Abstract
An outbreak of Streptococcus iniae occurred in the early months of 2008 among wild reef fish in the waters of the Federation of St Kitts and Nevis, lasting almost 2 months. Moribund and dead fish were collected for gross, histological, bacteriological, and molecular analysis. Necropsy findings included diffuse fibrinous pericarditis, pale friable livers, and serosal petechiation. Cytological and histological analysis revealed granulocytic and granulomatous inflammation with abundant coccoid bacterial organisms forming long chains. Necrosis, inflammation, and vasculitis were most severe in the pericardium, meninges, liver, kidneys, and gills. Bacterial isolates revealed β-hemolytic, Gram-positive coccoid bacteria identified as S. iniae by amplification and 16S ribosomal RNA gene sequencing. Results from biochemical and antimicrobial susceptibility analysis, together with repetitive element palindromic polymerase chain reaction fingerprinting, suggest that a single strain was responsible for the outbreak. The inciting cause for this S. iniae–associated cluster of mortalities is unknown.
Streptococcus iniae is an aquatic pathogen associated with significant losses in cultured fish worldwide and has the potential for human health risk. 1,5,19 In several cases, this potentially zoonotic microorganism has also been implicated in disease outbreaks in wild fish populations. 3,6,20
During January 2008, sporadic reports of large groups of floating dead fish were reported to national officials in several coastal areas in the Federation of St Kitts and Nevis, West Indies. Dead fish were noted on the ocean floor by recreational SCUBA divers, while moribund fish were swimming slowly and erratically at the water’s surface. Few clinical signs were observed, including obvious respiratory distress (fish gasping at the surface or demonstrating rapid opercular movements), erratic or slow swimming near the surface, and occasional areas of petechiation around the fins.
A number of fish were submitted for postmortem examination; however, many had been dead for some time, and postmortem autolysis made them unsuitable for diagnosis. Eight moribund or freshly dead fish were obtained from several locations from January to February 2008 (Fig. 1 and Table 1). Fifteen nonclinically affected fish, harvested by local fishermen from areas reported to have shown sick fish, were also examined (Table 2).
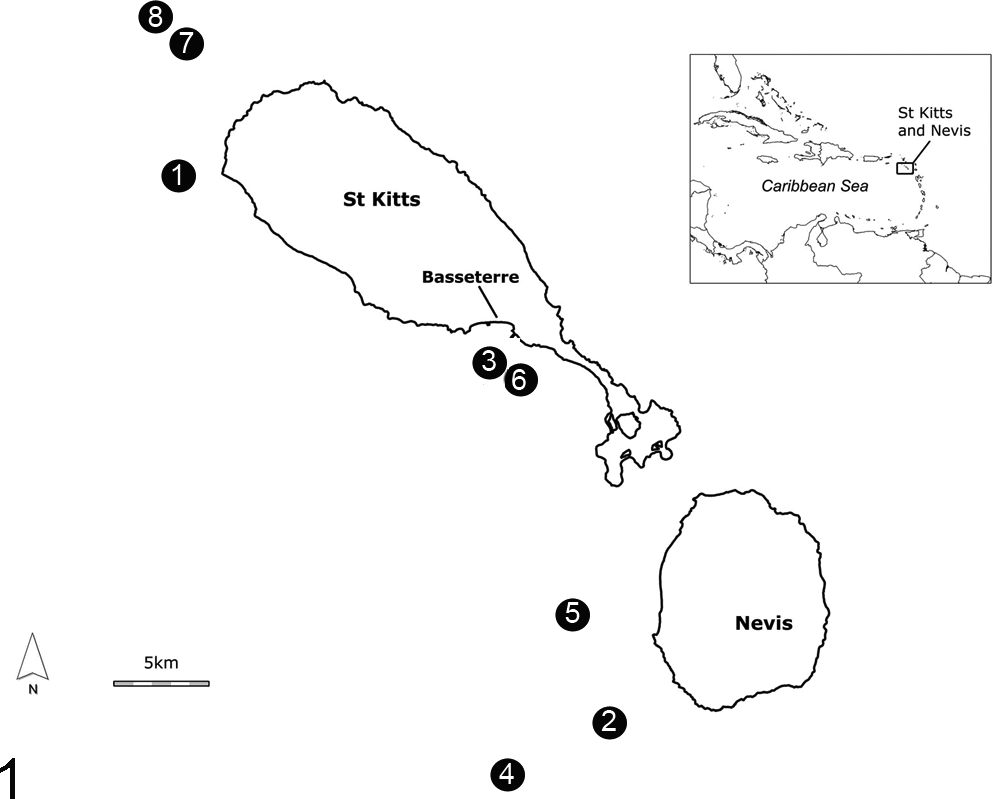
Map detailing harvest sites of fish confirmed to be infected with Streptococcus iniae.
List of Confirmed Streptococcus iniae Infections Throughout the 2008 Reef Fish Outbreak.
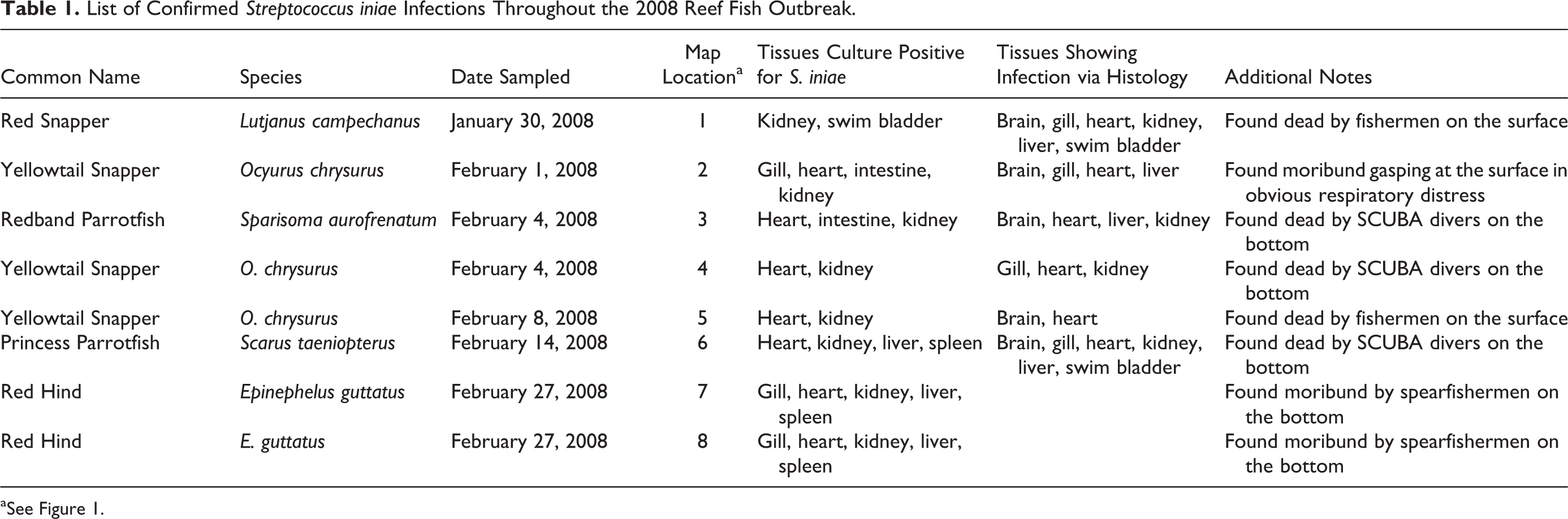
aSee Figure 1.
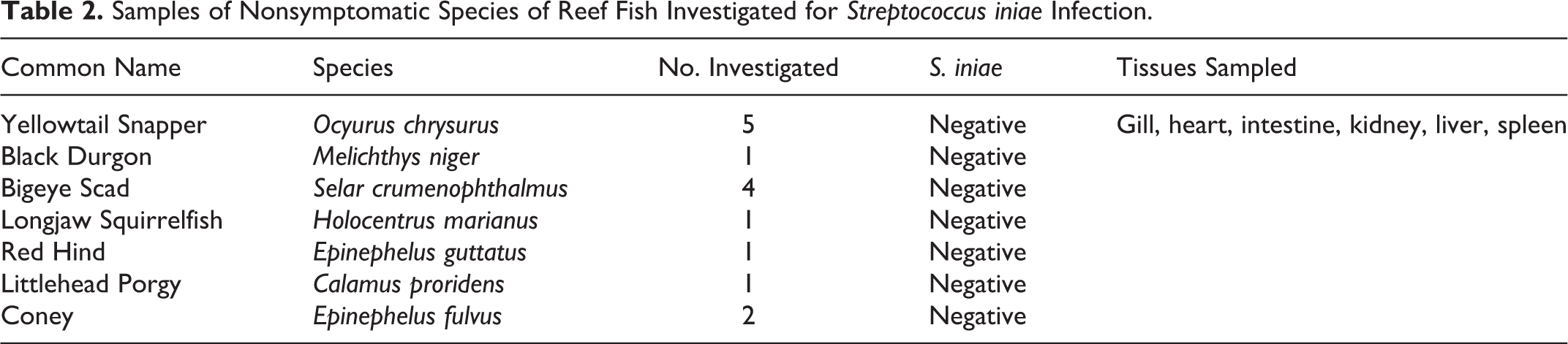
Samples of Nonsymptomatic Species of Reef Fish Investigated for Streptococcus iniae Infection.
Gross examination was performed, and samples from relevant tissues were collected for cytological, histological, and bacteriological analyses. Tissues were fixed in 10% neutral-buffered formalin and processed in a routine manner and stained with hematoxylin and eosin (H&E) or Brown and Brenn (Gram B&B) for histological examination.
Consistent necropsy findings in clinically affected fish included petechiation along the ventral abdomen, gills and/or fins (n = 2); gills were occasionally diffusely red and covered by a thick layer of clear mucus or multifocal raised areas composed of fibrin and exudate (n = 3; Fig. 2). The pericardium was covered by a multifocal to coalescing layer of white opaque material that was mostly not adherent (fibrin) but was focally adherent to the epicardial surface (fibrosis) (n = 5; Figs. 3, 4). Livers were typically diffusely pale and friable (n = 8), and gallbladders were prominent with empty intestinal lumina (n = 8), suggesting a period of inanition. Cytological examination from postmortem liver and pericardial imprints revealed degenerate cells due to autolysis. Frequent chains of bacterial cocci were observed extracellularly; in some cases, remnants of leukocytes contained intracytoplasmic coccoid bacteria, suggesting antemortem phagocytosis rather than postmortem bacterial overgrowth (Fig. 5).
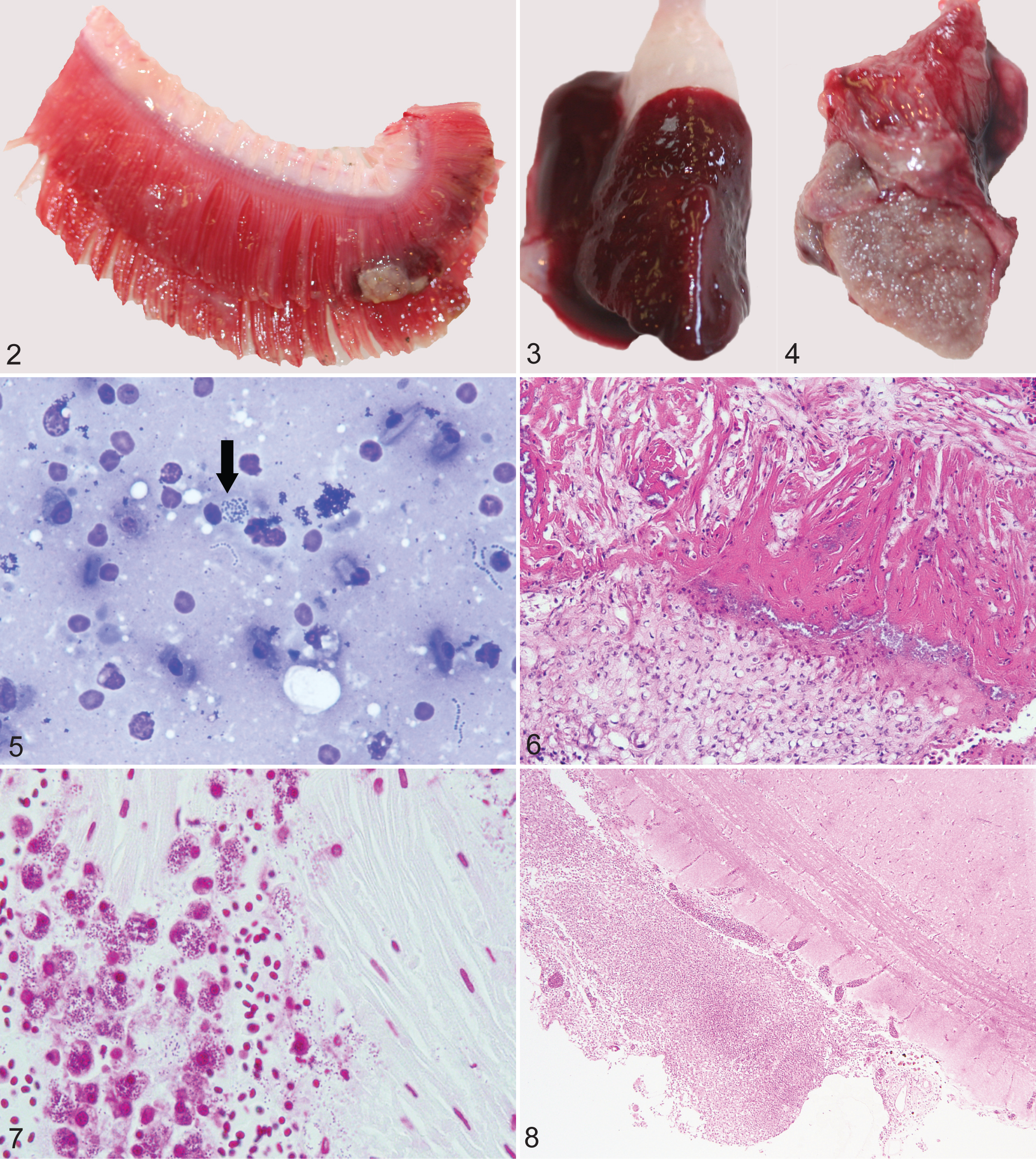
Fibrinous or fibrogranulomatous epicarditis and pericarditis with intralesional streptococci was the most prominent gross and histologic feature (n = 5), with marked bacterial colonization of the epicardium and infiltration of the underlying myocardium with hemorrhage and a diffuse infiltrate of granulocytes, plump fibroblasts, and macrophages that often contained Gram-positive bacterial cocci (Figs. 6, 7). Pericardial adipose tissues overlying the ventricular epicardium at the junction to the bulbus arteriosus were expanded by a marked diffuse infiltrate of macrophages, granulocytes, fibroblasts, and increased deposition of fibrous connective tissue, with cholesterol cleft formation (pericardial granulomatous steatitis; n = 1). Fibrinous perihepatitis was a frequent feature, with intralesional bacterial colonies, which were also prominent within sinusoids and portal veins. Histologic examination revealed atrophy and loss of the secondary gill lamellae, replaced by infiltrates of predominantly macrophages and granulocytes; Gram stains revealed Gram-positive coccoid organisms within the interstitium of the secondary lamellae, occasionally phagocytized by macrophages. Within the cerebrum, fibrinous meningitis (n = 3; Fig. 8) with mononuclear perivascular cuffing and intralesional diplococci were pronounced, with transmural inflammatory infiltrates and fibrinoid necrosis of the vessel walls (vasculitis; not shown). Within the gray matter, there were increased microglial cells (gliosis) and prominent perivascular inflammation. Multisystemic granulocytic vasculitis with intravascular diplococcal bacteria with marked bacterial colonization of endothelial linings occurred in the swim bladder, kidney, and spleen. All bacterial colonies were Gram-positive on Gram B&B. The asymptomatic fish collected as controls did not contain relevant gross or histologic lesions, nor were bacterial agents grown from a variety of tissues (Table 2).
Remel BactiSwab Amies Clear Gel Collection and Transport Swabs (Remel Inc, Lenexa, KS) were used to sample a variety of tissues at necropsy, including epicardium, swim bladder, kidney, gills, intestine, spleen, and liver (Table 1). Surface swabs were inoculated onto 5% sheep blood agar plates and incubated at 35°C in 5% CO2 for 18 to 24 hours. The API 20 Strep (bioMérieux, Inc, Durham, NC) was used to biochemically identify the isolates following the manufacturer’s instructions. Single β-hemolytic colonies were picked for Gram staining. Once single colonies were observed and purity of the isolate determined, the isolate was frozen at –80°C in 1× phosphate-buffered saline (PBS) containing 20% glycerol for later use. Bacterial swabs from various tissues (Table 1) readily produced pure cultures of catalase-negative, β-hemolytic streptococci, consistent with S. iniae. Although the API 20 Strep commercial kits do not have a code for S. iniae, the bacteria isolated during this outbreak gave essentially identical profiles, with most isolates being positive for esculin hydrolysis, pyrrolidonyl arylamidase, β-glucuronidase, arginine dihydrolase, alkaline phosphatase, and leucine aminopeptidase. All isolates could use ribose, trehalose, starch, and glycogen after a 24-hour incubation at 37°C.
All recovered isolates were used for molecular analysis. A loop of the bacterium was suspended in 500 μl of sterile 1× PBS, and genomic DNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Valencia, CA) following the manufacturer’s suggested protocol for Gram-positive bacteria. Extracted DNA was stored at –20°C until further use.
Molecular identification of the isolates was determined by amplification and sequencing of the 16S ribosomal RNA (rRNA) gene. 4 Amplicons for sequencing were purified with the QiaQuick PCR Cleanup Kit (Qiagen) according to the manufacturer’s suggested protocol and submitted to Davis Sequencing (Davis, CA) for sequence determination. The obtained sequences were compared with those stored in GenBank using a BLASTn search of the nonredundant nucleotide (nr/nt) database of the National Center for Biotechnology Information.
Molecular diagnosis provided by amplification and sequencing the 16S rRNA gene confirmed the isolates were S. iniae. A 99% identity was shown between the isolates recovered from wild reef fish in St Kitts and other S. iniae sequences stored in GenBank. The closest match was the partial 16S rRNA gene sequence of S. iniae strain CMS004, isolated from fish of Guangxi, China (GenBank: EU620577.1). This is not surprising due to the highly conserved nature of the 16S rRNA gene among members of the same bacterial species.
Genetic fingerprinting for each isolate was carried out using modifications to existing protocols, 8,15 using Escherichia coli (ATCC 25922) as an outlier. Repetitive sequence–based polymerase chain reaction (PCR) was carried out using either the BOX (5′-CTACGGCAAGGCGACGCTGACG-3′) or the GTG5 (5′-GTGGTGGTGGTGGTG-3′) primers, respectively. 17,18 For both primer sets, electrophoretic profiles demonstrated high levels of homogeneity for the 8 different S. iniae isolates, all of which were significantly different from the E. coli outlier (Fig. 9).
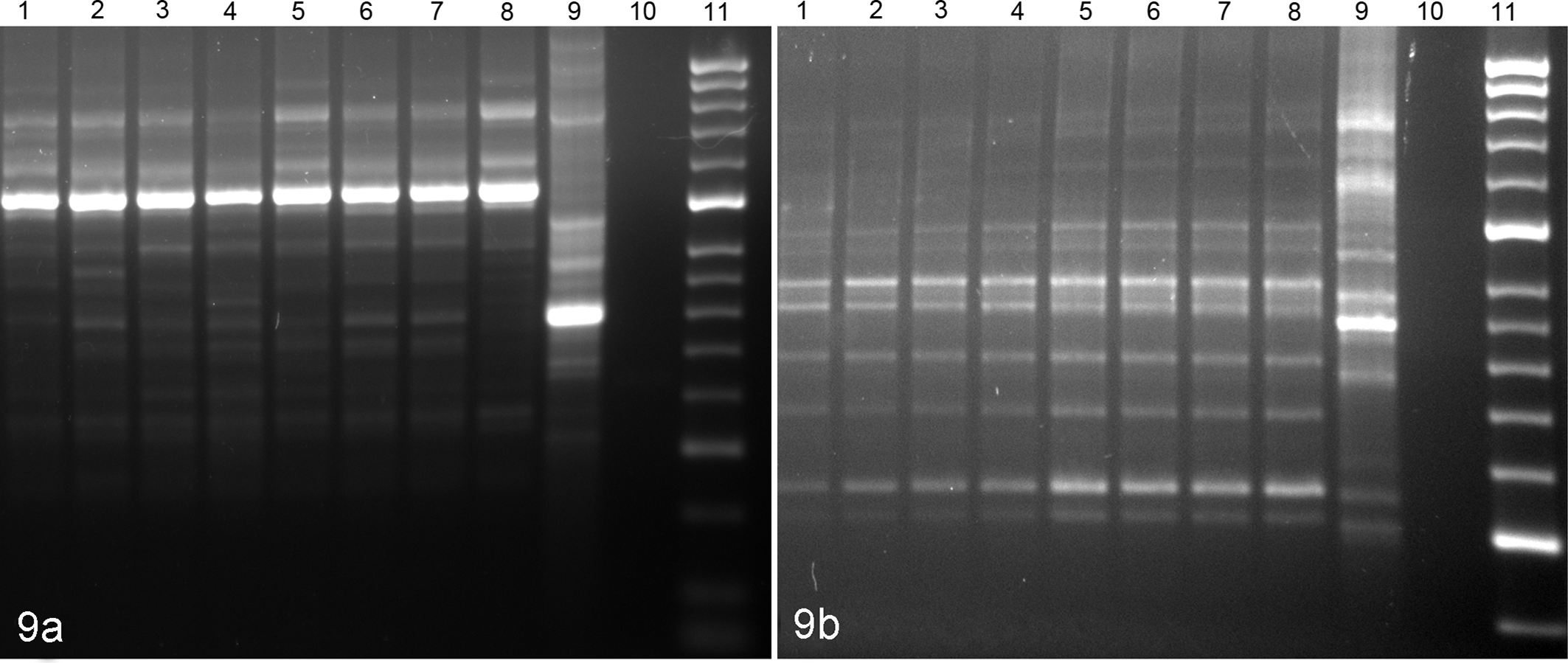
Electrophoretic profiles of Streptococcus iniae isolates recovered from an outbreak in reef fish and reference isolate generated by (a) BOX primer sets and (b) GTG5 primer sets. Lanes 1–8, S. iniae isolates; lane 9, Escherichia coli reference isolate (ATCC 25922); lane 10, negative control; lane 11, molecular weight ladder (HyperLadder 50 bp; Bioline USA, Taunton, MA).
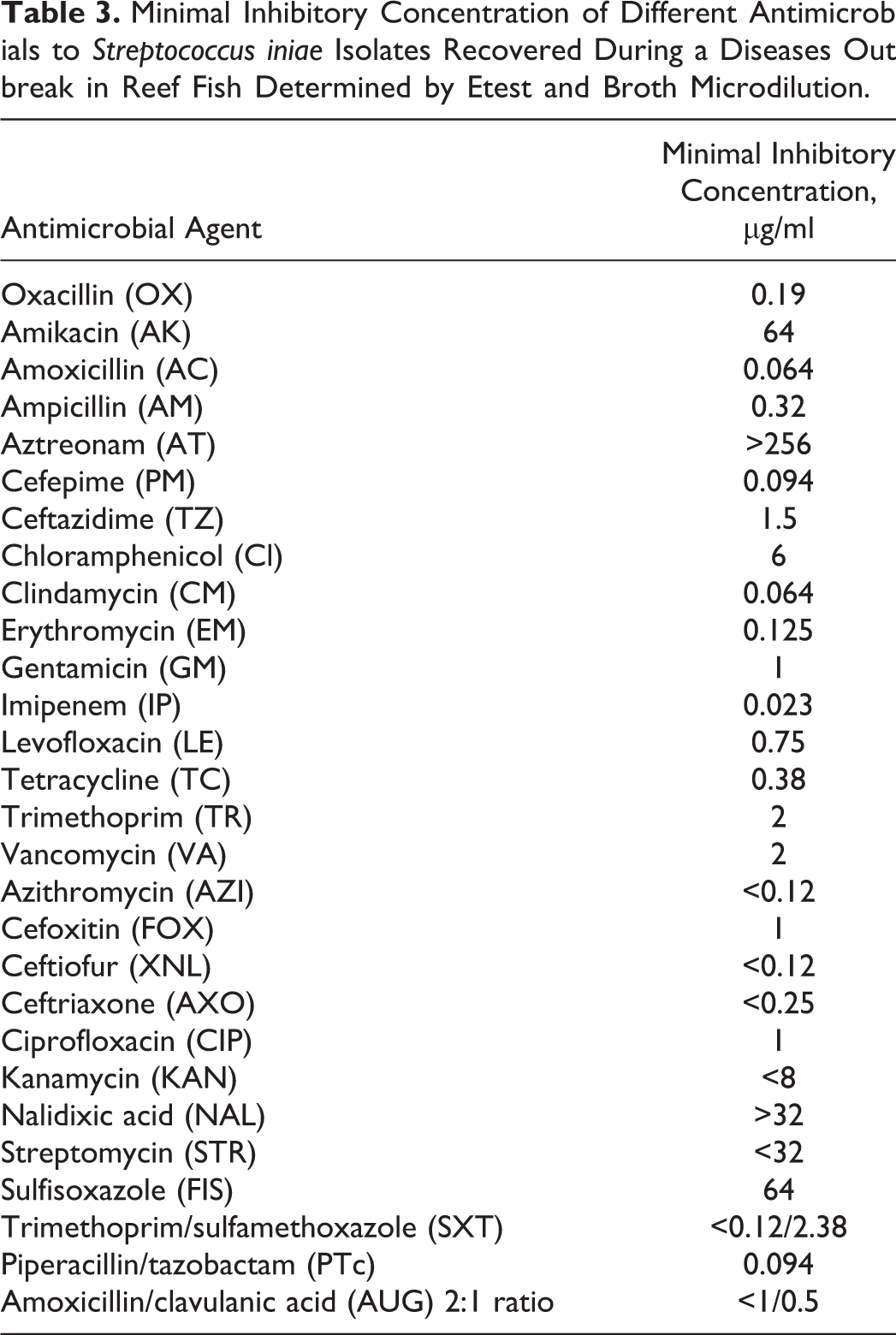
The minimal inhibitory concentrations (MICs) of 28 antimicrobial agents to S. iniae isolates from reef fish and quality controls (E. coli ATTC strain) were tested using the Etest and the Sensititres Gram Negative NARMS Plate format (Trek Diagnostic System, Cleveland, OH), using the manufacturer’s suggested protocol and the Clinical and Laboratory Standards Institute published protocols. 2 However, interpretation of results and use of antibiotics for these organisms are difficult since no criteria have been established for interpreting the MIC test results (Table 3).
Minimal Inhibitory Concentration of Different Antimicrobials to Streptococcus iniae Isolates Recovered During a Diseases Outbreak in Reef Fish Determined by Etest and Broth Microdilution.
Little is known about the nature and relative covariates or causes of S. iniae outbreaks in tropical reef fish. Mortality events in both freshwater and marine fish populations have been attributed to pollutants, harmful algal blooms, and a variety of natural and anthropomorphic causes and are more often reported in farmed fish populations, which are more carefully assessed. Factors leading to wild fish outbreak events often remain unclear, making their etiologies enigmatic and their occurrences difficult to predict. Panek 13 reviews epizootic events in the tropical western Atlantic and highlights the general lack of information on such sporadic events in the past.
Warm-water streptococcosis was associated with a Southeastern Caribbean wild fish outbreak and resultant mortality event in 1999. 6 During this particular outbreak, a number of possible inciting causes were suggested, including stress from a harmful algal bloom or some relation to a discharge of the Orinoco River during the rainy season. 14 During the 2008 St Kitts outbreak, no water samples were analyzed for the presence of toxic algae. However, there was higher than normal rainfall during that season that may have resulted in soil runoff as a factor. However, this is an unlikely cause since the reported fish kills had no correlation to areas of the islands that experienced increased runoff. The possibility of a bloom flowing from an adjacent or nearby island, as in the case of the aforementioned Orinoco bloom of 1999, cannot be entirely discounted. There is little support for such a hypothesis, however, since no known nearby islands, subject to the same ocean currents, experienced any mortality event. This is despite the country having several nearby islands, one within approximately 10 km.
Zoonotic infections by this bacterium requiring hospitalization are rare. 9,11 However, the risk to an immunocompromised population should not be disregarded. S. iniae infections in humans are considered an emerging complication and have been documented mainly in North America, the Middle East, and the Asian-Pacific region. 16,19 Infections are usually associated with processing of fresh fish; however, there is a general opinion that many cases are often unrecognized and therefore are underreported. 5,7,10,12,16
According to the St Kitts and Nevis chief medical officer, despite the unguarded handling of infected fish by fishermen and other members of the public sector, no human streptococcosis cases were reported during the 2008 outbreak. However, due to the zoonotic potential of these bacteria, in-depth antimicrobial susceptibility analysis was performed using the recovered isolates. These data can be used as a baseline of treatment in future human and/or animal outbreaks in the region (Table 3).
Genotypic (repetitive element palindromic PCR), phenotypic, and antimicrobial susceptibility analysis of recovered isolates indicated homogeneity among S. iniae recovered during the outbreak. A detailed comparison of these Caribbean isolates to similar isolates recovered from other geographical locations and commercial fish farms should add to the understanding on the homogeneity of this pathogen. Further research should focus on developing surveillance programs for the detection of Streptococcus sp in aquatic and terrestrial animals in the region. A One-Health approach should be targeted due to the zoonotic implications of these bacteria, and a better understanding of environmental conditions, human and animal interaction, and the relationship between them that allow outbreaks in wild fish populations is needed. This approach is necessary, not only for local communities that depend on tourism and wild fisheries as a source of income and protein but also for the growing Caribbean aquaculture industry.
Footnotes
Acknowledgments
We thank The St Kitts Department of Fisheries for their technical assistance and St Kitts and Nevis Department of Health for assisting with public awareness and information dissemination and access to information on potential zoonotic infection of humans presenting to the hospital with bacterial infections during the outbreak. We also thank Dr Gerry Johnson and Dr David Groman for technical assistance during the outbreak and Matthew McMillan and Cynthia Ware for their technical assistance in the preparation of this manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
