Abstract
Identification of fungal organisms often poses a problem for pathologists because the histomorphology of some fungal organisms is not specific, fresh tissues may not be available, and isolation and identification in culture may take a long time. The purpose of this study was to validate the use of panfungal polymerase chain reaction (PCR) to identify fungal organisms from formalin-fixed paraffin-embedded (FFPE) tissues. Formalin-fixed paraffin-embedded curls were tested from 128 blocks containing canine, feline, equine, and bovine tissues with cutaneous, nasal, pulmonary, and systemic fungal infections, identified by the presence of fungi in histologic sections. Quantitative scoring of histologic sections identified rare (11.9%), occasional (17.5%), moderate (17.5%), or abundant (53.1%) fungal organisms. DNA was isolated from FFPE tissues and PCR was performed targeting the internal transcribed spacer 2 (ITS-2) region, a segment of noncoding DNA found in all eukaryotes. Polymerase chain reaction products were sequenced and identified at ≥97% identity match using the Basic Local Alignment Search Tool and the NCBI database of ITS sequences. Of the 128 blocks, 117 (91.4%) yielded PCR products and high-quality sequences were derived from 89 (69.5%). Sequence and histologic identifications matched in 79 blocks (61.7%). This assay was capable of providing genus- and species-level identification when histopathology could not and, thus, is a beneficial complementary tool for diagnosis of fungal diseases.
Keywords
An exponential growth in molecular diagnostics for infectious diseases has produced opportunities for confirming culture results and can be combined with other methods, such as histopathology, to confirm a diagnosis when culture is not possible. 1,5,14 DNA-based assays have been particularly helpful in diagnosis of fungal infections due to the speed of these tests compared to fungal culture, which can take 2 to 4 weeks. 10 Many fungi are not readily cultivable in the laboratory and others require biosafety level 3 precautions. In addition to culture, fungal identification has relied on histopathology and cytology, although inherent challenges exist due to the subtle nuances in distinguishing fungal morphology. 41
Polymerase chain reaction (PCR) assays have been developed to target and quantify specific genera and species of fungi from clinical samples. 11,13 These assays, however, require prior suspicion of the etiologic agent and are not available for every fungal organism, including emerging pathogens. The greatest utility has come from panfungal PCR, which amplifies conserved genes that are present in all fungi such as the ribosomal RNA (rRNA) region, including the large (28S) and small (18S) subunits, 5.8S, and internal transcribed spacer (ITS-1 and -2) and intergenic spacer (IGS) regions. Primers targeting these conserved rRNA sequences were originally developed for phylogenetic studies. 45 The resulting PCR products have regions that are hypervariable among fungi, allowing for identification by comparison of their sequences to a reference fungal database. The ITS regions are currently considered the superior regions for species resolution. 42 Panfungal PCR can potentially identify any fungal agent without prior knowledge of its suspected identity, which can greatly reduce the time and cost spent on other molecular diagnostics.
Panfungal PCR on formalin-fixed paraffin-embedded (FFPE) tissues is useful when fresh tissue was not or could not be collected. It has been successfully applied in different human tissues and for a variety of fungal infections 30,36,39 ; however, less work has been performed in veterinary cases. Two of the human studies 30,36 have included small subsets of animal tissues (6 and 19, respectively) but were lacking in breadth of animal species and tissue types. Recently, there have been more extensive applications of this assay in cats, 3,8 but there is still a need for validation in other animals. Furthermore, there does not appear to be a general consensus on DNA extraction methods for FFPE tissues, 36,40 and many of the recommended kits are expensive or the protocols are time intensive.
Improving fungal diagnostics in veterinary medicine has the potential to not only improve animal healthcare but also enable proper selection of antifungal therapeutics, thereby reducing the development of resistance that has already been documented in poultry, 7 dogs, 37 and other animals. 12 In addition, improvement in the accuracy and specificity of identifying fungi to the genus and species level would allow for better epidemiologic studies of current and emerging fungal pathogens. The purpose of this study was to validate the use of panfungal PCR on FFPE specimens in a variety of veterinary cases from birds, cats, camels, cattle, dogs, horses, and sheep. Furthermore, we aimed to test and optimize a cost-effective DNA extraction kit for amplification of fungal DNA from FFPE animal tissues.
Materials and Methods
Case Selection
From the pathology archives at the Texas A&M University College of Veterinary Medicine and Biomedical Sciences, the Dermatopathology Specialty Service, and the Texas A&M Veterinary Medical Diagnostic Laboratory, an exhaustive query was performed, and 128 paraffin blocks from 96 animals were selected for inclusion in this study. The initial selection criteria included the presence of fungi observed histologically (Supplemental Fig. 1), a diagnosis of primary fungal infection, and an attempt to select representative numbers of animal species, tissue types, and fungal etiologic agents. Once the blocks were selected, a single pathologist reviewed all archived slides to confirm the histologic diagnosis and to categorize the cases based on the amount of fungal organisms. Separate guidelines were used for yeast and hyphal morphologies since hyphae are generally more prolific, and cases were classified according to the number of organisms seen in 20 fields at 200× magnification as either rare (<3 yeast or <8 hyphae), occasional (3-10 yeast or 8-20 hyphae), moderate (10-15 yeast or 20-50 hyphae), or abundant (>15 yeast or >50 hyphae). The cases included tissues of dogs (66), cats (25), horses (19), cattle (7), birds (6), camels (3), sheep (1), or wild cats (1). The tissue types included oral cavity, respiratory tract, gastrointestinal tract, urinary tract, skin, brain, heart, lymph node, and bone. Previous ancillary testing included 69 cases with special stains, 26 with culture, 4 with cytology, 3 with PCR performed at other institutions, and 1 with immunohistochemistry. Most cases without ancillary testing showed strong fungal identification based on histopathology, and additional tests were deemed unnecessary due to costs or unavailability of fresh samples.
Twenty-four blocks were used to compare 2 commercially available kits for extraction of fungal DNA from FFPE tissues, another 52 blocks were used to further optimize the superior kit’s protocol, and the remaining 52 blocks were processed following the optimized protocol (Supplemental Fig. 1). From each block, sections totaling 50 μm in depth were cut in 5-μm scrolls with a microtome, and the blade was cleaned with absolute alcohol between blocks. The initial scroll for each block was discarded to minimize environmental and carry-over contamination. The scrolls were stored in DNase-, DNA-, and RNA-free microcentrifuge tubes until DNA was extracted.
ITS-2 and IRBP Polymerase Chain Reactions and Sequence Analysis
Fungal DNA was amplified by PCR targeting the internal transcribed spacer-2 (ITS-2) region, located between the coding regions for the 5.8S and small ribosomal RNA subunits, using the following panfungal primers: ITS3-F (5’-GCATCGATGAAGAACGCAGC-3’) and ITS4-R (5’-TCCTCCGCTTATTGATATGC-3’).
45
To evaluate quality of host DNA in each sample, a PCR targeting the
All PCR products were separated on a 2% agarose gel made with 1× Tris boric acid ethylenediaminetetraacetic acid (TBE) buffer and GelRed® (Biotium, Inc., Hayward, CA). The sample was separated by electrophoresis for 1 hour at 120 V and visualized with a UV transilluminator (GelDoc EZ™ system; Bio-rad Laboratories, Inc., Hercules, CA). Every ITS-2 band was excised from the gel, and the DNA was isolated using the E.Z.N.A.® gel extraction kit (Omega Bio-Tek, Inc., Norcross, GA). This purified DNA was then submitted for sequencing (Eton Bioscience, Inc., San Diego, CA) using the ITS3-F and ITS4-R primers. Sequences were trimmed for quality by removal of the beginning and end of sequences, joined as contigs using Sequencher® 4.8 (Gene Codes Corporation, Ann Arbor, MI) and then queried against the GenBank database using the Basic Local Alignment Search Tool (BLAST, NCBI). 2 Published GenBank accession matches were given precedence over unpublished ones. Sequences were assessed both quantitatively with the quality score given in Sequencher and qualitatively by visual inspection of the electropherogram. Sequences were identified as poor quality for the remaining analyses when the overall quality score was less than 70%, where no or little sequence was detected, and when a majority of peaks overlapped for each base. A fungal identification was made when a sequence matched the reference at ≥97% identity (for genus and species levels). When the sequence matched to more than 1 taxon at ≥97%, the one with the highest identity score was chosen. When there were multiple matches at the highest identity score, all matches were reported. Also, when there were multiple matches from different bands, all matches were reported. Interphotoreceptor retinoid-binding protein bands were considered positive or negative based on the presence or absence of a band at the same level as positive controls and were not submitted for sequencing.
Comparison of DNA Extraction From Paraffin Blocks Using 2 Commercially Available Kits
Two commercially available kits, the QIAGEN QIAamp® DNA FFPE Tissue Kit and Deparaffinization Solution (QIAGEN, Valencia, CA) and MO BIO Bi
Optimization of the MO BIO BiO stic Kit Protocol
The MO BIO Bi
Phylogenetic Analysis of Aspergillus spp
All
Statistical Analysis
Contingency analysis was performed in JMP® Pro 12 (SAS Institute, Inc., Cary, NC) to determine whether an association existed between the number of organisms observed histologically and ITS-2 PCR results, and with the quality of sequences derived from the PCR products. First, the correlation between variables was assessed with a chi-square test. When a significant correlation was identified for variables with multiple categories, the samples were then scored (yes/no) for the category in each variable that was suspected to contribute to the correlation. A second chi-square test was then performed between 1 category from each variable, and significant
Results
Comparison of 2 DNA Extraction Kits
DNA extraction using the QIAGEN QIAamp DNA FFPE Tissue Kit and MO BIO Bi
Comparison of DNA Extraction Kits and Effect of Adding Lyticase, With Respect to DNA Quality, PCR Efficiency, and Sequence Quality.
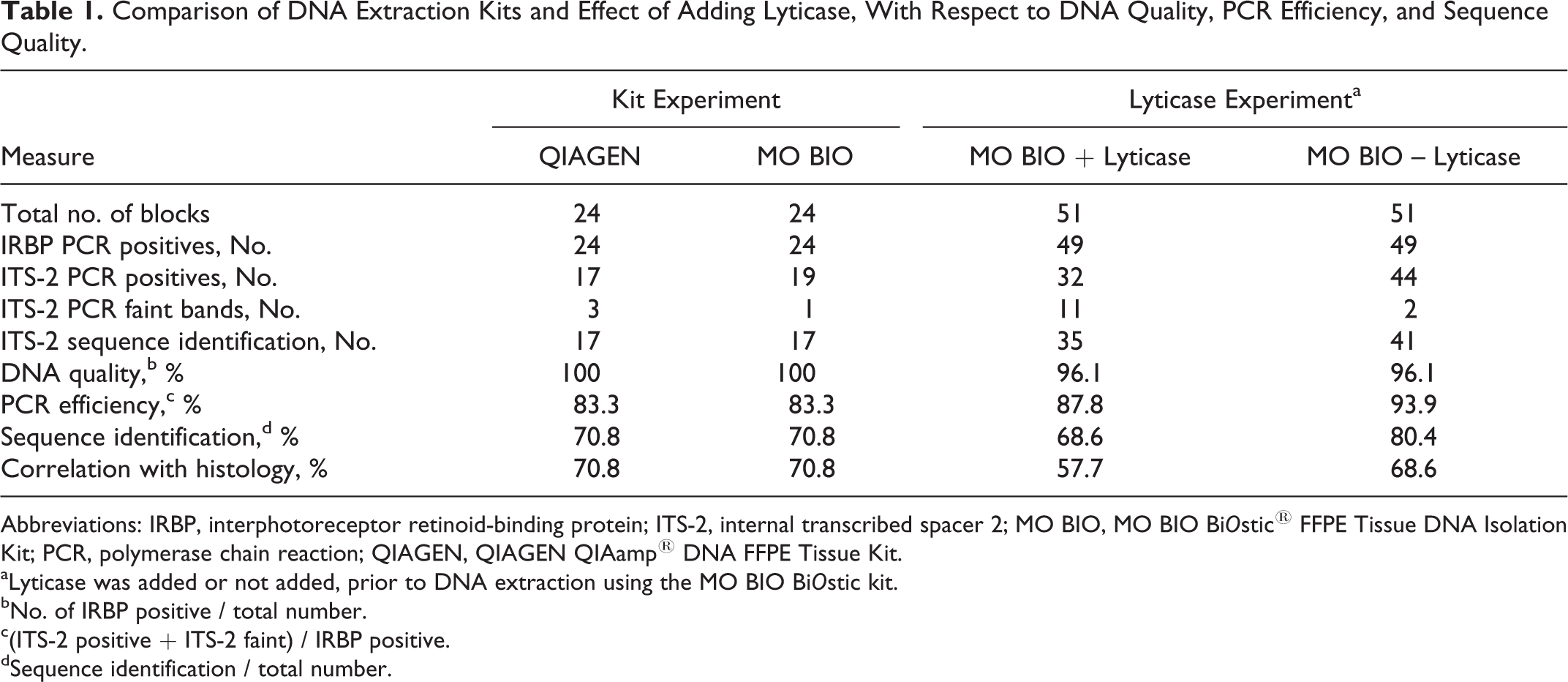
Abbreviations: IRBP, interphotoreceptor retinoid-binding protein; ITS-2, internal transcribed spacer 2; MO BIO, MO BIO Bi
aLyticase was added or not added, prior to DNA extraction using the MO BIO Bi
bNo. of IRBP positive / total number.
c(ITS-2 positive + ITS-2 faint) / IRBP positive.
dSequence identification / total number.
Optimization of the MO BIO BiO stic FFPE Tissue DNA Isolation Kit
The addition of lyticase digestion to the QIAGEN QIAamp kit protocol has been shown to improve PCR efficiency,
3,36
so we wanted to test whether this would also be true for the MO BIO Bi
Sequence Analysis of Archived Case Material
An additional 52 blocks were processed with the MO BIO Bi
To calculate sensitivity relative to histologic examination, the number of cases for which genus-level identification of fungi was possible through both histology and the panfungal PCR (
Number of Fungal Identifications Made Through Histopathology or Panfungal Polymerase Chain Reaction (PCR) for Calculation of Sensitivity.a
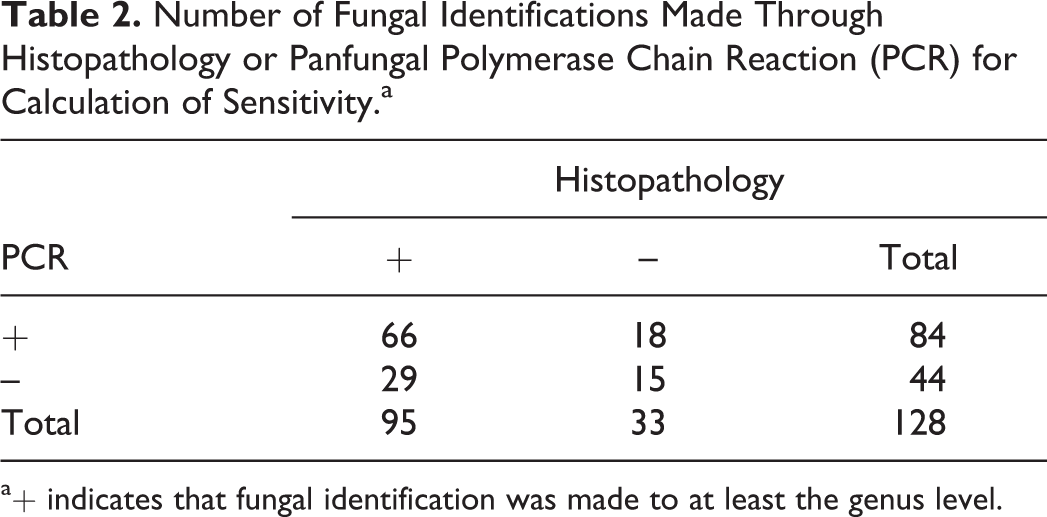
a+ indicates that fungal identification was made to at least the genus level.
A fungal identification was made through ITS-2 PCR and sequencing for 83 of 128 (65%) blocks tested. The vast majority of these sequence-derived identities (79/83, 96%) corresponded to those made histologically and only a small number (4%) identified as an organism that did not match the histologic descriptions. A genus- or species-level identification was made by BLAST analysis of sequences in 48 of 83 (60%) cases, whereas histologic evaluation was unable to do so. This included 24 samples for which sequencing provided a genus-level resolution, and 24 samples a species-level resolution. Three representative cases were selected to demonstrate the benefit of using panfungal PCR where histologic fungal morphology could not provide a genus-level identification (Figs. 1–3).
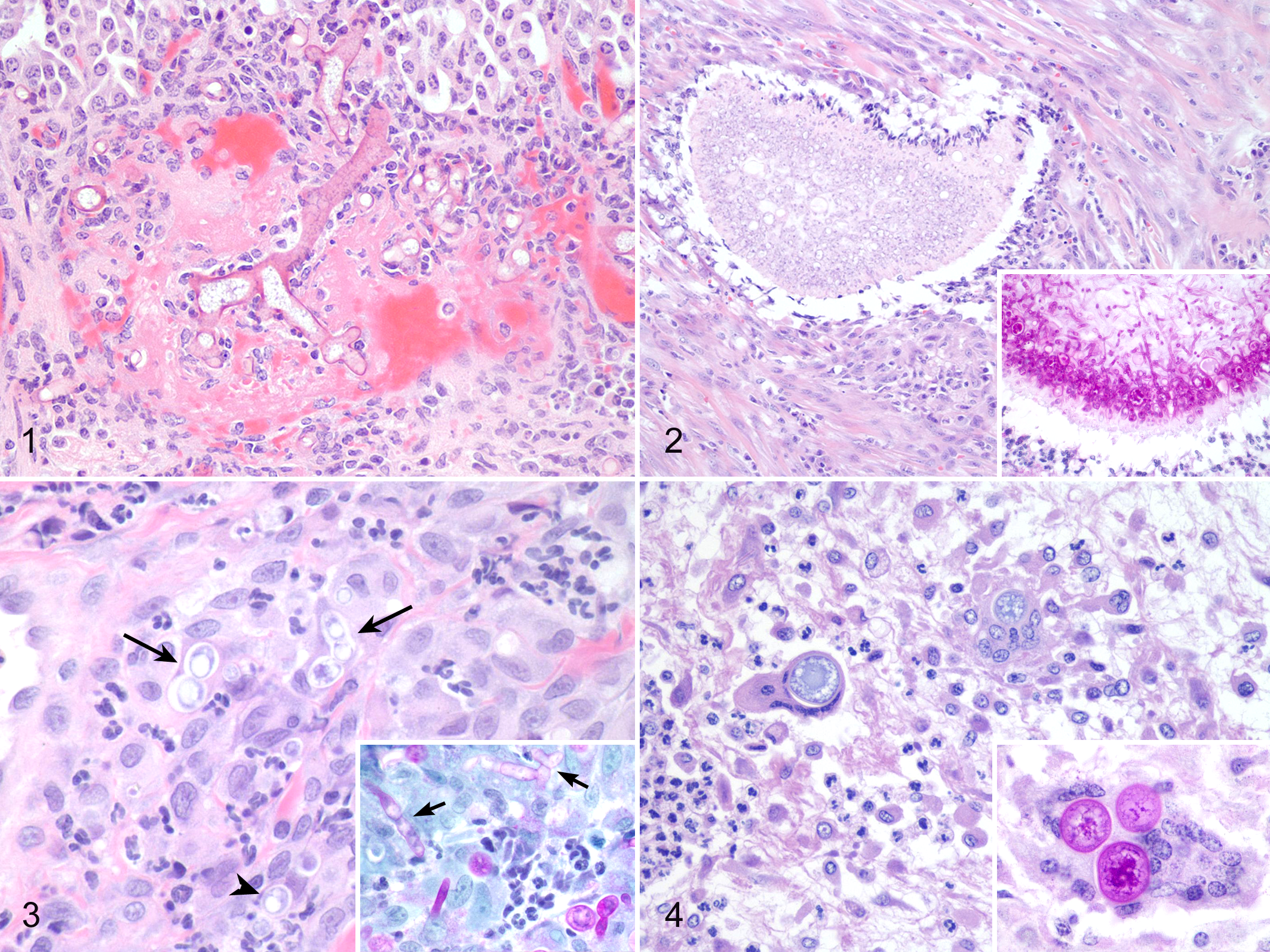
Various mycotic infections in domestic species that were classified using panfungal polymerase chain reaction.
Some of the cases that were included from the archives also had ancillary testing performed, which enabled further comparison with culture results. Fungal culture had been successfully performed on 25 cases. Panfungal PCR and sequencing results correlated with culture in 12 of 25 (48%) cases, whereas 7 of 25 (28%) did not.
Due to the high prevalence of
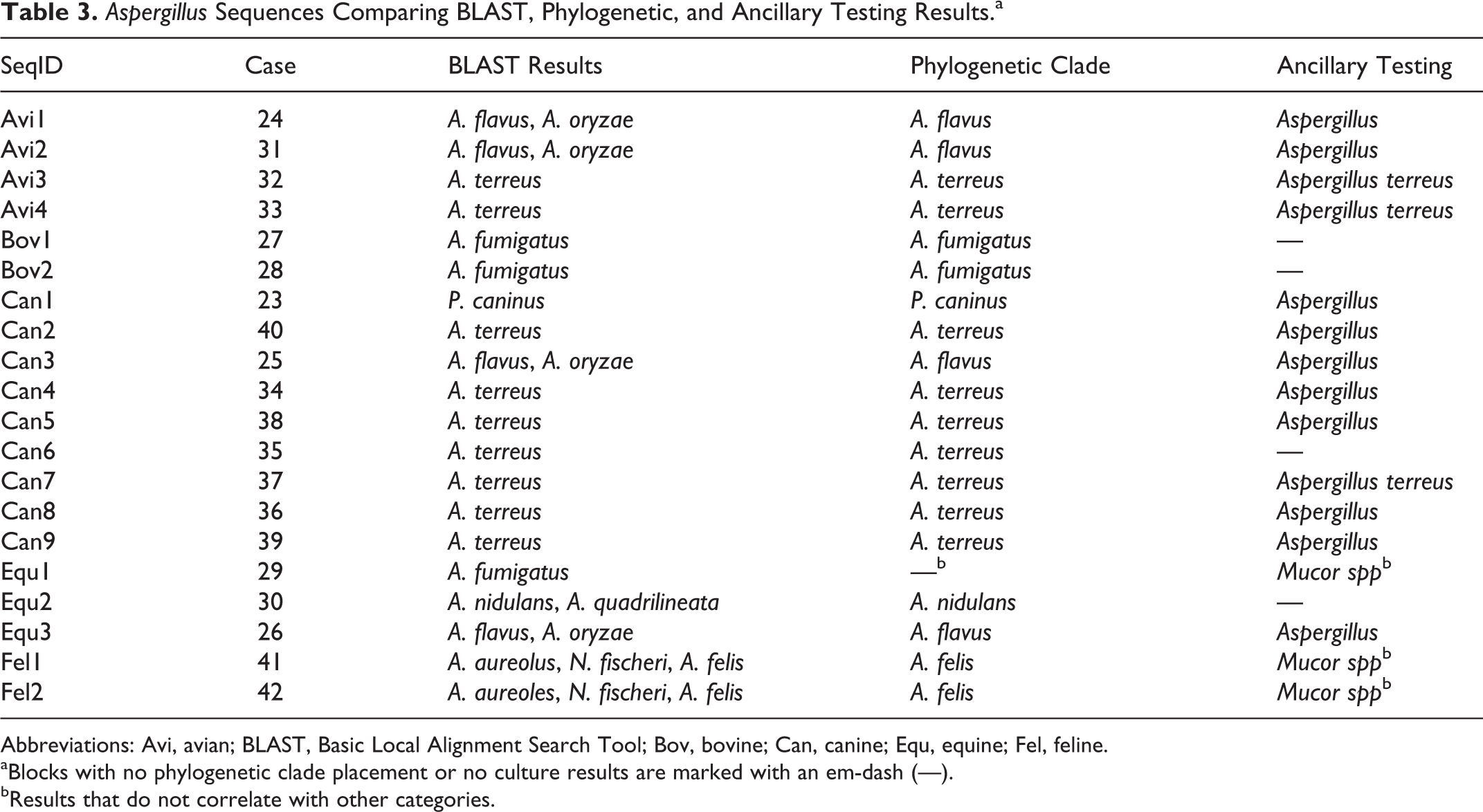

Abbreviations: Avi, avian; BLAST, Basic Local Alignment Search Tool; Bov, bovine; Can, canine; Equ, equine; Fel, feline.
aBlocks with no phylogenetic clade placement or no culture results are marked with an em-dash (—).
bResults that do not correlate with other categories.
Statistical analysis revealed no influence of host, tissue, or fungal type on PCR results (positive/negative). Respiratory tract samples in this study were significantly associated with an increased likelihood of having more than 1 band on gel electrophoresis (
Discussion
The panfungal PCR diagnostic assay presented here had a sensitivity of 69.5%, calculated as the percentage of cases for which ITS-2 sequencing provided a fungal identification. This sensitivity correlates well with previous studies, which also amplified the ITS-2 region from DNA extracts of FFPE tissues and showed sensitivities of 53.8%
4
and 64.3%.
30
These 2 studies with similar sensitivities both targeted the ITS-2 regions but used other methods of DNA extraction. Our work demonstrated that this assay was successful in a wide variety of animal hosts, tissues, and fungi and, thus, is a valuable test when used in conjunction with histopathology for identification of fungal infections in animals. The results presented here also show that the MO BIO Bi
Many fungi have characteristic histologic features, which can be used to identify fungi in typical presentations and representative sections. However, there are cases of atypical morphology, stemming from genotypic variation.
23
For example, some
There appears to be greater challenges in identifying hyphae compared to yeast, especially the hyaline septate molds that are indistinguishable from one another morphologically, including
Statistical analyses demonstrated that this assay is equally successful for both yeast and hyphae. There was not any influence of the type of tissue from which the DNA was isolated on PCR results (positive/negative); however, lung samples were more likely to have multiple PCR product bands, and skin samples were more likely to yield poor-quality sequences. The skin and respiratory tracts are known to have resident microbiota, which might influence PCR performed on these samples. It is also possible that the method of sample collection, or postmortem handling of the tissues prior to embedding, could have introduced contamination. Another limitation of this assay is that sensitivity decreases when organisms are rarely observed. More sensitive sequencing techniques, such as next-generation sequencing, might have greater success in amplifying fungal DNA when there are rare organisms in the embedded tissue. However, these techniques are costly and not yet suitable for diagnostics.
There was variable ability of the assay to derive genus- and species-level resolution. The sequence analysis provided genus- and species-level resolution in 36% of all cases where they could not be distinguished based only on histologic morphology. For some genera, such as those belonging to the phaeohyphomycoses, the ITS-2 region was unable to distinguish between genera such as
It is interesting that 28 of 128 (21.9%) of all FFPE blocks returned sequences that were of poor quality, in spite of being purified from a strong band on the gel. Similar reports of ambiguous or poor-quality sequences were given in 1 of the original applications of this assay,
30
but this was not described in another more recent work.
36
Studies have shown through cloning that ITS sequence heterogeneity exists in some fungal genera including
Another rarely reported finding 30 that was encountered in the present study was the presence of multiple bands in a sample identified with gel electrophoresis. Use of the IRBP PCR enabled confirmation of high-quality extracted DNA for these samples, and thus the presence of multiple bands is not likely due to fragmentation of the DNA. It is possible that these additional sequences were amplified from commensal or contaminating fungi present in the tissues. At least for the skin, 35 gastrointestinal tract, 18 and respiratory tract 15 tissues, recent next-generation sequencing studies have identified an enormous diversity of commensal fungi inhabiting these tissues. Although these commensal fungi are rarely seen histologically, their DNA could still be present in the tissues and amplified with this assay, possibly producing a false positive result. An additional possibility that could explain the presence of multiple bands is variation in the copy number of the ITS coding region, 9,32 which is present as tandem repeats in some species of fungi. 25,26 Another recent application of this assay on tropical mycoses 19 found that some of the amplified fungal ITS-2 sequences were not consistent with the histologic findings on those blocks. That study suggested that the amplified fungal DNA came from environmental contamination of the blocks. Although this is possible, it is also possible that commensal fungal DNA was amplified, as hypothesized here. This limitation indicates that the PCR and sequencing results provided by this assay need to be interpreted in the context of fungal morphology observed histologically.
In summary, this assay has been improved through reduction in cost and time, and it has been validated for a variety of animal hosts and fungal pathogens that commonly infect animals. Although some limitations still exist for the assay, it could provide a diagnosis when morphology based on histologic examination alone could not, and when fresh tissues were not available for fungal culture. Having a quick and accurate diagnosis enables the correct selection of antifungal therapeutics. Accurate identification of fungal infections in animals will enhance epidemiologic studies of current and emerging fungal threats not only to animals but to humans as well. Further development of genus- and species-specific PCR could help to resolve fungal identification for instances when the ITS region possesses sequence heterogeneity within species or when the ITS region is homogeneous across genera.
Footnotes
Acknowledgements
We thank Dr Xiaorong Lin, Department of Biology, Texas A&M University, for providing purified fungal genomic DNA, and Anna Blick, Department of Veterinary Pathobiology, Texas A&M University, for assistance with phylogenetic analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
