Abstract
Here we report a pig with amyloid A (AA) amyloidosis associated with Streptococcus suis infection and identification of a unique amyloid sequence in the amyloid deposits in the tissue. Tissues from the 180-day-old underdeveloped pig contained foci of necrosis and suppurative inflammation associated with S. suis infection. Congo red stain, immunohistochemistry, and electron microscopy revealed intense AA deposition in the spleen and renal glomeruli. Mass spectrometric analysis of amyloid material extracted from the spleen showed serum AA 2 (SAA2) peptide as well as a unique peptide sequence previously reported in a pig with AA amyloidosis. The common detection of the unique amyloid sequence in the current and past cases of AA amyloidosis in pigs suggests that this amyloid sequence might play a key role in the development of porcine AA amyloidosis. An in vitro fibrillation assay demonstrated that the unique AA peptide formed typically rigid, long amyloid fibrils (10 nm wide) and the N-terminus peptide of SAA2 formed zigzagged, short fibers (7 nm wide). Moreover, the SAA2 peptide formed long, rigid amyloid fibrils in the presence of sonicated amyloid fibrils formed by the unique AA peptide. These findings indicate that the N-terminus of SAA2 as well as the AA peptide mediate the development of AA amyloidosis in pigs via cross-seeding polymerization.
Keywords
Amyloid A (AA) amyloidosis is caused by the accumulation of amyloid fibrils derived from serum AA (SAA) protein in tissues throughout the body. An increase in circulating SAA levels under conditions of chronic inflammation or infectious diseases is required to induce AA amyloidosis. 10
There are only 3 published cases of porcine AA amyloidosis. 4,8,14 In pigs, bacterial diseases occur frequently, and circulating SAA concentrations increase during the infection. 13 Further, circulating SAA concentrations increase in pigs experimentally infected with Actinobacillus pleuropneumoniae and Staphylococcus aureus, 1 although there are no reports of concurrent AA amyloidosis. The low frequency of AA amyloidosis in pigs, despite the increase in SAA during inflammation, suggests that SAA is poorly amyloidogenic in pigs, although there are no relevant published studies to our knowledge. Pigs have 4 SAA genes, named SAA 1–4, that share high sequence similarity although their N-terminal sequences differ. The N-terminal domain of SAA is critical for the formation of amyloid fibrils in humans and mice. Therefore, porcine SAA isoforms may differ in their amyloidogenic properties. In pigs, SAA3 is the major isoform in circulation, 11 but amyloidogenesis of this isoform is not reported.
Niewold et al found a unique sequence of SAA that was highly similar to that of the N-terminus of SAA2. 4 This sequence was present in AA deposits in the tissue of a pig and might, therefore, play a role in the pathogenesis. However, this is the only published example. Here we identified the unique sequence of SAA reported by Niewold et al in the AA deposits in the tissues of a pig with AA amyloidosis associated with Streptococcus suis infection and characterized its biologic properties.
Materials and Methods
Characterization of the Amyloid Deposits
A 180-day-old underdeveloped pig was euthanized and tissues were fixed with 10% buffered formalin and embedded in paraffin. Paraffin sections were stained with hematoxylin and eosin and with Gram and Congo red stains. For transmission electron microscopy, renal tissue was fixed in 2.5% glutaraldehyde, postfixed in 1% Osmium Tetraoxide(OsO4), and embedded in epoxy resin. Ultrathin sections were mounted on a grid and stained with uranyl acetate and lead citrate. Sections were examined using a transmission electron microscope (JEM-2100; JEOL, Tokyo, Japan). An indirect immunoenzyme polymer method was used to conduct immunohistochemical analyses for SAA and S. suis in paraffin sections of the spleen, kidney, liver, and heart. Dewaxed sections were incubated with a monoclonal antibody against human SAA (Kyowa Medex Co., Ltd., Tokyo, Japan) at room temperature for 1 hour. Peroxidase-conjugated anti-mouse immunoglobulin G (IgG; Histofine Simple Stain MAX-PO (M); Nichirei, Tokyo, Japan) was used as the secondary antibody. For immunohistochemical detection of S. suis, dewaxed sections were microwaved at 90°C for 10 minutes and incubated with a rabbit antibody against S. suis serotype 2 for 1 hour at room temperature. A peroxidase-conjugated anti-rabbit IgG (Histofine; Nichirei) was used as the secondary antibody. Immunoreactions were visualized using 3-amino-9-ethylcarbazole containing hydrogen peroxide (3-Amino-9-ethylcarbazole + high-sensitivity substrate chromogen; DAKO, Tokyo, Japan).
Amyloid fibrils were extracted from formalin-fixed, paraffin-embedded spleen according to the method of Murphy et al. 3 In brief, 0.2 g of paraffin-embedded tissue was dewaxed twice in xylol at 65°C for 1 hour and washed overnight with absolute ethanol and then with 80% ethanol at room temperature. After washing in tap water for 3 hours, tissues were homogenized in 2 ml of 0.15 M sodium chloride using a Dounce homogenizer (Sansyo, Tokyo, Japan). After centrifugation at 15 000 × g for 30 minutes, the precipitate was suspended in 2 ml of 6 M guanidine hydrochloride in 0.1 M Tris buffer, pH 10.4. The suspension was incubated at room temperature for 12 hours and centrifuged at 50 000 × g for 1 hour. The supernatant (amyloid extract) was subjected to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) analysis or mass spectrometry. The amyloid extract (200 μl) was precipitated with methanol–chloroform solution. The protein sediment was dissolved in 20 μl of Laemmli’s sample buffer and heated at 100°C for 5 minutes. Protein components were electrophoretically separated through a 5%–20% gradient polyacrylamide gel, and the proteins were visualized using Coomassie Brilliant Blue R-250 (Bio-Rad, Hercules, California). The amyloid extracts (200 µl) were precipitated in methanol–chloroform solution, as described earlier. The precipitate was dissolved in 50 μl of 100 mM Tris-HCl buffer (pH 8.0) containing 1.2 M urea and digested with 0.5 μg of N-Tosyl-L-phenylalanine Chloromethyl Ketone (TPCK)-treated trypsin (Promega, Madison, Wisconsin) at 37°C for 16 hours. The tryptic digests were acidified using formic acid (<pH 2.0) and centrifuged at 21 500 × g for 15 minutes. The supernatants were subjected to high-performance liquid chromatography (Advance System; AMR, Tokyo, Japan) connected to an electrospray ionization-triple quadrupole mass spectrometer (4000QTRAP; AB Sciex, Framingham, Massachusetts). Extracts were injected into a reversed-phase column (electrospray ionization column [Octa Decyl Silyl (ODS)] 75 μm inner diameter × 100 mm long, 3-μm diameter particles; LC Assist, Tokyo, Japan) that was eluted with a 5%–45% gradient of acetonitrile containing 0.1% formic acid for 60 minutes at 300 nl/min.
The mass spectrometer was operated in selected reaction monitoring mode to analyze QWLSFLGEAYEGAK, that was an N-terminal sequence of SAA2 (XP_003122985.1), and QSFVGEAYEGAK, that was reported as a sequence of deposited AA in porcine AA amyloidosis. Selected reaction monitoring transitions of the peptides were calculated using Skyline Software (MacCoss Lab, Seattle, Washington). 9 Six transitions for each peptide (Supplemental Table 1) were collected in 1 cycle (20 milliseconds dwell time). Ionization was performed using an ion-spray voltage of 2 000 V at a capillary temperature of 200°C. The selectivity of Q1 or Q3 scan types was set to 0.7 Da in full width at half-maximum peak height.
In Vitro Testing of Amyloid Fibril Formation
Genomic DNA was extracted from paraffin sections of liver of the pig (DEXPAT kit; Takara Bio, Japan). Primers were designed to detect the SAA2 gene (Gene ID: 100525680) as well as the nucleotide sequences predicted from the amino acid sequence of the AA peptide determined using mass spectrometry. For polymerase chain reaction (PCR) analysis, the forward primers were designed to anneal within sequences that varied across the AA peptide-coding gene and the exon 2 of SAA2. The forward primers for the SAA2 gene and the gene predicted to encode the AA peptide were 5′-GAGTCCACAGTCAGTGGCTT-3′ and 5′-AGTCCACAGTCAGTCCTTCG-3′, respectively. The sequence of the common reverse primer for both SAA2 and the predicted gene was 5′-ACACCTGGCACATTCTACCC-3′, which includes the sequence corresponding to the intron region of the SAA2 gene. All components of the PCR mixture were purchased (KOD FX; Toyobo, Tokyo, Japan). After incubation at 94°C for 1 minute, genomic DNA was amplified using 35 cycles at 94°C for 1 minute, 57°C for 30 seconds, 72°C for 1 minute, and the reactants were stored at 4°C. The amplicons were resolved electrophoretically through a 1.5% agarose gel and visualized using Midori Green DNA stain (NIPPON Genetics, Tokyo, Japan). The PCR amplicons detected using the primers specific for the AA peptide were extracted from agarose gel and purified (High Pure PCR Cleanup Micro kit; Roche, Mannheim, Germany) and sequenced using a 3730xl DNA Analyzer (Applied Biosystems, Foster City, California).
The N-terminal peptide (QWLSFLGEAYEGAK) of SAA2 (SAA2 peptide) and the AA peptide (QSFVGEAYEGAK) were synthesized by Sigma Genosys (Hokkaido, Japan). Lyophilized peptides were dissolved in 0.5% ammonium chloride to a concentration of 5 mg/ml and stored at −80°C. The stock solutions of the peptides were dried and then dissolved in 10% acetic acid to obtain a concentration of 6 mM. Fibrillation was induced by sonication followed by incubation at 37°C for 24 hours without agitation. Incubated samples were dried on a glass slide, stained with Congo red, and viewed using polarization microscopy. For electron microscopy, samples were diluted 10-fold with distilled water and placed on copper grids covered with a collodion film (Nissin, Tokyo, Japan). The grids were stained with 2% uranyl acetate. The samples were examined using a transmission electron microscope (JEM-2100).
To prepare biotin-labeled amyloid cores, fibrils of the AA peptide prepared in 10% acetic acid solution (described above) were dissolved in 40 mM 4-(2-HydroxyEthyl)-1-PiperazineEthaneSulfonic acid (HEPES)-NaOH, pH 7.4, and disaggregated using sonication. The sonicated fibrils were added to a 3 mM solution of the AA peptide conjugated at its C-terminus to biotin (Sigma Genesys) at a 1:10 (w/w) ratio in HEPES buffer. After incubation at 37°C for 5 days, fibril formation was observed using an electron microscope. The fibrils were labeled with streptavidin conjugated to 5-nm diameter colloidal gold (nanogold) particles (Nanoprobes, Yaphank, New York) and then sonicated. After centrifugation at 20 000 × g, the sediments were dissolved in HEPES buffer in the same volume used for preincubation and then used as the biotin-labeled amyloid core. The biotin-labeled amyloid core was added to a 3 mM solution of SAA2 peptide in HEPES buffer at a ratio of 1:10 (w/w). Fibrillation was induced by incubation of the preparations at 37°C for 2 days. The samples were stained and examined using an electron microscope as described earlier.
Results
Clinical and Pathologic Findings
Suppurative inflammation with bacterial cocci was observed in the spleen (Fig. 1), heart (Supplemental Fig. 1), cerebrum (Supplemental Fig. 2), and kidney of the affected pig. A few small suppurative foci were also present in the liver, lung, and cortical interstitial tissues of the kidneys. The cocci were gram positive and were identified by immunohistochemistry as S. suis type 2 (Fig. 2).
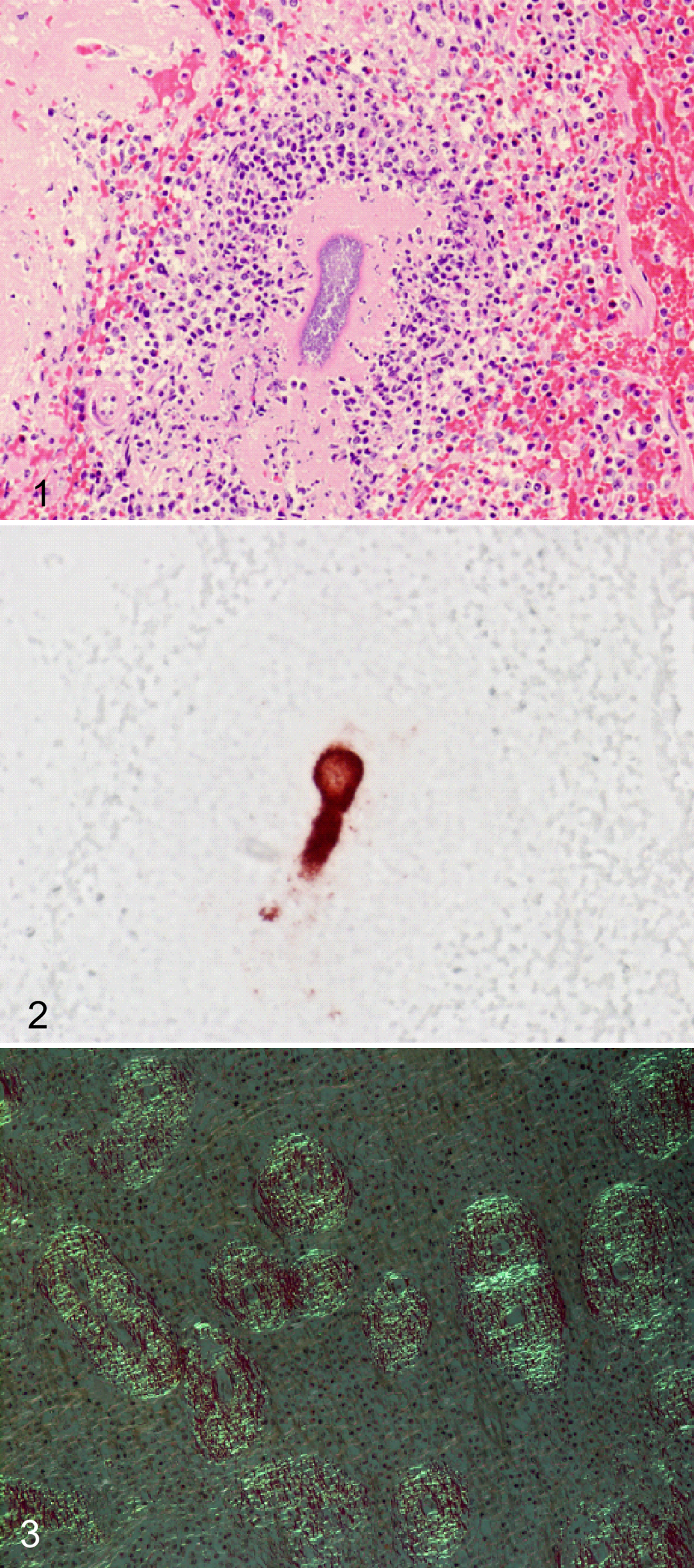
Eosinophilic proteinaceous material was extensively deposited around penicillar and sheathed arteries in the spleen (Supplemental Fig. 3). The same material was moderately deposited around the interlobular arteries in the liver, the glomeruli, and interstitial tissue of the kidney and arteries in the epicardium. The deposits were stained orange by Congo red when observed using brightfield microscopy and showed a characteristic birefringent green when observed using polarized light (Fig. 3).
Immunohistochemical analysis revealed that the deposits in the spleen reacted with an antibody against human SAA (Supplemental Fig. 4). Electron microscopy showed the accumulation of rigid fibrils (10-nm diameter) in the subendothelium and in the mesangial area of renal glomeruli (Supplemental Fig. 5). According to these histological and immunological findings, it was diagnosed as systemic AA amyloidosis with associated S. suis type 2 infection.
Mass Spectrometry and Bioinformatic Analysis
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis analysis of amyloid fibrils extracted from the spleen revealed a distinct band that migrated at approximately 14 kDa (Supplemental Fig. 6). This molecular mass is similar to those predicted for the porcine SAA proteins SAA2 (14.4 kDa) and SAA3 (14.5 kDa).
Mass spectrometry identified the peptide by detecting multiple transitions for each peptide eluted at the same retention time. The amino acid sequence of the peptide that eluted at 71.2 minutes (Fig. 4a and c) was QSFVGEAYEGAK and that of the peptide eluted at 74.4 minutes (Fig. 4b and d) was QWLSFLGEAYEGAK. Bioinformatic analysis showed that the peptide sequence QWLSFLGEAYEGAK (eluted at 74.4 minutes) is derived from SAA2 (XP_003122985.1) digested by trypsin. The other peptide sequence QSFVGEAYEGAK (eluted at 71.2 minutes) had been identified previously in AA deposits in a pig with amyloidosis. 4 This sequence is highly similar to that of the N-terminus of SAA2 (XP_003122985.1); however, this peptide lacks the second and third residues of SAA2 (2W and 3L) and V replaces the sixth residue (6L) of the SAA2 (Fig. 5a). The nucleotide sequence predicted using the detected amino acid sequence QSFVGEAYEGAK was highly similar to that of the 5′ end of the SAA2 gene (Gene ID: 100525680) but lacked fourth to ninth residues of TGGCTT and sixteenth base of C was replaced by G compared to the sequence encoding the N-terminus of SAA2 (Fig. 5b). The predicted nucleotide sequence was dissimilar to SAA3 (Gene ID: 733603) and SAA4 (Gene ID: 100526034).
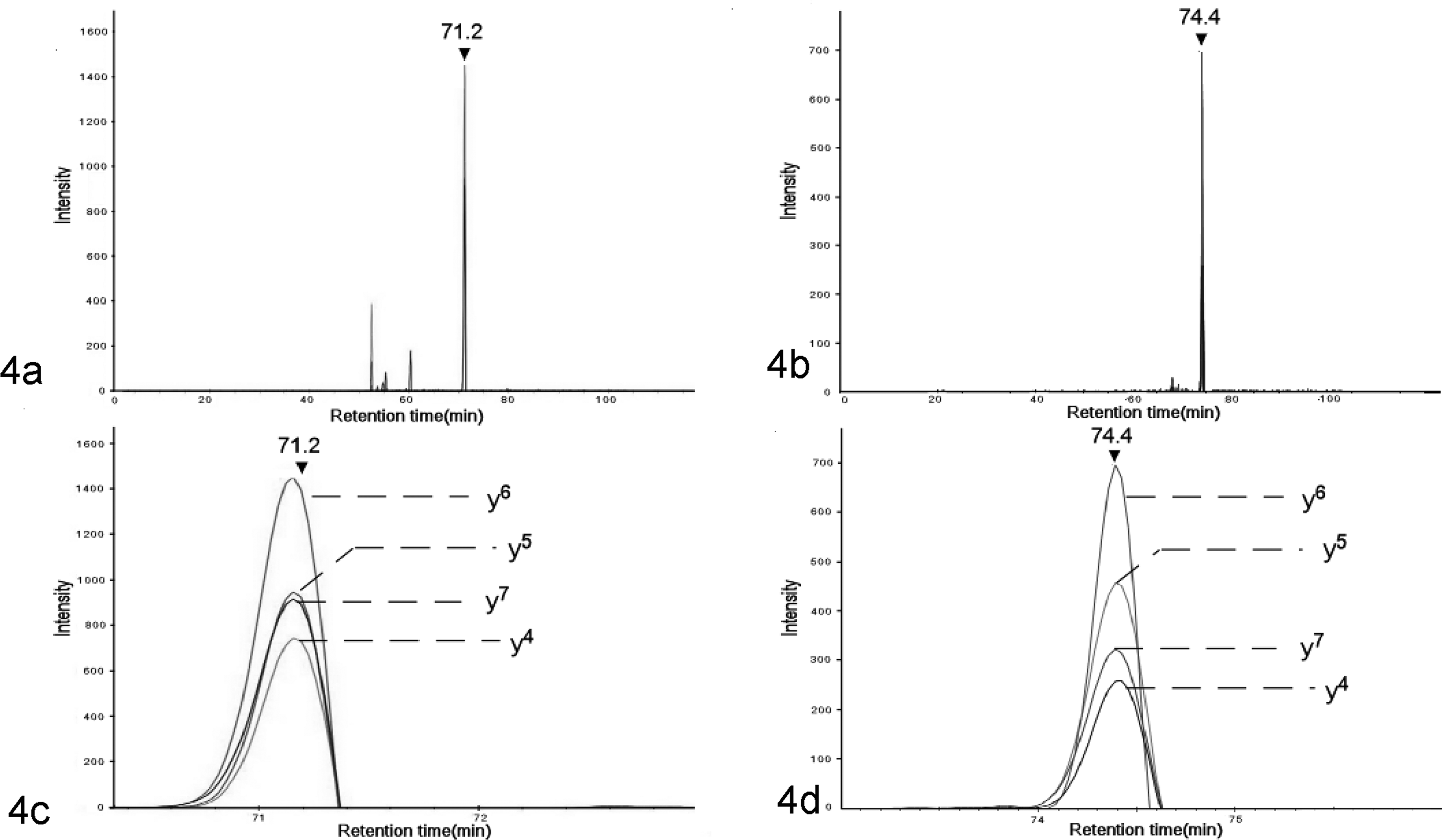
Mass spectrometry of the extracted amyloid from the spleen. Selected reaction monitoring (SRM). (a) Total ion chromatogram for the AA peptides. (b) Total ion chromatogram for N-terminus sequence of serum AA 2 (SAA2). (c) Expansion of chromatograms of the AA peptides. Four peaks derived from each transition for AA peptide were detected at 71.2 minutes. (d) Expansion of chromatograms of the N-terminus of SAA2. Four peaks derived from each transition for SAA peptide were detected at 74.4 minutes.
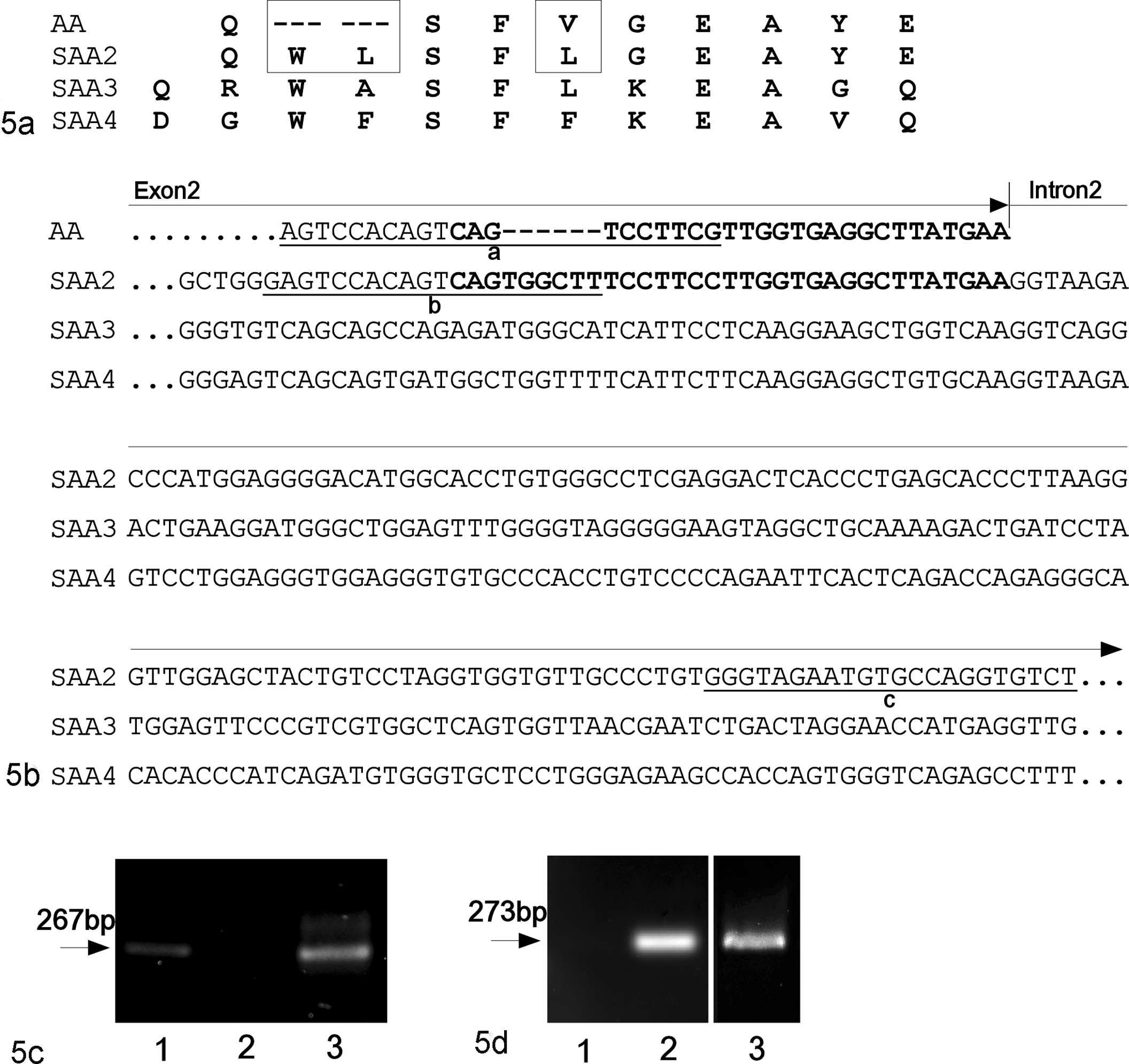
Polymerase chain reaction (PCR) analysis of DNA extracted from the liver of a pig to detect porcine serum AA 2 (SAA2) variants. (a) Amino acid sequence of N-terminus of SAA2, 3, and 4 and the unique sequence of SAA detected in the current amyloidosis case (labeled AA). Boxes indicate differing sequences between SAA2 and AA. (b) Partial genome sequence of exon 2 and intron 2 of SAA2, 3, and 4 and predicted sequence encoding the AA peptide (labeled AA). Boldface characters indicate the sequences encoding the AA peptide and the N-terminus of SAA2. Underlined characters indicate the primer sequences for PCR analysis as follows: sequence “a,” forward primer specific for the predicted nucleotide sequence encoding the AA peptide; sequence “b,” forward primer sequence specific for the SAA2 gene; sequence “c,” reverse primer specific for intron 2 of the SAA2 gene. (c, d) PCR analysis using (c) the forward primer specific for the predicted AA gene (5′-AGTCCACAGTCAGTCCTTCG-3′) and (d) the reverse primer representing the sequence shared by SAA2 (5′-ACACCTGGCACATTCTACCC-3′). AA-sequence-inserted plasmid vector (lane 1) and SAA2-sequence-inserted plasmid vector (lane 2) were analyzed as positive or negative control. The specific bands of AA gene and SAA2 gene were amplified from the DNA extracted from the pig with amyloidosis (lane 3).
Two PCRs were conducted using forward primers specific for exon 2 of SAA2 gene and for the predicted nucleotide sequence encoding the AA peptide, and a single reverse primer specific for intron 2 of SAA2 gene (Fig. 5c and d). Polymerase chain reaction amplicons of 267 bp were detected using the DNA extracted from the liver of the pig of this study and the primers specific for the AA peptide (Fig. 5c). The sequence of this amplicon corresponded to the predicted nucleotide sequence encoding the AA peptide and was registered in the DNA Data bank of Japan (accession No. LC131462). Polymerase chain reaction amplicons of 273 bp were detected using the same DNA extract and the primers specific for the exon 2 of the SAA2 gene (Fig. 5d). Polymerase chain reaction detection using forward primer specific for the AA peptide and reverse primer specific for the intron 2 of SAA2 gene suggested that the AA peptide sequence of the pig studied in this study was derived from the gene encoding the SAA2 variant.
In Vitro Fibrillation Assay
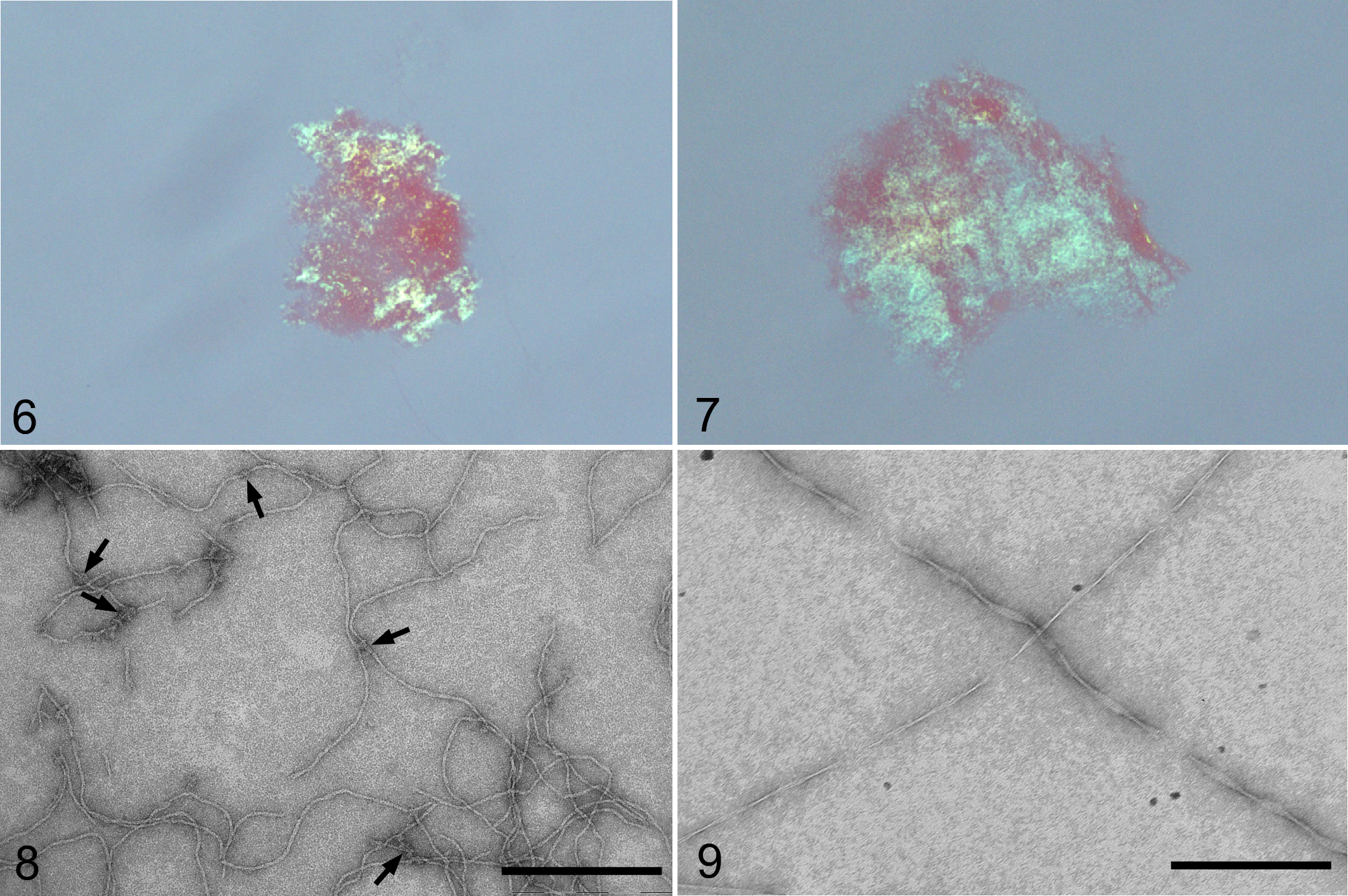
To clarify the amyloidogenic properties of the N-terminus of SAA2 and AA sequences, we conducted an in vitro fibrillation assay of each peptide. After incubation in 10% acetic acid, the synthesized peptides of SAA2 and AA exhibited green birefringence when illuminated separately with polarized light (Fig. 6 and 7). Electron microscopy revealed that the SAA2 peptide formed short, fine fibrils that were arranged in a zigzag pattern (Fig. 8). Further, some fibrils formed by the SAA2 peptide were branched with small crystal-like structures (Fig. 8). The AA peptide formed typical amyloid fibrils, which were long, rigid, and had twisted ribbon patterns (Fig. 9). The widths of the AA peptide fibrils ranged from 10.8 ± 0.8 nm to 2.2 ± 0.2 nm. The mean width of the SAA2 peptide fibrils was 7.0 ± 0.9 nm.
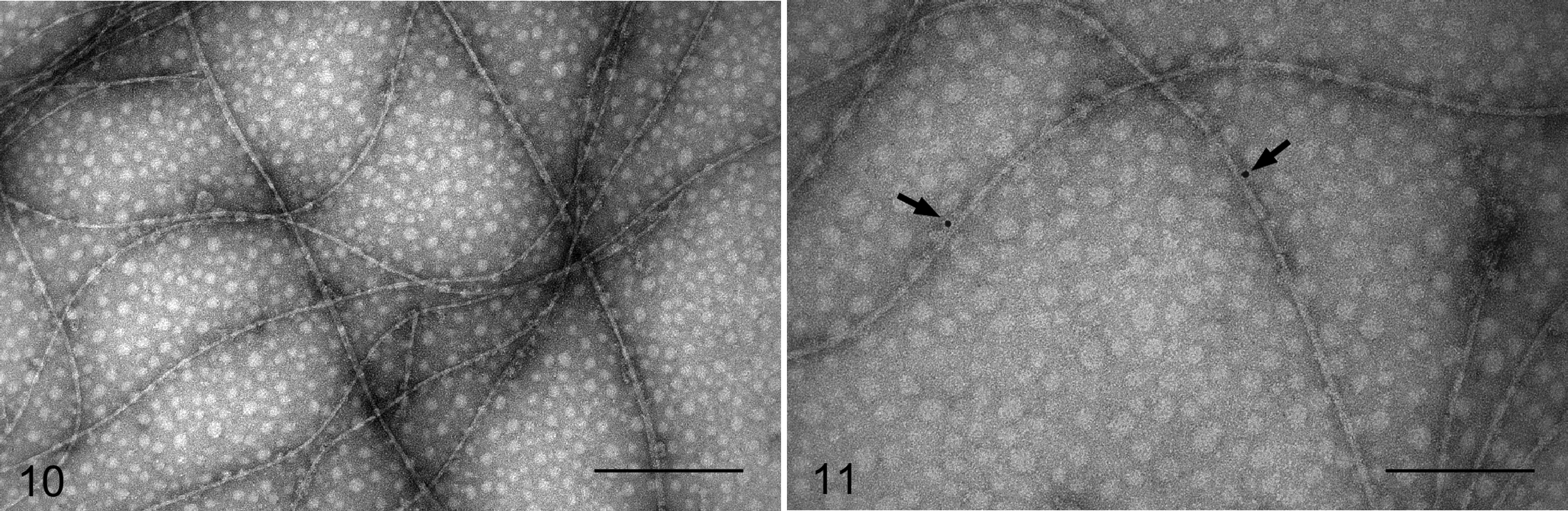
Disaggregated fibrils of the biotin-labeled AA peptide were used as an amyloid seed. In the presence of these disaggregated fibrils, the peptides formed long and rigid fibrils with a twisted ribbon-like structure, similar to those of the fibrils formed by the AA peptides (Fig. 10). Samples were probed with streptavidin conjugated to nanogold particles. Nanogold particles labeled fibrils within these samples, indicating the presence of the biotin-labeled AA peptide in the fibrils (Fig. 11). However, most parts of the fibrils were not labeled with nanogold particles, suggesting that the twisted ribbon-like fibrils were formed mainly from SAA2 peptides that were biotin free and not labeled with nanogold particles.
Discussion
In the present study, histopathological, immunohistochemical, and ultrastructural findings led us to identify an underdeveloped pig with AA amyloidosis associated with S. suis type 2 infection. 6 Increased levels of circulating SAA are present in pigs infected with S. suis 13 ; however, there are no reports of S. suis type 2 infection associated with AA amyloidosis. In this pig, the systemic inflammatory lesions caused by S. suis infection suggest a reactive increase in SAA, although the serum concentration of SAA was not evaluated.
We detected the N-terminus peptide of SAA2 as well as AA peptide in amyloid extracts. The AA peptide was previously identified as a unique sequence of porcine SAA in the tissue of a pig with amyloidosis. 4 The sequence of the AA peptide was similar to the N-terminal sequence of porcine SAA2, and the extracted materials from the spleen showed a single band of 14 kDa by SDS-PAGE analysis, suggesting the presence of an SAA isoform containing the AA peptide sequence. In this case, the detection of SAA2 and AA peptide in amyloid extract suggested that the deposited AA fibrils were formed by multiple SAA isoforms with N-terminal heterogeneity.
Polymerase chain reaction analysis using a specific primer corresponding to the intron region of the SAA2 gene suggested that the affected pig has 2 different gene sequences of SAA2, the sequence reported in NCBI database (Gene ID: 100525680) and a variant sequence that encodes the AA peptide.
An in vitro fibrillation assay showed that the AA peptide and the N-terminal peptide of SAA2 formed amyloid fibrils, which were stained by Congo red. Moreover, there was a morphological difference between the fibrils formed by the AA and SAA2 peptides. Thus, the AA peptide formed typical amyloid fibrils, whereas the SAA2 peptide formed atypical short, zigzagged fibrils.
Interestingly, SAA2 peptide formed long and rigid fibrils in the presence of the fibrils formed by the AA peptide. This observation may be explained by polymerization of amyloid that is induced by cross seeding. Seeding polymerization is postulated to explain the kinetics of fibrillation of amyloid proteins and the pathogenesis of amyloidosis. 2,7,12 Seeding polymerization occurs between different amyloid proteins. 5 For example, human and rat islet amyloid peptides 15 as well as Aβ amyloid and islet amyloid peptide 4 accelerate amyloid fibril formation by heterologous cross seeding. In the pig studied here, the deposited amyloid formed only rigid fibrils, not the zigzagged fibrils that were formed with the SAA2 peptide in vitro. Those findings raise the possibility that amyloid fibrils in the pig with AA amyloidosis were formed via cross-seeding polymerization between SAA2 and an amyloid core formed with the protein that contained the AA peptide sequence. These results indicate the importance of the AA peptide in the development of AA amyloidosis in pigs. The detection of the AA peptide here and by a previous study suggests that pigs with AA amyloidosis have an SAA2-variant gene encoding AA peptide, and SAA2 isoforms increase in serum under inflammatory conditions. It appears that pigs with SAA2 variant containing the AA peptide may be rare because of very low frequency of AA amyloidosis in pigs.
We report here the first example of a pig with AA amyloidosis associated with S. suis infection and presence of SAA2 and the unique AA peptide which was identified in a past case of porcine AA amyloidosis. Detection of this unique AA sequence in both porcine AA amyloidosis cases suggested that this sequence may be involved in the pathogenesis of AA amyloidosis in pigs. This hypothesis is supported by demonstration of in vitro cross seeding between AA peptide and an N-terminus peptide of SAA2. Interaction between AA peptide and SAA2 may be important for the development of AA amyloidosis in pigs.
Footnotes
Acknowledgements
The authors thank Dr. Seigo Ito and Dr. Sachiko Arai (Porcine Clinic Center and Veterinary Internal Medicine I, Department of Veterinary Medicine, Azabu University), Dr. Fujiko Sunaga (Laboratory of Infectious Disease, Department of Veterinary Medicine, Azabu University), and Dr. Munetaka Oi (Toyoura Veterinary Clinic) for supplying the pig with AA amyloidosis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was partially supported by a research project grant awarded by the Azabu University and JSPS KAKENHI, Grant Number 15K07730.
