Abstract
Objective
The relationship between periodontal disease and adipokines has been extensively investigated. Using a Mendelian randomization approach, this study aimed to determine whether circulating leptin and resistin levels are causally implicated in the etiology of periodontal disease and whether the genetic risk of periodontal disease influences circulating adipokine levels.
Methods
We obtained genetic data regarding circulating leptin and resistin and periodontal disease from comprehensive genome-wide association study meta-analyses. We selected single nucleotide polymorphisms associated with periodontal disease and adipokine levels at a stringent genome-wide significance threshold (P < 5 × 10−6) as instrumental variables. Linkage disequilibrium clumping was applied, and F-statistics were calculated for all single nucleotide polymorphisms included in the Mendelian randomization study. Data analyses were performed using the inverse variance–weighted (IVW) method implemented in R software.
Results
The genetic risk of periodontal disease showed a positive effect on circulating resistin levels (IVW: odds ratio = 1.061, 95% confidence interval = 1.013–1.111, P = 0.013). However, no evidence supported a causal influence of circulating leptin or resistin levels on the risk of periodontal disease (IVW: leptin, odds ratio = 0.989, 95% confidence interval = 0.869–1.124, P = 0.861; resistin, odds ratio = 0.947, 95% confidence interval = 0.854–1.051, P = 0.304). Furthermore, the genetic risk of periodontal disease showed no causal association with circulating leptin levels (IVW: odds ratio = 1.021, 95% confidence interval = 0.993–1.050, P = 0.141).
Conclusions
These findings suggest that periodontal disease may exert a causal effect on circulating resistin levels. In contrast, circulating leptin and resistin levels do not appear to causally influence the risk of periodontal disease.
Introduction
Periodontal disease, a multifactorial inflammatory disorder affecting the tissues surrounding the teeth, poses significant global health challenges. 1 Characterized by gingival inflammation, periodontal pocket formation, and alveolar bone loss, it is not only a leading cause of tooth loss but has also been linked to systemic conditions such as cardiovascular disease, diabetes, chronic renal disease, and metabolic syndrome as well as adverse pregnancy outcomes. 2 The prevalence of periodontal disease and its impact on oral and systemic health underscore the need for a deeper understanding of its pathophysiology and associated factors.
Recent studies have highlighted the potential role of adipokines—biologically active molecules secreted by adipose tissue—in periodontal disease. 3 Among these, leptin and resistin are of particular interest because of their involvement in inflammatory and metabolic processes. 4 Leptin, traditionally recognized for its role in energy homeostasis, has emerged as a key regulator of immune function. 5 It exerts its effects through the leptin receptor, which is expressed in various cell types, including periodontal fibroblasts, osteoblasts, and immune cells such as macrophages and T cells. 6 Upon activation, leptin triggers signaling pathways that promote the production of proinflammatory cytokines, including tumor necrosis factor-α (TNF-α), interleukin-6 (IL-6), and IL-12. 7 This dual function places leptin in a paradoxical position within periodontitis, potentially supporting host defense while, upon dysregulation, exacerbating inflammatory tissue destruction. 8 Similarly, resistin acts as a proinflammatory adipokine, contributing to periodontal inflammation by upregulating the expression of mediators such as IL-6, IL-12, and C-reactive protein (CRP) and by stimulating the production of matrix metalloproteinases that degrade the periodontal extracellular matrix. 9 Furthermore, resistin promotes the expression of endothelial adhesion molecules, facilitating immune cell infiltration into periodontal tissues and amplifying the inflammatory response. 10
The interplay between periodontal disease and adipokines suggests a complex relationship in which periodontal inflammation may influence systemic adipokine levels, while changes in adipokine levels could, in turn, modulate periodontal health.3,11 In 2017, the author of the current study conducted an evidence-based investigation into the association between leptin levels and periodontitis, reporting higher circulating leptin levels in affected individuals. 12 In the same year, a meta-analysis by Akram et al. identified resistin as a potential biomarker for periodontitis. 13 These meta-analyses integrated individual observational study outcomes into quantitative estimates, providing a comprehensive overview; however, observational studies have often yielded inconclusive results, primarily due to confounding variables such as diet, lifestyle, and socioeconomic factors, which may simultaneously affect periodontal health and adipokine levels. To address these limitations, the present study employed a Mendelian randomization (MR) approach, which uses genetic variants as proxies for modifiable risk factors—such as circulating leptin and resistin levels—to infer causal effects on health outcomes. As alleles are randomly segregated during meiosis, the genetic variants used as instrumental variables (IVs) in MR are not influenced by the confounding factors that affect traditional observational studies. This method thus provides an opportunity to assess causality in the absence of confounding and to clarify the directionality of observed associations.
The study aimed to elucidate the causal relationship between periodontal disease and circulating leptin and resistin levels using an MR framework. By analyzing data from large-scale genome-wide association studies (GWAS) regarding genetic variants associated with these adipokines, we assessed whether leptin and resistin are causally implicated in the etiology of periodontal disease or whether the genetic risk of periodontal disease affects their circulating levels. The findings are expected to contribute to a more comprehensive understanding of periodontal disease pathogenesis and may inform the development of targeted interventions and personalized treatment strategies.
Materials and methods
Study design
Our study employed a two-sample MR framework, using genetic variants as IVs to infer the causal effects of exposures—circulating leptin and resistin—on the outcome (periodontal disease). This design is advantageous because it reduces confounding and reverse causation, common limitations of observational studies.
Data sources
We obtained genetic data regarding circulating leptin and resistin and periodontal disease from comprehensive GWAS meta-analyses, available through an open-access database curated by the MRC Integrative Epidemiology Unit at the University of Bristol. This repository provides complete GWAS summary findings, with provisions for open-source data retrieval. To minimize potential bias from sample overlap between exposure and outcome datasets, GWAS summary statistics for circulating leptin/resistin and periodontal disease were derived from independent, large-scale consortia. The search codes for genetic data on leptin and resistin were “ebi-a-GCST90012076” and “ebi-a-GCST90012034,” respectively, encompassing 21,758 individuals of European ancestry. These data originated from the same cohort, including 30,931 participants across 14 studies, which identified and replicated 315 primary and 136 secondary protein quantitative trait loci for 85 circulating proteins. 14 For periodontal disease, we used data from “GCST90018897,” comprising 1740 cases and 347,186 controls, with single nucleotide polymorphisms (SNPs) linked to periodontitis identified through genome-wide association analyses. 15
Selection of genetic variants
We selected IVs based on the three core assumptions of MR analysis (Figure 1). SNPs associated with periodontal disease and circulating leptin and resistin levels that met a stringent genome-wide significance threshold (P < 5 × 10−6) were included.16,17 To ensure that these SNPs were not confounded by other variables, we employed linkage disequilibrium (LD) clumping, excluding SNPs in close proximity (r2 ≥ 0.001) to a more strongly associated SNP within a 1-megabase window. SNPs that were palindromic or had uncertain allele frequencies were removed to prevent potential biases. The strength and precision of the association between each SNP and the exposure were assessed using the F-statistic, calculated as F = β²/SE², where β represents the estimated effect of the SNP on the exposure per allele and SE is the standard error. SNPs with an F-statistic below 10 were excluded to reduce the risk of weak instrument bias, which could compromise the validity of the MR analysis.

Overview of the MR design. Assumption 1: The IVs are strongly associated with the exposure. Assumption 2: There are no unmeasured confounders between the IVs and the outcomes. Assumption 3: The IVs affect the outcome only through the exposure. MR: Mendelian randomization; IV: instrumental variables.
Statistical analysis
Data analyses were performed using R software, version 4.3.2 (R Foundation for Statistical Computing, Vienna, Austria). All estimates were reported with a two-tailed significance threshold of 0.05. The inverse variance–weighted (IVW) method was employed as the primary approach to estimate the causal effects of circulating leptin and resistin levels on periodontal disease risk, assuming that all genetic variants were valid instruments. To verify the robustness of our findings, we employed several supplementary MR methods, such as MR-Egger regression, which provide causal estimates even in the presence of pleiotropy, and the weighted median approach, which yields consistent estimates if at least 50% of the information originates from valid instruments. Statistical power for the MR estimates was assessed using the online tool mRnd (https://shiny.cnsgenomics.com/mRnd/).
Heterogeneity and sensitivity analyses
We conducted several sensitivity analyses to assess the robustness of the findings and the validity of the MR assumptions. MR-Egger regression was used to test for pleiotropy by examining the intercept term. Statistical heterogeneity among SNPs in the IVW and MR-Egger analyses was evaluated using Cochran’s Q test, with heterogeneity considered significant at P < 0.05. We also conducted a leave-one-out analysis to determine whether the results were influenced by a single SNP or a subset of SNPs.
Results
Applying the genome-wide significance threshold (P < 5 × 10−6), we initially yielded total of 190, 534, and 87 SNPs for leptin, resistin, and periodontal disease, respectively. Following LD-based clumping and evaluation of F-statistics, 164, 521, and 74 SNPs were excluded because they were palindromic or had low F-statistics. Ultimately, 26, 13, and 13 SNPs were included in the statistical analysis. The results of the MR analyses are presented in Tables 1 and 2.
Mendelian randomization estimates for the genetic association between periodontal disease and circulating leptin and resistin levels.

MR: Mendelian randomization; OR: odds ratio; CI: confidence interval.
Summary of bidirectional MR analysis between circulating adipokines and periodontal disease.
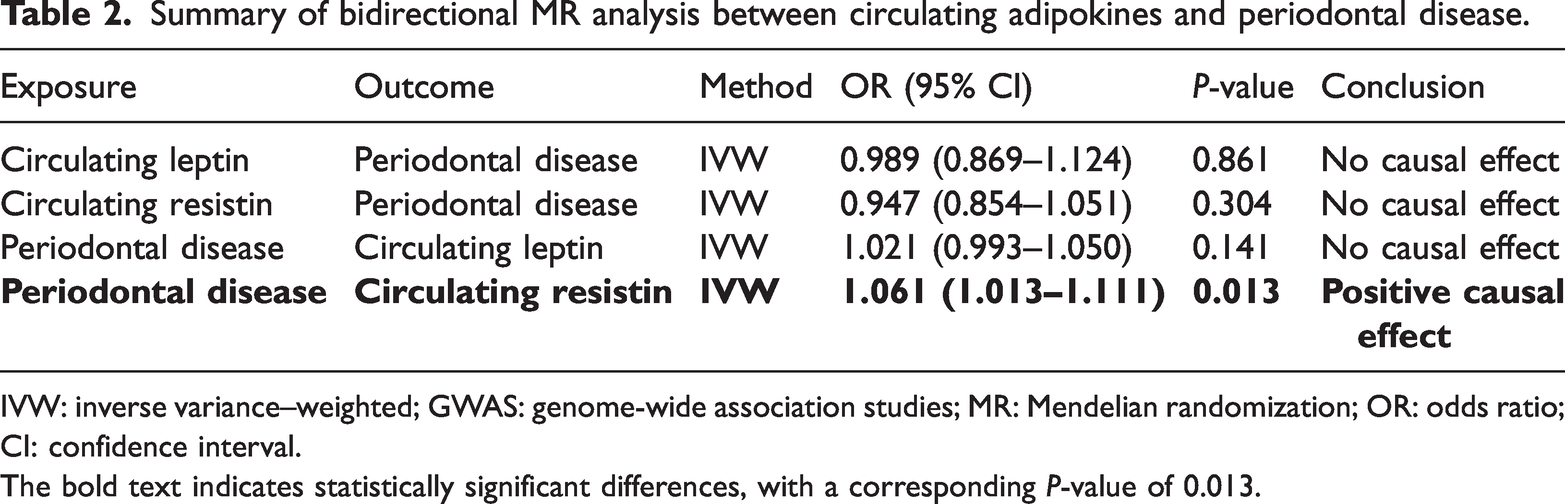
IVW: inverse variance–weighted; GWAS: genome-wide association studies; MR: Mendelian randomization; OR: odds ratio; CI: confidence interval.
The bold text indicates statistically significant differences, with a corresponding P-value of 0.013.
Circulating leptin level and periodontal disease
To investigate the potential causal relationship between circulating leptin and periodontal disease, we initially treated circulating leptin as the exposure and periodontal disease as the outcome. Forest plots and scatter plots are shown in Figures 2(a) and 3(a). IVW analysis yielded an odds ratio (OR) of 0.989 (95% confidence interval (CI): 0.869–1.124; P = 0.861), MR-Egger analysis yielded an OR of 0.759 (95% CI: 0.500–1.153; P = 0.209), and the weighted median method yielded an OR of 0.961 (95% CI: 0.807–1.144; P = 0.653). Heterogeneity was assessed using Cochran’s Q test, showing no significant heterogeneity for IVW (P = 0.169) or MR-Egger (P = 0.201). MR-Egger regression indicated no evidence of horizontal pleiotropy (P = 0.206). Leave-one-out analysis, performed by iteratively excluding each SNP, demonstrated that the overall effect estimates remained stable, supporting the robustness of the findings (Figure 4(a)).

Forest plots of the genetic association between circulating leptin levels and periodontal disease. (a) Exposure: circulating leptin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating leptin.

Scatter plots of the genetic association between circulating leptin levels and periodontal disease. (a) Exposure: circulating leptin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating leptin.

Leave-one-out sensitivity analysis for the genetic association between circulating leptin levels and periodontal disease. (a) Exposure: circulating leptin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating leptin.
Next, periodontal disease was treated as the exposure and circulating leptin as the outcome. The forest plots and scatter plots are shown in Figures 2(b) and 3(b). The IVW method yielded an OR of 1.021 (95% CI: 0.993–1.050; P = 0.141), MR-Egger yielded an OR of 0.993 (95% CI: 0.959–1.028; P = 0.680), and the weighted median method yielded an OR of 0.996 (95% CI: 0.957–1.036; P = 0.834). Cochran's Q test indicated no heterogeneity for IVW (P = 0.995) or MR-Egger (P = 1.000). MR-Egger analysis detected evidence of horizontal pleiotropy for IVW (P = 0.007). Leave-one-out analysis showed that excluding individual SNPs did not substantially alter the overall effect estimates, indicating the stability of the findings (Figure 4(b)).
Circulating resistin level and periodontal disease
To assess the potential causal relationship between circulating resistin and periodontal disease, resistin was first treated as the exposure and periodontal disease as the outcome. Forest plots and scatter plots are shown in Figures 5(a) and 6(a). The IVW method yielded an OR of 0.947 (95% CI: 0.854–1.051; P = 0.304), MR-Egger yielded an OR of 1.019 (95% CI: 0.857–1.212; P = 0.836), and the weighted median method yielded an OR of 0.972 (95% CI: 0.859–1.099; P = 0.647). Cochran’s Q test indicated no significant heterogeneity for IVW (P = 0.315) or MR-Egger (P = 0.323). MR-Egger regression revealed no evidence of horizontal pleiotropy (P = 0.326). Leave-one-out analysis confirmed the stability of the effect estimates, supporting the robustness of the findings (Figure 7(a)).
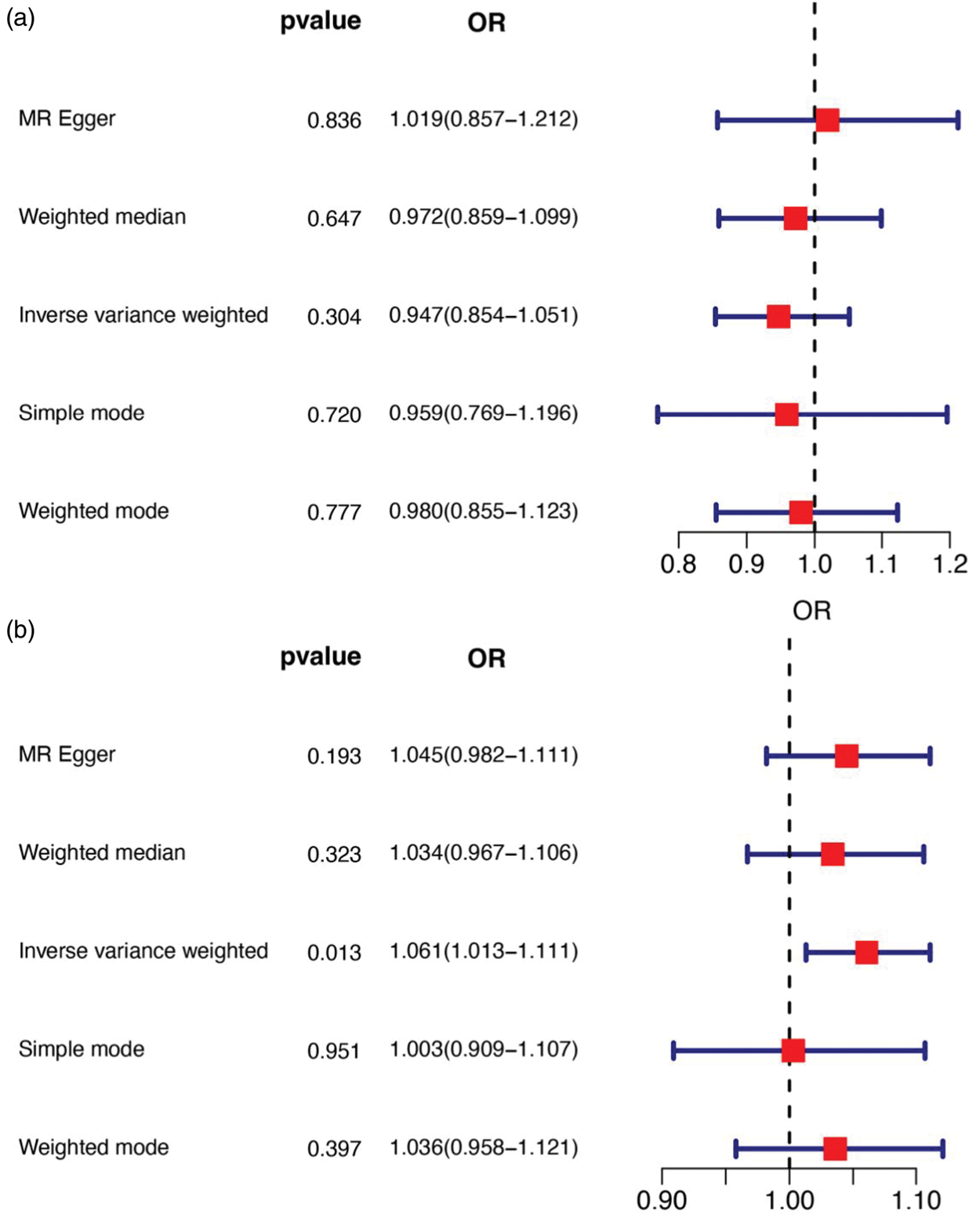
Forest plots of the genetic association between circulating resistin levels and periodontal disease. (a) Exposure: circulating resistin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating resistin.

Scatter plots of the genetic association between circulating resistin levels and periodontal disease. (a) Exposure: circulating resistin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating resistin.

Leave-one-out sensitivity analysis for the genetic association between circulating resistin levels and periodontal disease. (a) Exposure: circulating resistin; outcome: periodontal disease and (b) exposure: periodontal disease; outcome: circulating resistin.
Next, periodontal disease was treated as the exposure and circulating resistin as the outcome. Forest plots and scatter plots are shown in Figures 5(b) and 6(b). The IVW method yielded an OR of 1.061 (95% CI: 1.013–1.111; P = 0.013), MR-Egger yielded an OR of 1.045 (95% CI: 0.982–1.111; P = 0.193), and the weighted median method yielded an OR of 1.034 (95% CI: 0.967–1.106; P = 0.323). These results indicated that an increased risk of periodontal disease was associated with genetically predicted higher resistin levels. Cochran’s Q test indicated no significant heterogeneity for IVW (P = 0.587) or MR-Egger (P = 0.547), and MR-Egger regression revealed no evidence of pleiotropy (P = 0.487). Leave-one-out analysis further confirmed the robustness of the findings (Figure 7(b)).
Statistical power analysis
The analysis demonstrated substantially high statistical power for the effect of periodontal disease on circulating resistin, with the results indicating that the analysis was well-powered (>80%). In contrast, the power for detecting the effects of adipokines on periodontal disease was considerably lower, being very low (<10%) for leptin and low (25%) for resistin.
Discussion
This study employed a two-sample bidirectional MR approach to investigate the potential causal relationships between circulating leptin and resistin levels and periodontal disease. Using this design and a rigorous analytic strategy that leveraged genetic instruments, we aimed to minimize potential confounding inherent in observational studies and assess both the direction and robustness of the associations. MR analyses revealed no evidence that genetically predicted circulating leptin or resistin levels influence the risk of periodontal disease. In contrast, the genetic risk of periodontal disease was associated with increased circulating resistin levels, suggesting a potential causal effect that warrants further investigation.
In 2017, the author of the present study conducted a meta-analysis examining the correlation between circulating leptin levels and periodontitis, revealing a significant positive association. However, the present MR analyses did not provide strong evidence for a causal genetic association between circulating leptin levels and periodontal disease. The IVW, MR-Egger, and weighted median methods yielded nonsignificant results, which were supported by homogeneous outcomes in heterogeneity tests. Leave-one-out sensitivity analyses indicated that no single SNP disproportionately influenced the overall effect estimate, supporting the robustness of the negative findings. Notably, when the circulating leptin level was considered the outcome and periodontal disease the exposure, MR-Egger intercept tests suggested the presence of pleiotropy; therefore, these results should be interpreted with caution.
This MR study did not establish a genetic association between circulating leptin levels and periodontal disease. However, leptin may still play a context-dependent role in disease development, such as modulating local inflammatory responses, which cannot be fully assessed using genetic data.18–20 Additionally, leptin could serve as an indicator of periodontitis activity or treatment efficacy, consistent with previous reports.21,22 Therefore, despite the lack of evidence for a direct causal relationship in this study, the involvement of leptin in inflammation and oral health is well-recognized.23,24
Regarding circulating resistin levels, our research did not provide compelling evidence regarding a causal link to periodontal disease when resistin was treated as the exposure. The statistical tests used mirrored those used for leptin, with all P-values exceeding 0.05, indicating no causal association. Heterogeneity tests confirmed consistency among the genetic instruments, and pleiotropy assessments showed no significant deviations from the expected model.
Conversely, when periodontal disease was considered the exposure and circulating resistin the outcome, the IVW method yielded an OR of 1.061 (95% CI: 1.013–1.111; P = 0.013), indicating a statistically significant association and suggesting that periodontal disease may increase circulating resistin levels. No significant heterogeneity or pleiotropy was detected, supporting the validity of the SNPs as IVs in this context. This finding is clinically relevant, as it implies that periodontal inflammation could have systemic implications by altering circulating resistin levels.
Resistin acts as a proinflammatory cytokine, primarily produced by inflammatory cells such as monocytes, macrophages, and neutrophils. 25 Previous studies have demonstrated that resistin acts as an endogenous ligand for Toll-like receptor 4 (TLR4) and is involved in the activation of multiple inflammatory signaling pathways. 26 Consequently, systemic inflammation resulting from periodontal disease may be influenced by resistin release, which could partially explain the strong association between periodontal disease and systemic disorders. 27 The present study indicated that periodontal disease may have a causal influence on circulating resistin levels, whereas resistin itself does not appear to causally elevate the risk of periodontal disease. This suggests that systemic resistin elevation may represent a downstream consequence of periodontal inflammation rather than a primary etiological factor. Based on this causal direction, effective periodontal treatment could reduce circulating resistin levels by mitigating periodontal inflammation. Previous clinical studies have reported reductions in systemic inflammatory markers, including CRP and proinflammatory cytokines, following periodontal therapy. 28 Given that resistin is predominantly secreted by inflammatory cells, its expression is likely to decrease as the periodontal inflammatory burden is alleviated, highlighting a potential avenue for future research to investigate the biological mechanisms linking periodontal disease and systemic resistin levels.
Several limitations of the present MR analyses should be noted. Although the study was well-powered to detect the observed effect of periodontal disease on circulating resistin, it was underpowered to identify the very small causal effects of leptin and resistin on periodontal disease. Therefore, the null findings regarding the causal role of these adipokines should be interpreted with caution, as they may be subject to Type II error due to limited sample sizes in the underlying GWAS. Additionally, pleiotropic effects could be obscured by the small number of genetic instruments or limited sample sizes. Such pleiotropy could arise if genetic variants associated with periodontal disease also influence leptin levels through shared inflammatory pathways, metabolic conditions such as obesity, or other unmeasured confounders. Moreover, the GWAS summary data were primarily derived from individuals of European ancestry. Genetic architecture, including LD patterns, allele frequencies, and population-specific variants, can vary substantially across ancestral groups. 29 Environmental and lifestyle factors affecting both periodontal health and adipokine levels also differ globally. 30 Therefore, these MR estimates may not be directly generalizable to non-European populations. Future research should include larger and more diverse cohorts as well as experimental studies to investigate the mechanisms mediating the observed associations. Intervention studies targeting periodontal health and monitoring changes in circulating resistin levels could provide critical insights into the clinical relevance of these findings.
In summary, this MR study found no evidence supporting a causal effect of circulating leptin or resistin levels on the risk of periodontal disease. Conversely, a potential causal effect of periodontal disease on elevating circulating resistin levels was observed, suggesting that effective management of periodontal disease may help regulate resistin expression and thereby modulate systemic inflammation. Further studies are warranted to validate and extend these findings.
Footnotes
Acknowledgments
Not applicable.
Author contributions
Junfei Zhu: Conceptualization, methodology design, supervision of the research project, overall guidance, and review and approval of the final manuscript.
Yanan Jing: Data collection, data analysis, interpretation of results, drafting and revision of the manuscript.
AI tools
DeepSeek was used to check the grammar in the manuscript.
Data availability statement
All data generated or analyzed during this study are included in this article.
Declaration of conflicting interests
The authors declare that they have no competing interests.
Funding
This study was supported by the National High Level Hospital Clinical Research Funding (2025-NHLHCRF-PY-43).
