Abstract
Middle East respiratory syndrome coronavirus (MERS-CoV) was initially isolated from a Saudi Arabian man with fatal pneumonia. Since the original case in 2012, MERS-CoV infections have been reported in >1500 humans, and the case fatality rate is currently 35%. This lineage C betacoronavirus has been reported to cause a wide range of disease severity in humans, ranging from asymptomatic to progressive fatal pneumonia that may be accompanied by renal or multiorgan failure. Although the clinical presentation of human MERS-CoV infection has been documented, many facets of this emerging disease are still unknown and could be studied with animal models. Several animal models of MERS-CoV have been developed, including New Zealand white rabbits, transduced or transgenic mice that express human dipeptidyl peptidase 4, rhesus macaques, and common marmosets. This review provides an overview of the current state of knowledge on human MERS-CoV infections, the probable origin of MERS-CoV, and the available animal models of MERS-CoV infection. Evaluation of the benefits and limitations of these models will aid in appropriate model selection for studying viral pathogenesis and transmission, as well as for testing vaccines and antivirals against MERS-CoV.
Keywords
Middle East respiratory syndrome coronavirus (MERS-CoV) was first isolated from a man in Saudi Arabia in 2012, and MERS-CoV infections have now been reported in >20 countries. 76,79 Originally called human coronavirus-EMC/2012, the virus was renamed MERS-CoV by the International Committee on Taxonomy of Viruses. 20,67 In humans, MERS-CoV is 1 of 6 coronaviruses that cause respiratory disease; however, it is currently the only known pathogenic human lineage C betacoronavirus. 31,68 MERS-CoV infections in humans range from asymptomatic to progressive fatal pneumonia with occasional renal or multiorgan failure. 44,52 To date, published autopsy data from fatal human cases are not available, and the exact route of viral transmission to and among humans is unknown. Additionally, there are currently no approved MERS-CoV-specific countermeasures. Thus, development of animal models to study the pathology and pathogenesis of MERS-CoV, the routes of viral transmission, and the efficacy of treatments and vaccines is critical. Here, we review the current state of knowledge on MERS-CoV infections in humans and potential animal reservoirs for the virus and provide an overview and analysis of animal models of MERS-CoV infection.
Human MERS-CoV Infections
Since the initial case of MERS-CoV, a wide spectrum of disease severity, ranging from asymptomatic to severe fatal pneumonia, has been documented in humans infected with MERS-CoV. 44,52 More than 1500 human cases of MERS-CoV have been identified, with the majority occurring in the Arabian Peninsula. 72,75 Initially, MERS-CoV predominantly affected middle-aged males; however, as cases continue to be reported, the gap between males and females has narrowed, and all age groups have been affected, from children to the elderly. 4,8,9,14,52,56
The median incubation period from MERS-CoV infection to development of disease is approximately 5 days. 9 Individuals typically present with respiratory symptoms, including coughing and shortness of breath, often accompanied by nonspecific signs of disease, such as fatigue, myalgia, arthralgia, fever, headaches, vomiting, or diarrhea. 11,28,44,53,66 Lower respiratory tract disease is most often reported; however, some individuals present with upper respiratory tract symptoms, including a sore throat or rhinorrhea. 19,52 In patients who develop pneumonia, the disease can rapidly progress to acute respiratory distress syndrome that is sometimes associated with acute renal failure, multiorgan failure, and death. 13,59 The case fatality rate is currently 35%. 75 MERS-CoV infection has also rarely been associated with secondary complications, such as disseminated intravascular coagulation, hyperkalemia, or stillbirth. 3,55 Severe disease is most common in individuals with comorbidities such as hypertension, diabetes mellitus, renal or cardiovascular disease, obesity, smoking, or any disease that results in immunosuppression. 9,28,44,52,71 Alternatively, asymptomatic or mild MERS-CoV infections mainly occur in previously healthy immunocompetent individuals; these cases are being more frequently reported due to increased surveillance. 14,23,65
Ancillary tests have detected abnormalities in complete blood counts, biochemistry panels, and radiographs in MERS-CoV-infected patients. Lymphopenia, neutrophilia, thrombocytopenia, and increased levels of C-reactive protein have all been reported. 8,11,53,79 Increases in creatine kinase, alanine aminotransferase, lactate dehydrogenase, creatinine, and blood urea nitrogen have been identified in some individuals, suggesting the development of hepatic and renal disease. 11,28,53,79 Pulmonary interstitial infiltrates have been documented in radiographs from individuals with pneumonia. 9,79 MERS-CoV infections are commonly diagnosed by detecting viral RNA from several specimen types using real-time reverse transcription polymerase chain reaction. 58 Samples obtained from the lower respiratory tract, such as bronchoalveolar lavage samples or sputum, are considered the most reliable for testing purposes since the highest viral loads have typically been found in this location. 19,28 However, MERS-CoV has also occasionally been detected in nasal or oropharyngeal swabs, urine, feces, serum, and blood. 24,28,35,47
Humans acquire MERS-CoV through zoonotic transmission, and bats and dromedary camels are thought to serve as a source of infection. 29,43,51 Additionally, several reports of nosocomial transmission and limited human-to-human transmission in family clusters have been documented, with MERS-CoV potentially spreading through respiratory droplets, direct contact or fomites.* In fact, transmission of MERS-CoV in health care facilities resulted in infection of >180 people in a recent outbreak in South Korea. 37,73,74 The outbreak stemmed from an individual who had traveled to the Middle East, developed clinical signs upon his return to South Korea, and visited multiple hospitals before he was diagnosed with MERS-CoV.73, 74
Molecular Biology and Origin of MERS-CoV
MERS-CoV is an enveloped, nonsegmented, single-stranded, positive-sense RNA virus. The MERS-CoV genome is composed of 30 119 nucleotides with at least 10 open reading frames (ORFs). 67 Two of the proposed ORFs (ORF1a and ORF1b) encode 16 nonstructural proteins important for virus replication. 67 The remaining ORFs encode structural and accessory proteins, including the envelope, membrane, nucleocapsid, and spike proteins. 67
Evidence suggests that the ancestor of MERS-CoV, potentially MERS-CoV itself, may be harbored by bats. Several reports have described a genetic link between MERS-CoV and other coronaviruses detected in bats. The lineage C betacoronaviruses Pipistrellus bat coronavirus HKU5 and Tylonycteris bat coronavirus HKU4 are genetically related to MERS-CoV and likely all emerged from a distant common ancestor centuries ago. 39 Lineage C betacoronaviruses identified in European and African Pipistrellus and Nycteris bats were determined to be genetically similar and phylogenetically related to MERS-CoV. 7 Additionally, in Bisha, Saudi Arabia, a few months after the first human case of MERS-CoV infection was documented, a 190-nucleotide fragment of a coronavirus RNA-dependent RNA polymerase gene was amplified from bat feces and shared 100% sequence identity with the original MERS-CoV isolate from the same geographic region. 45
Although MERS-CoV may have originated in insectivorous bats, humans do not commonly interact with these species, thus limiting the opportunities for viral transmission. Conversely, direct contact between humans and dromedary camels occurs on a regular basis in regions with high numbers of reported human MERS-CoV cases and dromedary camels have been implicated as a source for human MERS-CoV infections. Neutralizing antibodies against MERS-CoV, or a closely related virus, have been detected in dromedary camel serum samples collected from several Middle Eastern and African countries, with seropositivity dating back to 1983. † Moreover, MERS-CoV RNA has been identified in nasal and conjunctival swabs, milk, and, rarely, rectal swabs from dromedary camels. ‡ Although MERS-CoV seropositivity is more common in adult camels, MERS-CoV RNA is most often detected in nasal swabs from juvenile camels, suggesting that active infections typically occur in younger camels. 5,48 MERS-CoV infections in camels appear to be acute and transient. Most infected camels are asymptomatic; however, some camels develop rhinorrhea and an increase in body temperature. 1,10,16,43 Experimental inoculation of young adult dromedary camels with MERS-CoV has been shown to cause mild disease consisting of degeneration and necrosis of the respiratory epithelium in the upper and lower respiratory tract in the absence of pneumonia. 1 Viral antigen and high levels of infectious virus were detected in affected tissues, while the nasal turbinate respiratory epithelium was identified as the main site of virus replication. 1 High amounts of infectious virus were isolated from nasal discharge despite development of only mild clinical disease, 1 which may result in MERS-CoV infections in camels being overlooked, leading to undetected viral transmission among camels and between camels and humans.
Epidemiologic data have suggested a link between MERS-CoV infections in camels and humans. Direct contact with camels or drinking of unpasteurized camel milk has been reported prior to MERS-CoV infection in some humans. 44,59 Genetically similar, or identical, viral sequences have been isolated from camels and individuals caring for these animals, with serologic data in one case providing evidence for camel-to-human transmission. 10,29,43 Although the exact mode of transmission between camels and humans is unknown, it is speculated that transmission may occur through direct contact, ingestion of unpasteurized camel milk, or contact with respiratory droplets or nasal secretions, which have been shown to carry high viral loads. 1,9,27,44 Despite the high prevalence of MERS-CoV in camels, viral transmission from camels to humans appears to be limited, and multiple countries with seropositive camels have yet to detect MERS-CoV infections in humans. 18,32,49
MERS-CoV Species Tropism and the Development of Animal Models of MERS-CoV Infection
Dipeptidyl peptidase 4 (DPP4; also known as CD26) has been identified as the receptor for the MERS-CoV spike protein (S) and is required for viral binding and entry into host cells. 60 DPP4 is a type II transmembrane glycoprotein that is expressed on epithelial and endothelial cells throughout the body. 38 Although DPP4 is evolutionarily conserved, differences in the amino acids present in its extracellular domain, which interacts with MERS-CoV S, have been noted among various animal species and humans. Specifically, 14 amino acids in DPP4 appear to be critical in determining whether the MERS-CoV S can bind to DPP4. 70 MERS-CoV S cannot bind to DPP4 in species that have significant differences in these 14 amino acids as compared with human DPP4 (hDPP4), such as ferrets, hamsters, and mice; thus, these species are resistant to infection. 21,40,62,69 Species with few or no differences in the 14 amino acids seem to be susceptible to MERS-CoV, including rhesus macaques, common marmosets, and camels. 25,50,69 In addition to analyzing the DPP4 amino acid sequence, evaluating the binding energies between MERS-CoV and DPP4 has proven useful in assessing whether MERS-CoV S can bind to DPP4 from different species. A low binding energy between DPP4 and MERS-CoV S has been associated with susceptibility to MERS-CoV infection, whereas high binding energies are reported between MERS-CoV S and DPP4 from nonpermissive species. 25,69 Determination of binding energies can guide the selection of animal species during development of animal models of MERS-CoV infection.
Several laboratory animal species that are often used to model viral diseases are not susceptible to MERS-CoV infection due to differences in critical amino acids in the S-binding domain of their DPP4. 17,21,62 Despite a lack of conventional small animal models, researchers have identified several other species that are susceptible to MERS-CoV infection, including rabbits, hDPP4-transduced or hDPP4-transgenic mice, rhesus macaques, and common marmosets (summarized in Tables 1 and 2). Each animal model of MERS-CoV infection is discussed in detail.
Inoculation Route and Dose, Clinical Disease, and Gross and Microscopic Lesions That Develop in Animal Models of Middle East Respiratory Syndrome Coronavirus Infection.
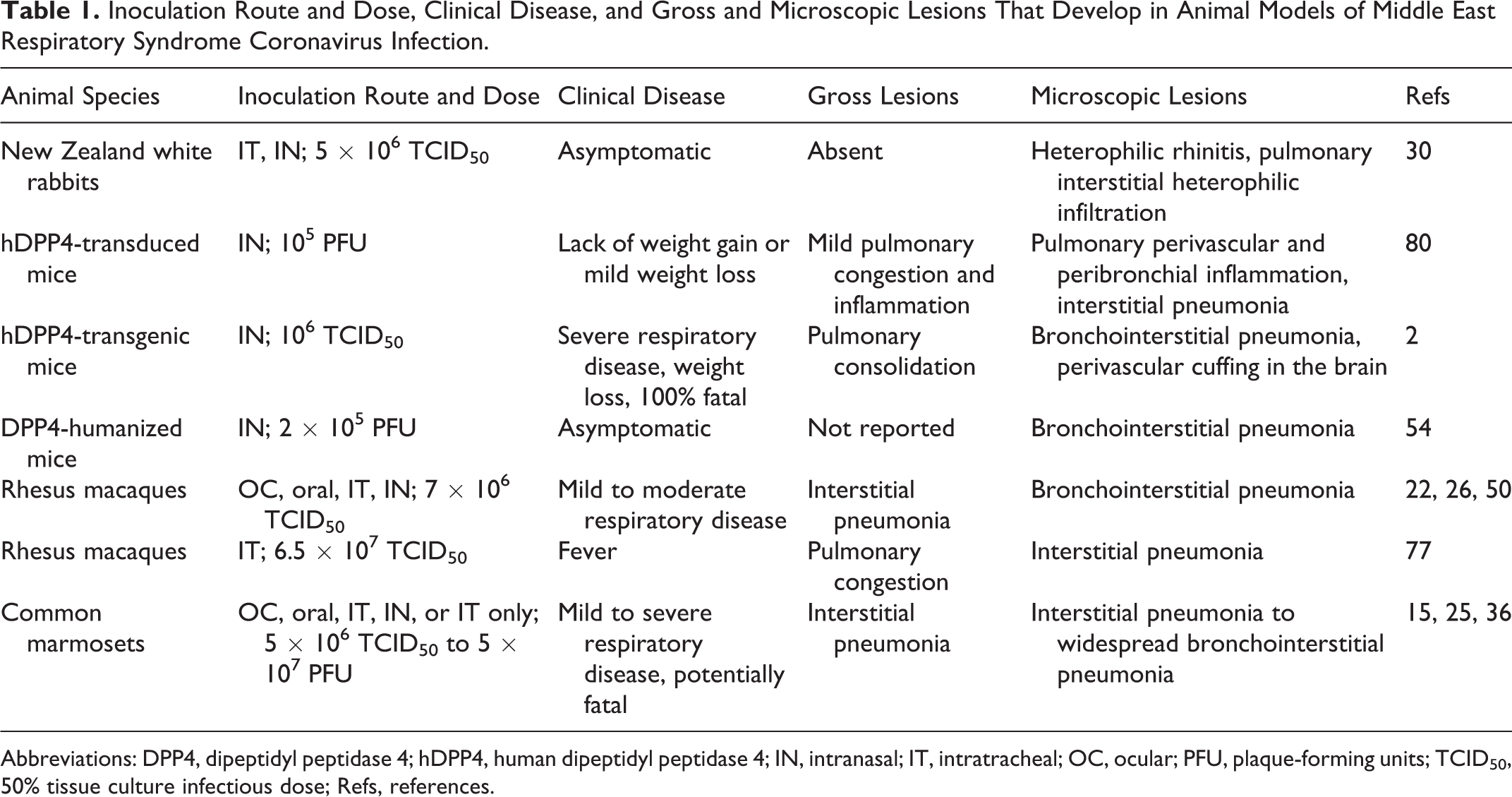
Abbreviations: DPP4, dipeptidyl peptidase 4; hDPP4, human dipeptidyl peptidase 4; IN, intranasal; IT, intratracheal; OC, ocular; PFU, plaque-forming units; TCID50, 50% tissue culture infectious dose; Refs, references.
Tissues and Cell Types Containing Viral RNA, Viral Antigen, or Infectious Virus in Animal Models of MERS-CoV Infection.
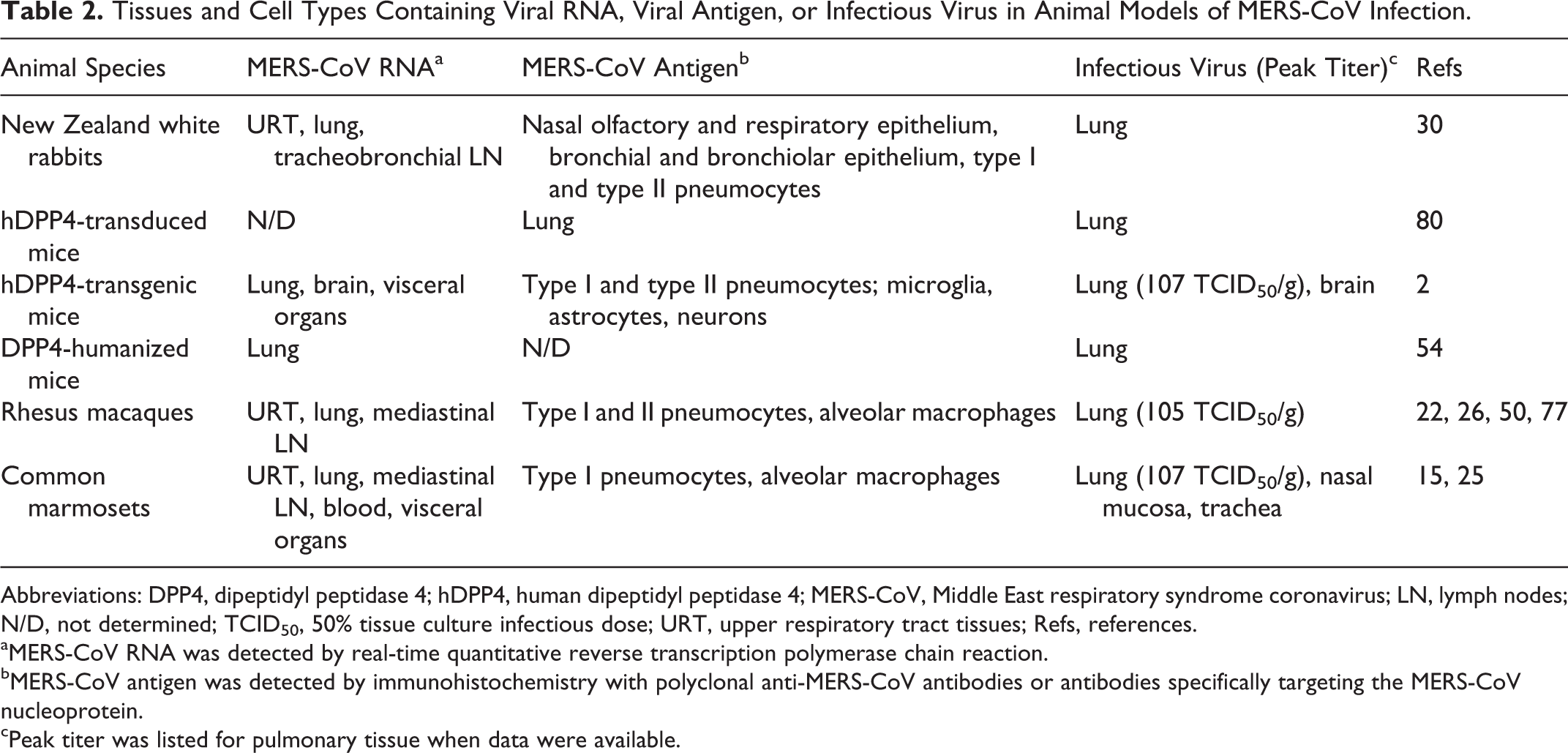
Abbreviations: DPP4, dipeptidyl peptidase 4; hDPP4, human dipeptidyl peptidase 4; MERS-CoV, Middle East respiratory syndrome coronavirus; LN, lymph nodes; N/D, not determined; TCID50, 50% tissue culture infectious dose; URT, upper respiratory tract tissues; Refs, references.
aMERS-CoV RNA was detected by real-time quantitative reverse transcription polymerase chain reaction.
bMERS-CoV antigen was detected by immunohistochemistry with polyclonal anti-MERS-CoV antibodies or antibodies specifically targeting the MERS-CoV nucleoprotein.
cPeak titer was listed for pulmonary tissue when data were available.
Rabbits
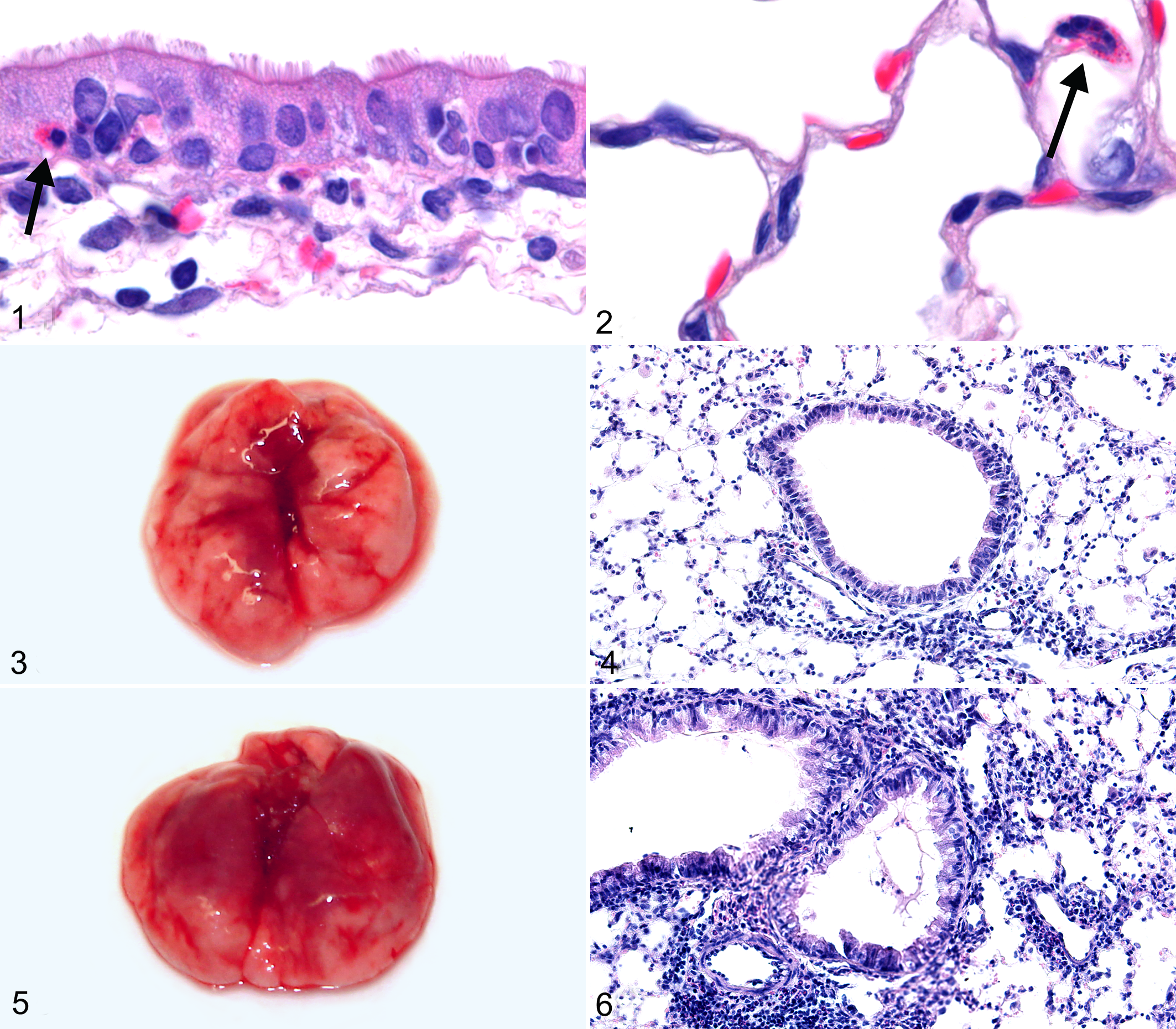
New Zealand white rabbits have been described as an animal model of asymptomatic MERS-CoV infection (Table 1). 30 Clinical signs and gross lesions were not observed in rabbits inoculated through intratracheal and intranasal routes. Microscopically, lesions were detected in the upper and lower respiratory tract at 3 and 4 days postinoculation (dpi). In the nasal cavity, there was a mild to moderate infiltration of heterophils that in some cases was accompanied by epithelial necrosis and regeneration (Fig. 1). The lungs exhibited mild heterophilic infiltration in alveolar septa and lumina, predominantly around terminal bronchioles (Fig. 2). Type II pneumocyte hypertrophy and hyperplasia of bronchus-associated lymphoid tissue were also observed.
Virus was detected in tissues in the upper and lower respiratory tract by real-time quantitative reverse transcription polymerase chain reaction, in situ hybridization, and immunohistochemistry (IHC; Table 2). Additionally, virus titration revealed the presence of infectious virus in nasal swabs up to 7 dpi. Although New Zealand white rabbits could be used to study MERS-CoV transmission, since they shed MERS-CoV from their upper respiratory tract, they are not suitable for studying clinical disease progression, given that they remained asymptomatic after MERS-CoV inoculation. Overall, New Zealand white rabbits seem to be of limited value for modeling MERS-CoV infections.
Mice
Although wild-type mice are not susceptible to MERS-CoV infection, 17 several mouse models of MERS-CoV infection that express hDPP4 have been developed (summarized in Tables 1 and 2). In the first model, an adenoviral vector was used to induce transient expression of hDPP4 in the lungs of BALB/c mice (National Cancer Institute, Frederick, MD, USA), C57BL/6 mice (National Cancer Institute), and multiple knockout mouse strains. 80 hDPP4 was expressed only by epithelial cells lining the airways and alveoli. After intranasal inoculation with MERS-CoV, clinical signs in the hDPP4-transduced wild-type mice were minimal and characterized by lack of weight gain in young mice and mild weight loss in older mice. Mice exhibited mild gross pulmonary lesions, which corresponded to peribronchiolar and perivascular inflammation that developed into interstitial pneumonia (Figs. 3, 4). IHC showed that MERS-CoV antigen was colocalized with hDPP4 expression in the lungs. Virus replication was detected in the lungs by 2 dpi; however, viral infection was transient. Clearance of MERS-CoV from the lungs was faster in younger mice. Transduced knockout mice—which had impaired immune systems, such as a lack of type I IFN signaling (IFNAR-/-; The Jackson Laboratory, Bar Harbor, ME, USA)—typically exhibited earlier, more severe clinical signs and gross and microscopic lesions than infected transduced wild-type mice (Figs. 5, 6). In all mouse strains, gross and microscopic lesions and virus replication were not present in organs outside the respiratory tract, which fits with the lack of detectable hDPP4 in these tissues.
Since hDPP4-transduced mice exhibited mild transient clinical disease and pulmonary lesions, these mice could be used to model mild MERS-CoV disease. However, this model cannot be used to analyze clinical respiratory disease, since it lacks respiratory clinical signs. An increase in clinical disease severity with earlier onset of gross and microscopic lesions was achieved by transducing immunodeficient knockout mice. Due to the ease of transducing different strains of knockout mice, this technique could be used to study moderate MERS-CoV disease and the impact of different aspects of the immune system on MERS-CoV infections. However, disease and hDPP4 expression are transient in both wild-type and immunodeficient hDPP4-transduced mouse models, which may interfere with antiviral and vaccine screening. Additionally, the expression of hDPP4 in these mouse models may not exemplify normal hDPP4 expression, as hDPP4 in these mice is solely expressed in the lungs with widespread expression by airway epithelial cells and pneumocytes and is not present in other cell types or tissues, including the upper respiratory tract. Thus, the physiologic response to MERS-CoV infection may not fully represent that in infected humans.
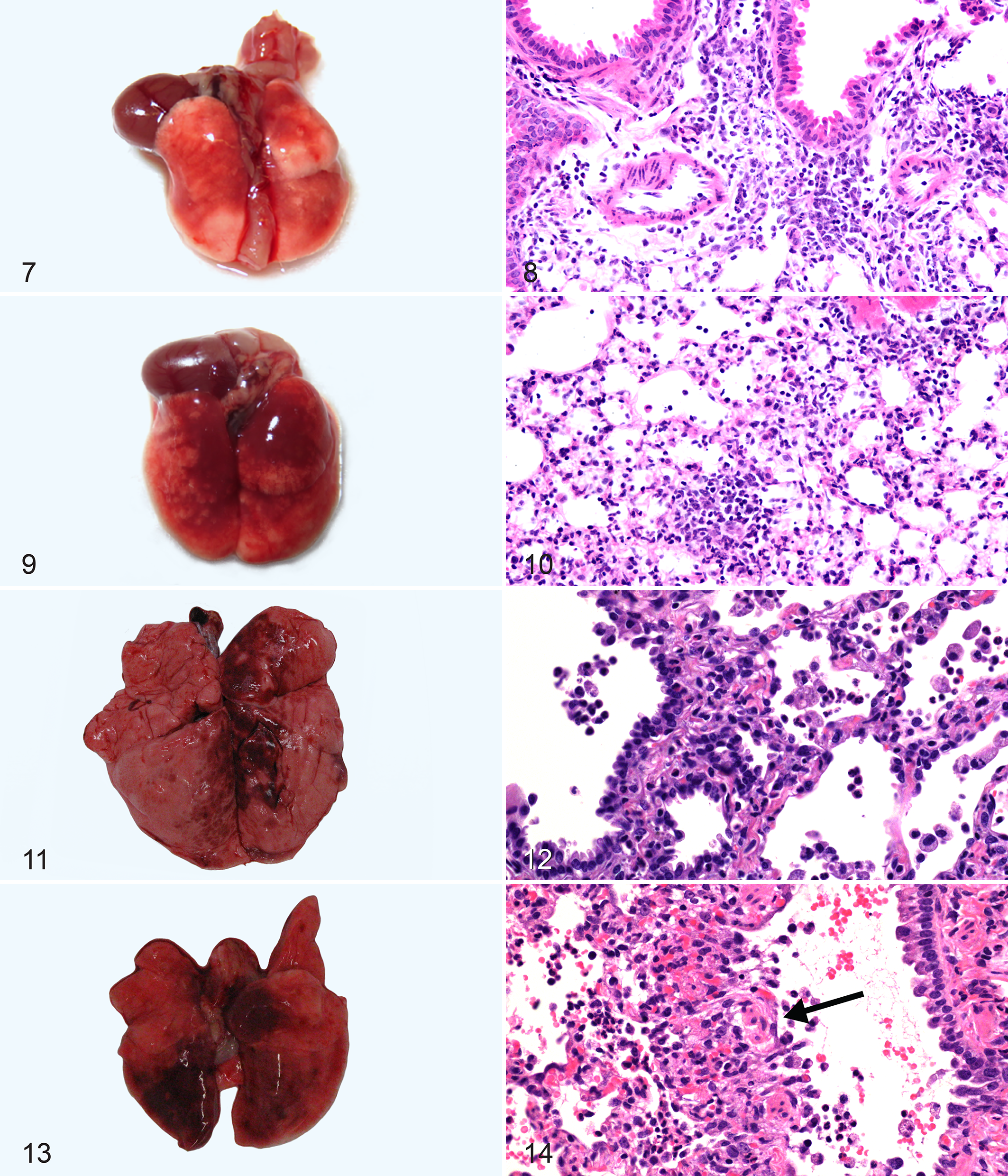
Transgenic mice expressing hDPP4 were created to model MERS-CoV infection. In one model, hDPP4-transgenic mice were developed on either a C57BL/6 J or C57BL/6 J × B6C3F1/J background and then crossed with C57BL/6 J or B6C3F1/J mice. 2 These hDPP4-transgenic mice globally expressed hDPP4—on their pulmonary pneumocytes, renal epithelium, neurons in the brain, and endothelial cells in multiple tissues—and additionally maintained normal expression of mouse DPP4. 2 Intranasal inoculation with MERS-CoV resulted in severe progressive respiratory disease that was fatal by 6 dpi in 100% of the hDPP4-transgenic mice. Gross lesions were noted only in the lungs, while microscopic lesions were observed in the lungs and brain (Figs. 7 –10). Red areas of pulmonary congestion and consolidation were more extensive at 4 dpi than 2 dpi. Histologically, mice exhibited bronchointerstitial pneumonia with perivasculitis. A single mouse exhibited perivascular cuffing in the brain.
In the hDPP4-transgenic mice, high levels of infectious virus were detected in the lung at 2 dpi, with lower levels detected at 4 dpi. Infectious virus was found only at 4 dpi in the brain. Viral RNA was also present in visceral organs, such as the heart and spleen, suggesting that widespread viral dissemination had occurred. Infectious virus and viral RNA were not present in the kidneys.
Currently, the hDPP4-transgenic mouse is the only available small animal model of severe, lethal MERS-CoV infection that develops respiratory disease. However, these mice globally express hDPP4 in all cell types, unlike normal DPP4 expression in humans. The widespread expression of hDPP4 in the transgenic mice resulted in the development of nervous system lesions as well as lesions in the respiratory tract, which is likely not representative of human MERS-CoV infections. Nevertheless, this model could still be used to screen the efficacy of antivirals and vaccines to mitigate or prevent MERS-CoV-induced respiratory disease.
In a different transgenic approach, VelociGene technology was used to replace the mouse DPP4 ORF with the hDPP4 ORF. 54 In these mice, DPP4 was expressed solely by cell types that would normally express mouse DPP4, thus retaining authentic DPP4 expression patterns. After intranasal MERS-CoV inoculation, DPP4-humanized mice remained asymptomatic yet developed pulmonary lesions consistent with bronchointerstitial pneumonia by 4 dpi. High levels of MERS-CoV RNA and infectious virus were detected in the lungs. Since the DPP4-humanized mouse model was asymptomatic, these mice could be used to model mild MERS-CoV infections, but they are not an appropriate model for severe MERS-CoV disease. Although both the DPP4-humanized and hDPP4-transgenic mouse models could be used to screen antivirals and vaccines, each model has its own advantages. The DPP4-humanized mouse model may be preferred since these mice do not develop brain lesions and may best simulate hDPP4 expression patterns, whereas the hDPP4-transgenic mouse model may be favored because it is a lethal model, which allows for easier determination of antiviral and vaccine efficacy.
Rhesus Macaques
Koch’s postulates for MERS-CoV were fulfilled in rhesus macaques, which were the first animal model developed for MERS-CoV infection. Rhesus macaques exhibited acute, transient, mild to moderate respiratory disease, including tachypnea, deep abdominal breathing, and coughing, in addition to fever and anorexia. 22,26,50,77 Clinical signs were observed by 1 dpi and resolved as early as 4 dpi. 22 Correspondingly, serial complete blood counts showed development of lymphopenia and leukocytosis accompanied by neutrophilia 1 dpi, which resolved by 3 dpi. 22
Rhesus macaques developed gross lesions only in the lungs, which consisted of firm, edematous, bright to dark red discolored foci (Fig. 11). 50,77 Histologic pulmonary lesions were described as either an interstitial pneumonia 77 or a bronchointerstitial pneumonia, which centered on terminal bronchioles, with development of hyaline membranes and type II pneumocyte hyperplasia (Fig. 12). 22,26 In pulmonary lesions and serum, transient upregulation of proinflammatory mediators, including interleukin 6 and chemokine (C-X-C motif) ligand 1, was identified. 22 Additionally, MERS-CoV-specific neutralizing antibodies were detected in serum samples as early as 7 dpi, indicating that an immune response directed at MERS-CoV infection had developed. 77
Infectious virus was isolated from the lungs, 50,77 and MERS-CoV RNA was detected in several upper and lower respiratory tract tissues (Table 2). 22 MERS-CoV RNA was also identified in nasal swabs, bronchoalveolar lavage samples, and a few oropharyngeal swabs. 22 Despite the presence of viral RNA and evidence for viral shedding from the upper respiratory tract, lesions and virus replication were observed only in lower respiratory tract tissues, with virus replication occurring in type I and type II pneumocytes. 22 Sites of virus replication and the location of viral antigen, as determined by IHC, correlated with the reported sites of cellular expression of DPP4 in the lungs of rhesus macaques. 22,69 Viral antigen in the lung was exclusively present in areas of pneumonia. 22,77 Viral RNA was not present in blood or any visceral organs, including the kidney. 22,77
The transient respiratory disease that developed in MERS-CoV-infected rhesus macaques models human cases of mild MERS-CoV infections. Macaques also exhibit complete blood count abnormalities similar to those reported in humans. As such, macaques can be used to study the pathogenesis of mild MERS-CoV infections. Additionally, rhesus macaques are a suitable model to confirm the efficacy of vaccines and antivirals.
Common Marmosets
MERS-CoV administered through a combination of ocular, oral, intratracheal, and intranasal routes caused severe, potentially lethal respiratory disease in common marmosets (Table 1). 25 Clinical signs included tachypnea, labored or shallow breathing, cyanosis, and hemorrhagic oral discharge, which necessitated early euthanasia of 2 of 9 marmosets. Clinical signs were first noted 1 dpi, peaked at 4 to 6 dpi, and resolved by 13 dpi. Correspondingly, radiographs showed pulmonary interstitial infiltration as early as 1 dpi; infiltrates were no longer evident by 13 dpi.
Gross lesions were present only in the lungs; affected lobes were dark red and edematous and failed to collapse (Fig. 13). Microscopically, gross lesions corresponded to foci of bronchointerstitial pneumonia that centered on terminal bronchioles and extended into adjacent parenchyma (Fig. 14). The pneumonia was widespread, moderate to severe, and in some cases diffusely affected the lung. Bronchointerstitial pneumonia was characterized by development of hyaline membranes, alveolar edema, hemorrhage, fibrin, and type II pneumocyte hyperplasia often with severe airway lesions affecting bronchi and bronchioles. In foci of pneumonia, there was upregulation of RNA for molecules involved in proinflammatory and antiviral pathways and fibrogenesis.
IHC detected abundant MERS-CoV antigen within foci of pneumonia in type I and type II pneumocytes and macrophages (Table 2). In situ hybridization indicated that type I pneumocytes and alveolar macrophages were the main sites of virus replication, and IHC showed that these cell types expressed DPP4. High levels of viral RNA were detected in the lung, while lower levels of RNA were present in upper respiratory tract tissues and nasal and oropharyngeal swabs. Infectious virus was isolated from both upper and lower respiratory tract tissues. MERS-CoV RNA was also detected in blood and several visceral organs, including the kidney. Identification of viremia and viral RNA in multiple organ systems suggested that MERS-CoV widely disseminated throughout the body of marmosets; however, lesions were present only in the respiratory tract. These findings were confirmed by Chan et al, who reported that intratracheal inoculation of marmosets with MERS-CoV caused similar clinical, pathologic, and virologic results. 15 However, intratracheal inoculation with MERS-CoV by Johnson et al resulted only in mild to moderate respiratory disease and interstitial pneumonia, which were attributed to experimental manipulations rather than virus replication. 36 Differences in disease development between these experiments could be due to the extended passage history of the MERS-CoV strain used by Johnson et al, which resulted in multiple amino acid changes, 36 or differences in the inoculum volume or the age and source of the common marmosets. Thus, MERS-CoV can result in severe, potentially lethal respiratory disease and associated high pulmonary viral loads in common marmosets, indicating that marmosets can be used to confirm the efficacy of vaccines and antivirals and to study the pathogenesis of severe human cases of MERS-CoV infection.
Advantages and Limitations of Animal Models of MERS-CoV Infection
Several animal models of MERS-CoV infection have been developed, each of which has its own set of benefits and limitations (summarized in Table 3). Rabbits, hDPP4-transduced and hDPP4-transgenic mice, rhesus macaques, and common marmosets were all susceptible to MERS-CoV infection, although disease severity was markedly different among the animal models. Compared with the other species, New Zealand white rabbits are of limited value for modeling MERS-CoV infections, as they remain asymptomatic and develop only mild histologic respiratory tract lesions. The main advantage to using hDPP4-transduced mice over the other models is that wild-type and knockout mouse strains can be rapidly generated and used to study the importance of select host factors in regard to MERS-CoV pathogenesis. In contrast, hDPP4-transgenic mice take longer to generate, but they are currently the only available small animal model of MERS-CoV that is 100% lethal, exhibits respiratory clinical disease, and develops viremia. Although hDPP4-transgenic mice globally overexpress hDPP4, lesions are observed only in the respiratory tract and rarely the brain. Although development of lesions in the nervous system is likely not representative of human MERS-CoV infections, the brain lesions were mild, present in only 1 mouse in the study, and did not result in neurologic signs. As such, the hDPP4-transgenic mouse model could be used to screen antiviral and vaccine efficacy. In comparison, the DPP4-humanized mice retain authentic DPP4 expression patterns, develop pulmonary lesions, and yet lack nervous system lesions. Thus, of the 3 available small animal models, the DPP4-humanized mouse may be the best choice for elucidating the pathogenesis of MERS-CoV infections and for studying the effect of comorbidities on the pathogenesis of MERS-CoV infections.
Advantages and Limitations of Animal Models of MERS-CoV Infection.
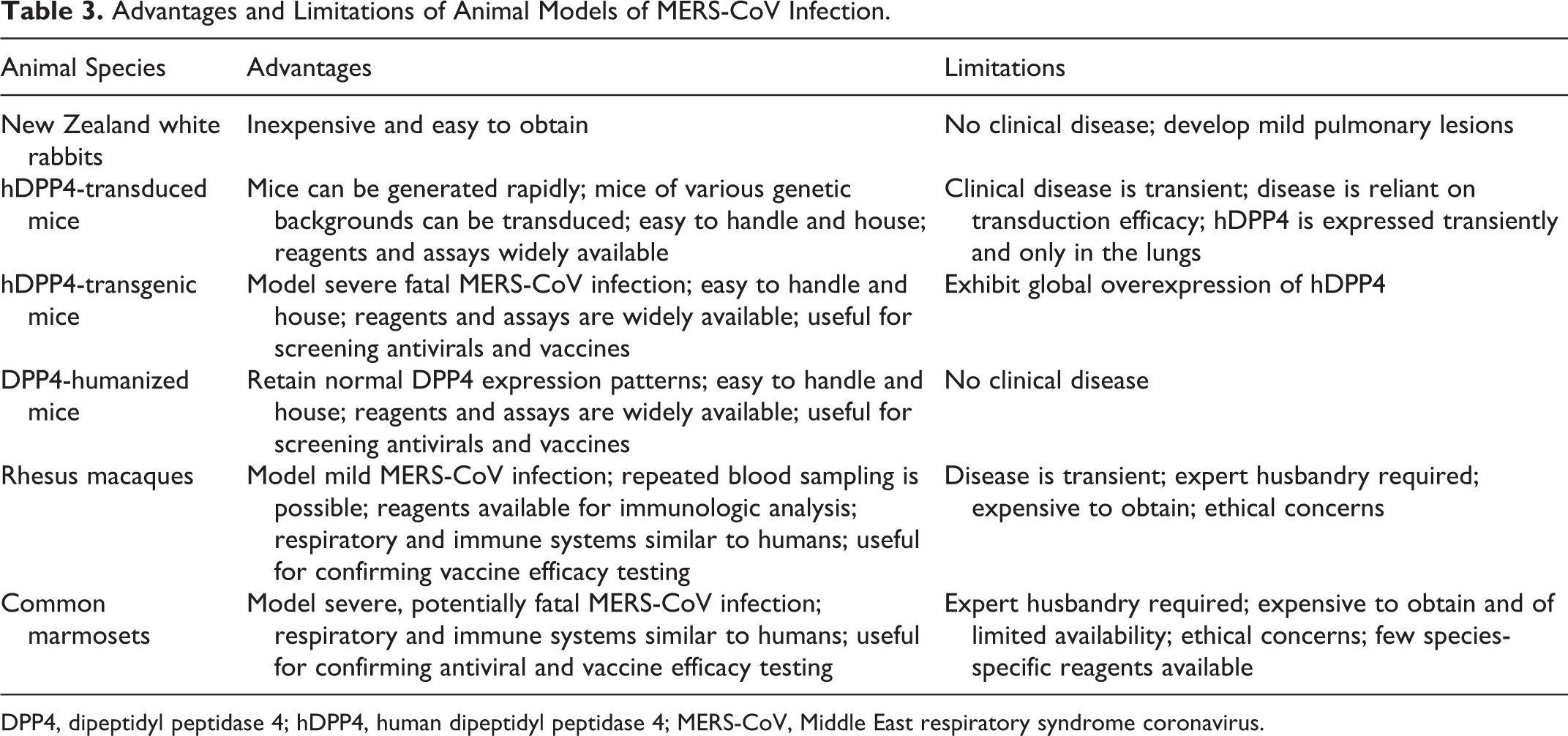

DPP4, dipeptidyl peptidase 4; hDPP4, human dipeptidyl peptidase 4; MERS-CoV, Middle East respiratory syndrome coronavirus.
In contrast to the New Zealand white rabbit, hDPP4-transduced mice, and DPP4-humanized mice, both nonhuman primate models developed respiratory disease and pneumonia, which corresponds to what has been reported in humans. Additionally, the nonhuman primates have respiratory and immune systems that are more similar to those of humans. In regard to clinical disease, rhesus macaques are more suited to model mild human cases of MERS-CoV infection, while common marmosets are a better model for severe MERS-CoV disease. In terms of hematologic parameters, macaques are preferred over marmosets since the larger size of the macaques allows repeated blood sampling over short periods. Although both nonhuman primate models can satisfy confirmatory efficacy testing requirements for vaccines and antivirals under the Food and Drug Administration animal efficacy rule, the lower body weight and development of more severe disease make common marmosets better suited for antiviral testing. Even though the nonhuman primate models recapitulate human MERS-CoV infections quite well, these models have several disadvantages versus the small animal models, including difficulties in handling larger animals in biocontainment, increased housing costs, and added ethical concerns surrounding the use of nonhuman primates in research.
Acute renal failure has been reported in some humans with MERS-CoV infections, although this has not been demonstrated in any of the currently available animal models. A lack of renal disease in the animal models of MERS-CoV infection may not indicate that MERS-CoV has a tropism for the kidney only in humans, since the acute renal failure in humans appears to develop late in the disease course and is thought to occur secondary to shock or in association with previous renal disease rather than as a result of viral-induced injury. In support of this, common marmosets exhibited viremia, and viral RNA was detected in their kidneys despite a lack of renal disease, suggesting that renal failure is not a direct consequence of MERS-CoV infection.
Conclusions and Future Directions
Taken together, the animal models reviewed here replicated the wide range of disease severity observed in MERS-CoV-infected humans, which spans from asymptomatic to severe fatal pneumonia. Although none of the currently available animal models are perfect and efforts should be made to improve them, these animal models can nevertheless be used to address gaps in our knowledge of MERS-CoV disease. Ideally, in the future, data will be collected from fatal human MERS-CoV infections and used to determine which animal model best represents the disease that develops in humans. Appropriate selection of the most suitable animal model will allow researchers to study the pathogenesis and transmission of MERS-CoV and aid in the development and testing of antivirals and vaccines.
Footnotes
Notes
Acknowledgements
We are extremely grateful to Drs Bart Haagmans and Judith van den Brand (Figs. 1, 2),
30
Stanley Perlman, David Meyerholz, and Jincun Zhao (Figs. 3
–6),
80
Chien-Te Tseng and Xinrong Tao (Figs. 7
–![]() ),
2
and Darryl Falzarano (Figs. 11, 13)
25,26
for providing gross and/or histologic images and to Ryan Kissinger for his help with editing figures. The images in Figures 3
–6 are copyrighted and have been reproduced with permission from Proceedings of the National Academy of Sciences of the United States of America. The images in Figures 7
–10 are copyrighted and have been reproduced with permission from the American Society for Microbiology.
),
2
and Darryl Falzarano (Figs. 11, 13)
25,26
for providing gross and/or histologic images and to Ryan Kissinger for his help with editing figures. The images in Figures 3
–6 are copyrighted and have been reproduced with permission from Proceedings of the National Academy of Sciences of the United States of America. The images in Figures 7
–10 are copyrighted and have been reproduced with permission from the American Society for Microbiology.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Intramural Research Program of the National Institutes of Health, National Institute of Allergy and Infectious Diseases. Dr Baseler was supported in part by the National Institutes of Health Comparative Biomedical Scientist Training Program.
