Abstract
Several markers of malignancy have been proposed for canine mammary tumors on the mRNA and protein levels. However, their association with tumor malignancy applies only for mean values of large groups of tumors, but no single marker identified to date can be used to reliably predict malignancy for individual tumors. A quantitative real-time reverse transcription polymerase chain reaction array was established to quantify the expression levels of 49 genes relevant to carcinogenesis in laser-microdissected tumor cells of 10 benign and 13 metastatic canine mammary tumors. Analysis of variance and discriminant analysis were used to identify relevant gene expression patterns that differentiate adenomas from metastatic carcinomas and their lymph node metastases. Seventeen genes with significant (P < .05) differences in gene expression levels between benign and malignant tumors were identified—including ERBB1, SLIT2, progesterone receptor, MIG6, SATB1, and SMAD6—but correct classification of each tumor as benign or malignant was impossible on the basis of any of these genes alone. However, the combined expression patterns of BMP2, LTBP4, and DERL1 (Derlin-1) correctly classified each individual tumor as benign or malignant. This pilot study identified a complex mRNA expression pattern of 3 genes that was able to identify malignancy in laser-microdissected tumor cells for each individual tumor, instead of group means as used in previous studies.
Canine mammary tumors are commonly diagnosed on the basis of their histologic features according to the World Health Organization tumor classification. 18 These features are still the most robust indicators of survival and the gold standard against which new prognostic indices are compared. Although several proteins have been identified to be differentially expressed between large groups of benign and malignant tumors, the immunohistochemical detection of biomarkers has not yet become a diagnostic routine in the diagnosis of canine mammary tumors. Previous studies on the supplementation of the histopathologic prognosis of the clinical outcome of canine mammary tumors focused on the establishment of immunohistochemical markers. Several attempts have been reported on the use of differential expression of single genes for an improved prognosis of canine mammary tumors, including p53, ERBB1, and BRCA1. 21,22,29 In these studies, a significant correlation was established between protein expression and malignancy when group means of benign and malignant tumors were compared. However, none of these markers was able to correctly classify each individual tumor as benign or malignant. In contrast, the clinical outcome of several human cancers is highly correlated with specific complex mRNA expression patterns that yield additional clinically relevant information that cannot be obtained by conventional histopathology. 6,30 –32,35 Recent studies on global gene expression profiles in canine mammary tumors indicate that similar complex gene expression profiles may exist in canine mammary tumors. 26
The goal of this study was to identify a set of molecular marker genes for the correct classification of each individual malignant tumor. Gene expression levels of 49 relevant genes were quantified and statistically analyzed from laser-microdissected nonneoplastic benign and malignant canine mammary epithelial cells. Surprisingly, a set of 3 genes was found sufficient to identify malignancy in each tumor analyzed.
Methods
Dogs and Tissue Processing
Thirteen simple carcinomas with lymph node metastases at the time of tumor resection and 10 simple adenomas were selected from 130 snap-frozen canine mammary tumors collected over a period of 4 years. Nonneoplastic mammary gland was obtained from the mammary complex next to the complex with the tumors. All subjects had no radiographically detectable pulmonary metastases. Selection criteria for adenomas included absence of lymph node metastases and a postoperative tumor-free interval of 2 or more years. Tissue specimens of tumors, nonneoplastic glands, and lymph node metastases were sliced at 5-mm thickness; every other slice was immersion fixed in neutral buffered 4% formaldehyde or snap frozen in liquid nitrogen within 15 minutes after resection and stored at –80°C until further use.
Representative samples from directly adjacent tumor tissue were formaldehyde fixed and routinely embedded in paraffin. Sections of 2-μm thickness were mounted on adhesive glass slides and stained with HE. Two board-certified pathologists independently evaluated the tumors histologically, following the criteria of the World Health Organization classification of canine mammary tumors. 18
Laser-Capture Microdissection and Quantitative Real-Time Polymerase Chain Reaction
Pure tumor cells or nonneoplastic mammary epithelial cells were subject to laser-capture microdissection without contaminating stromal or inflammatory cells. Five consecutive sections of 6- to 8-μm thickness from the frozen tissue samples were mounted on glass slides covered with a polyethylene naphthalate membrane (PALM Microlaser Technologies, Bernried, Germany) for laser microdissection. Sections were fixed for 2 minutes in 95% ethanol at –20°C and stained with HE solubilized in diethylpyrocarbonate-treated water. Sections were subsequently dehydrated in ascending graded ethanol and air-dried at room temperature. At least 25 × 106 μm2 were excised, consisting of nonneoplastic epithelial cells, adenomas, carcinomas, or lymph node metastases, followed by laser pressure catapulting into caps of 0.5-ml reaction tubes containing 30 μl of lysis buffer (NucleoSpin RNA XS, Macherey-Nagel, Düren, Germany). Total RNA was extracted and purified using a commercial system (NucleoSpin RNA XS) and reverse transcribed using the iScript cDNA synthesis system (Bio-Rad, Munich, Germany).
Samples were examined for RNA integrity in an Agilent Bioanalyzer 2100 (Agilent, Palo Alto, Calif), as previously described. 28 Samples with an RNA integrity number below 5 were excluded from the analysis.
Table 1 lists the primer sequences for the 49 target genes and the 3 housekeeper genes: HPRT (hypoxanthine–phosphoribosyl transferase), A5B (ATP-synthase subunit 5B), and RP32 (ribosomal protein L32). Target genes were selected on the basis of their confirmed pathogenetic or diagnostic relevance in human breast cancer or canine mammary tumors. 1,5,8 –10,12,17,20,22,23 Real-time reverse-transcription quantitative polymerase chain reaction (RT-qPCR) and data analyses were performed using the MX 3000P Quantitative PCR System and MX Pro software (Stratagene, La Jolla, Calif). Reactions were run as triplicates of each cDNA sample. The RT-qPCR efficiency was between 95% and 105% in all assays and all yielded products of the expected sequences. Cycling conditions were 10 minutes at 95°C, followed by 40 cycles of 30 seconds at 95°C, 1 minute at 58°C, and 30 seconds at 72°C. The cDNA of all samples were amplified on the same plate for each primer pair to ensure equal amplification conditions. Specificity of amplification products was confirmed by melting curve analyses.
Sequences of Primers Used for Quantitative Polymerase Chain Reaction
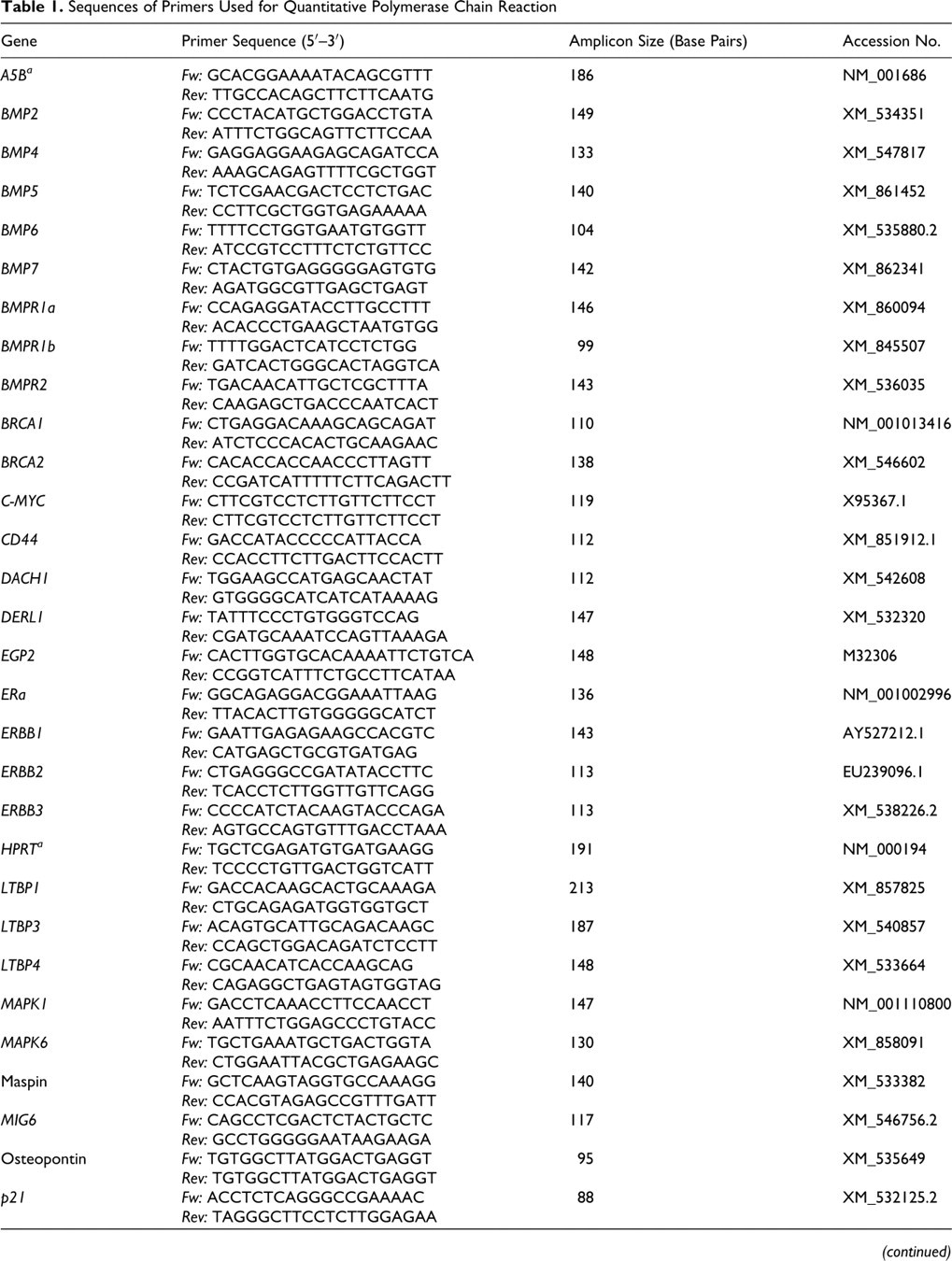
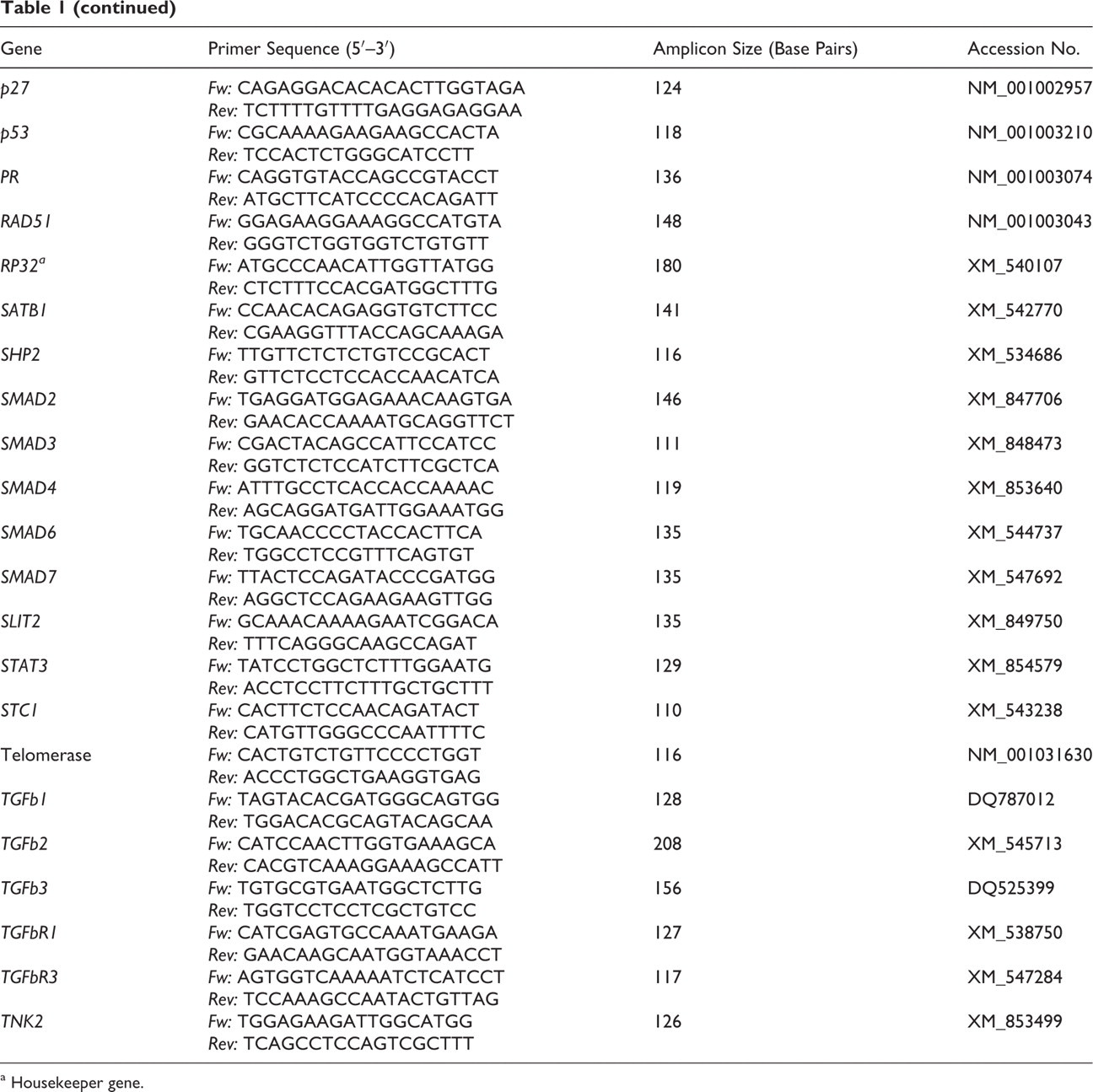
a Housekeeper gene.
Quantification of Target Gene Expression
Relative expression of the target genes was calculated with the comparative ΔΔCT method, with multiple housekeepers as previously described. 16,33 The housekeeper genes were selected from a panel of reference genes according to the GeNorm algorithm. 4 Data are presented as fold change in gene expression levels in the sample of interest normalized to the housekeepers and relative to the sample control. First, the normalization factor was calculated with the geometric mean of 2ΔCT of the 3 reference genes and the gene of interest (GOI) in the sample of interest and sample control. Finally, the fold change (2ΔΔCT) of GOI expression was calculated with the ΔCTGOI multiplied by the normalization factor. Specifically, gene expression levels of all adenomas, metastatically active carcinomas, and lymph node metastases were normalized to reference gene expression and compared with nonneoplastic gland epithelium from the same dog.
Statistical Analysis
Analysis of variance (ANOVA) and discriminant analysis were used to identify relevant genes that discriminate adenomas, carcinomas, and lymph node metastases. Fold change values (2ΔΔCT) were logarithmized to the base 2 (log2). Shapiro–Wilk test was used to confirm normal distribution, and only genes with normally distributed gene expression levels in all tissue groups were included in statistical analyses. Genes with significant differential mRNA expression (P ≤ .05) between at least 2 tissue types were preselected by ANOVA for discriminant analysis. Most relevant genes for the correct classification of tumors according to their histologic diagnosis were selected out of the ANOVA preselected genes by discriminant analysis. To this end, the gene with the least relevance to the classification of the tumors in the discriminant analysis with the ANOVA preselected genes was excluded, as displayed by the smallest structure matrix coefficient in both discriminant functions. A consecutive discriminant analysis with the remaining ANOVA-preselected genes was then conducted. This procedure was repeated until carcinomas and metastases were no longer classified correctly to 100% from adenomas. The remaining genes were then accepted as most relevant to discriminate defined adenomas, carcinomas, and lymph node metastases.
Results
Expression Levels of the Bone Morphogenetic Protein Genes
Gene expression levels of 5 members of the bone morphogenetic protein (BMP) family compared adenomas, carcinomas, and lymph node metastases (Table 2 ). Only BMP2 was significantly differentially expressed in adenomas, carcinomas, and metastases. Carcinomas and lymph node metastases had a markedly increased mean BMP2 expression of 21.6-fold and 18.4-fold, respectively. In contrast, adenomas had a reduced mean expression of 0.6-fold when compared to the normal gland of the same dog. No significant expression differences were observed for BMP4, BMP5, and BMP6. BMP7 receptor expressions BMPR1a, BMPR1b, and BMPR2 varied markedly in the different tissue samples, and no significant expression differences were found. Significant differences in mean mRNA expression levels were found for SMA/SMADs (mothers against decapentaplegic), which is the signal transducer and transcription factor associated with the BMP binding to its receptor. Most notably, SMAD6 was significantly overexpressed in carcinomas and lymph node metastases when compared to adenomas.
Fold Change in Mean Gene Expression Levels in Adenomas, Carcinomas, and Lymph Node Metastases for 49 Preselected Carcinogenesis-Relevant Genes a
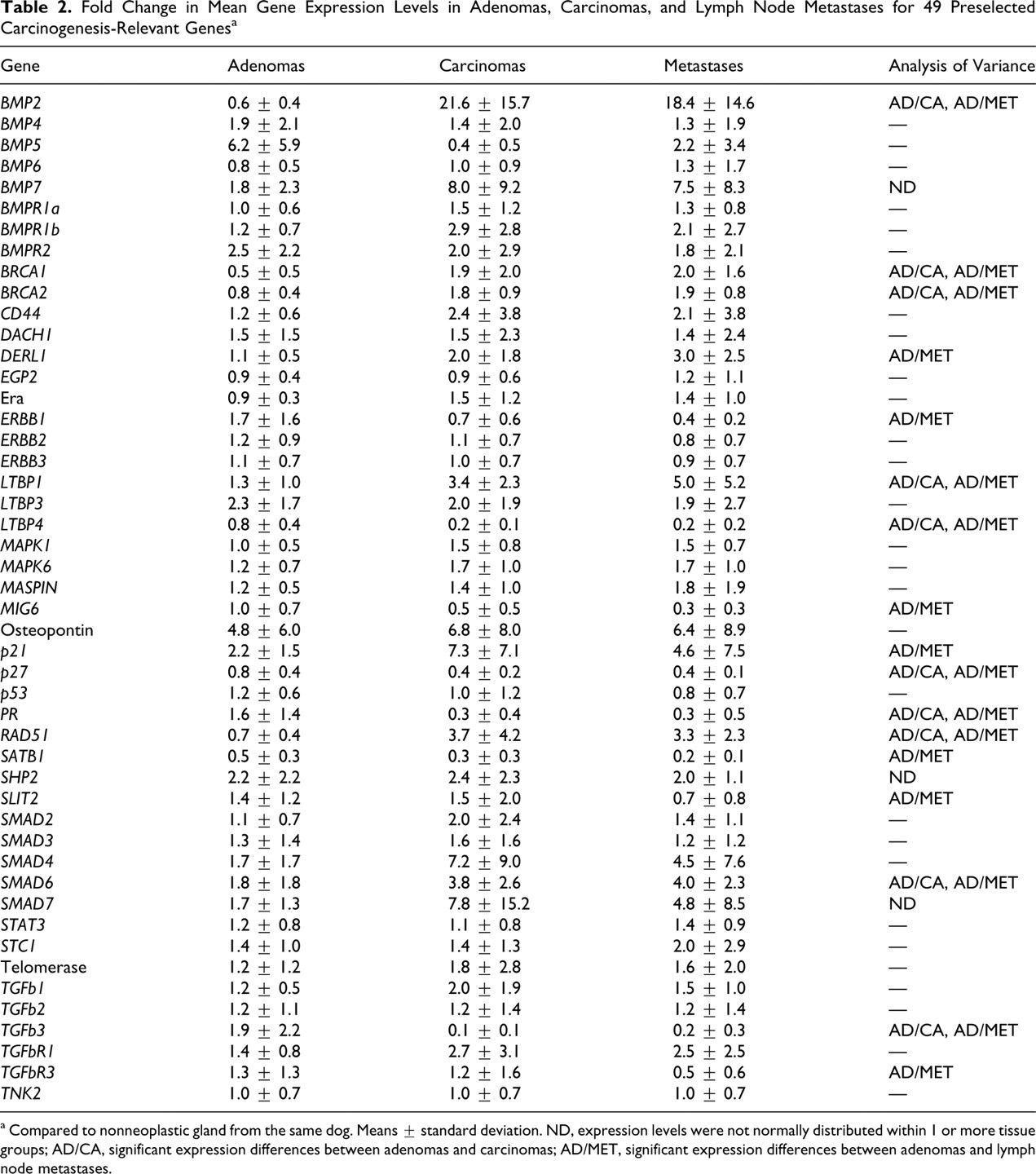
a Compared to nonneoplastic gland from the same dog. Means ± standard deviation. ND, expression levels were not normally distributed within 1 or more tissue groups; AD/CA, significant expression differences between adenomas and carcinomas; AD/MET, significant expression differences between adenomas and lymph node metastases.
Expression Levels of the Epidermal Growth Factor Receptor Family Genes
Expression of all 4 members of the epidermal growth factor receptor family of receptor tyrosine kinases and their downstream signal transducers were analyzed (Table 2). ERBB1 expression levels were significantly reduced in lymph node metastases when compared to adenomas. In contrast, no significant changes in expression were found for ERBB2 and ERBB3. Furthermore, expression of members of the ERBB signaling pathway—namely, MAPK1, MAPK6, STAT3, and TNK2—did not differ significantly among adenomas, carcinomas, and lymph node metastases.
Expression Levels of SATB1 and SLIT2 Genes
The chromatin organizer and transcription factor SATB1 had decreased expression in the majority of adenomas, carcinomas, and lymph node metastases (Table 2). A significantly decreased SATB1 expression was observed in lymph node metastases when compared to adenomas. The tumor suppressor SLIT2 had significantly reduced expression in metastases when compared to adenomas. No significant expression differences were observed between adenomas and carcinomas.
Expression of Estrogen and Progesterone Receptor Genes
Estrogen receptor alpha expression was variable in adenomas, carcinomas and lymph node metastases when compared to normal gland (Table 2). No significant expression differences were found when adenomas, carcinomas and lymph node metastases where compared with each other. Progesterone receptor expression demonstrated a 10-fold reduction in carcinomas and lymph node metastases when compared to normal gland. Similarly, mean progesterone receptor expression significantly decreased when carcinomas and metastases were compared with adenomas.
Expression of the Transforming Growth Factor Beta Superfamily Genes
Expression levels of the genes encoding transforming growth factor beta 3 (TGFb3), transforming growth factor beta receptor 3 (TGFbR3), and the latent transforming growth factor beta binding proteins 1 and 4 (LTBP1 and LTBP4) had significantly different expression levels in adenomas, carcinomas, and metastases. 12 TGFb1, TGFb2, TGFbR1, and LTBP3 were not differentially expressed in adenomas and carcinomas and metastases. In brief, TGFb3 and LTBP4 had significantly decreased expression levels in carcinomas and metastases when compared to adenomas. TGFbR3 had decreased expression in metastases when compared to adenomas (Table 2). In contrast, LTBP1 expression was significantly increased in carcinomas and metastases when compared to adenomas.
Expression of the BRCA–RAD51 Complex Genes
Expression levels of BRCA1, BRCA2, and RAD51 were significantly increased in carcinomas and lymph node metastases when compared to adenomas. 10
Expression of the p21, p27, and p53 Genes
Expression of p21 was significantly increased in lymph node metastases when compared to adenomas. In contrast, p27 expression was significantly reduced in carcinomas and metastases when compared to adenomas. Finally, p53 expression did not differ significantly among the 3 groups. 9,11
Expression of DERL1, Maspin, EGP2, STC1, Osteopontin, and Telomerase Genes
Expression levels of DERL1 (Derlin-1) was significantly increased in lymph node metastases when compared to adenomas. 8 No significant differences in expression among any of the groups tested were observed for maspin, STC1 (stanniocalcin-1), EGP2, osteopontin, and telomerase.
ANOVA
ANOVA identified 10 genes with significant gene expression differences between adenomas and carcinomas and between adenomas and metastases: BMP1, BRCA1, BRCA2, LTBP1, LTBP4, p27, progesterone receptor, RAD51, SMAD6, TGFb3. Significant differential expression levels between adenomas and lymph node metastases were only identified for 7 genes: DERL1, ERBB1, MIG6, p21, SATB1, SLIT2, and TGFbR3. In contrast, the remaining 24 genes had no significant differences in their expression levels among adenomas, carcinomas, and lymph node metastases.
Discriminant Analysis
All 17 genes with significant differences in the ANOVA were included in a discriminant analysis to identify genes most relevant to distinguish between benign tumors (adenomas) and malignant tumors (carcinomas, metastases). In a first discriminant analysis with all 17 genes, 97.2% of all tissue samples were correctly classified as adenoma, carcinoma, or lymph node metastasis by their molecular phenotype. The smallest set of genes that correctly distinguished all malignant from benign tumors consisted of 3 genes—namely, LTBP4, BMP2, and DERL1 (Fig. 1). With these 3 genes, 100% of the adenomas were classified as adenomas. Furthermore, 84.6% of the actual carcinomas were classified as carcinoma and 15.4% as lymph node metastases; 76.9% of the actual lymph node metastases were classified as lymph node metastases and 23.1% as carcinomas. Correct discrimination of malignant from benign tumors was impossible with fewer than 3 genes or with any of the LTBP4, BMP2, and DERL1 expression levels alone.
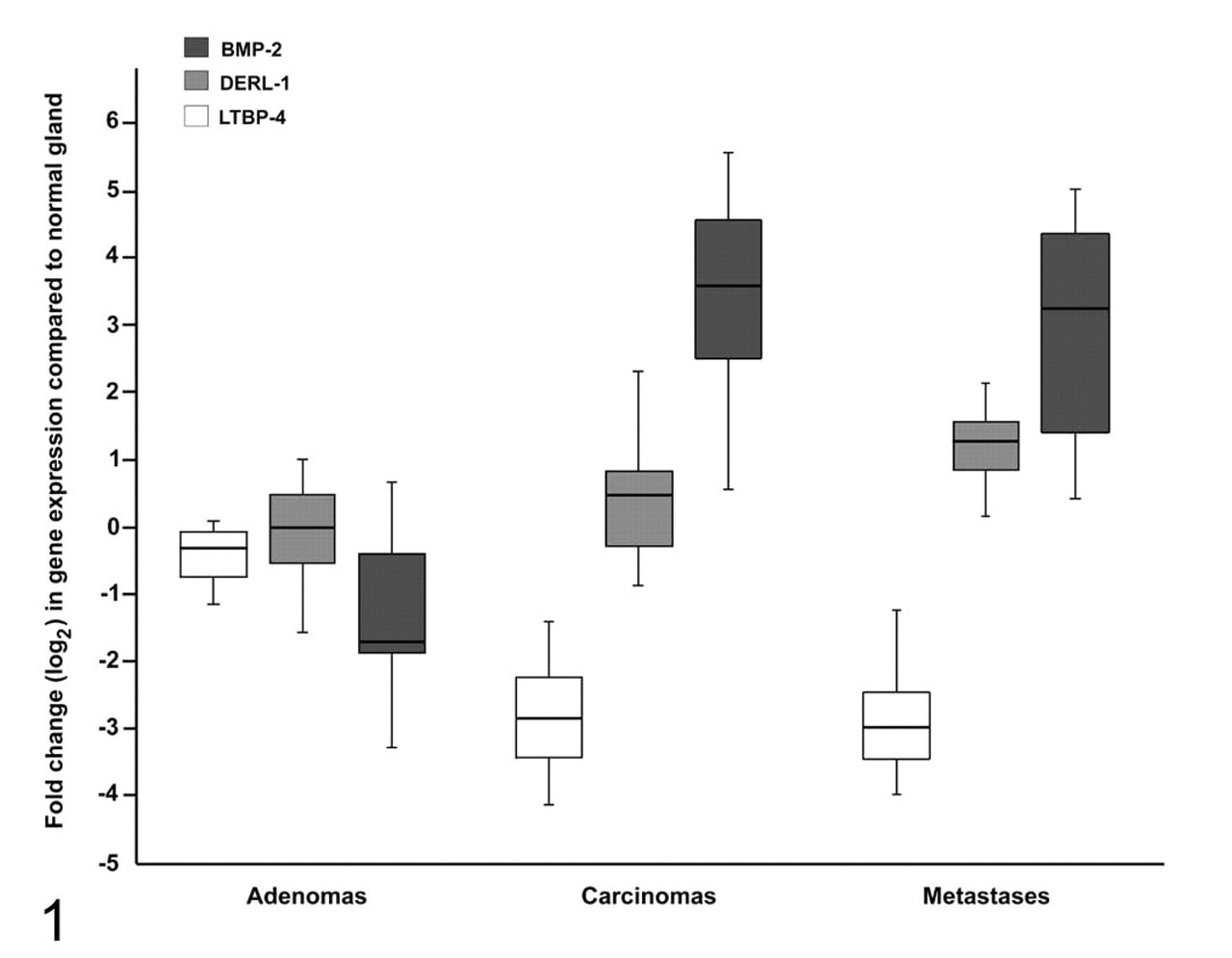
Malignancy-associated BMP2, DERL1, and LTBP4 expression pattern in adenomas, carcinomas, and metastases. When compared to normal gland of the same dog, each malignant tumor was characterized by increased BMP2 and DERL1 expression and markedly decreased LTBP4 expression; in contrast, adenomas had decreased BMP2 and LTBP4 expression, and their DERL1 expression was similar to that of the normal gland.
Discussion
Several previous studies have identified tumor markers that were differentially expressed between benign and malignant canine mammary tumors when entire groups of tumors were compared by their group means. 5,21 –23 In contrast, this pilot study identified a set of molecular markers that, for the first time, allows the identification of malignancy for each tumor and not only for means of larger groups of tumors. Thus, a combination of increased BMP2 and DERL1 expression and reduced LTBP4 expression allowed each tissue to be identified as a benign or malignant tumor. In contrast, the single-gene expression level of any of the 3 aforementioned genes or other 49 genes analyzed here was unable to classify each tumor correctly. The combined gene expression pattern of BMP2, DERL1, and LTBP4 was therefore able to confirm the histopathologic categorization, whereas all single-gene expression analyses were inferior to histopathology. Surprisingly, correct discrimination of lymph node metastases from their primary tumors was impossible, even when all 17 ANOVA preselected genes were included in the analysis. This observation was confirmed by the results of the ANOVAs of group differences. Here, no statistically significant differences were found between carcinomas and their metastases for all 49 genes. Primary tumors and their lymph node metastases therefore seem to be similar in the majority of their gene expression levels. However, only a fraction of all canine genes were analyzed here, and other genes not included here may be involved, which leads to 2 conceivable scenarios: First, metastasis of canine mammary tumors is a random process, and the average gene expression of metastatic cells is similar to that of the primary tumor. Alternatively, primary and metastatic tumor cells differ in their expression levels of genes not included in this study. Clearly, future analyses of the global gene expression patterns of canine mammary tumors are needed to identify genes that are differentially expressed between primary tumors and their lymph node metastases.
Whether BMP2, DERL1, and LTBP4 are primarily involved in the carcinogenesis of malignant canine mammary tumors or only differentially regulated as part of a complex, malignancy-associated molecular phenotype is unclear at this time. However, a direct interaction or molecular pathway shared by all 3 genes is unknown so far. Furthermore, genetic aberrations have not been reported for any of these 3 genes in breast cancer patients. BMP2 is a member of the TGF superfamily, and it plays an important role in human breast cancer neovascularization, regulation of tumor cell motility, and inhibition of hypoxic cell death. 2,14,24,25 The significance of canine BMP2 in tumor progression and carcinogenesis is unclear, but its strong overexpression in the malignant canine mammary tumors is in sharp contrast to reduced BMP2 expression in highly malignant human breast cancer. 3 LTBP4 is a modulator of TGFb processing and activity, and the TGFb–LTBP complex functions as a depot for quickly available but latent TGFb in the extracellular matrix. 7,19,27 Because of its effect on TGFb availability, LTBP4 plays an indirect but important role in the pathogenesis of mammary neoplasms, and it has decreased expression in both human breast cancer and canine mammary tumors. 12,17 DERL1 serves as a transporter protein for the export of misfolded proteins from the endoplasmic reticulum and as an inhibitor of endoplasmic reticulum stress-induced apoptosis. 15 Increased DERL1 expression is a feature of canine mammary tumors and human breast cancer. 8,13,34 Overexpression of DERL1 in human breast cancer suggests that regulation of the endoplasmic reticulum stress response pathway may be critical in the development and progression of breast cancer, and a similar role has been proposed for canine mammary tumors. 8,13,34
The number of tumors in the present pilot study was limited because of the strict selection criteria for adenomas and carcinomas and the use of laser-capture microdissection to obtain pure, high-quality tumor cell mRNA. The time-consuming and laborious method of laser microdissection had to be employed here because it is the only approach to reliably quantify tumor cell mRNA from histologic specimen without contamination by stromal and immune cells. Thus, given the temporal efforts required to laser dissect sufficient tumor cells, the number of patients that can be analyzed in such a study is limited, irrespective of the subsequent molecular analysis. The results of this pilot study show that a combination of increased BMP2 and DERL1 expression and decreased LTBP4 expression may be used to discriminate between benign and metastatic canine mammary gland tumors in each tumor, in contrast to group comparisons of previous studies. However, the routine use of this diagnostic pattern and studies with large patient numbers are still of concern owing to the time-consuming and technically demanding laser microdissection deemed critical to obtain pure tumor cell mRNA.
Footnotes
Acknowledgements
We thank Leo Brunnberg, Helge Linzmann, Martin Köhle, and Thomas Lottermoser for providing tumors samples and Monika Schärig for excellent technical assistance.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
