Abstract
Equine penile papillomas, in situ carcinomas, and invasive carcinomas are hypothesized to belong to a continuum of papillomavirus-induced diseases. The former ones clinically present as small grey papules, while the latter 2 lesions are more hyperplasic or alternatively ulcerated. To test the hypothesis that these lesions are papillomavirus-induced, samples of 24 horses with characteristic clinical and histologic findings of penile papillomas or in situ or invasive squamous cell carcinomas were collected. As controls, 11 horses with various lesions—namely, Balanoposthitis (6 cases), melanoma (3 cases), follicular cyst (1 case), and amyloidosis (1 case)—were included. DNA was extracted and polymerase chain reaction applied to amplify papillomavirus DNA. The respective primers were designed to amplify DNA of the recently discovered equine papillomavirus EcPV2. All tested papilloma and squamous cell carcinoma samples were found to contain DNA of either of 2 previously published EcPV2 variants. Among the other samples 6 of 11 were found to contain EcPV2 DNA. To further support the findings and to determine where the papillomavirus DNA was located within the lesions, an in situ hybridization for the detection of EcPV2 DNA was established. The samples tested by this technique were found to clearly contain papillomavirus nucleic acid concentrated in the nucleus of the koilocytes. The findings of this study support previous data and the hypothesis that papillomaviruses induce the described penile lesions in horses.
Keywords
Papillomaviruses (PVs) are small double-stranded DNA viruses that play an important role in the development of various benign and malignant neoplasms of the skin and mucous membranes in humans and animals. 6 In horses, PV-associated skin disorders are common and occur in different forms, including the bovine PV-associated equine sarcoids. 1,11 Yet 3 actual equine PVs (EcPVs) have been identified in lesions that had been suggested to be potentially PV associated. 4,9,12,14 Those are the so-called classical equine viral papillomas, which are typically found in young horses (EcPV1), genital papillomas (EcPV2), and equine aural plaques (EcPV3). 9 One of the viruses, EcPV2, has recently been identified in genital papillomas and carcinomas independently by different groups, and it was shown that at least 2 EcPV2 variants exist. 8,13,15 EcPV2 DNA was also identified in ocular squamous cell carcinomas (SCCs) and in swabs of clinically unaffected skin. 15
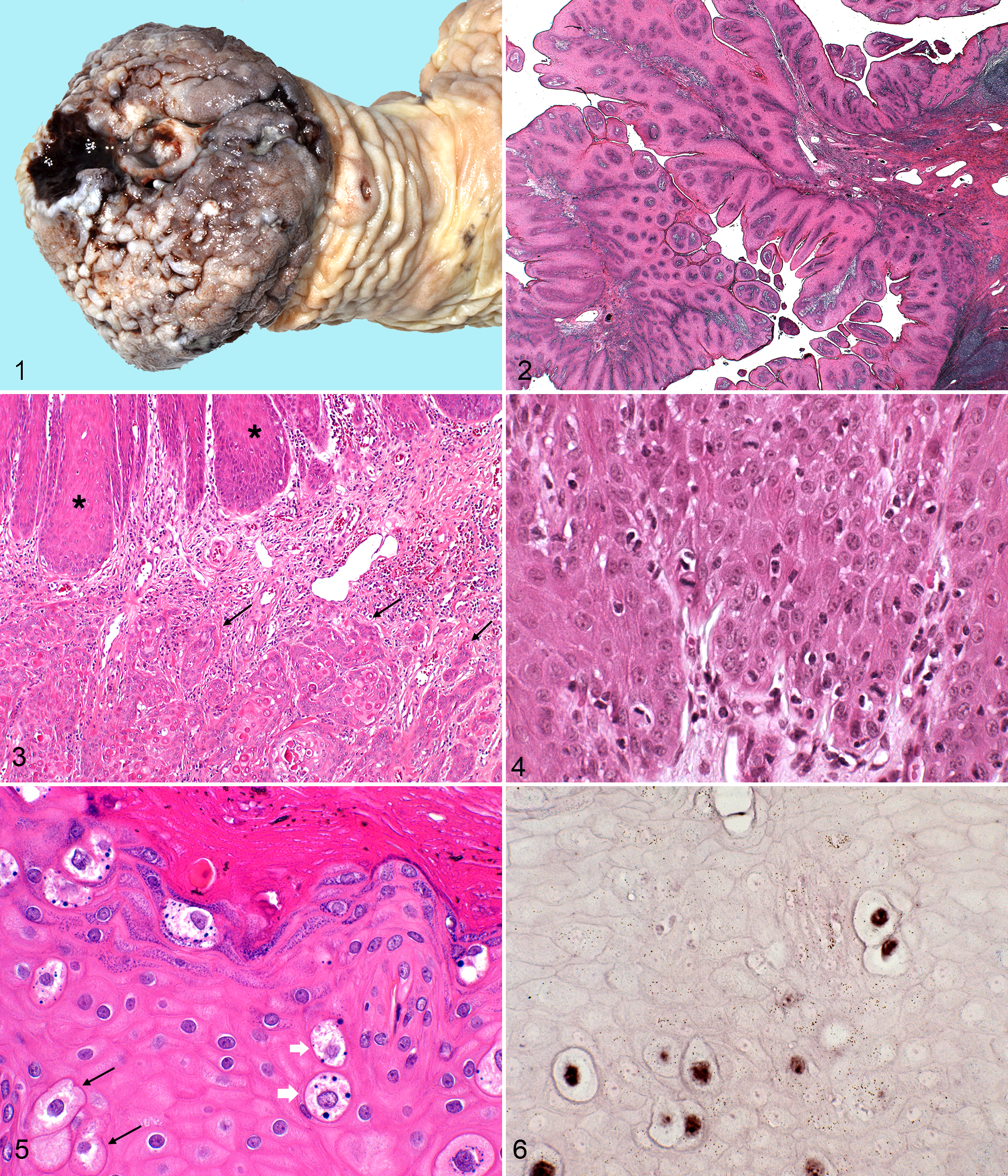
Equine penile papillomas, in situ carcinomas, and invasive carcinomas are hypothesized to belong to a continuum of PV-induced diseases. The former lesions clinically present as small grey papules (Fig. 1), while the 2 latter ones are more hyperplasic or alternatively ulcerated. Histologic changes range from hyperplasia of the epithelium with sparse koilocytes to in situ or invasive SCCs.
The finding of EcPV2 DNA in penile swabs of clinically healthy horses raises the question about the pathogenic role of the virus in lesions which contain PV DNA. 15 To establish detection methods and to address the question where the PV DNA is located in the penile skin, 2 polymerase chain reaction (PCRs) and a specific in situ hybridization (ISH) were evaluated, and the techniques were applied on samples of penile papilloma and penile SCC.
Materials and Methods
Samples and Histopathology
Samples were obtained from the Institute of Veterinary Pathology in Zurich and the Institute of Veterinary Pathology in Bern and were chosen according to the original clinical and pathologic diagnosis. In summary, 10 cases with invasive SCC, 7 cases of in situ carcinomas, and 7 cases with papilloma were selected for this study (Table 1). Eleven cases with other penile lesions were included as controls (Table 2). If more than 1 sample per case was available, those other samples were included as well but not counted separately. Tissue samples were fixed in 4% buffered formalin. After embedding in paraffin, 4-μm sections were stained with hematoxylin and eosin for histopatholgic examination. Papilloma was diagnosed based on finger-like projections of squamous epithelium with normal maturation and small central cores of connective tissue. Fibropapillomas were diagnosed if extensive central cores of connective tissue were present. Carcinoma in situ was diagnosed if a thickened epithelium with loss of uniform maturation of keratinocytes, loss of normal architectural orientation, and atypical keratinocytes was detected. The latter were either swollen with pale cytoplasm and a normal nucleus, swollen with intracytoplasmic clear vacuoles and a pyknotic nucleus (= koilocytes), or swollen with pale basophilic cytoplasm and intracytoplasmic blue granules. Diagnosed SCCs consisted of islands and trabeculae of squamous cells originating from the epithelium and infiltrating the connective tissue (Fig. 2).
Papilloma and Squamous Cell Carcinoma Samples
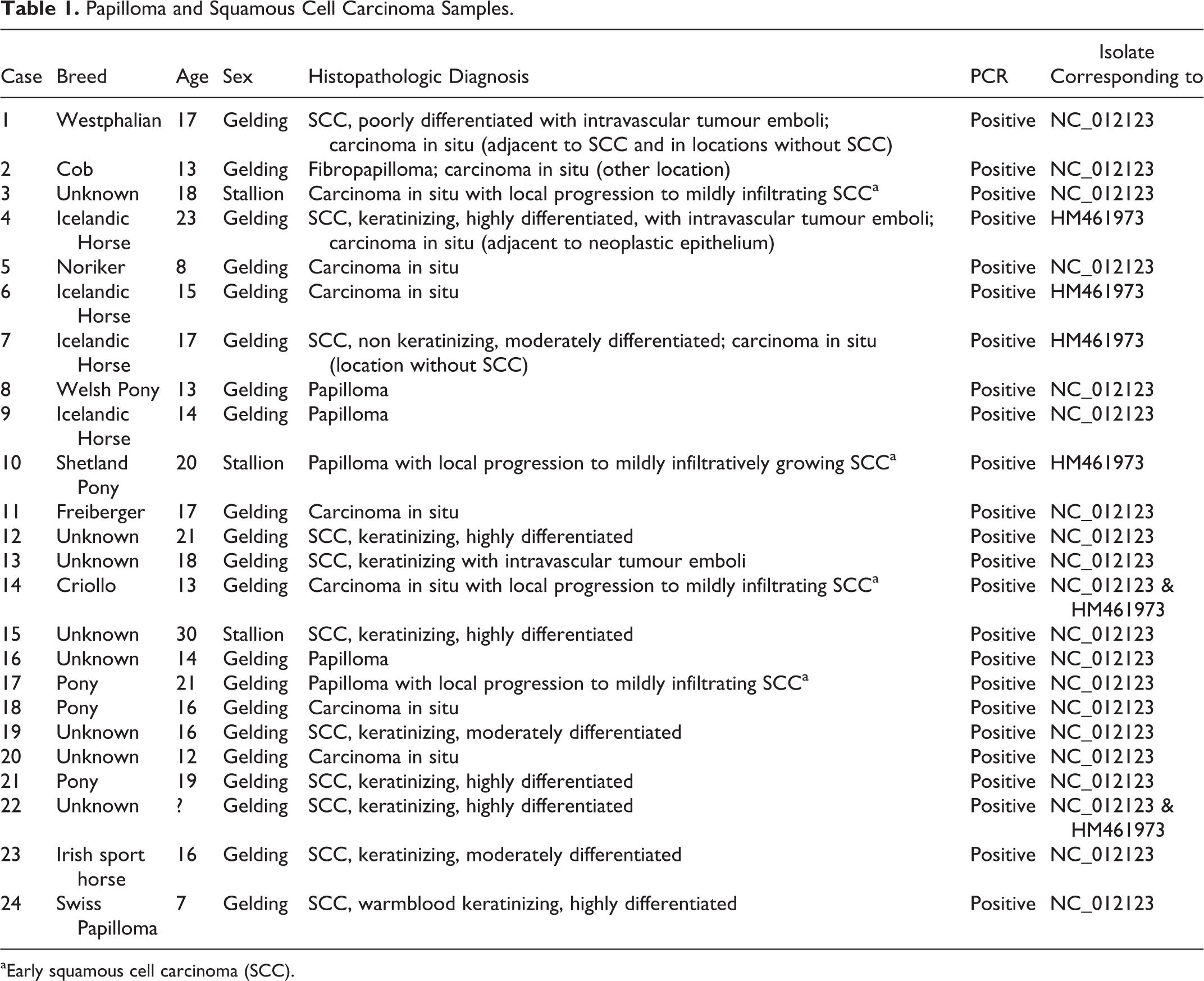

aEarly squamous cell carcinoma (SCC).
Control Samples
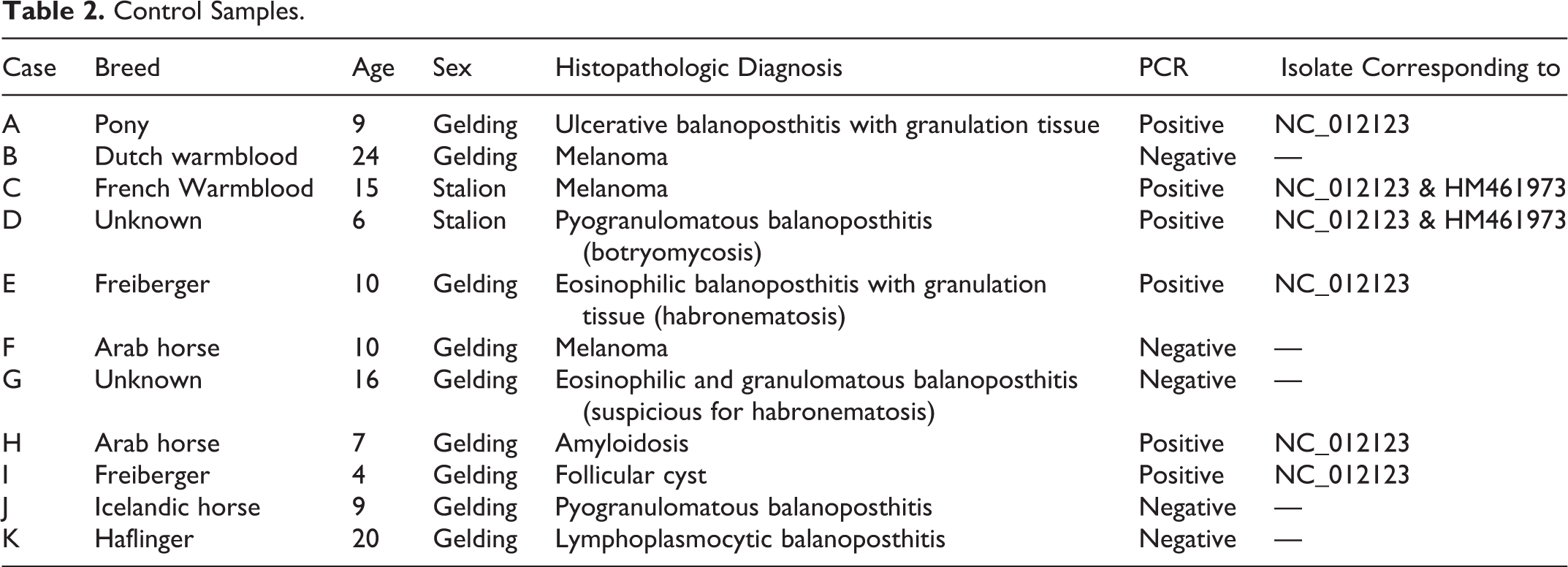

In Situ Hybridization
To generate an ISH probe, forward (5′-GAG CTG TGC AGT GTC ACG TT-3′) and reverse (5′-TCT CCT GGA CAA GCC ACT CT-3′) primers were designed based on the L1 gene sequence of the published sequence of EcPV2 (NC_012123). The 294nt Digoxygenin (DIG)-labeled probe was generated using the PCR DIG Probe Synthesis Kit (Roche) according to the manufacturer’s protocol. The following PCR conditions were applied: 95°C for 3 minutes, followed by 35 cycles of 95°C for 30 seconds, 60°C for 30 seconds, and 72°C for 30 seconds. Labeled PCR products were extracted from an agarose gel using a MinElute gel extraction kit (Qiagen).
A DIG-labeled probe targeting the green fluorescent protein (GFP) gene was generated in an analogous way employing following forward (5′-AGG ACG ACG GCA ACT ACA AG-3′) and reverse (5′-CTG GGT GCT CAG GTA GTG GT-3′) primers. The same PCR conditions were used; a pHSV-eGFP amplicon vector served as template. 2 This probe served as negative control throughout the hybridization experiments.
Tissue sections (3–4 µm) were obtained using a microtome (HYRAX M55, Zeiss); samples were placed onto positively charged slides (Thermo Scientific); and a modified standard ISH was performed as described before. 10
PCR Evaluation
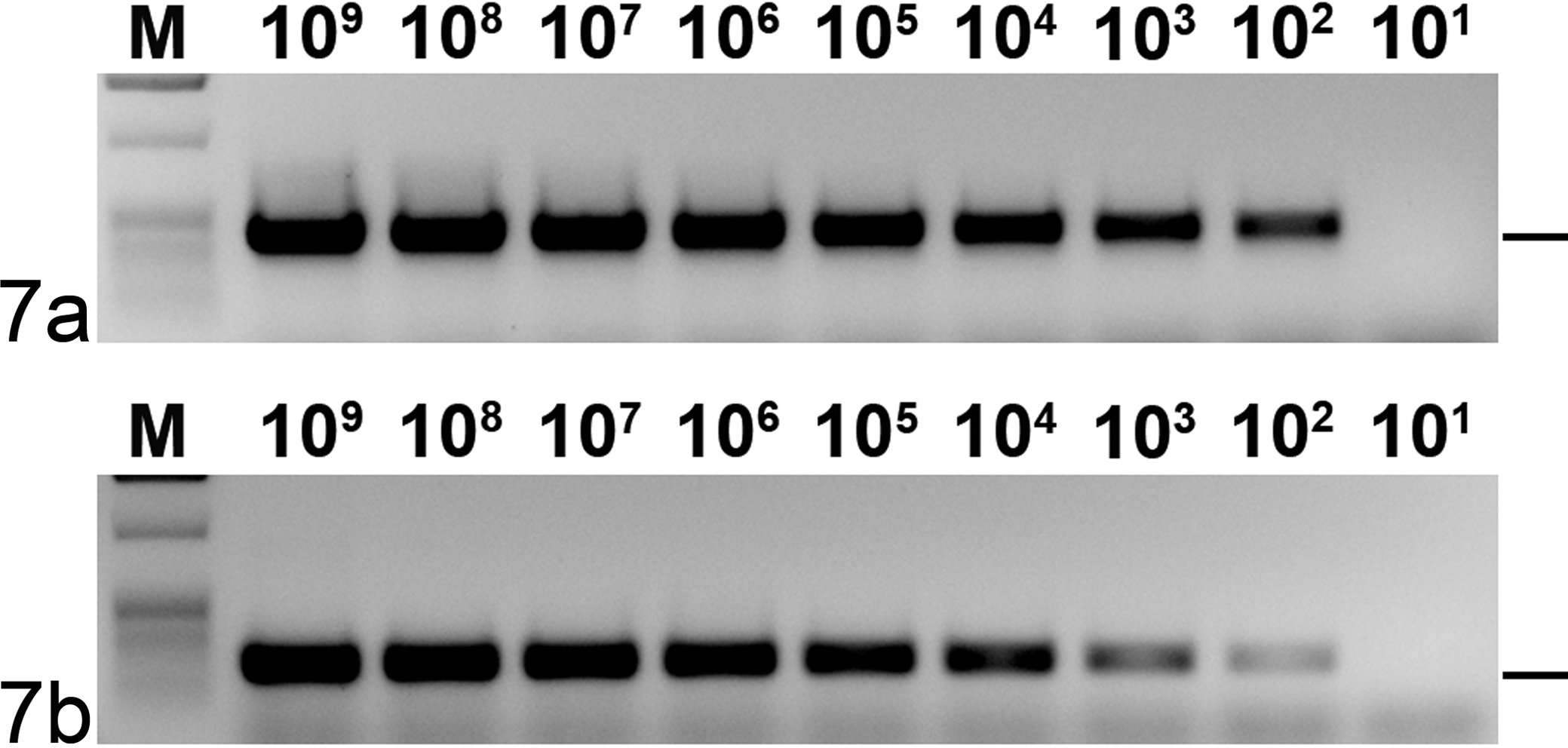
Two primer combinations were evaluated for PCR. One was the primer combination EcPV2probe f / EcPV2probe r, which amplifies a 294nt stretch of the L1 open reading frame (ORF) of EcPV2 and served also to generate the ISH probe. The second one was the primer combination EcPV2d f (5′-CAG ACT TGT CTG GGC TCT CC-3′) and EcPV2d r (5′-TCC CGC CTA GCA TAG AAG AA-3′), which had been designed to amplify a 474nt stretch of the L1 ORF of EcPV2. To determine the approximate detection levels of the PCRs, serial dilutions were made from 109 down to 101 molecules of an entire genomic clone of EcPV2. For that purpose the DNA concentration of the plasmid stock solution was measured using a Spectrophotometer (ND-1000, Thermo Scientific). Taking into account the plasmid size, the amount of molecules per microliter was calculated. RedTaq ready reaction mix (Sigma) was used, and conditions were chosen for both PCRs as follows: 94°C for 3 minutes, followed by 39 cycles of 94°C for 30 seconds, 50°C for 30 seconds, and 72°C for 30 seconds.
DNA Amplification and Analysis
Total DNA of fixed tissue samples was isolated using a DNeasy extraction kit (Qiagen) according to the manufacturer’s recommendations for paraffin-embedded tissues. EcPV2probe f / EcPV2probe r PCR and 2 broad range PCRs with primer combinations binding within the E1 ORF (CP4/CP5) or within the L1 ORF (FAP59/FAP64) were used to test the samples. 5,7 Electrophoresis in a 1% agarose gel containing ethidium bromide was used to detect the amplified fragments. PCR products were extracted from the gel using QIAEX II DNA extraction kit (Qiagen).
The nucleotide sequences of both strands of the PCR products were determined (Microsynth) by cycle sequencing using an ABI 377 sequencer (Applied Biosystems). Positive and negative strand of each isolate were aligned, and a consensus sequence of 169 nucleotides was generated for each of the PCR positive samples. The sequences were compared with the NCBI database (BLAST X) and were all aligned with the according L1 stretches of the 2 available sequences of EcPV2.
Statistical Analysis
To statistically compare the PCR results of the different groups, Fisher exact test was applied. A t test was used to determine the significance of age differences.
Results
Histopathology and ISH
Twenty-four male horses (21 geldings, 3 stallions) with depigmented plaques (Fig. 1), nodules, and/or ulcerations of the penis were included in the study group. The mean age in this group was 16.4 years, ranging from 7 to 30 years. Different breeds were represented (Table 1). The mean age of the 11 horses in the control group was 11.8 years, ranging from 4 to 24 years. They also were of different breeds (Table 2). The age difference between the 2 groups was statistically significant (P = .05).
Histologic examination revealed 9 keratinizing and 1 nonkeratinizing SCC of variable differentiation with intravascular tumour emboli in 3 of these cases (Table 1). In 3 of these cases, areas with carcinoma in situ were found adjacent to the SCC or in tissue sections from other locations. These were characterized by hyperplasia of the epithelium, mild loss of normal architectural orientation, and mild loss of uniform maturation of keratinocytes and by the presence of atypical keratinocytes that were swollen with pale cytoplasm (and normal nucleus), swollen with intracytoplasmic clear vacuoles and pyknotic nucleus (= koilocytes), or swollen with pale basophilic cytoplasm and intracytoplasmic blue granules (Figs. 3, 4). In 7 cases, only carcinoma in situ was diagnosed; however, very mild infiltrative growth was observed in 2 cases (Nos. 3 and 14), and a final diagnosis of early SCC was made. Six cases with finger-like projections of squamous epithelium with normal maturation typically seen in papillomas (Fig. 5) were included. Among them was one case with concurrent invasive SCC (No. 24); in 2 cases (Nos. 10 and 17), local progression to mildly infiltrative growth was detected, and a final diagnosis of papilloma with early SCC was made. One case with finger-like projections of hyperplastic squamous epithelium combined with extensive central cores of connective tissue was detected, and a diagnosis of fibropapilloma was made (Fig. 5; No. 2).
Two cases of in situ SCC were selected to be tested by ISH. Blocks from case Nos. 1 and 3 were chosen, as they displayed all characteristic changes described and contained large respective areas. Upon ISH a strong signal was evident in many of the koilocytes (Fig. 6), while no signal was present when a control probe was used. In case No. 1 numerous positive cells were visible. Primarily nuclei of koilocytes were labeled in all levels of the epithelium. In case No. 3, small to moderate number of positively labeled cells was found, and no labeling of cells was present in adjacent normal epithelium.
DNA Amplification and Analysis
To use PCR for PV detection, 2 primer combinations amplifying DNA from the L1 ORF of EcPV2 were evaluated. Both combinations performed equally well when tested on serial dilutions of EcPV2-plasmid DNA (Fig. 7). The detection level in both cases was about 100 copies of template. However, as fixed samples were used in this study, the primer combination leading to a smaller product was chosen. DNA could be amplified from all 24 papilloma and SCC samples (100%) using PCR with the primer combination EcPV2probe f/EcPV2probe r and in 6 of the 11 control samples (55%). Statistically, the difference between the groups is highly significant (P = .001). No DNA was amplified by PCR with primer combinations CP4/CP5 or FAP59/FAP64, respectively, in any of the tested samples. In all cases, where more than 1 sample of the same case was tested, the results were consistent.
Sequencing of the PCR products confirmed the amplified DNA to belong to EcPV2. While the sequences obtained from 18 cases were identical, those of 6 cases were different in 2 positions (6921 and 7002 relative to EcPV2-HM461973). Aligning the 24 sequences with the according stretches of the 2 EcPV2 sequences revealed that the sequences of cases Nos. 1–3, 5, 8, 9, 11–13, 15–21, 23, and 24 were identical with EcPV2-NC_012123, while the sequences of cases Nos. 4, 6, 7, and 10 were identical with EcPV2-HM461973. In case Nos. 14 and 22, the signal in positions 6921 and 7002 was not corresponding clearly to either of the 2 variants but was equally strong, indicating both respective bases in the same positions. EcPV2 DNA was amplified from 6 of the 11 control samples—namely, cases A, C, D, E, H ,and I with diagnoses of balanoposthitis, melanoma, amyloidosis, and follicular cyst. EcPV2-NC_012123 DNA was clearly identified in 4 of these 6 cases, while in the other 2, the obtained sequences were equally indicating EcPV2-NC_012123 and EcPV2-HM461973 DNA. Sequencing results from cases where more than 1 sample was tested were consistent.
Discussion
Equine penile papillomas and in situ and invasive SCCs are hypothezised to belong to a continuum of PV-induced disorders. They are common diseases and in case of invasive SCCs may require aggressive treatment (amputation) when diagnosed in late stages. The involvement of PVs had been suggested and a putatively involved PV had been identified. 8,9,13,15 Here we report the establishing of a PCR assay detecting EcPV2 DNA in 24 of 24 papilloma in situ SCC or SCC lesions and in 6 out of 11 other penile lesions. The sequencing of the PCR products revealed the sequences of 2 EcPV2 variants, and ISH in 2 cases revealed EcPV2 DNA in koilocytes.
Some recent independent studies identified EcPV2 DNA in penile SCCs in Australia, Austria, Belgium, the UK and the USA, using different techniques, including various PCRs, rolling circle amplification, ISH and immunohistochemistry. 8,9,13,15 However, the results of these studies regarding the incidence of EcPV2 vary significantly, ranging from 10% to 100% of tested SCCs. 8,15 Vanderstraeten et al report 100% (n = 15) of cases with penile lesions (penile SCCs, penile papillomas, and penile intraepithelial lesions) and 43% (n = 40) of penile control samples to contain EcPV2 DNA. Scase et al report in total 70% (n = 30) of penile lesions (penile SCCs, penile in situ SCCs, and penile papillomas) and 5% (n = 41) of penile control samples to contain EcPV2 DNA. Knight et al report 45% (n = 20) of penile lesions (penile SCCs) and 5% (n = 20) of penile control samples to contain EcPV2 DNA, all according to PCR results. The reason for these significant variations might be manifold. Differences regarding the samples including breeds, age of the horses, and environmental and other factors may play a role. The process of sampling, sample handling, and processing as well as the inclusion criteria for the individual studies could have an influence on the results as well. Finally the PCR assays applied in the different studies are not alike, since different primer pairs were used and no detection limits are reported. This makes comparison and interpretation PCR results difficult, especially in negative cases. Summarizing the cases of all studies mentioned above, EcPV2 DNA was detected in 80% of the lesional and in 23% of the control samples.
The PCR chosen here for EcPV2 detection amplified DNA from all 24 tested samples of papillomas and in situ and invasive SCCs and also from 6 of 11 controls. The presence of EcPV2 DNA in skin without the typical PV-associated lesions is not surprising, as PV-associated disorders are often latent and such findings have been reported before. The incidence is, however, much lower (55%) than in supposedly EcPV2-induced lesions (100%), and this difference is highly significant. Thus, regarding the incidence of EcPV2 infection, the results are similar to those of Vanderstraeten et al but differ to some extent from those of Scase et al and significantly from the results published by Knight et al. 8,13,15 We believe that the reasons for these differences are mainly the different PCR assays used, which probably do not have the same sensitivities. An actual difference based on the different geographical regions or slightly different inclusion criteria can, however, not be excluded.
Sequencing of the PCR products obtained here confirmed the DNA to be identical with EcPV2. Interestingly, the short stretches corresponded exactly to the 2 entirely sequenced and published EcPV2 variants. In 2 SCC and 2 control samples, however, the sequence was in positions 6921 and 7002, not corresponding clearly to either of the 2 variants but equally strong regarding the respective nucleotides of EcPV2-NC_012123 and EcPV2-HM461973. Sequencing errors are unlikely to be an explanation for this, as the finding was consistent and the electropherograms were otherwise of good quality, without noteworthy interferences. This consequently indicates that both variants may be found in the same tissue.
In previous studies, a variation in the sequences had been observed but not been analyzed in more detail, except for 2 cases where EcPV2-HM461973 was identified in Icelandic horses. 9,13 This current data also suggest that Icelandic horses may be more prone to EcPV2-HM461973 than to EcPV2-NC_012123 infection, but the sample size of this study is too small to draw definite conclusions. It is however obvious that EcPV2-NC_012123 is more prevalent in the tested samples than EcPV2-HM461973, as it was detected in 87% of the EcPV2 positives compared to 27% in case of EcPV2-HM461973.
ISH was established to determine whether the EcPV2 DNA, which had been amplified from the samples, is indeed located within the cells indicting a putative causal involvement or if it is just confined to the superficial epidermis, suggesting rather the role of a bystander. For this purpose, an EcPV2-specific probe and an unrelated probe of about equal size were generated and applied in the same way. Signal was only present in the sample when EcPV2 probe was applied. As DNA was detected within the koilocytes, it can be ruled out that the DNA is only present superficially. As koilocytes are regarded as typical indicators for a PV infection, finding PV DNA within their nucleus supports the hypothesis that EcPV2 is involved in the development of the respective lesions. However, it remains open if EcPV2 may contribute to the development of SCCs—and, if so, by which mechanisms. The intensity of the signal detected upon ISH indicates a high concentration of PV nucleic acids. The absence of an evident signal upon ISH with a GFP probe indicates that no unspecific binding occurs. In the target region of the probe, identities of up to 60% with corresponding sequences of other PVs can be found; thus, it might putatively bind not only to variants of the same but maybe also to other PVs. As no other PV DNA was detected in the samples by the generally more sensitive PCR, hybridization with the DNA of other PVs in the samples is not to be expected.
The mean age of the horses in the papilloma, in situ SCC, and invasive SCC group was 16.4 years, significantly higher than in the control group, where it was 11.8 years. A correlation between age and the required time for SCCs to develop may be reflected in this difference. The development of PV-associated malignancies is also described to be correlated with age. 6,16 In humans the presence of PV DNA is in this context of high predictive value, as, for example, almost all women who eventually will develop cervical carcinomas are positive for PV DNA, while almost no PV-negative one develops such a tumor. However, only a small percentage of those who were once tested positive for PV DNA will later develop cancer. 6,16 Some of the results published before and those presented here may indicate analogies to this in the horse. 15 The high percentage of DNA containing clinical cases of papillomas and in situ and invasive SCCs compared to those without such lesions at least points in that direction.
The histologic findings of the cases included here support the previous hypothesis of causal correlation, as characteristics of a viral infection like koilocytes are present. They needed, however, to be placed into a context with results derived from other diagnostic methods. To confirm and determine the viral DNA in fixed samples, PCR as an easy and sensitive method has been chosen. Primers binding in the L1 ORF of EcPV2 were designed. The L1 ORF is the most conserved region in the PV genome, and it is also used for classification. 3 Thus, primers binding within this ORF are likely to bind on the DNA of variants and close relatives of the targeted virus. Sequencing of the products may give some indication of the putative classification of the detected virus.
The evaluation of the 2 primer combinations designed to amplify EcPV2 DNA revealed almost equal sensitivity tested on plasmidic DNA as template. However, we did not expect intact PV DNA in the samples. Therefore, we chose the primer combination amplifying the shorter stretch of DNA, as the samples available for testing were extracted from formalin-fixed, paraffin-embedded tissues. Since formalin is known to induce DNA damage, the chances of successfully amplifying DNA from such a sample may be higher if the expected PCR product is small. Additional evidence from our own experiments supports this (data not shown).
Two broad-range PCRs that were applied failed to amplify DNA in the samples, which can probably be explained by a large proportion of mismatches. Upon alignment with the EcPV2 sequence there are 3 mismatches close to the 5′ and 3 close to the 3′ end of the primer FAP59. In case of the primer CP5, 5 mismatches are located around the middle of the primer sequence.
In conclusion, 2 PCRs suitable for EcPV2 detection in fixed samples have been evaluated, and 1 of them successfully applied. DNA of 2 EcPV2 variants was found in 24 cases, including 1 fibropapilloma, 6 papillomas (1 with concurrent invasive SCC, 2 with early SCC), 7 in situ carcinoma (2 with concurrent early SCC), and 10 SCC (3 with concurrent in situ carcinoma). In control samples, 6 out of 11 were containing EcPV2 DNA. Thus, a spread of EcPV2 in Switzerland has been demonstrated, with the EcPV2-NC_012123 variant showing up more often than the EcPV2-HM461973 variant. Strong indications for coinfections with both variants in the same tissue were found. Additionally, an alternative ISH specifically designed for EcPV2 DNA detection was established, and the association of viral DNA with koilocytes has been shown. These findings taken together support a causal context of equine penile papilloma, in situ carcinoma, and invasive carcinoma with EcPV2.
Footnotes
Acknowledgements
We thank Ines Krähenbühl of the Institute of Veterinary Pathology, Vetsuisse Faculty Zurich, for support with the ISH; Elisabeth Schraner of the Institute of Virology, Vetsuisse Faculty Zurich, for assistance with microscopy; Marco Geisseler of the Institute of Virology for his work in the lab; and also Meret Wehrli Eser and Michelle Jackson of the Equine Department, Vetsuisse Faculty Zurich, for their support. Finally, we like to thank Joan E. Lange for English editing.
Declaration of Conflicting Interests
The authors declare that they have no conflicts of interest with respect to their authorship or the publication of this article.
Funding
Financial support for this study was provided by the Krebsliga, Switzerland.
