were performed in biological safety cabinets, and standard anticontamination protocols were followed and RNase-free reagents used at all times to minimize contamination.
Sequence Alignment and Tree Construction
The coding sequence of 51 NDV full fusion genes were aligned using ClustalW as implemented in the BioEdit Sequence Alignment Editor (http://www.mbio.ncsu.edu/BioEdit/bioedit.html) and edited manually based on the amino acid sequence. Maximum likelihood (ML) phylogenetic analysis with bootstrap values for n = 100 replicates was performed using PHYML 2.4.4 software and using the general time reversible 7 model of nucleotide substitutions, ML estimates of base frequencies, estimated transition/transversion ratio, proportions of invariable sites with 4 categories of substitution rates estimated γ-shape parameter 1.136, and estimated proportion of invariant sites 0.292. 13
Results
Pathogenicity Index—ICPI
The ICPI values were 1.88 for Long Bien, 1.88 for Australia, and 1.53 for Nevada cormorant. Therefore, all the viruses could be considered velogens according to OIE standards and notifiable to the international community. 2,31
Clinical Disease, Virus Isolation, and Serology
Results for observed clinical signs are presented in Table 1 . Results for virus isolation are presented in Table 2 . For all 3 strains, all infected birds with the 3 strains developed clinical disease, and virus was successfully isolated from at least 1 oral and cloacal sample from all 3 groups of animals. Birds inoculated with the Long Bien isolate had severe illness, characterized by marked depression, reluctance to move, and high mortality, with all of the birds dying or being euthanatized by 3 dpi. Two birds inoculated with the Australia isolate showed depression and open-mouth breathing between days 3 and 5. Mild neurological signs, such as head tremors and twitch, were noted in this group between 4 and 9 dpi, but all became clinically normal by 10 dpi. Birds inoculated with the Nevada cormorant isolate had marked neurological signs by 4 dpi consisting of head tremors, twitching, recumbency, and paralysis; by 10 dpi all these animals were euthanatized with severe neurologic signs.
Clinical and Pathological Findings of Chickens Infected With Newcastle Disease Virus Strains
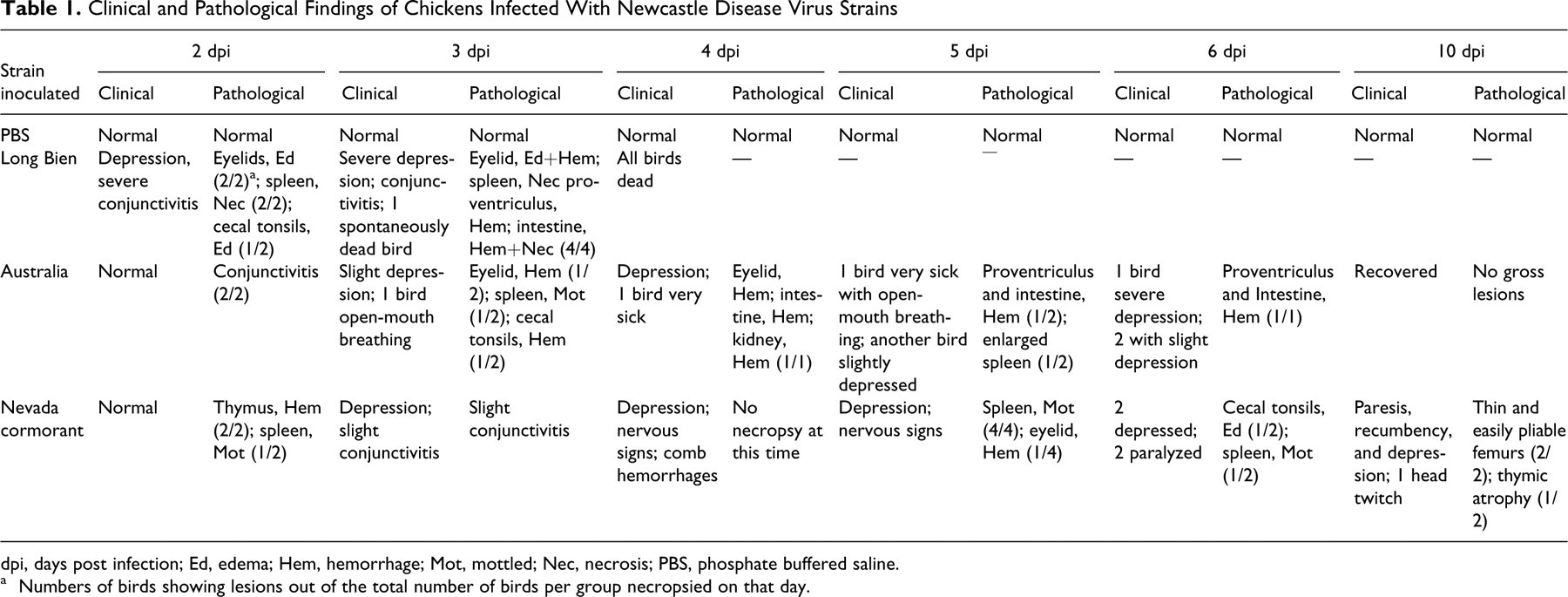
dpi, days post infection; Ed, edema; Hem, hemorrhage; Mot, mottled; Nec, necrosis; PBS, phosphate buffered saline.
a Numbers of birds showing lesions out of the total number of birds per group necropsied on that day.
Virus Isolation and Titration of Oral (O) and Cloacal (C) Swab Samples
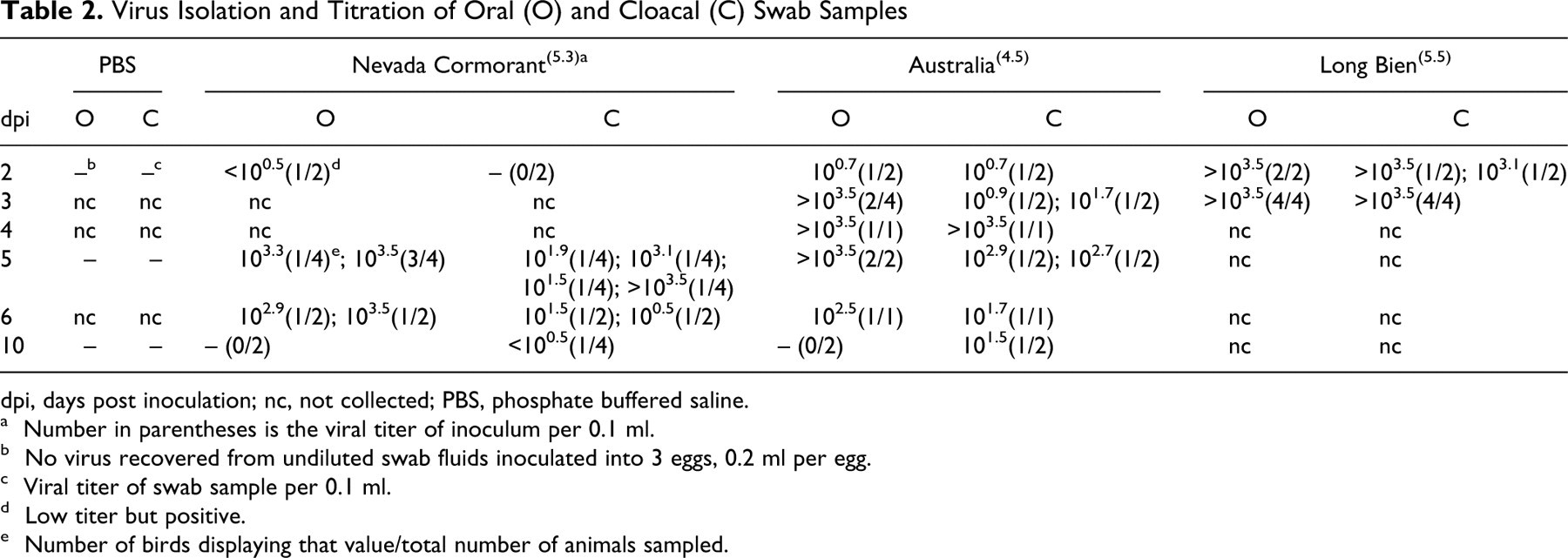
dpi, days post inoculation; nc, not collected; PBS, phosphate buffered saline.
a Number in parentheses is the viral titer of inoculum per 0.1 ml.
b No virus recovered from undiluted swab fluids inoculated into 3 eggs, 0.2 ml per egg.
c Viral titer of swab sample per 0.1 ml.
d Low titer but positive.
e Number of birds displaying that value/total number of animals sampled.
NDV was isolated from all the birds inoculated with Long Bien and Australia isolates starting from 2 dpi. In the group inoculated with Nevada cormorant virus, at 2 dpi NDV was isolated only from 1 bird, whereas all the sampled birds were positive by 5 dpi. By day 10, virus isolation was positive (Australia and Nevada cormorant isolates only, because all Long Bien birds were dead by this time) but at very low titers and only for the cloacal swabs. Seroconversion (≥:128) occurred in birds infected with Australia and Nevada cormorant isolates at 10 dpi (data not shown). Serology was not performed on animals infected with Long Bien because they were all euthanatized by day 3.
Pathology
Gross findings are presented in Table 1. The most striking findings were in birds inoculated with the Long Bien isolate. By 2 dpi, the birds had severe edema and petechial hemorrhages of the eyelid and mottled spleens. On day 3, the spleens were uniformly enlarged and had multifocal to coalescing white stippling (necrosis), both on the surface and in cut section (Fig. 1 ). Other lesions were thymic hemorrhages, proventricular hemorrhages, multiple foci of necrosis in the small intestine (Fig. 2 ), and edema, necrosis, and hemorrhage of the cecal tonsils.
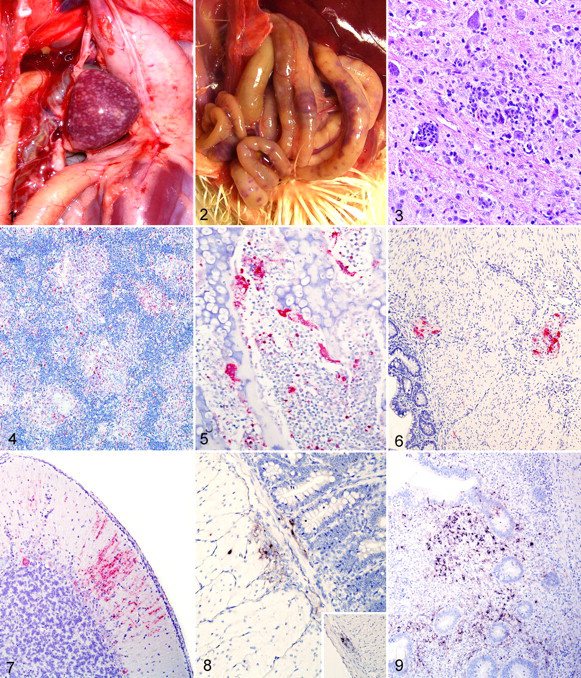
The birds infected with the Australia isolate had minimal to moderate petechiation of eyelids and moderately enlarged spleens by days 2 and 3. At days 5 and 6, the spleens were enlarged, and proventricular hemorrhages were present in 2 birds.
The birds infected with the Nevada cormorant isolate had minimal gross lesions. There was mild conjunctivitis by 2 dpi, moderately congested and mottled spleen at 5 dpi, and moderate thymic atrophy in 1 bird at day 9. There were no abnormal gross findings in the noninfected control birds.
Histology
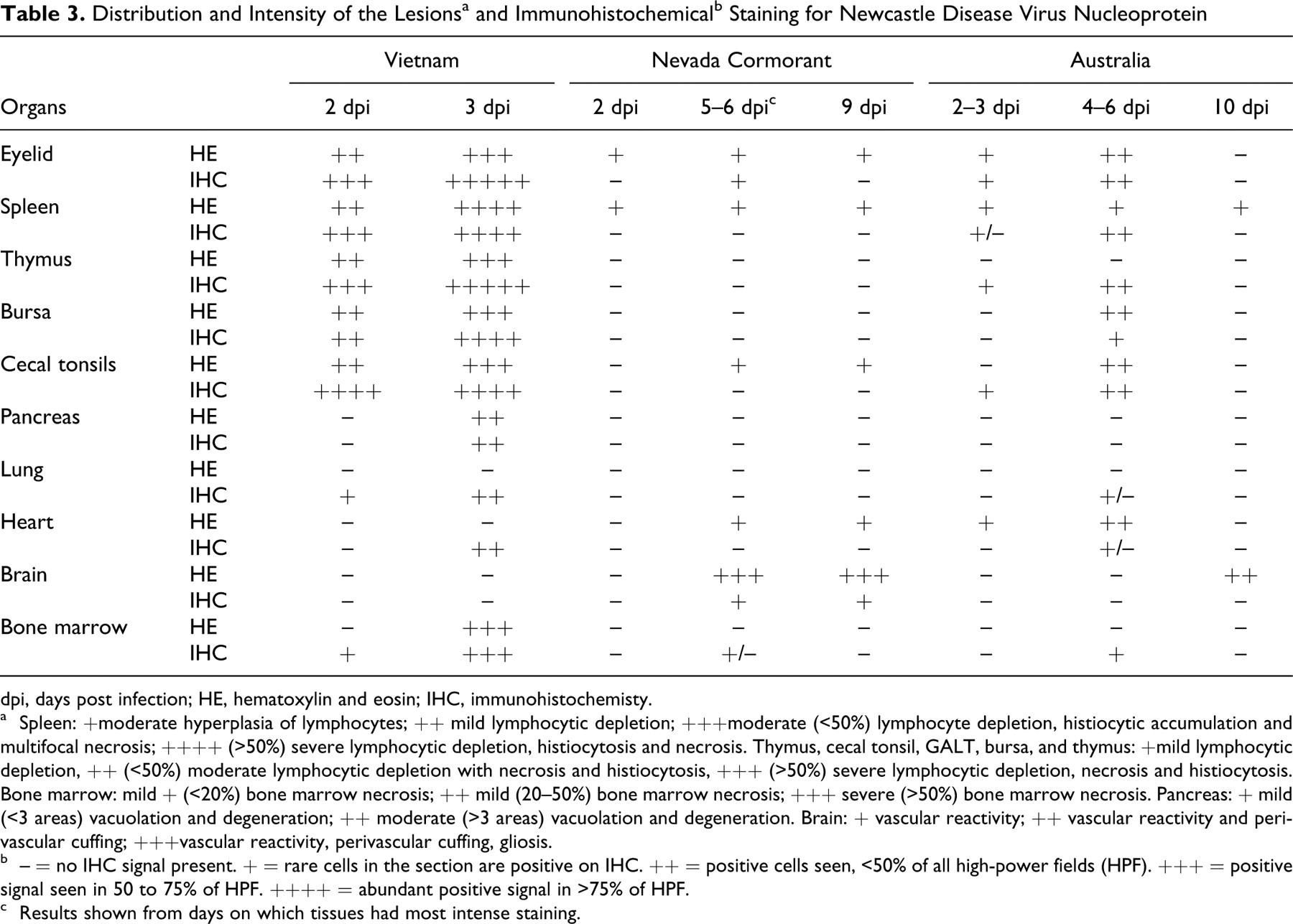
Histological as well as IHC findings are presented in Table 3 . The most severe lesions were observed in birds infected with the Long Bien isolate, starting at day 2 and culminating at day 3. These lesions were mainly confined to the site of inoculation (eyelid), the lymphoid organs (spleen, thymus, bursa), or the lymphoid aggregates of the intestines (cecal tonsils and other gut-associated lymphoid tissue). In the eyelid, the lesions were characterized by severe edema, hemorrhage, multifocal areas of necrosis associated with fibrin exudation, and pleomorphic inflammatory infiltrate, mainly composed of heterophiles and macrophages. In the lymphoid organs (thymus, bursa, spleen, and intestinal lymphoid patches, especially cecal tonsils), lesions consisted of severe necrosis, marked lymphocyte depletion, infiltration of numerous macrophages, and moderate heterophilic infiltrate. In the intestines, the severe necrosis of the lymphoid-dependent areas was associated with focal to locally extensive ulceration of the epithelium and accumulation of necrotic material within the intestinal lumen. Multifocal areas of necrosis were observed within the pancreas. By 3 dpi, at the level of the physis and metaphysis, especially in the areas beneath the articular cartilage, there were multifocal areas of bone marrow necrosis.
dpi, days post infection; HE, hematoxylin and eosin; IHC, immunohistochemisty.
a Spleen: +moderate hyperplasia of lymphocytes; ++ mild lymphocytic depletion; +++moderate (<50%) lymphocyte depletion, histiocytic accumulation and multifocal necrosis; ++++ (>50%) severe lymphocytic depletion, histiocytosis and necrosis. Thymus, cecal tonsil, GALT, bursa, and thymus: +mild lymphocytic depletion, ++ (<50%) moderate lymphocytic depletion with necrosis and histiocytosis, +++ (>50%) severe lymphocytic depletion, necrosis and histiocytosis. Bone marrow: mild + (<20%) bone marrow necrosis; ++ mild (20–50%) bone marrow necrosis; +++ severe (>50%) bone marrow necrosis. Pancreas: + mild (<3 areas) vacuolation and degeneration; ++ moderate (>3 areas) vacuolation and degeneration. Brain: + vascular reactivity; ++ vascular reactivity and perivascular cuffing; +++vascular reactivity, perivascular cuffing, gliosis.
b – = no IHC signal present. + = rare cells in the section are positive on IHC. ++ = positive cells seen, <50% of all high-power fields (HPF). +++ = positive signal seen in 50 to 75% of HPF. ++++ = abundant positive signal in >75% of HPF.
c Results shown from days on which tissues had most intense staining.
In birds infected with the Australia strain, similar lesions were observed within the eyelid and lymphoid organs, but these were much less intense than those with Long Bien and reached their peak at 4 dpi. The most severe lesions were in the bursa and cecal tonsils and consisted mainly of necrosis, lymphocyte depletion, and accumulation of macrophages. The heart, at 5 and 6 dpi, had multifocal myonecrosis with accumulation of macrophages. At day 10, there was moderate nonsuppurative encephalitis, consisting of multifocal areas of lymphoplasmacytic perivascular cuffing.
Birds infected with the Nevada cormorant isolate had minimal lesions within the lymphoid tissues but showed severe nonsuppurative encephalitis and moderate myocarditis. The encephalitis was confined between day 5 and 10 and was characterized by perivascular accumulations of lymphocytes, macrophages, rare plasma cells, and hypertrophy of the endothelial cells (Fig. 3 ). The myocarditis was characterized by multifocal necrosis associated with moderate macrophage infiltration. One bird infected with Nevada cormorant at day 5 had severe laryngitis, with necrosis of the epithelial layer (ulceration), accumulation of fibrin, and infiltration with heterophiles and macrophages.
Immunohistochemistry
In the infected cells, the immunolabeling for NDV was intracytoplasmic and finely to coarsely granular. Long Bien–infected birds had the broadest viral distribution (23/25 different tissues were positive) and the most intense signal, starting from day 2 and culminating at day 3. The organs with the strongest signal were the eyelids, the lymphoid organs, and the mucosa-associated lymphoid aggregates in multiple organs (Fig. 4 ). In these tissues, the positive cells consisted mainly of lymphocytes and macrophages. In the spleen, the immunoreactivity was confined to the fixed-macrophage–dependent areas around the penicillary arteries, whereas the lymphocyte-dependent areas were devoid of signal. In the respiratory system, the positive signal was confined to the nasal and tracheal mucosa, scattered lymphoid aggregates closely associated with the secondary and tertiary bronchi (bronchus-associated lymphoid tissue), and the squamous epithelium of the atria, air capillaries, and air sacs. In the digestive tract, intense positivity for NDV was observed only within the submucosal lymphoid aggregates, and no staining was observed associated with the epithelial lining. Scattered Kupffer cells in the liver and several epithelial cells in the distal tubules of the kidneys had positive signal for NDV. In the bone marrow, numerous osteoclasts and mononuclear cells just beneath the growth plate or surrounding areas of necrosis showed intense immunolabeling (Fig. 5 ). In the heart, the staining was limited to the epicardium. Birds infected with the Australia isolate had a more limited viral distribution (15/25 positive tissues) and lesser amounts of signal even in the positive tissues when compared with Long Bien, but the pattern of staining and the type of infected cells was the same as described above. The viral distribution was mainly limited to the lymphoid organs and lymphoid aggregates and reached its peak at 5 dpi. In the intestines of these animals, immunolabeling was also present within the submucosal and myenteric plexuses in cells (Fig. 6 ) consistent with neurons and glial cells. Tissues from birds infected with the Nevada cormorant isolate had minimal signal for NDV, which was primarily confined to the portal of entry (eyelid), with the exception of moderate immunolabeling of scattered neurons within the brains of birds at day 6 and 10 dpi (Fig. 7 ).
In Situ Hybridization
In selected sections, the negative sense riboprobe for NDV matrix gene labeled the same structures and cells immunolabeled with IHC. The type of staining was cytoplasmic and evenly distributed (Figs. 8 and 9).
Sequencing and Phylogeny
Results of sequencing and phylogenetic trees are presented in Fig. 10. (Detailed correspondence between names used in the tree and NCBI accession numbers are available in the online supplement. To view the online supplement, please go to http://www.vet.sagepub.com/supplement.)
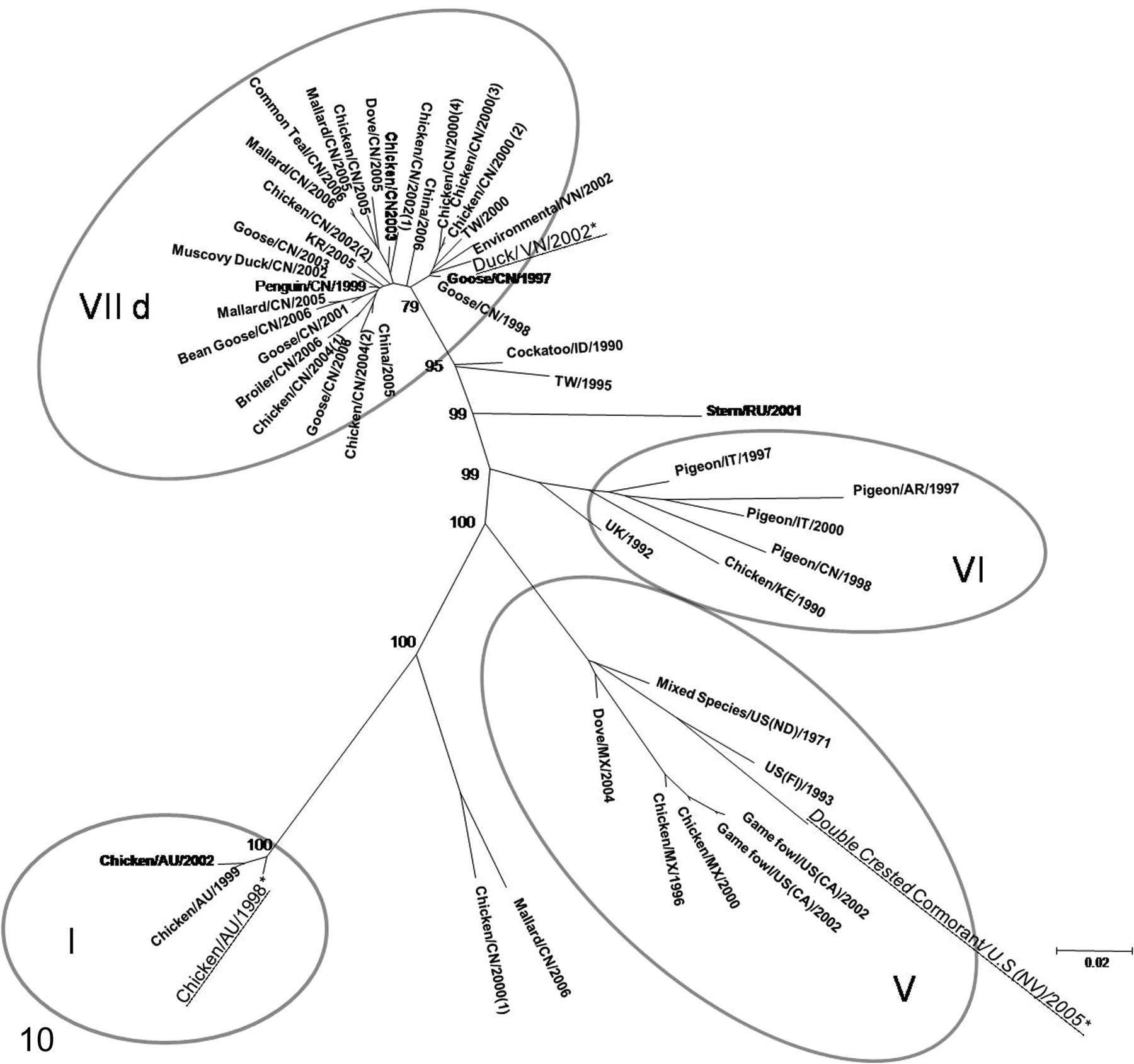
Evolutionary relationships of 3 selected NDV isolates in comparison to reference genotypes. The evolutionary history was inferred using the maximum likelihood method. The optimal tree with the sum of branch length is shown, and bootstrap values greater than 70 are included in the nodes. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. Correspondence between tree names and GenBank accession numbers are presented in Table 4 as online supplement. The name of the viruses described in the article are followed by an asterisk and underlined. Roman numbers within circles indicate the genotype.
The Genome sequences of the viruses were submitted to Gene Bank under the following names: APMV-1/duck/Vietnam, Long Bien/78/2002: APMV-1/chicken/Australia/9809-19-1107/1998: APMV-1/double-crested cormorant/USA, Nevada/19529-04/2005:
Phylogenetic analysis with full fusion sequences for the 3 selected strains revealed that the Nevada cormorant isolate belongs to a new branch of genotype V that is clearly separated from other viruses that caused outbreaks in cormorant in 2008.
35
The Australia isolate belongs to genotype I viruses and groups with other contemporary Australian viruses. The Long Bien isolate falls within genotype VIId and is most closely related to isolates from China in 2000 from chickens and geese (Fig. 10). All 3 viruses, Australia, Long Bien, and Nevada cormorant, have a virulent fusion cleavage site motif.
31
The –R R K K R F– motif was present in Long Bien, –R R Q R R F– in the Australia isolate, and –R R Q K R F– in the Nevada cormorant virus.
Discussion
Although all 3 viruses were considered virulent, by both ICPI and fusion protein amino acid sequence, these strains showed differing clinicopathological pictures, both in severity and in distribution of the virus through various tissues of the body.
The Long Bien isolate belongs to genotype VIId, and to the authors' knowledge this is the first full clinicopathological characterization of an isolate belonging to this genotype. Clinicopathologically, the phenotype of Long Bien was similar to other previously described virulent NDV strains belonging to genotypes II, V, VI, and VIIa. 7,19 –21,37 Long Bien strain caused severe disease, with birds succumbing rapidly after infection. By IHC, the viral tropism was mainly for lymphoid tissues, but viral protein was also very evident in the respiratory epithelium of trachea, lung, and air sacs and in the renal epithelium. Within the lymphoid tissues, and particularly in the spleen, the majority of the immunolabeled cells were morphologically compatible with macrophages, specifically those surrounding the penicillary arteries in the spleen. This tropism for cells belonging to the monocytes–macrophage series was supported by visualization of positive osteoclasts within the bone marrow, as these cells are derived from a subset of circulating monocytes. 6,40
The Australia isolate, another velogen, had a less severe clinicopathological presentation. Lesions were milder, and viral distribution, as detected by IHC, was more limited. Lymphoid tissue appeared to be the primary target, but damage in these areas and associated IHC signal were much less than those seen with Long Bien. An unexpected finding was the presence of abundant viral nucleoprotein and ISH staining within the myenteric plexuses of the intestines, which was most notable at 5 dpi and was not associated with any inflammatory changes. The positive ISH signal using a negative riboprobe indicates that the virus is actively replicating in these cells, as the probe will bind only to positive sense RNA, which would be the replicative intermediate of the virus (which is negative sense) or the viral messenger RNA. Although some NDV strains are known for their ability to invade nervous tissue, it is usually the central nervous system that is affected, making this tropism for the peripheral nervous system unusual. Only 1 report in the literature, concerning an outbreak in Japan in vaccinated flocks, describes immunohistochemical staining of the intestinal nerves, but the genotype of the investigated virus was type VII, whereas the Australia virus is genotype I. 30
The marked differences in the clinicopathological assessment between the 2 velogenic strains with the highest ICPI, Australia and Long Bien, is also mirrored by the phylogenetic distances. Genotype VIId (Long Bien) encompasses numerous markedly virulent strains that are currently circulating in southeast Asia, Africa, and the Middle East and have also been reported to have increased host range. 15,16,26,27,38 On the other hand, the Australia isolate belongs to genotype I, a clade that contains mainly nonvirulent strains that often circulate in wild birds and waterfowl. It has been known that on rare occasions these strains can mutate and become virulent. This has been demonstrated for the 1998–2000 NDV outbreaks in Australia (of which our strain is an isolate) and the 1997 outbreaks in Ireland. In these cases, the virulent isolate viruses have been traced back to previously circulating nonpathogenic strains that acquired virulence by mutation of the F protein cleavage site. 4,8,12 Even though both Australia and Long Bien strains had the same ICPI value (1.88) and a virulent F gene cleavage site, they displayed strikingly different clinicopathological syndromes when inoculated into 4-week-old birds. Long Bien behaved as a full-fledged VVND (Velogenic Viscerotropic Newcastle Disease), acting very similarly to other virulent strains even if genetically distant, whereas Australia had a much milder pathotype. The reason for this is unknown, but it might be linked to the fact that class I viruses are usually adapted to waterfowl and not usually to chickens. 12 In addition, it should be considered that the F gene is not the only determinant of virulence but that other genes may also be involved in the disease-inducing ability of NDV strains. For example, the HN protein has been shown to influence tissue tropism and to trigger apoptosis in vitro, 14,33 the P gene has been demonstrated to inhibit the interferon pathway through the production of the V protein, 32 and the L gene has been associated with virulence through RNA processing ability. 34 The work presented here confirms that neither ICPI nor fusion protein cleavage is sufficient to fully predict the outcome of virus infection in poultry. 9
Phylogenetic analysis revealed that our third strain (Nevada cormorant) belongs to a distinct subgroup of genotype V recently isolated from cormorants. 35 Although the ICPI of this virus was 1.53, which makes this virus technically a velogen (>1.5), 2 and its fusion protein has a virulent cleavage sequence, Nevada cormorant isolate behaved similarly to many previously described mesogens of genotype V, 7 although the degree of neurologic involvement was more marked than what has been described. 7 Mesogens do not usually produce severe clinical signs in SPF chickens but can occasionally cause neurological signs. 1 The virus was detected by IHC only at the inoculation site (eyelid) and in the brain (only 4 birds). The presence of viral nucleoprotein and RNA, as revealed by IHC and in situ hybridization, was mainly in neurons of the cerebral cortex, the nuclei of the brainstem, and the cerebellum, thus accounting for the neurological signs. The involvement of the respiratory system was minimal. As described for the cormorants infected with NDV, 5,10,22 –24 clinicopathological findings in the chickens involved mainly the nervous system, suggesting that the tropism and the behavior of the virus were similar in different hosts. The development of clinical disease in chickens inoculated with a strain rescued from wild birds (cormorant) underlines the important role that wild birds could have in the natural or accidental transmission of virulent strain to commercial flocks.
This article is the first to fully characterize through sequencing and clinicopathological observation the recently isolated strains Australia, Nevada cormorant, and Long Bien. The constant characterization of newly isolated strains that circulate within and outside the United States will allow for assessment of eventual variation in their pathogenicity for poultry species. Such information will be helpful in devising appropriate prevention and control measures. This study reinforces the concept that standard indices and sequence analysis of NDV isolates may not fully predict their pathogenicity in susceptible animals, therefore making animal experiments important tools for a complete characterization of new isolates.
Footnotes
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.