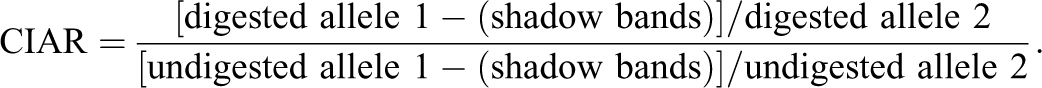Abstract
X-chromosome inactivation pattern (XCIP) analysis has been widely used to assess cell clonality in various types of neoplasms in humans. In the present study, a polymerase chain reaction–based feline XCIP analysis using the feline androgen receptor gene was developed. To construct the system of the analysis, polymorphism in CAG tandem repeats within the feline androgen receptor gene was explored using somatic DNAs from 50 male and 103 female cats. CAG tandem repeats in exon 1 of the feline androgen receptor gene were found to be polymorphic, containing 15 to 22 CAG repeats. Of the 103 female cats, 70 (68%) were heterozygous for the number of CAG repeats, indicating the possible usefulness of XCIP analysis in cats. Application of the feline XCIP analysis to 3 feline mammary gland adenocarcinoma cell lines revealed distinctly skewed XCIPs in these cell lines, indicating their clonal origins. Twelve (80%) of the 15 primary tissue/cell samples obtained from cats with various neoplastic diseases showed skewed XCIPs. Moreover, bone marrow samples from 3 cats with myelodysplastic syndrome were also found to have skewed XCIPs. The polymerase chain reaction–based XCIP analysis developed in this study can provide information on cell clonality in female cats, potentially facilitating the differential diagnosis of various disorders in cats.
Differentiation between neoplastic and nonneoplastic proliferation of cells is sometimes difficult in cats when diagnosing diseases such as hypereosinophilic syndrome, 16 chronic myeloproliferative disorders, 22 and histiocytic disorders. 1 In such disorders, assessment of the clonal origin of the cells would enable a more accurate diagnosis.
Assessment of the clonality of cell populations has been recently developed for lymphoid tumors in cats by detection of antigen receptor gene rearrangements. 9,19,20,26,28 However, to date, there is no assay system for evaluating the clonality of cells other than lymphoid cells in cats. In human medicine, X-chromosome inactivation pattern (XCIP) analysis has been used for the assessment of the clonal origin of the cells in various organ systems in female individuals.
In mammalian females, either the paternal or maternal X chromosome generally undergoes random inactivation in each cell through the epigenetic changes during embryogenesis. This event is stably inherited by the daughter progeny of each cell, resulting in the theoretical expectation of a 1:1 inactivation ratio between the paternally and maternally derived X chromosomes in normal or reactive tissues. By contrast, most neoplastic tissues are composed of a population clonally expanded from a single cell, generally exhibiting a uniform XCIP and resulting in a significant deviation from the expected 1:1 ratio. Therefore, demonstration of a nonrandom XCIP (skewed XCIP) infers the clonal origin of cells. 5
Although XCIP analysis is applicable only in female patients, it has been widely used for assessment of clonality in several neoplastic and preneoplastic conditions in humans (eg, chronic myeloid leukemia, 7 acute myeloid leukemia, 29 myelodysplastic syndrome [MDS], 3 lymphoma, 21 and certain nonhematopoietic tumors). 12,21,31 A major advantage of the XCIP clonality test is its potential application to any neoplastic diseases, even where a tumor-specific marker has not been identified.
A widely used method in humans for determining clonality using the X-chromosome inactivation principle is the human androgen receptor (AR) gene assay (HUMARA). 5 Transcription of human AR gene on the inactive X chromosome is strictly inhibited by methylation of the CpG island containing its promoter. 11 A similar epigenetic event is known on a region containing HpaII sites near the polymorphic CAG tandem repeat (CAGr) in exon 1 of the AR gene. 2 The HpaII sites on the inactive X chromosome are methylated, whereas those on the active X chromosomes are unmethylated. Differences in the methylation status can be used to differentiate the inactive X chromosomes from the active one. A difference in the number of CAG repeats in the AR gene between 2 X chromosomes enables their distinction from each other.
Similar to the human AR gene, the feline AR gene is known to be located on the X chromosome and contains 2 clusters of CAGr in its exon 1 (GenBank accession number: AJ893545.1 and ACBE01496690.1). However, as yet, there has been no report of polymorphism of the CAGr in the feline AR gene. In this study, to develop a polymerase chain reaction (PCR)–based XCIP assay in cats, polymorphism of the CAGr in the feline AR gene was characterized by using nonneoplastic somatic DNA samples from male and female cats. By using the sequence data obtained in the present study, an XCIP assay to assess the clonal origin of cells in female cats was constructed, and its ability to detect cell clonality was examined in cultured tumor cell lines and primary tissue/cell samples obtained from cats with various diseases.
Materials and Methods
Nonneoplastic Somatic DNA Samples From Cats
Peripheral blood samples were obtained from 50 male and 103 female cats; they included 24 healthy blood donors and 129 patients referred to the Veterinary Medical Center of the University of Tokyo for the diagnosis and treatment of various diseases unrelated to hematopoietic neoplasms. Use of the blood samples from the patients for research purposes was approved by the owners in written statements.
The breeds of the 153 cats were mixed (115 cats), American Short Hair (11), Scottish Fold (6), Maine Coon (4), Persian (4), Abyssinian (3), British Short Hair (2), Himalayan (2), Norwegian Forest Cat (2), Somali (2), Ragdoll (1), and Russian Blue (1).
Feline Tumor Cell Lines
Three feline mammary gland adenocarcinoma cell lines—FMCp2, FMCm, and FYMp 24 —were used. FMCp2 and FYMp were established from primary mammary gland masses resected from 2 female cats with mammary gland adenocarcinoma. FMCm was established from a metastatic lymph node of the same cat from which FMCp2 was obtained. These cells were cultured in RPMI1640 (Invitrogen, Carlsbad, CA) supplemented with 10% fetal bovine serum (Biowest, Nuaille, France) and antibiotics (100 IU/mL penicillin and 0.1 mg/mL streptomycin) (Sigma-Aldrich, St. Louis, MO) at 37°C in a humidified atmosphere containing 5% CO2.
Primary Neoplastic Tissue/Cell Samples
Twenty-one neoplastic tissue or cell samples diagnosed by histopathology or cytology were used. The neoplastic tissue/cell samples were obtained from female cats referred to the Veterinary Medical Center of the University of Tokyo between 2009 and 2011. Diagnosis in these cats included lymphoma (7 cats), epithelial tumors (6; ceruminous adenocarcinoma [2], lung adenocarcinoma [2], mammary gland adenocarcinoma [1], and meningioma [1]), mesenchymal tumors (4; leiomyomas [2], fibrosarcoma [1], and leiomyosarcoma [1]), mast cell tumor (2), acute lymphoblastic leukemia (ALL) (1), and chronic lymphocytic leukemia (CLL) (1). Of the 21 cats, 5 cases with hematological malignancies (lymphoma [3], ALL [1], and CLL [1]) were diagnosed by cytology, and the remaining 16 cats were diagnosed by histopathology. The ages of the 21 cats with neoplastic diseases ranged from 2 to 20 years (median, 10 years). Eighteen cats were spayed and 3 cats were intact. The breeds were mixed (18 cats), American Shorthair (2), and Maine Coon (1).
Three female cats with MDS and 5 female cats with immune-mediated hematological disorders (immune-mediated neutropenia [3], immune-mediated hemolytic anemia [1], and pure red cell aplasia [1]) were diagnosed by the hematological examination, including bone marrow cytology. Specimens of the bone marrow aspirates were subjected to the feline XCIP analysis. These 8 cats were mixed-breed spayed female cats with ages ranging from 2 to 13 years. The 3 cats with MDS could be further subclassified into refractory cytopenia with multilineage dysplasia (RCMD) (1) and refractory anemia with excess blasts (RAEB) (2) according to the diagnosis and classification system of MDS in cats. 27
DNA Isolation
Genomic DNA samples were isolated from cells or tissues by using the DNeasy Blood and Tissue Mini Kit (QIAGEN, Hilden, Germany). Concentration of the extracted genomic DNA samples was calculated on the basis of absorbance at 260 nm measured with a spectrophotometer.
Sequencing of CAGr in the Feline AR Gene in Male Cats
Primer pairs were designed according to the sequence of the feline AR gene (GenBank accession number: AJ893545.1) by using Primer3Plus (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) to amplify a DNA fragment spanning both the first (GenBank accession number: AJ893545.1; nucleotide [nt] 91–165) and second CAGr (AJ893545.1; nt 475–531) in its exon 1 (AR-P1F: 5′CGCGAAGTGATCCAGAACC-3′, nt 18–37; AR-P1R: 5′-AACTGTCCTTGGAGGAGGTG-3′, nt 588–607). Each reaction mixture contained 50 ng genomic DNA, 2.0 mM MgCl2, 0.2 mM of each dNTP, 0.5 μM of each primer, 1× PCR buffer, and 0.5 units of ExTaq DNA polymerase (Takara Bio, Inc, Otsu, Japan) in a total volume of 20 μL. Cycle conditions consisted of an initial denaturation step at 95°C for 1 minute, followed by 30 cycles of denaturation at 94°C for 30 seconds, annealing at 60°C for 30 seconds, extension at 72°C for 1 minute, and a final extension at 72°C for 7 minutes. PCR products were purified using a DNA purification kit (SUPREC PCR; Takara Bio, Inc). Sequencing analyses of the PCR products were carried out with BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA) and the Genetic Analyzer 3130xl (Applied Biosystems) by using the primers AR-P1F and AR-P1R. The repeat number of the 2 CAGr in each cat was counted with Sequence Scanner Software v1.0 (Applied Biosystems).
Examination of the Heterozygosity of CAGr in the Feline AR Gene in Female Cats
Primer pairs were designed using Primer3Plus to amplify a region encompassing 3 HpaII digestion sites and the polymorphic CAGr, as shown in Fig. 1 (AR-P2F: 5′-CTGGAGAGAGGCTGTGTTCC-3′, nt 271–290; AR-P2R: 5′-CTTGTTCTCCCGCTGCTG-3′, nt 552–569). The PCR mixture consisted of 50 ng genomic DNA, 2.5 mM MgCl2, 0.2 mM of each dNTP, 1× PCR buffer, 0.2 μM FAM-labeled forward primer (AR-P2F), 0.2 μM reverse primer (AR-P2R), and 0.5 units of AmpliTaq Gold DNA polymerase (Applied Biosystems) in a total volume of 20 μL.
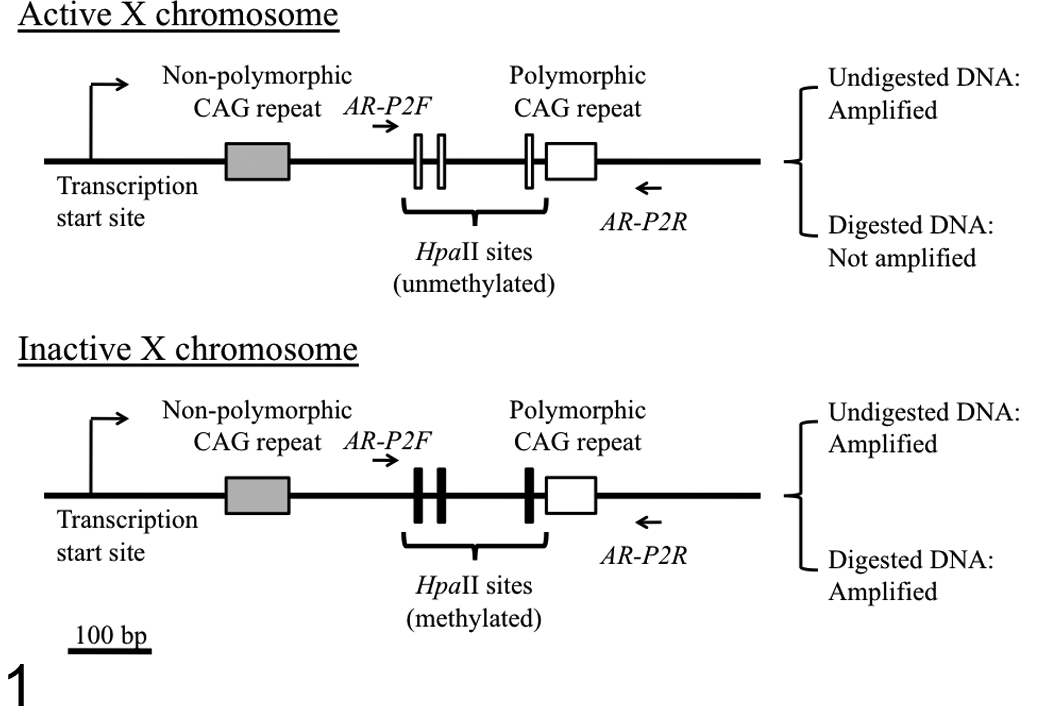
Schematic structure of the feline androgen receptor (AR) gene exon 1 and a polymerase chain reaction (PCR) strategy to distinguish between active and inactive X chromosomes by amplification of a polymorphic CAG tandem repeat (CAGr) within its exon 1. The schematic structure is based on the registered sequences of feline AR gene (GenBank accession numbers: AJ893545.1 and ACBE01496690.1). A DNA fragment can be amplified by PCR using AR-P2F and AR-P2R primers from both active (unmethylated) and inactive (methylated) X chromosomes. After HpaII digestion, the DNA fragment on the active (unmethylated) X chromosome cannot be amplified, whereas that on the inactive (methylated) X chromosome can be amplified because of failure of cleavage by HpaII.
Cycle conditions consisted of an initial denaturation step at 95°C for 5 minutes, followed by 28 cycles of denaturation at 95°C for 15 seconds, annealing at 64°C for 15 seconds, extension at 72°C for 20 seconds, and a final extension at 72°C for 7 minutes. One microliter of the 10-fold diluted PCR product was mixed with 8.5 μL formamide and 0.5 μL 600LIZ size standard (Applied Biosystems) in a 96-well plate. These samples were denatured at 95°C for 5 minutes and immediately placed on ice for 15 minutes. The length of the PCR product was determined by capillary electrophoresis using the Genetic Analyzer 3130xl and evaluated by GeneMapper software (Applied Biosystems).
XCIP Analysis
DNA samples (500 ng) were incubated with 5 units of a methylation-sensitive endonuclease, HpaII (New England Biolabs, Inc, Ipswich, MA), in 1× buffer 1 in a total volume of 50 μL. Undigested controls of each sample were incubated in 1× buffer 1 (total volume, 50 μL) without HpaII. Both the digested and undigested samples were incubated for 16 hours at 37°C followed by inactivation for 20 minutes at 65°C.
A 50-ng aliquot of the digested and undigested DNA samples was subjected to subsequent PCR amplification with the primers AR-P2F and AR-P2R.
As shown in HUMARA, because HpaII is a methylation-sensitive endonuclease and its cleavage sites near the polymorphic CAGr in the AR gene on the inactive feline X chromosome DNA are considered methylated, the HpaII sites would not be cleaved by this enzyme. Therefore, the DNA with methylation of the sites can be conceivably amplified with the AR-P2F and AR-P2R primers (spanning the HpaII sites) even after HpaII digestion. In contrast, the HpaII sites on the active feline X chromosome are considered unmethylated; therefore, HpaII digestion of the DNA of the active X chromosome leads to failure of PCR amplification because of the cleavage of the HpaII sites. As a control for HpaII digestion, the same amount of DNA from a male nonneoplastic peripheral blood sample was used because male somatic cells contain only 1 active X chromosome in which the HpaII sites are not methylated.
Results of the XCIP analysis were evaluated after confirming the absence of gene amplification from the completely digested male somatic DNA control. The degree of skewness (imbalanced inactivation) was determined by comparing the area under the curve (AUC) for each allele of the digested and undigested samples using the results of the capillary electrophoretogram. The corrected inactive allele ratio (CIAR) was defined as the allele ratio of the digested sample divided by the allele ratio of the undigested sample, to avoid the influence of the potential preferential amplification of either allele.
Stutter peaks (shadow bands), generated by slippage synthesis of short tandem repeats,
25
are observed as minor peaks 3 bp and 6 bp shorter than the real peaks. When the CAGr in one allele is longer than that in another allele by 3 bp (1 CAG repeat) or 6 bp (2 CAG repeats), the longer allele (Fig. 2C, allele 2) affects the measured AUC of the shorter allele (Fig. 2C, allele 1). To avoid the influence of the shadow bands, the percentage of shadow bands for each repeat length was calculated from the data obtained from DNA samples of male cats. When the size of allele 1 was shorter than that of allele 2 by 1- or 2-repeat lengths, 27% or 7%, respectively, of the AUC of allele 2 was subtracted from that of allele 1. Thus, the CIAR was calculated as follows:
Representative capillary electrophoretograms after polymerase chain reaction (PCR) to amplify the polymorphic CAG tandem repeat in the feline androgen receptor (AR) gene. (A) A nonneoplastic blood DNA sample from a female cat homozygous for the repeat number of CAG trinucleotides (16 CAG repeats). (B) A nonneoplastic blood DNA sample from a male cat (20 CAG repeats). (C, D) Nonneoplastic blood DNA samples from female cats heterozygous for the repeat number of CAG trinucleotides (C: 18 and 19 CAG repeats, D: 18 and 22 CAG repeats). Two major peaks are observed in heterozygous female cats (C, D), whereas only 1 major peak is observed in a homozygous female cat (A) and a male cat (B). Stutter peaks (shadow bands, indicated as S) are observed as minor peaks shorter than the major peaks by 3 or 6 bp.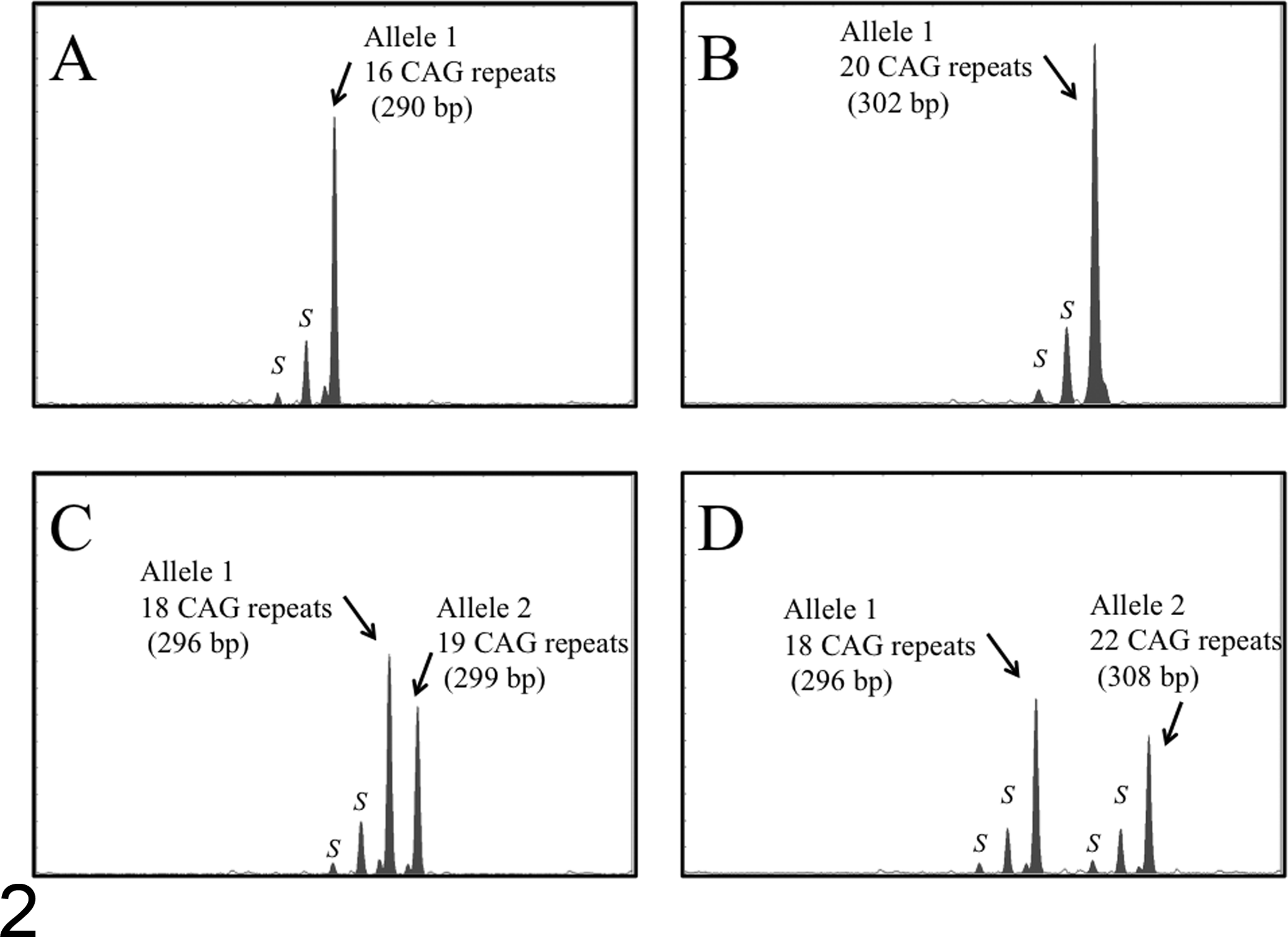
As proposed in previous reports on XCIP analysis in humans, 14,18 the sample population was judged to be clonal when the CIAR >3.0. For ease of comparison, where CIAR <1, the corrected ratio was calculated as the inverse value of the CIAR.
Statistical Analysis
Pearson’s χ 2 test was used for comparing the allele frequency between male and female sample populations. The significance level was set at P < .05. Statistical testing was performed using JMP version 8.0.1 (SAS Institute, Cary, NC).
Results
Sequencing of the First and Second CAGr in Exon 1 of the Feline AR Gene
The number of CAG trinucleotide repeats in 2 CAGr in exon 1 of the feline AR gene was examined in 50 male cats. There was no polymorphism in the first CAGr; however, the number of CAG trinucleotide repeats in the second CAGr varied between 16 and 20. Of the 50 male cats, 9 (18%), 5 (10%), 12 (24%), 20 (40%), and 4 (8%) cats had 16, 17, 18, 19, and 20 repeats of CAG, respectively.
Heterozygosity of the Second CAGr of the Feline AR Gene in Female Cats
PCR amplification of the second CAGr followed by capillary electrophoresis was performed in 103 female cats. In homozygotes for the second CAGr, a single major peak was observed (Fig. 2A), similar to the result for male cats (Fig. 2B). Heterozygotes for the second CAGr were identified by the occurrence of 2 major peaks of different lengths after PCR amplification with the AR-P2F and AR-P2R primers (Fig. 2C, D). In all of the samples, stutter peaks (shadow bands indicated as S) were observed as minor peaks, shorter than the major peaks by 3 or 6 bp (Fig. 2A–D).
Of the 103 female cats, 70 (68%) were found to be heterozygous for the second CAGr. Heterozygosity of the second CAGr was shown in 57 of 79 (72%) mixed-breed cats and 13 of 24 (54%) purebred cats. The CAG repeat numbers in the second CAGr were 15 (2 alleles; 1%), 16 (37 alleles; 18%), 17 (30 alleles; 15%), 18 (37 alleles; 18%), 19 (79 alleles; 38%), 20 (18 alleles; 9%), and 22 (3 alleles; 1%). Allele frequency with respect to the number of CAG repeat was compared between females and males, and there was no significant difference between the 2 sample populations in this study.
Accuracy, Reproducibility, and Specificity of XCIP Analysis Using the Feline AR Gene
Before analyzing the samples from cats with neoplastic diseases, the performance of the feline XCIP analysis constructed in this study was evaluated. First, to confirm that the PCR amplification ratio corresponds to the amount of template DNA, DNA-mixing experiments were performed. Two DNA samples from male cats containing 16 or 19 CAG trinucleotides in the second CAGr were mixed in various proportions as follows: 1:10 (the ratio of the amount of DNA containing 16 CAG oligonucleotides to the amount of DNA containing 19 CAG oligonucleotides), 1:5, 1:3, 1:1, 3:1, 5:1, and 10:1. PCR was performed in triplicate using the same conditions as the feline XCIP analysis without digestion with HpaII, and the allele ratios were calculated. To compensate for amplification bias toward the shorter allele, all allele ratios were divided by the allele ratio obtained in a sample of 1:1 DNA mixture. The allele ratios calculated from the capillary electrophoretic analysis were 1:10.0 (for the 1:10 mixture), 1:5.23 (1:5), 1:2.94 (1:3), 1:1.00 (1:1), 3.08:1 (3:1), 5.31:1 (5:1), and 12.3:1 (10:1).
Second, blood samples from healthy female cats heterozygous for the second CAGr were used to determine the reproducibility of the analysis. The assay was performed in triplicate on the same DNA samples of 10 healthy female cats, resulting in a coefficient of variation for CIAR of 13%.
Finally, specificity of the feline XCIP analysis was examined using DNA samples of peripheral blood from 10 healthy female cats. CIAR values obtained from these 10 samples ranged from 1.01 to 2.62 (median: 1.18) (Fig. 3B and Fig. 4), less than the threshold value 3.0 (indicative of clonal origin) reported in XCIP analysis in humans. 14,18
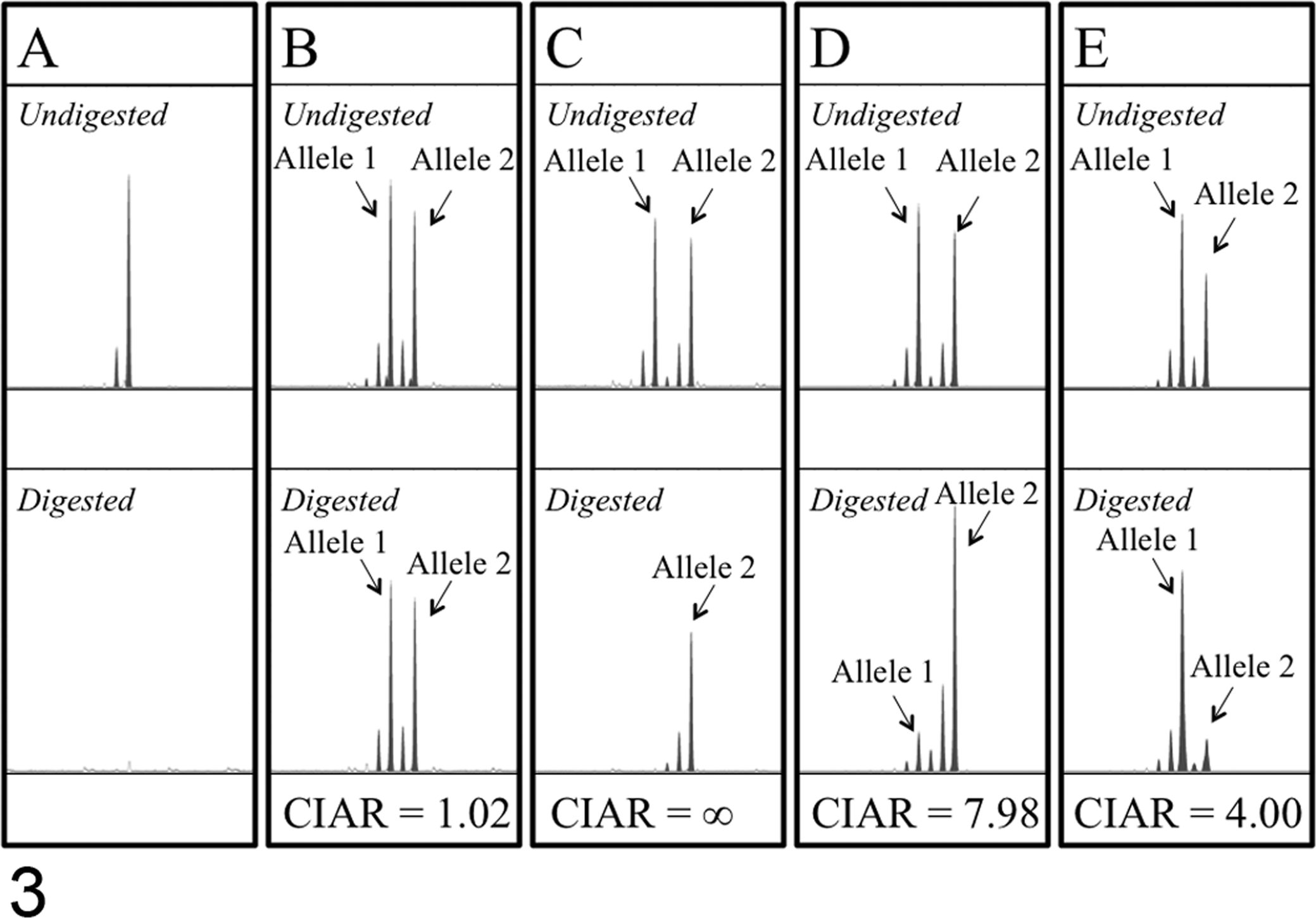
Representative results of X-chromosome inactivation pattern (XCIP) analysis of the androgen receptor (AR) gene in nonneoplastic and neoplastic cell samples obtained from female cats. (A) A nonneoplastic peripheral blood cell sample from a male cat. A peak corresponding to the polymerase chain reaction (PCR)–amplified AR gene is observed in an undigested sample but not in the HpaII-digested sample, indicating complete digestion with HpaII. (B) A nonneoplastic peripheral blood cell sample from a female cat heterozygous for the CAG repeat number of the second CAGr in the AR gene. Because the cells are polyclonal, the AR gene on both alleles (allele 1 and allele 2) was almost equally amplified in the undigested and HpaII-digested samples (corrected inactivated allele ratio [CIAR]: 1.02). (C) A feline mammary gland adenocarcinoma cell line, FMCp2. One of 2 X chromosomes (allele 1) was not amplified in the HpaII-digested sample, whereas 2 X chromosomes were almost equally amplified in the undigested sample. The CIAR of FMCp2 was calculated as ∞, indicating clonal origin of the cells. (D, E) Primary neoplastic tissue samples (D: case 6, ceruminous adenocarcinoma; E: case 4, lung adenocarcinoma). The AR gene on both alleles (allele 1 and allele 2) was similarly amplified in the undigested samples; however, the amount of PCR product from one allele (allele 1 in D, allele 2 in E) was markedly reduced in the HpaII-digested samples. The CIARs of these samples were 7.98 (D) and 4.00 (E), exceeding the threshold value 3.0, indicating clonal cell proliferation in these tissue samples. Stutter peaks are observed as peaks 3 or 6 bp shorter than the major peaks in all of the electrophoretograms.
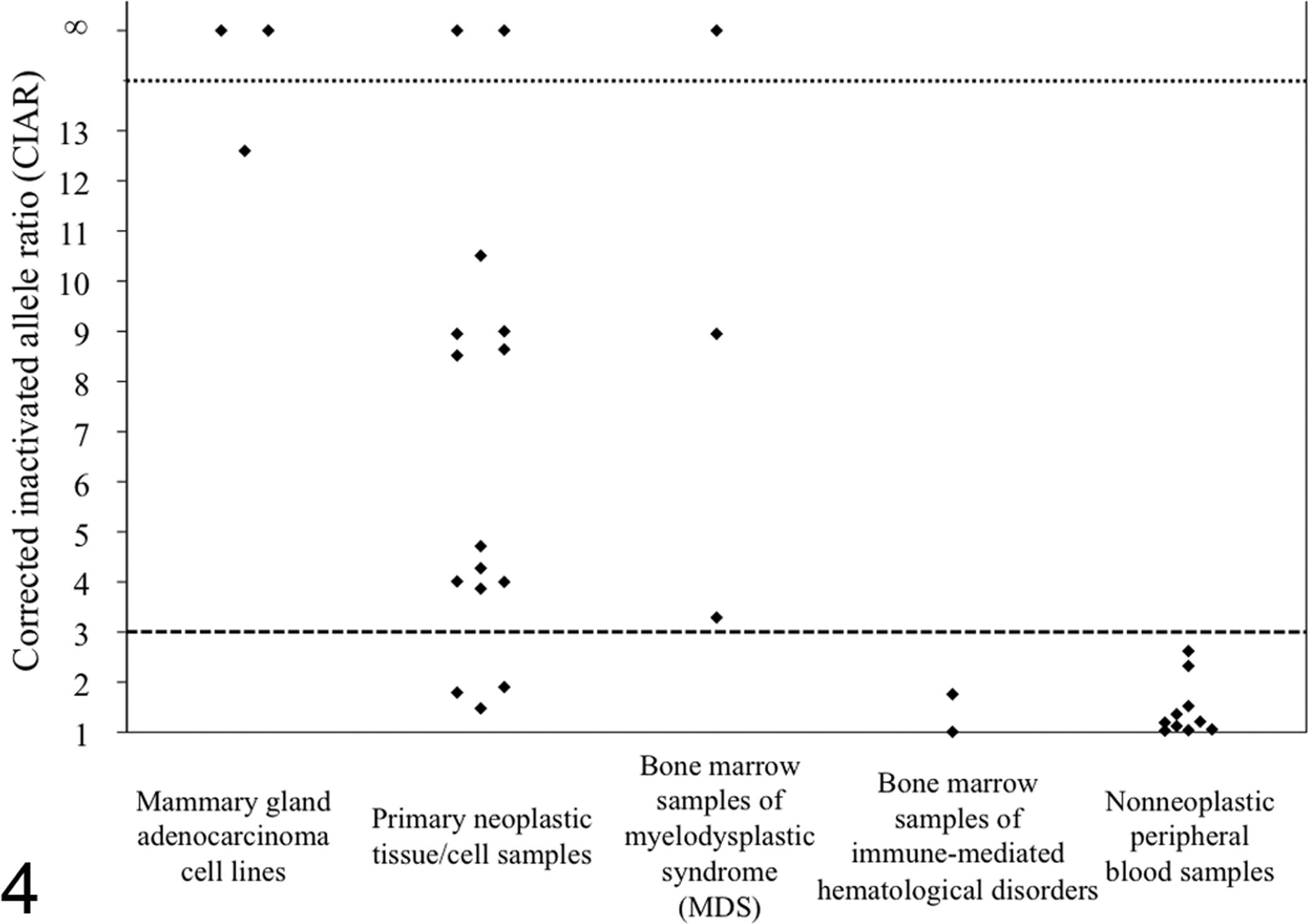
Corrected inactivated allele ratios (CIARs) obtained from the X-chromosome inactivation pattern (XCIP) analysis in 3 mammary gland adenocarcinoma cell lines, 15 primary neoplastic tissue/cell samples, 3 bone marrow samples of myelodysplastic syndrome, 2 bone marrow samples of immune-mediated hematological disorders, and 10 nonneoplastic peripheral blood samples. The threshold value of CIAR was set as 3.0.
Feline XCIP Analysis in Feline Mammary Gland Adenocarcinoma Cell Lines
DNA samples from 3 feline mammary gland adenocarcinoma cell lines (FMCp2, FMCm, and FYMp) were analyzed by XCIP analysis using the feline AR gene. FMCp2 and FMCm, which are derived from the same cat, showed identical results in XCIP analysis (Fig. 3C). CIARs of FMCp2 and FMCm samples were calculated as ∞ because HpaII digestion completely inhibited PCR amplification of the active (unmethylated) allele with a shorter CAGr, causing the AUC of the shorter allele to approach 0. FYMp also showed a distinctly skewed XCIP (CIAR: 12.6).
Feline XCIP Analysis in Clinical Samples
Prior to the feline XCIP analysis, 21 primary neoplastic tissue/cell samples from cats with neoplastic disorders and 8 bone marrow aspirates from cats with MDS or immune-mediated diseases were examined for the heterozygosity of the second CAGr in AR gene. Of the 21 primary tissue/cell DNA samples from cats with neoplastic disorders, 15 samples (71%; lymphoma [5], leiomyoma [2], ceruminous adenocarcinoma [1], lung adenocarcinoma [1], mammary gland adenocarcinoma [1], meningioma [1], leiomyosarcoma [1], fibrosarcoma [1], ALL [1], and CLL [1]) were shown to be heterozygous for the second CAGr, being appropriate for feline XCIP analysis. Similarly, of the 8 bone marrow DNA samples, 5 samples (63%; MDS [3], immune-mediated hemolytic anemia [1], and pure red cell aplasia [1]) were shown to be heterozygous for the second CAGr and could be evaluated by feline XCIP analysis.
Of the 15 samples of neoplastic disorders, CIARs in 12 samples exceeded 3.0 (3.87–∞, median: 8.58), indicating skewed XCIPs (Fig. 3D, E); however, those of the remaining 3 samples (lymphoma [2] and mammary gland adenocarcinoma [1]) were <3.0 (1.48, 1.79, and 1.90) (Fig. 4).
Bicytopenia (case No. 16: neutropenia and thrombocytopenia and case No. 17: anemia and thrombocytopenia) or pancytopenia (case No. 18) was a characteristic feature in the peripheral blood of the 3 cats with MDS. Bone marrow specimens of these cats were characterized by varying degrees of dysplastic features in myeloid, megakaryocytic, and erythrocytic cell lineages, including neutrophils with a ring-shape nucleus, small megakaryocytes with a single nucleus, and megaloblastoid change of erythroblastic cells (Figs. 5 and 6, case No. 1). Cats with immune-mediated hematological disorders were devoid of such dysplastic changes. Of the 5 bone marrow samples, 3 bone marrow samples from cats with MDS showed skewed XCIPs (CIARs: 3.29, 8.95, and ∞) (Fig. 7, case No. 1), whereas 2 samples from cats with immune-mediated hematological disorders did not show skewed XCIPs (CIARs: 1.01 and 1.76).
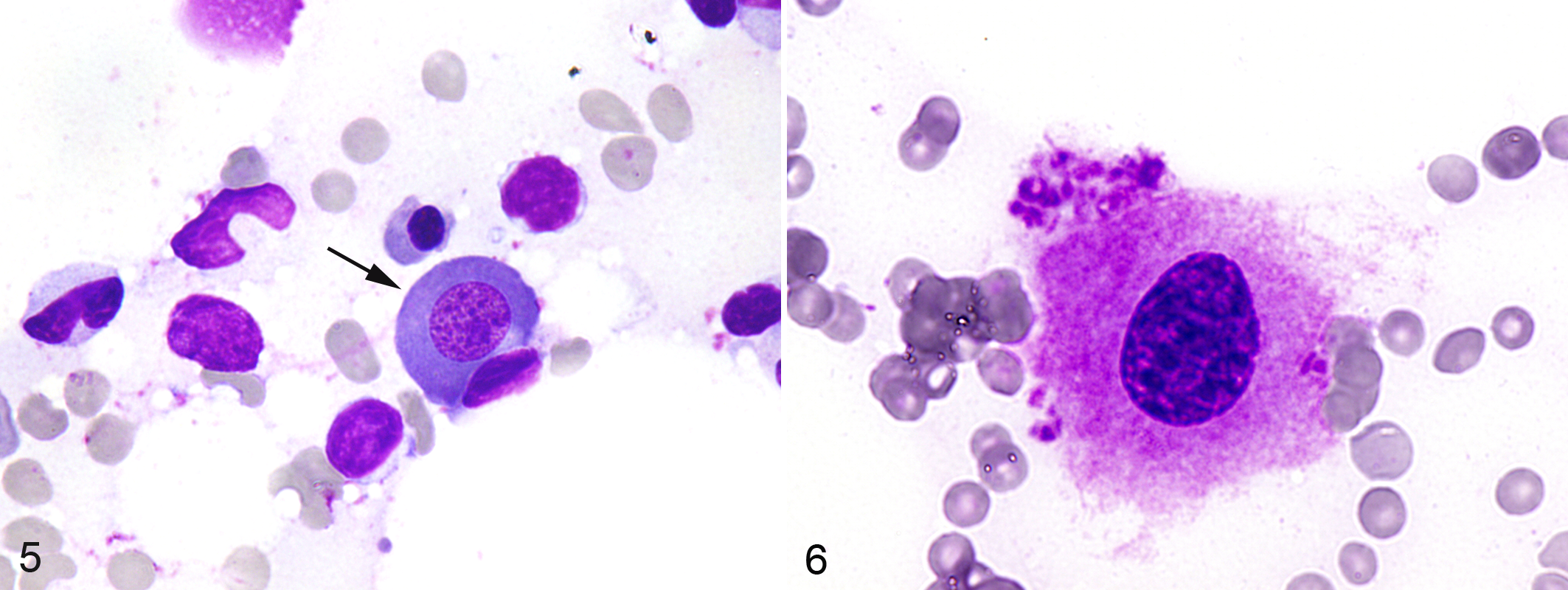
Bone marrow, cat, case No. 1, cytology of bone marrow aspirates from a cat with myelodysplastic syndrome (MDS)–refractory anemia with excess blasts. Megaloblastoid change of a prorubricyte (arrow). Wright’s Giemsa stain.
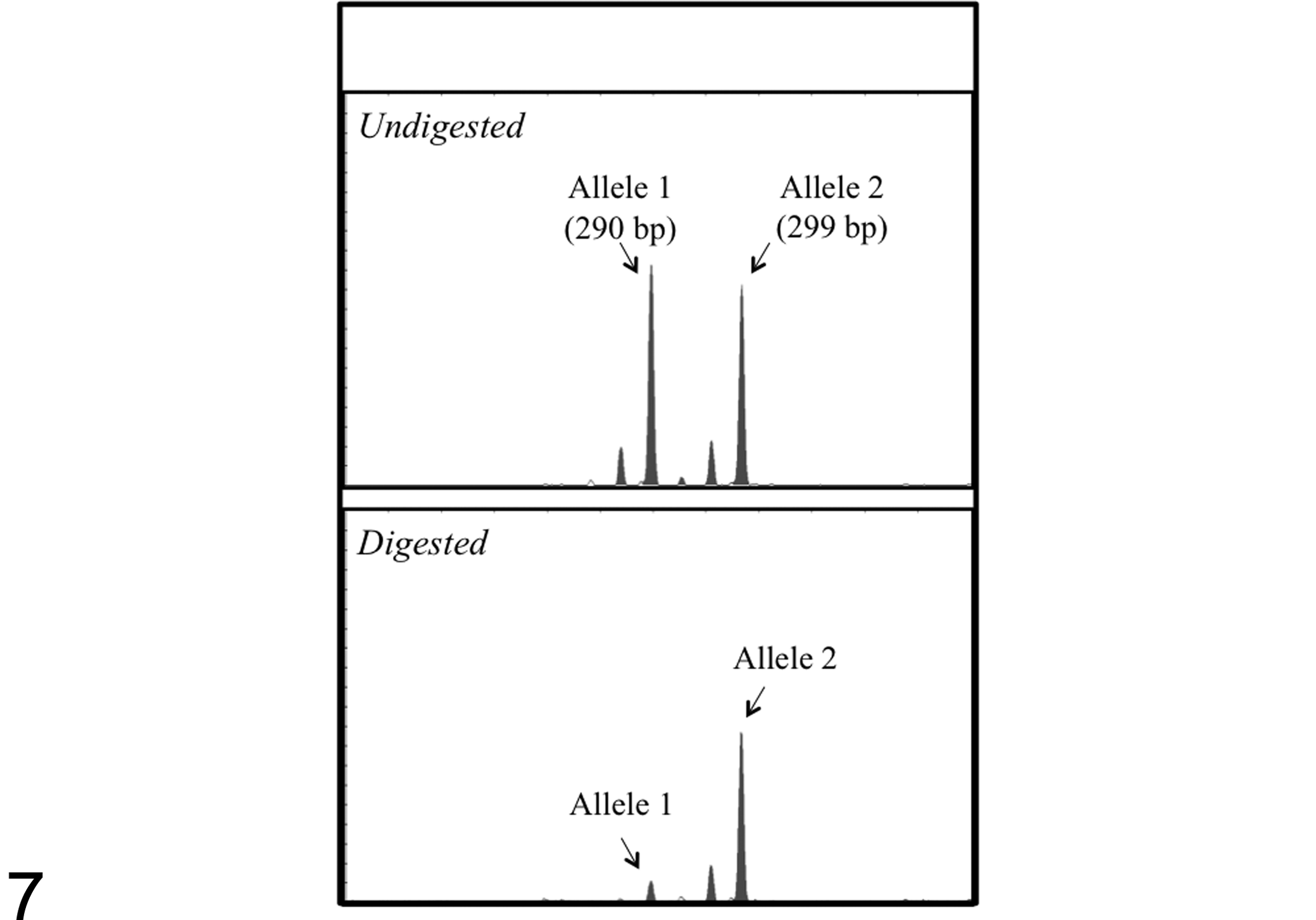
X-chromosome inactivation pattern analysis of bone marrow aspirates from a cat with myelodysplastic syndrome (MDS)–refractory anemia with excess blasts (case No. 1). The androgen receptor gene on both alleles (allele 1 and allele 2) was similarly amplified in the undigested samples; however, the amount of polymerase chain reaction product from allele 1 was markedly reduced in the HpaII-digested sample (corrected inactive allele ratio: 8.95).
Discussion
To construct the feline XCIP analysis, we first investigated the repeat-number polymorphism of the 2 CAGr in exon 1 of the feline AR gene and revealed that 70 of 103 (68%) female cats were heterozygous for the polymorphic second CAGr, which contains 15 to 22 CAG trinucleotides. It has been reported that the AR genes of other mammals, such as humans and dogs, contain CAGr. 6,13,23 One of the CAGr in the human AR gene is polymorphic and contains 11 to 31 repeats. 6 Canine AR gene contains 2 polymorphic CAGr (the first CAGr containing 10–12 repeats and the second CAGr containing 10–13 repeats). 13,23 Heterozygosity for the CAGr in the AR gene was shown in 90% of human female individuals 2,8 and 40% of female dogs. 23 In the present study, 68% of female cats were demonstrated to be heterozygous. Therefore, XCIP analysis for the AR gene would be applicable in approximately two-thirds of female cat patients in clinical settings.
The performance of the feline XCIP analysis constructed in this study was evaluated in terms of accuracy, reproducibility, and specificity. The allele ratio after PCR corresponded well with the amount of DNA template used for PCR. In addition, the coefficient of variation was shown to be relatively low. None of the nonneoplastic blood cell samples examined from female cats showed a skewed XCIP, and all 3 of the mammary gland adenocarcinoma cell lines showed distinctly skewed XCIPs. From these results, it is conceivable that the feline XCIP analysis using the AR gene developed in this study can be used as an assay to examine the clonality of cells with reasonable accuracy.
In primary cell/tissue samples from cats with neoplastic diseases, 12 of 15 neoplastic samples (80%) showed skewed XCIP, indicating clonal cell proliferation in these samples. However, 3 of 15 neoplastic samples (20%) did not show skewed XCIPs. There are several possible explanations for these results. First, success in the detection of unbalanced XCIPs largely depends on the proportion of tumor cells in the samples used for the assay. The tissue or cell samples usually contain certain numbers of nonneoplastic cells of the original organs, inflammatory cells, fibroblasts, and blood cells. When the percentage of the number of nonneoplastic cells in the neoplastic tissue/cell samples is high (theoretically more than 50%), false-negative results would be obtained. Tumor cell isolation methods such as microdissection and flow cytometric cell sorting may help to improve the sensitivity. Second, because the present assay is based on differences in the methylation status between active and inactive X chromosomes, the CIAR would not reflect the actual ratio of the inactivation pattern of the 2 X chromosomes when abnormal methylation occurs in the AR locus. Abnormal hypermethylation of the AR locus is reported in human lymphoma patients. 17
MDS is a group of diseases characterized by peripheral blood cytopenia due to dysplasia of hematopoietic cells. Occurrence of MDS in cats is associated with feline leukemia virus infection in cats. 10 Although the pathogenesis of MDS has not been elucidated, it is thought to be caused by clonal expansion of bone marrow cells from aberrant pluripotent hematopoietic stem cells in humans. Progression to acute myeloid leukemia from MDS has been reported in both humans 15 and cats. 10 Although the number of feline MDS patients examined in this study was small, distinctly skewed XCIPs were demonstrated in bone marrow obtained from cats hematologically diagnosed with MDS, indicating the clonal nature of bone marrow cells in cats with MDS. The present study supports the hypothesis that MDS is a preleukemic state resulting from the clonal expansion of abnormal hematopoietic stem cells in cats as well as humans.
In humans, although application of XCIP analysis is restricted to females, it has several advantages over other clonality assessment procedures such as cytogenetic analysis, mutation analysis of specific genes, or viral integration analysis. Most important, XCIP analysis can be used in a variety of diseases, even where no disease-specific marker has been identified. By using the XCIP analysis, Langerhans cell histiocytosis and hypereosinophilic syndrome in humans were shown to be disorders resulting from clonal proliferation of the respective cell lineages. 4,30 As feline counterparts of these diseases, feline progressive histiocytosis 1 and hypereosinophilic syndrome 16 have been identified; however, the clonal origin of these diseases has not been revealed. Feline XCIP analysis could be a useful tool for evaluating clonality of these disorders as well as other conditions where clonal origins are suspected, thereby allowing precise understanding of their pathogenesis and accurate diagnosis.
In conclusion, polymorphic CAGr in the feline AR gene were identified and a simple PCR-based feline XCIP clonality assay was constructed. This assay can be used as a practical tool for the assessment of cell clonality in various feline diseases.
Footnotes
Acknowledgements
The authors thank Drs Nobuo Sasaki and Takayuki Nakagawa for providing the feline mammary gland adenocarcinoma cell lines, Dr Naoya Yuhki for providing information regarding the feline genome, and Drs Kunio Shiota and Jun Ohgane for their helpful discussions.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship and/or publication of this article: This study was supported by the Japan Society for the Promotion of Science, KAKENHI 23381082.