Abstract
This article reviews the roles that laterally transferred genes (LTG) play in the virulence of bacterial pathogens. The features of LTG that allow them to be recognized in bacterial genomes are described, and the mechanisms by which LTG are transferred between and within bacteria are reviewed. Genes on plasmids, integrative and conjugative elements, prophages, and pathogenicity islands are highlighted. Virulence genes that are frequently laterally transferred include genes for bacterial adherence to host cells, type 3 secretion systems, toxins, iron acquisition, and antimicrobial resistance. The specific roles of LTG in pathogenesis are illustrated by specific reference to Escherichia coli, Salmonella, pyogenic streptococci, and Clostridium perfringens.
Bacterial genomes frequently contain a significant amount of foreign DNA, which is DNA that originated from another organism and has inserted into the genome of a bacterium. 29,38,42,50,98 The exotic nature of this acquired DNA is indicated by a difference in G+C percentage compared with that of the core genome, whose genes tend to have a G+C percentage characteristic of the species. 98 A plot of G+C content will frequently identify multiple areas of foreign DNA in a genome. Difference in codon usage is another marker of foreign DNA. 98 Furthermore, regions adjacent to a segment of foreign DNA tend to be marked by genetic elements associated with transfer of DNA, such as insertion sequences (ISs), tandem or inverted repeats, integrase genes, or associated integration hotspots. At the population level, genes received through transfer often show a restricted phylogenetic distribution when large numbers of strains are examined. 98 Depending on the frequency of such transfers and the time at which they occurred, this can be a restriction to only one or few species within a genus, to a single lineage within a bacterial species, or to multiple unrelated lineages.
Transfer of foreign DNA is also called horizontal or lateral gene transfer (LGT), and the DNA mobilized into the host bacterium is referred to as mobile genetic elements (MGEs). 37,42,115,124 MGEs have a huge impact on bacterial genomes, including causing marked differences in genome size and pathogenicity. Although bacteria have several mechanisms for resisting LGT, 104 MGEs play a major role in bacterial evolution and contribute greatly to adaptation to new and changing ecological niches. 134
For pathogens, the animal host is one of the environments that they have evolved to exploit—sometimes a specific site 9,26 in the host and sometimes several sites. 47 MGEs often carry fitness genes associated with entry and survival in the host. 50,133 In Escherichia coli, for example, pathogenic E. coli O157:H7 has a genome size of 5.5 Mb; uropathogenic E. coli CFT073, 5.23 Mb; and nonpathogenic E. coli K-12, 4.6 Mb. 50,133 The gain and loss of MGEs is responsible for the versatility of E. coli as a commensal and as a pathogen able to survive in a range of environments inside and outside the host.
Mechanisms of Transfer of DNA
The MGEs are transferred by 3 methods: conjugation, transduction, and transformation (Fig. 1). 42,98 Conjugation requires contact between donor and recipient bacteria (Fig. 1). 98 Typically, conjugation involves a type IV–like secretion system, with a sex pilus in Gram-negative bacteria or with a direct adhesin-based cell-to-cell attachment and translocation apparatus in Gram-positives, through which DNA is transferred. 6,98 Although this seems to be the main mechanism of conjugative transfer of MGEs, other mechanisms have been recently identified, which include transfer mediated by a single protein multimer 10 and by nanotubes. 33
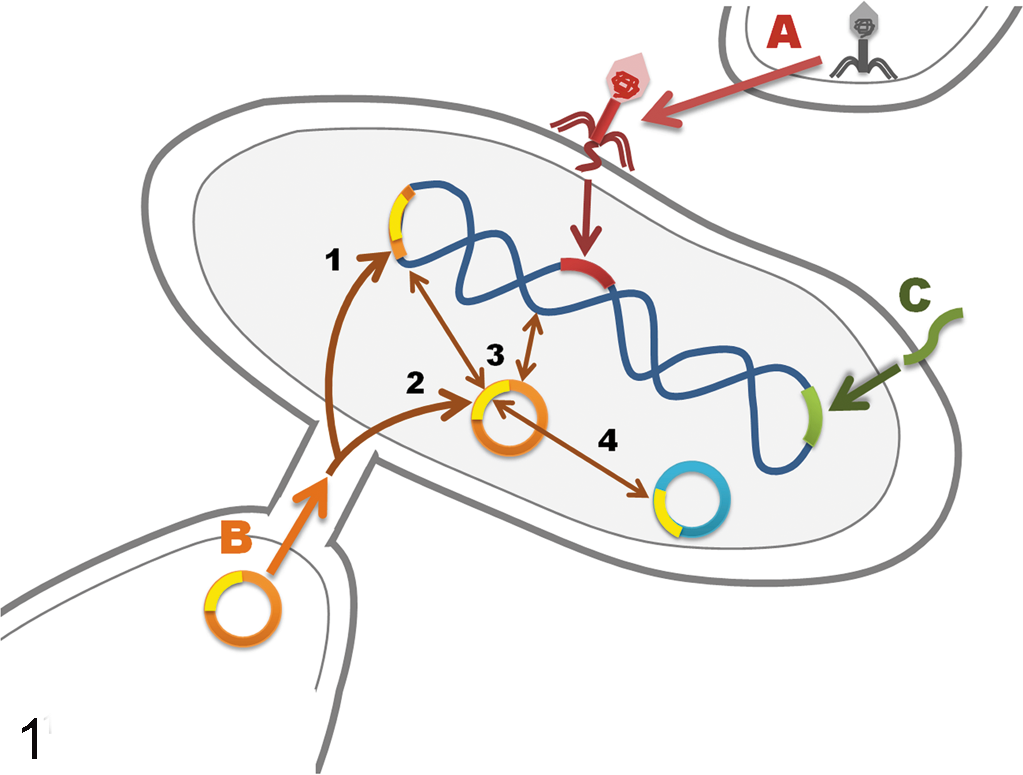
Mechanisms of DNA transfer between and within bacteria. (A) Transduction: injection of DNA into a bacterium by a phage. (B) Conjugation: plasmid in a donor bacterium is transferred through a pilus into a recipient bacterium; plasmid may integrate into the chromosome (1) or remain in the cytoplasm (2); plasmid may be transferred between cytoplasmic and chromosomal locations (3); plasmid may exchange insertion sequences or transposons with other plasmids (4) or the chromosome. (C)Transformation: uptake of naked DNA from the environment.
Transduction is the process by which bacterial viruses, called bacteriophages or phages, mediate transfer of DNA into bacteria (Fig. 1). 16,98 In one mechanism, an infecting phage lyses the bacterial chromosome and uses the host machinery to replicate its own DNA. The occasional accidental incorporation of bacterial DNA into a phage head allows this DNA to be transferred to another bacterium in a subsequent round of infection. This is called generalized transduction, as any region of the bacterial chromosome may be transferred. Specialized transduction occurs with phages that are integrated into the bacterial chromosome, a typical feature of the so-called temperate or lysogenic phages and prophages. Imperfect excision allows a region of the chromosome adjacent to the attachment site of the phage to be transferred. Some phages carry virulence genes as an integral part of their genomes, and these virulence factors may be expressed at high levels when prophages leave their dormant stage in the chromosome and start replicating in the course of a lytic cycle. 16
Transformation is the third mechanism of DNA transfer (Fig. 1). 59,98 This involves the uptake of naked DNA by bacteria that are in a state of natural competency, a physiologic state in which the bacteria are able to take up DNA and become transformed. 71 Single-stranded DNA is passed through the cell wall and cell membrane into the host through complex energy-requiring processes and enters the bacterial chromosome, mainly by homologous recombination 68 but also by the transient expression of nonhomologous recombination mechanisms encoded by the invading foreign DNA. 30 This mechanism has the potential to allow transfer of DNA between bacteria that are distantly related. Some bacterial species or strains within a bacterial species are more prone than others to be naturally competent for foreign DNA uptake. 86 Natural competency has also been shown to be regulated by external signals, such as bacterial pheromones. 59
The DNA that is transferred into a recipient bacterium becomes established through incorporation into the chromosome or as a self-replicating element in the cytoplasm. 138 Selective pressure then tends to determine which of the acquired genes remain in the population. Within a population, DNA acquired by LGT changes over time as a result of spontaneous mutations, changes in fitness, and interactions with the environment. Loss of sections of DNA also occurs under the same forces associated with modifications of acquired DNA, and such gene loss or silencing is sometimes associated with increased virulence. 8
Regulation of the incoming foreign DNA is critical. 29,98 Foreign DNA often comes in the form of virulent bacteriophages, and bacteria have several mechanisms for preventing the expression of entering DNA. 104 One of the major global negative transcriptional regulators involved in this process in Gram-negative bacteria is H-NS (histone-like nucleoid structuring protein). Not surprising, many virulence genes are negatively regulated by H-NS. There are antisilencing mechanisms that can overcome the effects of H-NS and permit expression of foreign DNA. To be effective, not only must the acquired virulence gene be expressed, but it must also be regulated in coordination with other genes involved in the host–pathogen interactions. 104
MGEs: Vectors of LGT
The transferred genetic elements have various characteristics that have resulted in different names (Table 1). 42,54,63,84 The elements discussed here are IS, transposon, integrative and conjugative element (ICE), plasmid, bacteriophage, and pathogenicity island (PAI). Table 2 presents a summary of the association of these elements with various bacterial pathogens.
Mobile Genetic Elements Involved in Horizontal Gene Transfer of Virulence Determinants.
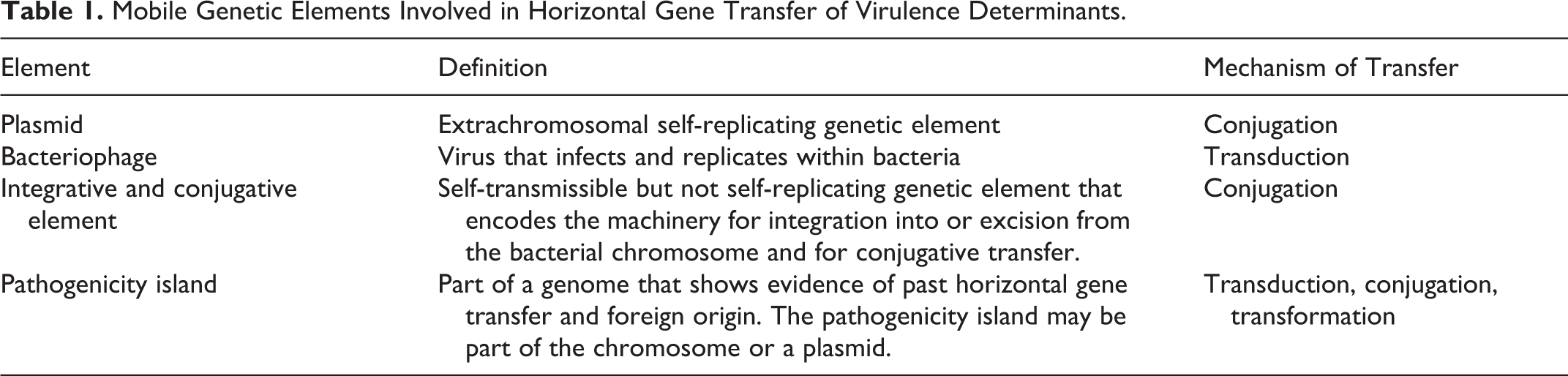

Mobile Genetic Elements in Selected Bacterial Pathogens.
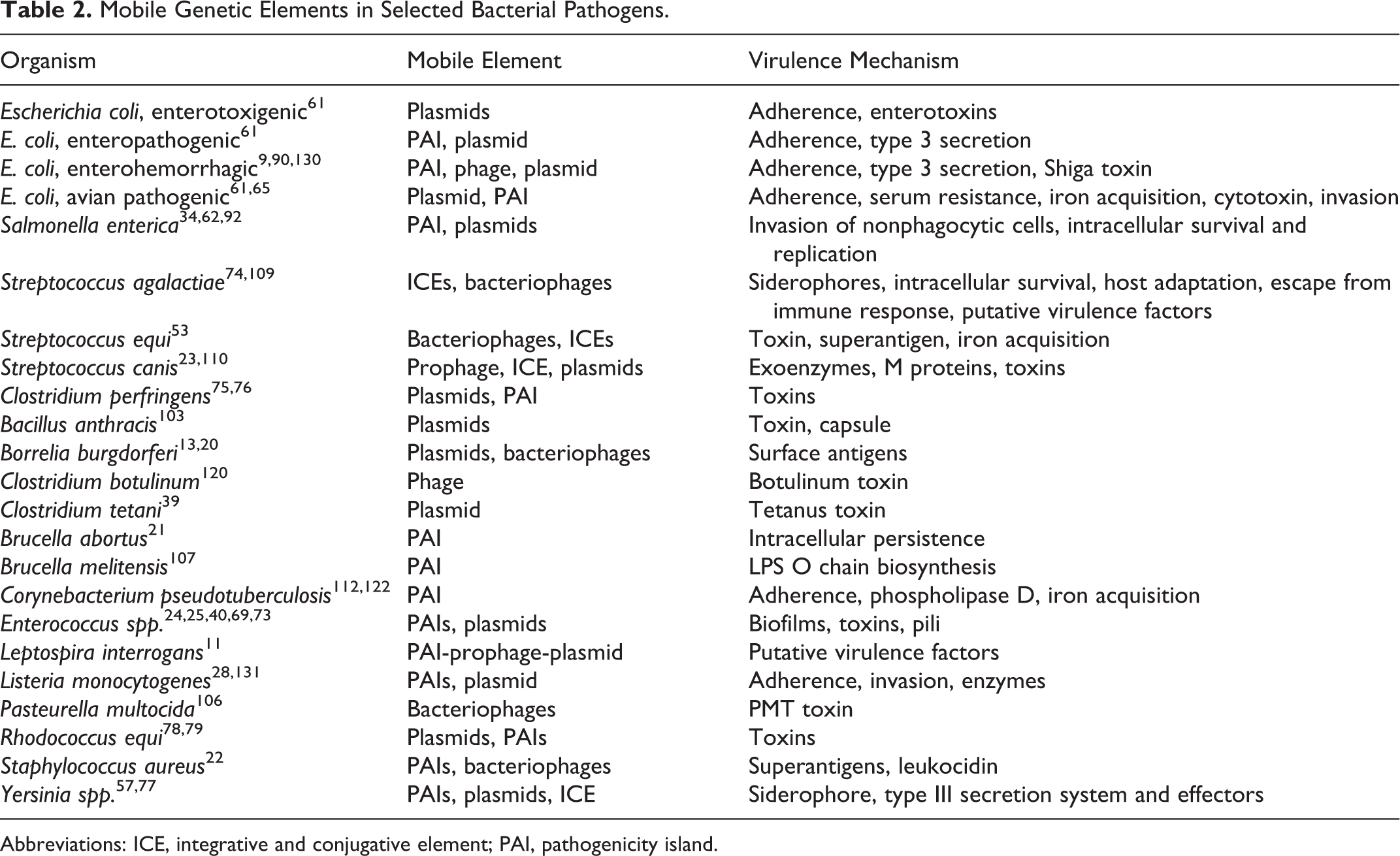
Abbreviations: ICE, integrative and conjugative element; PAI, pathogenicity island.
ISs and Transposons
An IS is a small segment of DNA (< 2.5 kb) with a simple organization, which carries no genes except those that allow it to insert at numerous sites in DNA. 84 It is the simplest of the MGEs and is widely distributed in bacteria. Typically, an IS consists of an open reading frame, which encodes a transposase enzyme, flanked by an inverted repeat sequence of 10 to 40 base pairs at each end. The enzyme, in concert with the inverted repeat sequences, cuts target DNA and inserts the IS. ISs have a preference for AT-rich areas of DNA but can insert in a range of sequences.
Insertion of an IS into DNA is mutagenic; it inactivates genes by disruption and can cause deletion or upregulation of adjacent genes. 84 Homologous recombination can also occur when multiple copies of an IS are present in a chromosome or a plasmid, with a variety of consequences, such as deletions, inversions, and duplications. A good example is the deletion of the region encoding the type 3 secretion system (T3SS) in a plasmid in the fish pathogen Aeromonas salmonicida. 125 The T3SS is essential for virulence, and its loss results in an avirulent bacterium. Also, when 2 identical IS elements flank a region of DNA, a so-called composite transposon is created, and the entire intervening DNA between the terminal inverted repeats can be mobilized by one or both of the IS-encoded transposases. ISs also affect virulence genes by virtue of their mutagenic properties and their ability to mobilize genes. For example, Ochi et al 97 noted that the genes for the E. coli heat labile enterotoxin (LT) and for the E. coli heat stable enterotoxin I (STIa) were present on a plasmid in which they were separated by 13.5 kb of DNA and were surrounded by 4 IS elements, 3 of which were identical. The authors suggested that the enterotoxin genes may have been mobilized into the plasmid by the IS elements. IS elements may also play a role in insertion and excision of plasmids into and out of the bacterial chromosome.
Transposons in general and composite transposons in particular are MGEs that, in addition to transposition function, carry other genes, such as genes for drug resistance, fitness, and virulence. 42 Transposons are involved in the intrabacterial movement of DNA. 84 They may, however, be transmitted to other bacteria by transfer of plasmid or chromosomal DNA.
Integrative and Conjugative Elements
ICEs are MGEs that share features of transposons, bacteriophages, and plasmids. 18,138 The ICEs include conjugative transposons. These elements are able to integrate into bacterial chromosomes, to excise from the chromosome, and to transfer themselves from one bacterium to another by conjugation but are not able to replicate by themselves. Integration of the ICE is usually site specific and frequently occurs at the 3′ end of a tRNA gene. ICEs are important in transfer of large amounts of genetic material, including virulence-related genes in Streptococcus (see below).
Plasmids
Plasmids are extrachromosomal DNA molecules that are usually circular, can self-replicate, and are distributed to daughter cells when the bacterium divides. 42,117 Some plasmids (conjugative or self-transmissible plasmids) carry genes that permit their transfer by conjugation 121 ; others lack such genes and rely on transfer function of other plasmids to enable their transfer. Plasmids are usually not part of the core genome of a species and encode a range of genes for a variety of nonessential functions, including drug resistance, virulence factors, fitness, and adaptation.
Many bacterial toxins are plasmid encoded. 15,36,39,48,60,61,67,75,81,103 Bacillus anthracis is an example of a pathogen in which the major virulence factors—the anthrax toxin and an antiphagocytic polyglutamic acid capsule—are plasmid encoded. 103,141 Genes for the anthrax toxin and for adhesins are found on a plasmid called pXO1, and genes for the capsule are present on a plasmid called pXO2. 103 Interestingly, the genome of B. anthracis is very similar to that of the less pathogenic Bacillus cereus, and the 2 are considered to be part of the same species, Bacillus cereus sensu lato, together with Bacillus thuringiensis, a well-known insect pathogen. 141 The major differences between these pathogens causing very different diseases are due to their plasmids; B. anthracis carries 2 plasmids that confer resistance to phagocytosis and lethal toxicity, whereas B. cereus has a plasmid that is similar to pXO1 of B. anthracis but encodes an emetic toxin. 141
Rhodococcus equi is another pathogen in which a plasmid is critical for virulence. 78,79 This bacterium has a large conjugative plasmid that encodes virulence associated proteins that are essential for intramacrophage growth and virulence in foals. 79 The vap genes are part of a PAI that encodes a family of secreted proteins. As for many other pathogens, recent research has demonstrated cross-talk and coregulation of plasmid-borne virulence factors with other chromosomal factors in R. equi, 78 thus illustrating the adaptation and integration of horizontally acquired genes into complex regulatory networks.
Yersinia is an example of a genus in which plasmids contribute to dramatic evolutionary, lifestyle, and virulence changes. 52 Yersinia enterocolitica and Yersinia pseudotuberculosis are mammalian enteropathogens that are found widely in the environment. Yersinia pestis, the cause of plague, has evolved only recently (15 000 to 20 000 years) from Y. pseudotuberculosis and is a blood-borne systemic pathogen with a complex life cycle involving a rodent reservoir and a flea vector. Much of this adaptation involves the acquisition of 2 plasmids by Y. pestis; one encodes a toxin and a capsule (and is essential for survival in the flea), and the other encodes a plasminogen activator, which facilitates dissemination following the flea bite. 52 There have been many other changes, as Y. pestis has discarded unnecessary chromosomal genes. 8
Bacteriophages
As noted earlier, a number of important bacterial toxins are found within the genomes of phages that incorporate into the bacterial chromosome. 3,15,16,26,78,106,120 The tox-positive phage of Corynebacterium diphtheriae, which encodes the diphtheria toxin (DT), is a well-studied example of this association between phages and bacterial virulence genes. 3,16 Interestingly, DT is also produced by strains of Corynebacterium ulcerans that carry a tox+ phage, which appears to have an origin distinct from that of the phage in C. diphtheriae. 116 The phage-encoded DT is also sometimes found in strains of Corynebacterium pseudotuberculosis. 85 Genes for botulinum toxins C and D, associated with botulism in animals, are carried by phages that circularize and behave like plasmids. 15,120 In Vibrio cholerae, the genes for cholera toxin (CT) are carried by a filamentous phage, CTX, that inserts its single-stranded DNA into one or both V. cholerae chromosomes. 35,98,132 Phage-encoded toxin-coregulated pili, which are important in colonization of the intestine by V. cholerae, serve as the receptor for the CTX phage. Strains of Staphylococcus aureus that cause staphylococcal scalded skin syndrome in humans carry either a phage-encoded exfoliative toxin 139 or a plasmid-encoded toxin. 58 Exfoliative toxin is also produced by the pig pathogen Staphylococcus hyicus, but there is no information on the location of the gene in this organism. 1
Pathogenicity Islands
Large segments (10–200 kb) of foreign DNA inserted into bacterial chromosomes are often referred to as genomic islands (GEIs) 63 ; segments less than 10 kb are called islets. 96 These regions of DNA are often flanked by direct repeats or IS elements, usually have mobility genes (eg, integrase or transposase), and are frequently inserted in or near tRNA genes. The GEIs are present in some genomes but absent from other, closely related ones and have been found on plasmids as well as chromosomes. Ho Sui et al 54 analyzed 631 bacterial genomes and determined that the percentage of genes in GEIs that encoded virulence factors was approximately 4 times that of genes outside GEIs. Those GEIs that carry virulence genes are called PAIs. 98 The mechanisms of PAI mobilization seem to be multiple and include conjugation, transduction, and possibly also transformation. 63,96 Some PAIs are stable and not easily transferable, while others are highly mobile.
In S. aureus, for instance, there are 12 PAIs that encode a variety of superantigen toxins, including toxic shock syndrome toxin and a number of enterotoxins. 96 These PAIs, now also dubbed phage-related chromosomal islands, can be very efficiently mobilized by helper phages and transduced into new S. aureus at high frequencies, 96 thus explaining the diversity of superantigen and toxin profiles found in S. aureus. The pXO1 plasmid of B. anthracis includes a PAI with genes for production and regulation of the anthrax toxin and illustrates another potential type of mobility for PAIs. 103
A PAI in Corynebacterium pseudotuberculosis includes genes for phospholipase D and genes for iron utilization. 122 Phospholipase D is considered to be the major virulence factor in this organism. It increases vascular permeability by attacking sphingomyelin in the membrane of endothelial cells. Fifteen other PAIs have been reported in C. pseudotuberculosis based on genome analyses. 122
In Listeria, the distribution of the PAIs LIPI-1 and LIPI-2 illustrates the gain and loss of PAI, paralleling host and environment adaptations. 131 LIPI-1 encodes for a cluster of virulence factors, including hemolysins and actin-polymerase factors necessary for the intracellular survival and spread of the organism. This PAI seems to have been acquired early in the evolution of the genus Listeria but lost by nonpathogenic species in the course of their adaptation to environmental conditions. 131 LIPI-2 encodes a sphyngomyelinase and a number of internalins, which enhance the intracellular survival of the organism and its ability to cause apoptosis in host cells. This PAI is specific to Listeria ivanovii and thought to be responsible for restriction of its host specificity to ruminants only and for its lack of the typical tropism for the central nervous system observed in Listeria monocytogenes. 31
MGEs and LGT in the Virulence of 4 Major Pathogens
E. coli
E. coli has a core of essential genes and an accessory genome that consists of adaptability genes, which appear to have aggregated over time as a result of gain and loss of genes from the environment. 29 The genome has a mosaic structure representing core DNA (G+C, 50.8%) and areas with DNA with markedly different G+C ratios. Genome size ranges from 4.6 to 5.6 Mb, with a core of about 1700 genes (present in 100% of 186 genomes) and a pan-genome of over 16 000 genes identified in a pool of 186 strains of the species. 64 The genome of pathogenic E. coli O157:H7 Sakai strain encodes over 1600 proteins and 20 tRNAs that are not found in nonpathogenic E. coli strain K-12, and most of the O157-specific proteins are encoded by prophages and prophage-like elements. 50 Complexity of gene structure and organization is increased by deletions, insertions, and rearrangements that have occurred over time and with spread of new and altered DNA within the species due to recombination involving homologous sequences that flank inserted DNA regions. Plasmids, phages, and PAIs transferred to nonpathogenic E. coli have created a range of pathogens that cause a variety of diseases. 26,61,115 These pathogens have been assigned to various pathotypes based on their virulence attributes and type of disease. 26
Enterotoxigenic E. coli possess plasmids that encode the critical virulence elements: pilus adhesins that mediate adherence to the intestinal epithelium and protein or peptide enterotoxins that cause enterocytes to hypersecrete fluid, resulting in diarrhea. 47,49 As examples, fimbrial adhesins K88 (F4), K99 (F5), 987P (F6), and F18 and enterotoxins LT, STa, and STb in porcine and bovine enterotoxigenic E. coli are all located on plasmids. 47,95,119 Some E. coli enterotoxin plasmids also carry genes for antimicrobial resistance. 36,45,48
Enteropathogenic E. coli (EPEC) and enterohemorrhagic E. coli (EHEC) are both characterized by their ability to form attaching and effacing lesions in the intestine. EPEC cause diarrhea in humans 90,99,115,137 and a range of animal species 47 ; EHEC cause bloody diarrhea and toxemia in humans and have their primary reservoir in the intestine of ruminants. 9,44,90 Most information on EHEC virulence is based on studies of EHEC serotype O157:H7, which evolved from an EPEC by acquisition of a bacteriophage that carries the stx gene. Recently, an enteroaggregative O104:H4 E. coli that had previously acquired a plasmid with genes for fimbrial adherence and colonization became infected with a phage carrying the stx gene, resulting in a highly virulent enteroaggregative hemorrhagic E. coli with a spectrum of diseases similar to that of EHEC O157:H7. 4,5 Whereas the EHEC have their reservoir in the intestine of ruminants, 44 the enteroaggregative hemorrhagic E. coli have humans as the only known reservoir, and they appear to be more proficient at causing severe disease and being transmitted from person to person. 4,5
Colonization of the intestine by EPEC and EHEC requires the locus of enterocyte effacement (LEE) PAI, in which genes necessary for the attaching and effacing lesion are encoded. 9,26,90,99,115,130,137 This PAI encodes proteins that form a T3SS, which includes a “molecular syringe” that injects effector proteins into the cytoplasm of the enterocyte. 26,90,99,137 T3SSs appear to have evolved from the basal portion of flagella and are used by a number of Gram-negative bacteria to transfer bacterial proteins into eukaryotic cells. DNA involved in regulation of the LEE genes and DNA for chaperones are also present in the LEE, which is inserted into the E. coli chromosome at one of a few sites associated with genes for tRNAs and is flanked by prophage DNA and IS elements. 115 The LEE is itself a mosaic with a core of 38.4% G+C and elements with different G+C ratios. The LEE T3SS is responsible for transfer of effectors encoded in the LEE as well as effectors encoded by genes outside the LEE. 137 The LEE is central to a complex colonization process that involves initial contact by a pilus adhesin, transfer of effector proteins into the enterocyte, reorganization of the enterocyte cytoskeleton, effacement of microvilli, and intimate adherence of the bacteria to the enterocyte. 115,137
Only 7 or 8 effector genes are encoded by the LEE; the majority (19 in EPEC and 55 in EHEC) are non-LEE-encoded (nle) genes. 137 In EHEC O157:H7, many of these effectors are encoded by chromosomal, defective lambdoid prophages (bacteriophages related to phage lambda, which can exist as DNA in the bacterial chromosome). The effectors injected into enterocytes are multifunctional proteins that manipulate host cell signaling pathways. Effectors are responsible for reorganization of the host cell cytoskeleton, effacement of microvilli, tight adherence of the bacteria to the host cell membrane, and changes in the metabolism of the cell. 115,137 One of the LEE-encoded effector proteins, Tir (translocated intimin receptor), associates with the enterocyte membrane and binds to intimin (a LEE-encoded outer membrane protein) on the surface of the bacterium, creating an intimate adherence. Diarrhea is attributed to the loss of absorptive microvillus surface, inflammation, disruption of enterocyte tight junctions, and alterations in fluid metabolism. 90,137
Whereas EPEC cause watery diarrhea, 99 EHEC cause watery diarrhea, bloody diarrhea, and life-threatening hemolytic uremic syndrome. 90,115,137 Greater severity of disease is attributable to phage-encoded Shiga toxin (Stx) (also called verotoxin), whose cytotoxicity for vascular endothelium is central to the tissue damage leading to hemorrhagic colitis and hemolytic uremic syndrome in humans. 12,140 The stx gene is carried in a lambdoid prophage embedded in the chromosome of EHEC. The phage transfers the stx gene from bacterium to bacterium and also regulates expression of stx in the host EHEC. The prophage is quiescent in the chromosome but can be induced to escape from the chromosome, form large numbers of phage particles, massively express the stx gene, lyse the host bacterium, and escape into the surroundings. A number of antibiotics (at subinhibitory concentrations) 88 and other noxious stimuli (eg, hydrogen peroxide 83 ) can induce the phage. Expression of the stx gene is under control of the phage and is enhanced following induction. Thus, the phage provides mechanisms for mobilization of the stx gene and for synthesis and release of large amounts of Stx. Little is known about the factors that affect phage induction in the intestine of the host, but these may play a role in the varying severity of disease in populations exposed to this organism. There has been much debate about the value to the host of a prophage that causes its lysis and death. A recent theory is that this represents bacterial altruism, in which a small percentage of the population sacrifices itself to produce a toxin that kills threats such as protozoan predators and neutrophils, allowing the population to survive. 82
In contrast to human isolates of EHEC O157:H7, which usually produce high amounts of Stx, bovine isolates produce variable amounts of Stx. 101,126 Differences in inducibility of the Stx phage and in regulation of expression of the stx gene may contribute to this situation. In some cases, low or no production of Stx in bovine strains has been attributed to inactivation of the stx gene by an IS element. 101
Stx may also play a role in adherence. 111 The outer membrane protein intimin, encoded by the LEE, binds not only to Tir (also encoded by the LEE) but also to surface-expressed nucleolin. 111 Stx increases expression of nucleolin at the surface of the epithelial cell and thereby promotes adherence. 111
In EHEC O157:H7, virulence genes, often associated with IS elements, are also carried on a 92-kb plasmid, pO157. 17 These include genes for a hemolysin, an extracellular serine protease, a catalase/peroxidise, and a metalloprotease. Typical EPEC strains also carry a virulence plasmid that encodes adherence properties.
Extraintestinal pathogenic E. coli is a pathotype that includes avian pathogenic E. coli (APEC), bovine septicemic E. coli, and human uropathogenic E. coli. 47,136 Plasmid-encoded virulence factors are important in this group. 61,100 In APEC, high virulence is associated with possession of large plasmids that carry genes for iron scavenging, proteases, serum resistance, adherence, and hemolysin. 61,100 These genes are found in strains that are highly heterogeneous. Similar virulence genes are carried in PAIs by human uropathogenic E. coli. 133,136 Some bovine septicemic E. coli carry a large transmissible plasmid called Vir, which possesses genes for cytotoxic necrotizing factor 2, a cytolethal distending toxin, fimbrial adhesins, and genes associated with bacterial biofilm formation. 60
Salmonella
Salmonella, a facultative intracellular pathogen that can invade intestinal epithelial cells and macrophages, and E. coli diverged from a common ancestor approximately 100 million years ago 27,32,63 but still share a substantial amount of their genomes. As with E. coli, Salmonella possess a core genome plus a flexible accessory genome consisting of Salmonella PAIs (SPIs), bacteriophages, and plasmids. The critical factor in divergence of Salmonella and E. coli was the acquisition of SPIs, which allowed a noninvasive commensal bacterium to become invasive and pathogenic. 92
There are at least 21 PAIs in Salmonella, but most attention has been paid to SPI-1 and SPI-2, which are critical for invasion of nonphagocytic cells and for replication in nonphagocytic and phagocytic cells, respectively. 27,56,72,124 Variations in presence and sequence of MGEs contribute to differences among serovars and among strains within a serovar in Salmonella. 92 Salmonella are responsible for enteric disease as well as systemic disease, but the disease process varies depending on which of the more than 2500 serovars of Salmonella is involved and the host with which it is interacting. 27,34 For example, Salmonella Typhimurium typically causes food-borne gastroenteritis in humans but is the cause of a severe systemic typhoid-like disease in mice. 37,38 Salmonella Typhimurium contains virulence genes in 5 SPIs, namely SPI-1 to SPI-5, several phages, and a virulence plasmid. 34 Rychlik et al 113 used Salmonella Enteritidis mutants with each of or all of 5 SPIs (1–5) deleted and determined their virulence in day-old chickens. They showed that SPI-1 and SPI-2 were both required for systemic infection, that SPI-1 was responsible for proinflammatory signalling, but that bacterial colonization of the cecum occurred in the absence of all 5 SPIs. These observations highlight the extent to which these virulence factors play different roles in various host species.
SPI-1 and SPI-2 encode T3SS-1 and T3SS-2, respectively, and play major roles in virulence. 27,37,56,124 These systems directly deliver more than 30 proteins into the cytoplasm of the mammalian cell. One study reported that there are 400 proteins secreted by Salmonella. 114 Some of the proteins secreted by SPI-1 are encoded within SPI-1, but others are encoded by prophages or phage remnants or by other SPIs. Most effectors are secreted through one or the other of the 2 secretion systems, but several effectors can be secreted by either system. 19 Attribution of function to the various effector proteins is complicated by multifunctionality and redundancy. 56
SPI-1 permits Salmonella to invade nonphagocytic cells in the intestine and is therefore critical for the intestinal phase of disease. 37,56,124 Following contact of the bacteria with intestinal epithelial cells, effectors secreted by SPI-1 use host signal transduction pathways to rearrange the cytoskeleton and induce membrane ruffling, allowing the bacteria to be taken up by these nonphagocytic cells and enveloped in a phagosome. 34 Expression of SPI-1 genes is regulated by a variety of environmental cues, such as pH, osmolarity, contact with epithelial cells, and nutrients. Regulation of the genes involved in secretion of effectors is critical, and there is a master regulator encoded on SPI-1. 34 There is evidence that proteins secreted through T3SS-1 end up both extracellularly and intracellularly; the former is attributed to secretion by bacteria that are in the intestinal lumen. T3SS-1 effectors also activate transcription factors NFκB and activator protein-1, leading to release of interleukin-8 (IL-8) through the basal and lateral borders of the cell. 124 IL-8 leads to recruitment of polymorphonuclear leukocytes that stream between cells into the intestinal lumen.
Although SPI-1 is primarily involved with the intestinal phase of infection, it plays a role in later stages. 72 Recently, Kyrova et al 72 provided evidence that the SPI-1 T3SS of Salmonella Typhimurium strain DT104 was able to downregulate macrophage antibacterial activity by steering macrophages toward a less bactericidal phenotype.
Interestingly, following uptake of Salmonella, SPI-1-encoded effectors reverse the cytoskeletal and cell membrane changes and restore the architecture of the epithelial cell. 56,124 The earlier stimulation of the inflammatory response is reversed and replaced by suppression.
SPI-2 genes are activated in the phagosome, and about 30 T3SS-2 effectors are secreted through the Salmonella-containing vacuole (SCV) inside the macrophage, allowing the ingested Salmonella to withstand the barrage of antibacterial activity of the macrophage and to multiply. 19,27,34,37,56,124 Conditions inside the SCV that trigger activation of the T3SS-2 include acidity, low concentrations of Ca++ and Mg++, poor nutrition, and antimicrobial peptides. 37,38,124 Regulation of SPI-2 activity is also critical and is effected by 3 two-component regulatory systems. 124 Effectors secreted by T3SS-2 are responsible for maturation of the SCV, alterations in the membrane, migration of the SCV along microtubules to the Golgi apparatus in a perinuclear location, and formation of Salmonella-induced filaments as well as actin filaments that surround the SCV. Bacteria in the cytoplasm are killed quite rapidly, but the SCV provides a safe environment in which Salmonella replicates. The T3SS-2 is important for intracellular replication of Salmonella. The SCV does not fuse with lysosome, and respiratory burst and nitric oxide production are inhibited in the Salmonella-infected macrophage. However, there is debate regarding the role of T3SS-2 in these activities. 37
SPI-3 encodes an inner membrane protein called MgtC, which contributes to intramacrophage survival, and an autotransporter protein, MisL, which is an adhesin that promotes adherence to intestinal epithelial cells. 113 SPI-4 contains 6 genes, 3 of which encode a type 1 secretion system; 1 of the genes encodes a large protein that is secreted by the type 1 secretion system. 34 In Salmonella Typhimurium, SPI-4 is involved in colonization of bovine but not chick intestines 93 and in oral virulence for mice. 70 SPI-5 encodes effectors secreted by T3SS-1 and T3SS-2. 34 SPI-6 encodes a type 6 secretion system, whose regulation is integrated into the virulence network of Salmonella Typhimurium and which appears to be involved in the late stages of macrophage infection. 94 SPI-7 is a large island (134 kb) found in Salmonella Typhi, the agent of typhoid fever. 118 It includes genes for the Vi capsular polysaccharide antigen, a cryptic phage encoding an effector secreted by the T3SS-1, and pili. Fragments of SPI-7 are found in several bacteria, including a few other serovars of Salmonella and Haemophilus influenzae, Pseudomonas aeruginosa, and Y. enterocolitica. 118 Little is known about the roles of the other SPIs in disease.
Prophages also play important roles in virulence of Salmonella. 38,92 For example, prophages Gifsy-1 and Gifsy-2 in Salmonella Typhimurium encode important virulence factors, including superoxide dismutase.
A 90-kb virulence plasmid is found in serovars of Salmonella that are highly invasive. The serovars that contain this plasmid include Salmonella Typhimurium, Salmonella Dublin, Salmonella Choleraesuis, Salmonella Pullorum, and Salmonella Gallinarum. 34,105 The region of the virulence plasmid that imparts virulence is an operon known as spv (Salmonella plasmid virulence) RABCD and a second operon known as pef (plasmid encoded fimbriae). 34,46,56,80 SpvB is secreted by T3SS-1 and has an effect on actin polymerization. 34,38,56,127 SpvC is secreted by T3SS-1 and T3SS-2 and has an anti-inflammatory effect, acting on MAP kinases ERK and JNK to inhibit release of IL-8 and TNF-α. 34,37,38,56,87
In 2009, Fricke et al 41 reported that a ColV plasmid in a Salmonella Kentucky isolate from chicken breast carried antimicrobial resistance genes and virulence genes. The plasmid possessed genes for resistance to streptomycin and tetracycline, associated with genes for transposases, and genes for iron acquisition, serum resistance, and iron/manganese transport. The plasmid has a backbone that is over 90% identical to that of certain virulence plasmids found in APEC and carries the same virulence genes present in the APEC virulence plasmids.
Shortly thereafter, Johnson et al 62 reported that emergence of Salmonella Kentucky as a significant pathogen in broiler chickens in the United States appears to have coincided with the presence of a ColV plasmid in strains of this serovar. They found that 72.7% of 293 Salmonella Kentucky isolates that were examined had the plasmid and belonged to a single clone and that its presence was associated with enhanced cecal colonization and invasion in chickens.
Pyogenic Streptococci
Streptococci represent an important group of human and animal pathogens, 43 including species such as Streptococcus pyogenes, 108 Streptococcus canis, 7 Streptococcus equi (ie, S. equi subsp. equi and S. equi subsp. zooepidemicus), 129 and Streptococcus agalactiae. 66,89 Recent comparative genomics studies discussed below have shed some light on the important role of LGT in the evolution of virulence and host adaptation of these pathogens, implicating in particular ICEs and prophages.
S. pyogenes is a major human-specific pathogen causing a variety of diseases ranging from local pyogenic infections to septicemia, toxic shock syndrome, necrotizing fasciitis, and postinfection rheumatic fever. The genomes of S. pyogenes, S. dysgalactiae, and S. equi are very closely related, and evolutionary relationships among these 3 species are difficult to reconstitute. However, a recent study of the core genomes of these species by Lefébure and collaborators 74 shows that S. pyogenes is a direct sister group of S. dysgalactiae. It has become host adapted to humans by acquisition of new virulence and colonization factors through LGT. Close to half of the 113 genes acquired by S. pyogenes during its separate evolution are located in prophages, including 5 virulence genes encoding for pyrogenic exotoxins and hyaluronidases involved in the spread of the organism and a regulator of DNA recombinase. Interestingly, numerous subsequent LGT events occurred with these bacteriophage-associated genes, and the majority happened between S. pyogenes and S. equi. The authors of the study suggested that these transfers were part of the events leading to the separation of the strictly horse-adapted S. equi subsp. equi from its more polyvalent parent S. equi subsp. zooepidemicus and may reflect the tight association between humans and horses.
The evolution of the 2 S. equi pathogens provides another illustration of the important role of LGT in host adaptation and evolution of virulence in streptococci. Previous investigations 53 had shown extensive gene function losses and gains paralleling the transition from the multihost S. equi subsp. zooepidemicus to the strictly host-adapted S. equi subsp. equi. The loss of functions and genes that became unnecessary seems to have been accelerated by the accumulation of multiple ISs in S. equi subsp. equi that facilitated deletions and gene knockout. Concomitantly with gene loss, 4 prophages related to S. pyogenes prophages were acquired by S. equi subsp. equi that encode virulence genes for a phospholipase and several superantigens. Interestingly, S. equi subsp. equi seems to lack the protective effects against bacteriophage infections provided by CRISPR elements, 123 thus explaining why this pathogen was able to integrate so many bacteriophages in its genome while S. equi subsp. zooepidemicus remained free of prophages. ICEs play an important role in the evolution of streptococci, and S. equi subsp. equi is no exception. Its genome contains an ICE encoding for a new siderophore called equibactin, 51 not found in S. equi subsp. zooepidemicus.
The results of the study by Lefébure and collaborators also suggest important LGT between S. agalactiae and S. pyogenes, including putative virulence factors implicated in survival at elevated temperatures, in production of siderophores, and in survival within macrophages. 74 The main hosts of S. agalactiae are cattle and humans, and different strains of this bacterium seem to have developed specific adaptations to these 2 hosts. Numerous ICEs have been identified in S. agalactiae, 14,49 some of which form GEIs containing virulence genes or genes important for the adaptation to the mammary gland and bovine environment found in cattle isolates only. 109 These islands contained, for instance, genes for a nisin operon (probably originating from Streptococcus uberis), genes involved in the metabolism and utilisation of fructose and lactose (probably from S. dysgalactiae), ABC transporters, a modified salivamicin immunity and production system (possibly involved in intracellular survival), and several putative virulence-associated proteins with an LPXTG motive for cell surface anchoring. Some of these GEIs were not related to ICEs but to prophages, and one of them even seems to represent a plasmid that integrated into the chromosome. Overall, the comparative genomics study of Richards and collaborators 110 suggests that LGT between organisms sharing the mammary ecosystem is an important determinant in the evolution of virulence of S. agalactiae, which could even extend in the future to the emergence of antimicrobial resistance in an organism that has remained until now surprisingly susceptible to antibiotic treatment. 109
S. canis is an important pathogen of dogs but also a variety of other animal species, including cattle. Similarly to the situation in the other streptococci, the recent analysis of the genome sequence from a bovine isolate demonstrated the presence of multiple MGEs in S. canis involving virulence genes. 109 As mentioned above, many of the related LGT events involved S. pyogenes. Beside the presence of prophages and an ICE carrying a number of putative virulence factors, one of the prominent features of the S. canis genome is the presence of an integrated plasmid similar to the one mentioned above for S. agalactiae. Some of the genes from this plasmid are homologous to virulence genes from L. monocytogenes, Enterococcus faecalis, and S. pyogenes, and the authors of this study suggest that this plasmid may have a common origin with the one from S. agalactiae and play an important role in adherence of S. canis to host cells. 109
More comparative genomics studies on isolates from a variety of sources are needed to test some of the hypotheses discussed above on the origins and transfer of virulence genes, but the data presented illustrate the prominent role of bacteriophages and ICEs and LGT in general in the host adaptation and acquisition of new virulence properties in pyogenic streptococci from human and animals.
Clostridium perfringens and Necrotic Enteritis
Extrachromosomal elements play a major role in diseases caused by clostridia, and many clostridial toxin genes are located on plasmids. 15 For instance, the gene for tetanus toxin is present on a 74-kb plasmid in Clostridium tetani, 39 and genes for Clostridium perfringens enterotoxin (cpe), beta, beta-2, epsilon, iota, large clostridial cytotoxin, and NetB toxins in C. perfringens are all carried on large conjugative plasmids. 2,15,67,75,91,102,128 Interestingly, the cpe gene is chromosomal in some strains and plasmid borne in others. There are differences in the genetics, physiology, disease spectrum, and epidemiology of strains with chromosomal versus plasmid cpe genes. 81
Necrotic enteritis in poultry is caused by a specific subset of C. perfringens type A strains. 128 It represents another example of the role of MGEs in the pathogenesis of an economically important disease. The disease had been controlled until recently through the preventive use of antimicrobials in feed. 55,135 However, restrictions on this practice led to increased prevalence of necrotic enteritis and triggered molecular investigations to identify possible targets for vaccine development. One of the major breakthroughs in this direction has been the discovery of NetB, a pore-forming toxin that seems to be one of the key factors in the development of necrotic enteritis. 67 Further studies on NetB and comparative genomics have subsequently demonstrated that the netB gene is part of a highly conserved 42-kb plasmid-borne locus (NELoc-1) encoding a number of putative virulence factors, such as an internalin-like protein coregulated with NetB, a ricin domain-containing protein, 2 leukocidins, and a signaling system known to regulate the expression of virulence factors in other pathogens. 2,75,76 Multiple recombinase genes or remnants thereof are present within and beside this locus, which attest to the repeated recombination and gene transfer events leading to the emergence of this locus. Although less conserved, a second smaller plasmid-borne locus (NELoc-3) was strongly associated with C. perfringens isolates from necrotic enteritis cases. This locus encodes a hydroxysteroid hydrogenase also found in Clostridium difficile, and it is thought to degrade bile acids into toxic by-products and provide an advantage to its producer when competing with other microorganisms in the gut environment. 76 A third virulence-associated locus (NELoc-2) encoding mainly membrane- and cell wall–associated factors was also found among clinical isolates. However, despite being the apparent result of LGT, this locus was chromosomal. Interestingly, NELoc-1 and NELoc-3 do not usually reside on the same plasmid. 76 However, these plasmids are mobile and share the same conserved backbone necessary for conjugative transfer. 102 These investigations and those of others 1,73 have started to shed some light on the central role of a family of transferable plasmids, called pCPF5603-like, in the pathogenesis of diseases caused by a variety of C. perfringens types. New models based on differences in the partitioning systems that allow plasmids to be distributed equally among daughter cells during cell division are emerging, 102 which explain how similar virulence plasmids are able to congregate, persist together, and recombine to shape very competitive pathogens such as the C. perfringens type A strains causing necrotic enteritis.
Conclusions
Bacteria acquire genes from their environments by bacterial conjugation, by infection with bacteriophages (transduction), and by direct uptake of naked DNA (transformation). These genes are referred to as horizontally or laterally transferred genes and may be incorporated into the bacterial chromosome or into chromosomal genetic elements called plasmids. Genes implicated in bacterial virulence, notably those that encode toxins, mechanisms for bacterial adherence to host cells, and systems for manipulating host signal transduction or extracting iron from low iron environments of the host, have frequently been acquired by horizontal transfer. These virulence genes are sometimes located in clusters of genes referred to as PAIs. To be effective, the acquired genes must fit into the complex regulatory networks of their bacterial hosts. Enteric bacterial pathogens reside in environments that are rich in mobilizable DNA and often use combinations of acquired DNA to achieve 2 main features of enteric pathogenesis—namely, colonization and alterations in host metabolism.
The study of horizontally transferred genes has contributed much to our understanding of bacterial adaptability, pathogenesis, and evolution. Because of their unique features, it is now simpler to conduct genome searches for foreign genes that affect adaptation to the animal or human environment. Knowledge of their behavior has allowed us to look at changes in the genomes of pathogens over millions of years of evolution and to recognize likely developments in the years ahead.
Increasing recognition of the coexistence of antimicrobial resistance genes and virulence genes in the same pathogens and, sometimes, on the same MGE 36,41,45,62 highlights the risk that antimicrobial use in agriculture may coselect for antimicrobial resistance and virulence factors. Such selection may accelerate the emergence of new virulent pathogens continuously created through LGT and reduce our capacity for treating the diseases they cause.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
