Abstract
Objective
This study aimed to systematically examine the causal relationships between 19 common dietary factors and the risk of periodontal disease, thereby providing reliable evidence for targeted nutritional interventions.
Methods
Genetic instruments for dietary exposures (e.g. alcohol, meats, dairy, fruits, vegetables, and beverages) were derived from the UK Biobank Genome-Wide Association Study database (N = 421,764–462,630). Outcome data included 97,830 periodontal disease cases and 272,252 controls from FinnGen biobank. Inverse-variance weighted regression served as the primary analysis, supplemented by weighted median and Mendelian randomization–Egger methods for the assessment of pleiotropy and heterogeneity. Sensitivity analyses included Mendelian randomization–Pleiotropy Residual Sum and Outlier for outlier correction and false discovery rate adjustment.
Results
After false discovery rate correction and Mendelian randomization–Pleiotropy Residual Sum and Outlier adjustment, inverse-variance weighted analysis identified the following three significant causal associations for periodontal disease risk: higher cheese intake reduced the risk (odds ratio = 0.807, 95% confidence interval: 0.714–0.912); higher lamb/mutton consumption increased the risk (odds ratio = 1.463, 95% confidence interval: 1.138–1.880); and habitual added-salt intake elevated the risk (odds ratio = 1.198, 95% confidence interval: 1.081–1.327). No significant associations were found for the remaining 16 dietary factors.
Conclusion
This Mendelian randomization study provides robust evidence showing that cheese consumption may protect against periodontal disease, whereas lamb/mutton and high added-salt intake elevate the risk.
Introduction
Periodontal disease is an inflammatory disorder caused by periodontopathic bacteria; over time, it leads to the breakdown of alveolar bone tissue. 1 The global burden of periodontal disease is extremely high; more than 1 billion people worldwide suffer from severe forms of the disease. 2 The primary modifiable risk factors—poor oral hygiene and tobacco use—are well-established; however, emerging evidence suggests that dietary factors may independently influence disease susceptibility and progression.2–4 Conventional risk factors such as smoking and poor oral hygiene constitute the major periodontal disease etiology, while the role of diet remains underexplored despite plausible biological mechanisms. Notably, although dietary factors and conventional risk factors remain central to understanding periodontal disease, recent research has shed new light on the role of chlorhexidine and systemic health, two aspects that hold significant potential to further deepen our comprehension of the disease’s pathogenic pathways, in modulating mediators within oral and periodontal tissues.5,6 Observational studies face methodological challenges, including reverse causation (e.g. periodontal pain alters dietary choices) and residual confounding (e.g. socioeconomic factors influence both diet and oral hygiene). 7 Mendelian randomization (MR), by leveraging genetic variants as instrumental variables for exposures, offers a robust approach to infer causality while minimizing these biases. 8 Given the underexplored causal role of diet in periodontal disease and biases in observational studies, a robust approach such as MR is urgently needed to address this gap. The present study aimed to systematically examine the causal relationships between 19 dietary factors and periodontal disease using MR analytical methods. By employing MR to overcome the key limitations of observational studies, this research provides novel insights into the causal role of diet in periodontal disease, which could inform targeted nutritional interventions for prevention and management.
Methods
Data sources and instrumental variable selection
The exposure data for this study comprised 19 dietary-related factors derived from the UK Biobank 9 through the Integrative Epidemiology Unit (IEU) Open Genome-Wide Association Study (GWAS) Project. 10 These exposures covered various aspects of food and beverage consumption patterns, including alcohol intake frequency; intake of different meat types (processed, poultry, beef, pork, and lamb), fish (non-oily and oily), dairy products (cheese), grains (bread and cereal), fruits and vegetables (fresh fruit, cooked vegetables, and raw vegetables), and beverages (tea, coffee, and water); and other dietary factors (dried fruit and salt addition). The sample sizes for these exposure GWAS datasets ranged from 421,764 to 462,630 individuals (Table 1). The outcome data for periodontal diseases were obtained from the 10th release of the FinnGen biobank, 11 comprising 97,830 cases and 272,252 controls. To ensure the validity of our MR analysis, we implemented strict criteria for instrumental variable selection. First, we identified single-nucleotide polymorphisms (SNPs) significantly associated with each exposure at the genome-wide significance threshold (p < 5 × 10−8). To ensure independence among instrumental variables, we performed linkage disequilibrium clumping using a stringent threshold (r2 < 0.001) with a 10,000-kb clumping window. Furthermore, we excluded any SNPs showing significant associations with the outcome (periodontal diseases) at p < 5 × 10−8 to minimize potential bias. This study utilized publicly available summary-level data from established databases (UK Biobank and FinnGen) and therefore did not require additional ethics committee approval. This study was performed in accordance with the ethical principles outlined in the 1975 Declaration of Helsinki, as revised in 2024. We declare that the reporting of this study complies with the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE)-MR checklist. 12
Overview of GWAS datasets for exposure and outcome.

The exposure dataset comprised 19 dietary-related factors, derived indirectly from the UK Biobank (https://www.ukbiobank.ac.uk/) through the IEU OpenGWAS project (https://gwas.mrcieu.ac.uk/). The outcome dataset focuses on gingivitis and periodontal disease risk, obtained from the 10th release of the FinnGen biobank (https://r10.risteys.finngen.fi/endpoints/K11_GINGIVITIS_PERIODONTAL).
GWAS: Genome-Wide Association Study.
Statistical methods
We employed a comprehensive MR framework to assess causal relationships. As primary methods for causal inference, we employed inverse-variance weighted (IVW) regression as our main analysis method, complemented by weighted median (WM) and MR-Egger approaches. The WM method provides consistent estimates when up to 50% of the instrumental variables are invalid, while MR-Egger regression assesses potential directional pleiotropy and provides alternative causal estimates that are robust to certain types of pleiotropy. We interpreted the MR-Egger intercept test (p < 0.05 indicating significant horizontal pleiotropy) in conjunction with its causal estimate to evaluate potential biases. To assess heterogeneity across instrumental variables, we applied Cochran’s Q-test, where significant heterogeneity (p < 0.05) suggested variability in causal estimates. To minimize false positives in our primary analysis, we applied false discovery rate (FDR) correction to the p-values obtained from the IVW method. Additionally, we conducted outlier detection and correction using MR–Pleiotropy RESidual Sum and Outlier test (PRESSO), which identifies and removes potentially influential SNPs that may bias causal estimates. All statistical analyses were performed using R (version 4.3.2), with the TwoSampleMR package (version 0.6.2) for MR analyses and MR-PRESSO for outlier detection.
Results
MR analyses evaluated the causal effects of 19 dietary factors on periodontal diseases using the following three complementary methods: IVW (primary analysis), WM, and MR-Egger regression. The complete results before outlier removal are presented in Supplementary Table 1, while Supplementary Table 2 displays the findings after excluding outliers identified using MR-PRESSO. In our analyses, horizontal pleiotropy was detected in only one exposure–outcome association: the relationship between tea intake and gingivitis/periodontal diseases after outlier removal (MR-Egger intercept p = 0.0202; Supplementary Table 2). For all other dietary exposures (18/19), no significant horizontal pleiotropy was observed (MR-Egger intercept p > 0.05; Supplementary Tables 1 and 2). The scatter plot results illustrating the associations between each of the 19 dietary exposures and periodontal disease from MR analyses are presented in Figure 1.

Scatter plots of dietary exposures and periodontal disease risk. Scatter plots depict the associations of different dietary exposures with periodontal disease risk based on Mendelian randomization analyses: (a) alcohol intake frequency, (b) processed meat intake, (c) poultry intake, (d) beef intake, (e) non-oily fish intake, (f) oily fish intake, (g) pork intake, (h) lamb/mutton intake, (i) bread intake, (j) cheese intake, (k) cooked vegetable intake, (l) tea intake, (m) fresh fruit intake, (n) cereal intake, (o) salad/raw vegetable intake, (p) coffee intake, (q) dried fruit intake, (r) added-salt intake, and (s) water intake. Each plot shows SNP-specific estimates of the exposure–outcome association, with the lines representing the Mendelian randomization estimates. SNPs: single-nucleotide polymorphisms.
After outlier correction, IVW analyses (Table 2) revealed three significant dietary associations for gingivitis/periodontal disease risk: protective effects of cheese intake (odds ratio (OR) = 0.807, 95% confidence interval (CI): 0.714–0.912, q = 0.00607), detrimental effects of lamb/mutton (OR = 1.463, 95% CI: 1.138–1.880, q = 0.0197), and detrimental effects of added-salt intake (OR = 1.198, 95% CI: 1.081–1.327, q = 0.00607). No significant associations were found for alcohol (OR = 1.051, 95% CI: 0.982–1.125), processed meat (OR = 0.968, 95% CI: 0.781–1.199), poultry (OR = 0.988, 95% CI: 0.664–1.471), beef (OR = 0.780, 95% CI: 0.557–1.093), non-oily fish (OR = 0.892, 95% CI: 0.595–1.335), oily fish (OR = 0.956, 95% CI: 0.834–1.095), pork (OR = 0.956, 95% CI: 0.590–1.549), bread (OR = 0.851, 95% CI: 0.711–1.018), cooked vegetable (OR = 1.140, 95% CI: 0.778–1.671), raw vegetable (OR = 1.028, 95% CI: 0.706–1.498), fresh fruit (OR = 0.912, 95% CI: 0.756–1.100), dried fruit (OR = 0.873, 95% CI: 0.712–1.071), cereal (OR = 0.859, 95% CI: 0.707–1.043), coffee (OR = 1.000, 95% CI: 0.815–1.226), or water (OR = 0.970, 95% CI: 0.804–1.171) intake (all q > 0.05).
Results of the MR analyses assessing the influence of dietary factors on gingivitis and periodontal disease risk.
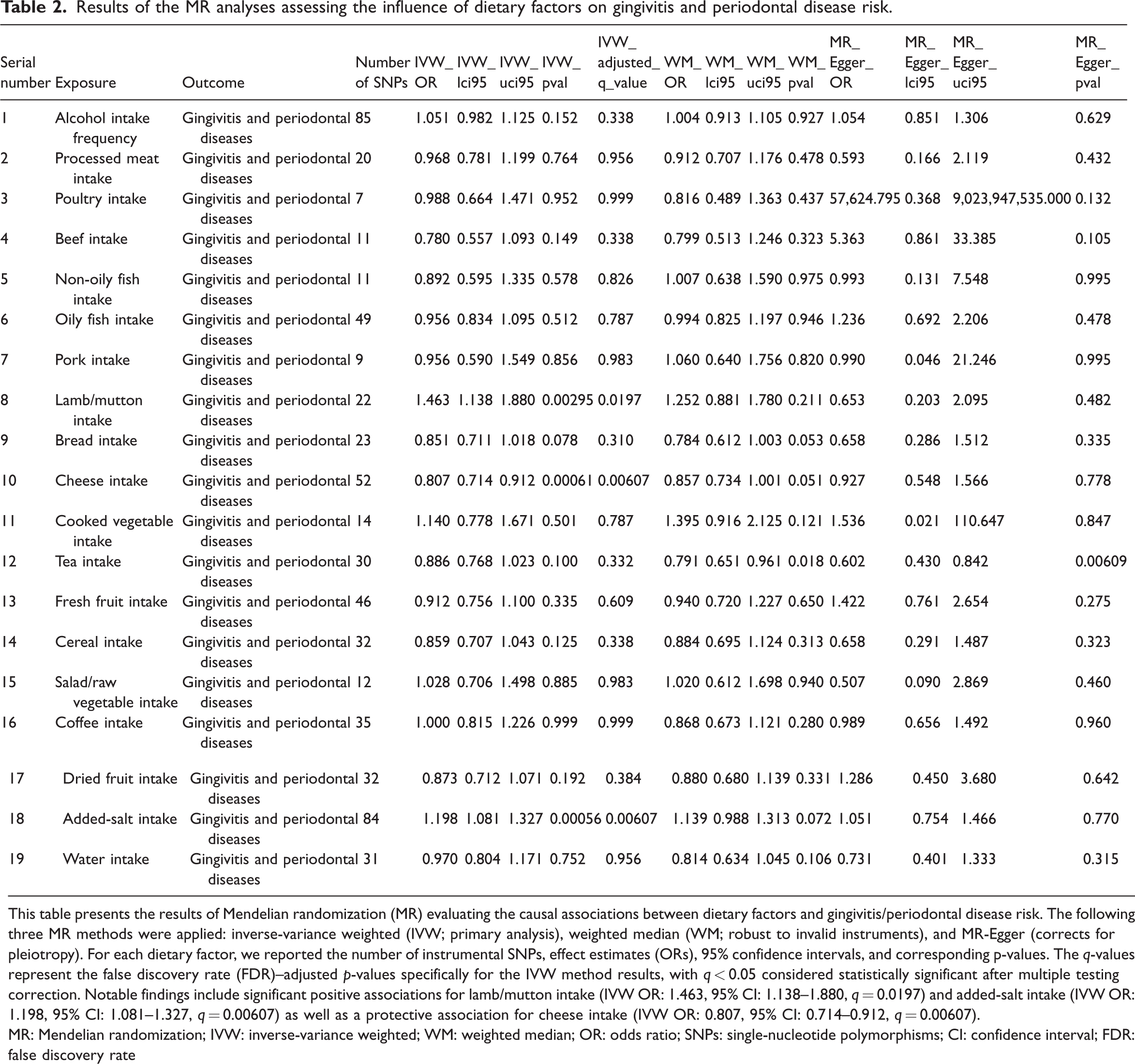
This table presents the results of Mendelian randomization (MR) evaluating the causal associations between dietary factors and gingivitis/periodontal disease risk. The following three MR methods were applied: inverse-variance weighted (IVW; primary analysis), weighted median (WM; robust to invalid instruments), and MR-Egger (corrects for pleiotropy). For each dietary factor, we reported the number of instrumental SNPs, effect estimates (ORs), 95% confidence intervals, and corresponding p-values. The q-values represent the false discovery rate (FDR)–adjusted p-values specifically for the IVW method results, with q < 0.05 considered statistically significant after multiple testing correction. Notable findings include significant positive associations for lamb/mutton intake (IVW OR: 1.463, 95% CI: 1.138–1.880, q = 0.0197) and added-salt intake (IVW OR: 1.198, 95% CI: 1.081–1.327, q = 0.00607) as well as a protective association for cheese intake (IVW OR: 0.807, 95% CI: 0.714–0.912, q = 0.00607).
MR: Mendelian randomization; IVW: inverse-variance weighted; WM: weighted median; OR: odds ratio; SNPs: single-nucleotide polymorphisms; CI: confidence interval; FDR: false discovery rate
Discussion
Our analysis revealed three significant causal associations between dietary factors and periodontal disease risk. First, higher genetically predicted cheese intake was associated with a reduced risk of periodontal disease, suggesting a protective effect. In contrast, increased consumption of lamb/mutton and habitual addition of salt to food were linked to an elevated risk of periodontal disease. Notably, no significant associations were observed for other dietary factors, including the intake of other meat types, fish, fruits, vegetables, or beverages, highlighting the specificity of these three dietary exposures in relation to periodontal health. The results derived from MR-Egger regression warrant cautious interpretation, particularly regarding the association between tea intake and periodontal disease. Although horizontal pleiotropy was detected for this specific exposure–outcome pair post-outlier removal (MR-Egger intercept p = 0.0202), the potential implications of this pleiotropy for the reliability of causal inferences about the role of tea in periodontal health remain underexplored in the current analysis. This pattern of findings, specifically the protective effects of cheese intake, detrimental effects of lamb/mutton intake and added-salt intake, and absence of meaningful associations for other dietary factors highlights that not all common dietary components exert a uniform impact on the risk of periodontal disease; instead, only a select subset plays distinct causal roles.
Current evidence regarding the association between cheese consumption and periodontal disease remains inconsistent. A cross-sectional study 13 involving 135 older Danish adults found that while milk and fermented dairy intake was associated with a reduced risk of periodontitis, and cheese consumption showed no significant protective association. Another national survey 14 involving 9798 Korean adults revealed that frequent dairy consumption (≥7 servings/week), particularly milk, was associated with a 24%–26% lower prevalence of periodontal disease, although cheese-specific effects were not reported. A Japanese community-based study 15 involving 942 adults aged 40–79 years reported that although a higher intake of lactic acid foods (≥55 g/day) was significantly associated with better periodontal health (40%–50% lower risk of severe periodontitis), no protective association was observed for cheese consumption. A recent MR analysis utilizing UK Biobank GWAS data demonstrated a significant inverse association between cheese consumption and periodontitis risk (OR = 0.545, 95% CI: 0.364–0.816). 16 Our study employed a similar methodology; however, we included a significantly larger sample size (97,830 cases of periodontal diseases vs. 4120 cases in the previous study); our findings validated their conclusions in that cheese consumption was inversely associated with the risk of periodontal diseases. Cheese is a nutrient-dense dairy product rich in high-quality protein, calcium, phosphorus, and vitamins, which also provides essential fatty acids and probiotics. 17 The consumption of cheese reportedly results in a statistically significant increase in the mean plaque concentrations of calcium and phosphorus. 18 Elevated concentrations of calcium and phosphate in the saliva are found to suppress the formation of bacterial biofilms.19,20 The association between salt intake and periodontal disease is not yet fully understood. In our study, excessive dietary salt intake was associated with an increased risk of periodontal disease. Research has demonstrated that high salt intake can alter gut microbiota composition, consequently impairing immune function. 21 The interaction of oxidative stress with inflammation may act as a crucial pathway linking dietary factors, including protective foods such as cheese and risk-associated foods such as lamb/mutton and added salt identified in this study, to periodontal health. This is because diet can modulate oxidative stress levels22,23 as well as the subsequent inflammatory responses that cause damage to periodontal tissues. Our study provides robust genetic evidence supporting causal relationships between specific dietary factors and periodontal health, addressing key gaps in the existing observational research on this topic. The identification of cheese intake as a protective factor, alongside the adverse effects of lamb/mutton and added-salt consumption, highlights modifiable dietary targets for periodontal disease prevention.
Several limitations of this study should be acknowledged. First, despite employing rigorous MR methods, residual confounding due to horizontal pleiotropy cannot be entirely ruled out, as exemplified by the detected pleiotropy in the tea–periodontal disease association. Second, data on dietary exposure and outcome were derived from European populations (UK Biobank and FinnGen), limiting the generalizability of the findings to non-European populations. Third, the dietary data lacked sufficient granularity (e.g. differentiation between cheese varieties, meat preparation techniques, or precise sodium quantification), which may obscure subtle associations. Fourth, although MR leverages genetic variants as proxies for exposure, it may not fully capture dynamic dietary changes or cumulative effects over time. Future studies incorporating diverse populations and detailed dietary assessments could address these limitations.
Conclusion
This study confirms that higher cheese intake is causally linked to a lower periodontal disease risk, while greater lamb/mutton and added-salt intake increase the risk. These findings highlight the causal role of diet in periodontal disease and guide actionable preventive targets. Future research should on diverse populations are required to validate the current findings.
Supplemental Material
sj-xlsx-1-imr-10.1177_03000605251389392 - Supplemental material for Dietary intake and periodontal health: Evidence from a Mendelian randomization analysis
Supplemental material, sj-xlsx-1-imr-10.1177_03000605251389392 for Dietary intake and periodontal health: Evidence from a Mendelian randomization analysis by Fulin Yan, Yongji Feng, Xin Dai and Hui Li in Journal of International Medical Research
Footnotes
Acknowledgments
We sincerely thank the UK Biobank (UKB), the FinnGen consortium, and the Integrative Epidemiology Unit (IEU) Open Genome-Wide Association Study (GWAS) project for providing publicly available genetic data and summary statistics, which were essential for this Mendelian randomization analysis.
Author contributions
Fulin Yan and Yongji Feng contributed equally to this work as co-first authors. Fulin Yan and Yongji Feng conceived the study, designed the methodology, conducted experiments, and wrote the original draft. Xin Dai performed data analysis, contributed to data interpretation, and participated in manuscript revision. Hui Li supervised the research project, provided critical guidance, acquired funding, and finalized the manuscript for submission. All authors reviewed and approved the final version of the manuscript.
Data availability statement
The datasets analyzed in this study were obtained from the following publicly available sources: the IEU OpenGWAS Project (https://gwas.mrcieu.ac.uk/) and the FinnGen consortium (![]() ).
).
Declaration of conflicting interests
None.
Funding
The authors declare that no funds, grants, or other financial support were received for this research.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
