Abstract
Objectives
We aimed to examine the significance of ephrin receptor A2 (EphA2) expression in pancreatic adenocarcinoma (PAAD) and its associated mechanism.
Methods
EphA2 mRNA expression patterns were compared in pancreatic cancer and normal tissues using GEPIA. Kaplan–Meier analysis was used to examine the correlation between EphA2 expression and PAAD patient prognosis. EphA2 gene methylation and associations with tumor immune cell infiltration were analyzed with UALCAN and TIMER, respectively. EphA2-interacting proteins were investigated with GeneMANIA, while STRING helped predict potentially relevant signaling pathways. EphA2 protein expression was examined with immunohistochemistry (IHC) in PAAD patient tissues.
Results
EphA2 was highly expressed in pancreatic cancer tissues and associated with pathological stage. PAAD patients with high EphA2 expression had shorter overall survival and disease-free survival times. EphA2 expression levels were significantly and positively associated with CD4+ T cell infiltration. EphA2 can interact with ENFNA1, ACP1, and CDC42. High EphA2 mRNA expression was enriched for regulation of cell size and cell proliferation. IHC assays suggested that pancreatic cancer tissues had higher EphA2 protein levels than normal pancreatic tissues.
Conclusions
EphA2 is highly expressed in PAAD and closely related to poor patient prognosis, and is therefore a potential biomarker and target for PAAD diagnosis and treatment.
Keywords
Introduction
Pancreatic adenocarcinoma (PAAD) is a highly malignant tumor type that is associated with rapid progression and poor prognosis. Because of the retroperitoneal location of the pancreas, pancreatic cancer does not have typical early clinical manifestations. This results in most patients having advanced tumors once they seek medical attention. The incidence and mortality of PAAD are increasing each year, mainly because of inadequate diagnostic and treatment options for this disease. 1 The specific mechanism controlling PAAD occurrence and development remains unclear, and there is a lack of specific diagnostic and therapeutic targets in clinical practice. There has been increasing interest in identifying novel specific and sensitive biomarkers for PAAD in recent years. Ephrin receptor A2 (EphA2) belongs to the ephrin receptor subfamily of the protein tyrosine kinase family. Previous studies have demonstrated that EphA2 is expressed in a variety of tumor types, with its expression being sufficient to support the malignant transformation and tumorigenic potential of cells.2–4 At present, there are few reports on the function, effect on patient survival, or related signaling pathways of EphA2 in PAAD. Therefore, this study aimed to clarify the mRNA expression levels of EphA2 in PAAD using data from publicly available databases. We further verified EphA2 expression levels in PAAD tumor and normal tissue samples. Bioinformatics methods were then used to analyze the relationships between EphA2 expression patterns and PAAD patient clinicopathological features and prognosis, as well as to explore the relevant signaling pathways involved. This work provides new insights for studying the early diagnosis, disease monitoring, and prognosis of PAAD.
Materials and methods
Gene Expression Profiling Interactive Analysis (GEPIA)
GEPIA (http://gepia.cancer-pku.cn/) is an online tool for gene expression and survival analysis using The Cancer Genome Atlas (TCGA) database and the Genotype-Tissue Expression (GTEx) portal. In the database, the conditions were set as “gene: EphA2”, “expression diy: boxplot”, and “cancer type: PAAD” to obtain the EphA2 expression differences between PAAD and normal tissues. The “Box Plot” module was used to analyze the correlation between EphA2 gene expression and clinical stage.
Kaplan–Meier (KM) plotter
The KM Plotter tool (http://KM-plot.com/analysis) can be used to assess the correlation between gene expression levels (mRNA, microRNA, or protein) and patient survival rates, including overall survival (OS) and disease-free survival (DFS). Sources for the databases include Gene Expression Omnibus (GEO), European Genome-phenome Archive (EGA), and The Cancer Genome Atlas (TCGA). The KM plotter tool was used to examine the relationship between EphA2 expression and pancreatic cancer patient survival time. The KM Plotter analysis module was selected and “EphA2” was input as the gene symbol. “Pancreatic cancer” was entered for cancer type. The patients were split by the automatically generated cut-off value. The following samples were included: all histology, any tumor grade, any tumor stage, and any TNM stage. OS and DFS analyses were performed.
The University of Alabama at Birmingham Cancer (UALCAN) Data Analysis Portal
The UALCAN data analysis portal (http://ualcan.path.uab.edu/analysis.html) is an effective cancer data online analysis and mining site that is mainly based on TCGA-related cancer data. Here, we selected the “TCGA analysis” module, then input EphA2 for Scan by gene and Pancreatic adenocarcinoma for TCGA dataset in the “TCGA gene analysis” interface. After selecting “Explore”, we chose “Expression” and “Methylation” to respectively analyze the expression levels and methylation status of the EphA2 gene in tumor and normal tissues.
Tumor Immune Estimation Resource (TIMER)
TIMER (https://cistrome.shinyapps.io/timer/) is an online tool for analyzing immune cell infiltration in tumors. The TIMER Gene analysis module was selected, then “EphA2” was entered for “Gene Symbol”, “PAAD” was entered for Cancer Types, and “B cell CD8+ T cell CD4+ T cell Macrophage Neutrophil Dendritic cell” was entered for “Immune Infiltrates”. A multivariate Cox proportional hazards model was constructed.
Search Tool for the Retrieval of Interacting Genes/Proteins (STRING)
STRING (https://string-db.org/) provides a confidence score for protein–protein interaction (PPI) using literature reports, data included in the database, and evidence of co-expressed proteins. A score closer to 1 indicates greater confidence in the interaction between molecules. A score closer to 0 indicates lower confidence from lack of currently available evidence. “EphA2” was searched for as the protein name in STRING, with “Homo sapiens” selected as the species to obtain EphA2-interacting proteins.
GeneMANIA
GeneMANIA (http://www.genemania.org) is a powerful website for genomics and proteomics analyses. The EphA2 PPI network was constructed as a supplementary analysis. GeneMANIA was used to explore the potential interactions of EphA2 with other proteins using pathway, physical, and genetic interactions to generate prediction networks.
Metascape
Metascape (http://metascape.org) is a powerful gene function annotation tool that enables users to apply common bioinformatic analysis methods to batch gene and protein analyses to understand gene or protein function. Gene annotation and functional enrichment analysis were performed for EphA2 and its neighbor genes using the “Express Analysis” module.
Tissue specimens and immunohistochemistry (IHC)
A retrospective analysis was conducted in this study. Tissue samples from 15 pancreatic cancer patients who received radical resection in Lishui Central Hospital from January 2021 to December 2022 were analyzed. EphA2 protein expression levels in pancreatic and para-cancer tissues were detected by IHC assays. The Ethics Committee of Lishui Municipal Central Hospital approved this study, and written informed consent was obtained from each patient. All patient details have been de-identified. Paraffin sections (4 μm thick) of pancreatic tissue specimens were prepared for analysis. IHC staining was performed in all cases using the streptavidin-peroxidase (SP) method. 5 A mouse anti-EphA2 antibody (MAB3035, 1:50, R&D Systems, Boston, MA, USA) was used.
Statistical analysis
Statistical analysis was performed using SPSS 20.0 (IBM Corp., Armonk, NY, USA). EphA2 expression in pancreatic and para-cancer tissues were analyzed using the χ2 test. P-values <0.05 were considered statistically significant.
Results
Pan-cancer expression of EphA2
EphA2 belongs to the ephrin receptor subfamily of the protein tyrosine kinase family, which are involved in mediating tumor events.2–4 EphA2 is expressed in various tumor tissues. Our analysis suggested that the tumor types with the highest EphA2 expression levels are esophageal carcinoma (ESCA), head and neck squamous cell carcinoma (HNSC), and PAAD (Figure 1).

EphA2 mRNA expression profiles of various tumor samples and paired normal tissues. The height of the bar represents the median expression level (in transcripts per million (TPM)) in each tumor or normal tissue type.
EphA2 expression patterns in PAAD patients
We next compared EphA2 expression levels in PAAD and normal pancreatic tissues. The data were acquired from TCGA database and GTEx portal. The results showed that EphA2 mRNA levels were significantly elevated (P < 0.05) in PAAD samples compared with normal pancreatic tissues (Figure 2a). Next, we used UALCAN to quantitatively evaluate EphA2 transcription. As shown in Figure 2b, we found that EphA2 showed high expression patterns in PAAD primary tumors (P = 1.68 × 10−12). Moreover, we used GEPIA to examine the association between PAAD clinical pathological stage and EphA2 expression. We found a significant correlation between EphA2 expression and pathological stage (P = 0.000281) (Figure 3). Collectively, these data suggest that EphA2 is substantially expressed in PAAD tissues and is involved in tumor formation and progression.
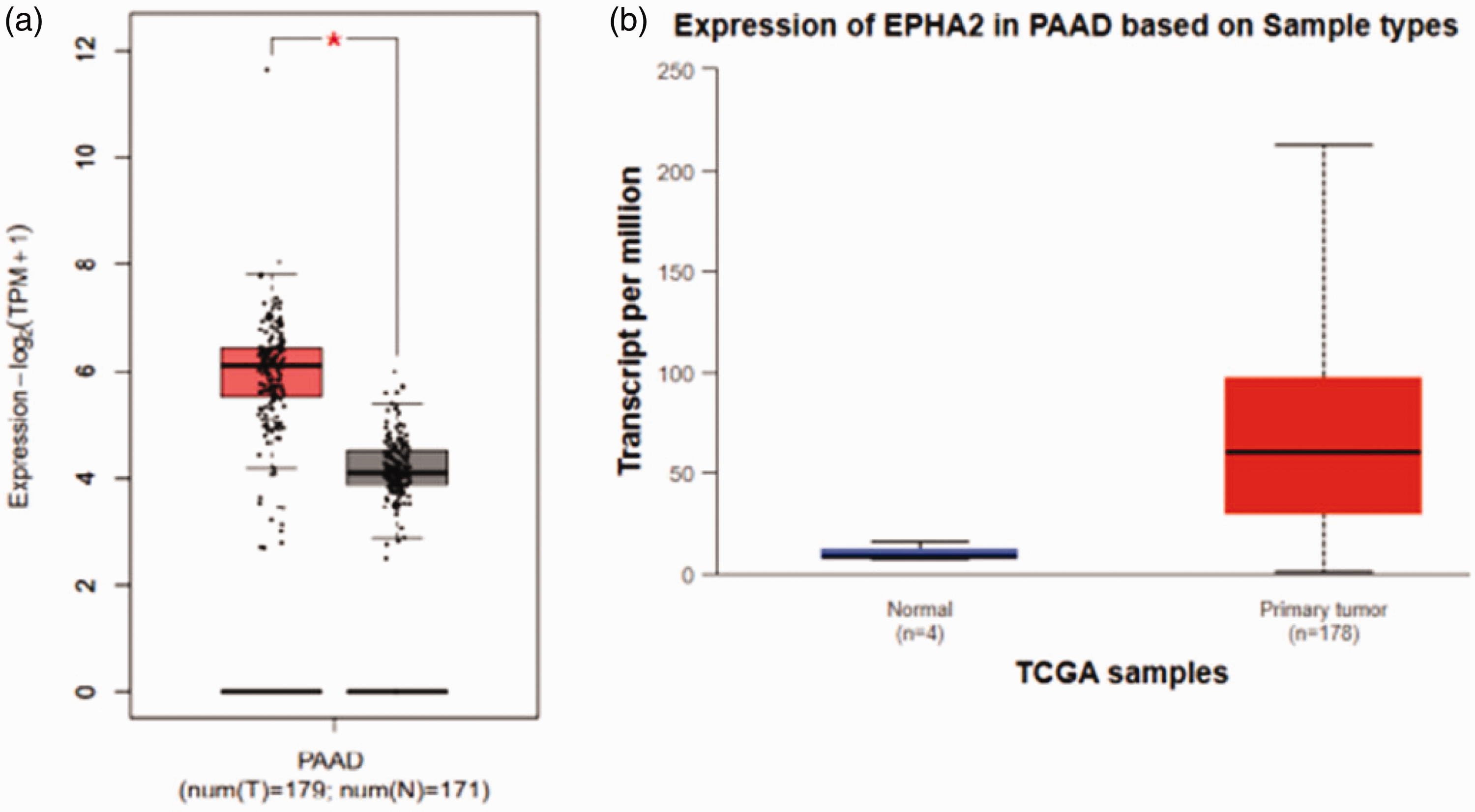
EphA2 expression patterns in pancreatic adenocarcinoma (PAAD) patients. (a) EphA2 transcriptional levels in PAAD and normal pancreatic tissues and (b) EphA2 mRNA levels in PAAD primary tumors and normal tissues.

Associations between pancreatic adenocarcinoma (PAAD) clinical pathological stage and EphA2 expression.
The prognostic value of EphA2 in PAAD patients
To further study the role of EphA2 in PAAD progression, survival correlation analysis was performed using the KM Plotter tool. Using the selected parameters, 1,189 patients were included in the OS analysis and 278 patients were included in the DFS analysis. PAAD patients with high EphA2 expression levels had significantly worse OS (P = 7.52 × 10−4, Figure 4a). Next, we estimated the role of EphA2 in prognostic stratification using DFS rates. We found that high EphA2 expression levels were significantly associated with poor DFS (P = 0.048, Figure 4b).

The prognostic value of EphA2 expression in pancreatic adenocarcinoma (PAAD) patients. Kaplan–Meier plots are shown for (a) overall survival and (b) disease-free survival.
EphA2 promoter DNA methylation in PAAD patients
To investigate the EphA2 methylation status in PAAD patients, we used the UALCAN database to perform methylation analysis. The Beta value indicates the level of DNA methylation ranging from 0 (unmethylated) to 1 (fully methylated). We observed a trend of higher EphA2 gene methylation levels in PAAD samples compared with normal tissues (Figure 5), but the difference was not statistically significant. We therefore excluded EphA2 methylation from further analysis.

EphA2 promoter DNA methylation patterns in pancreatic adenocarcinoma (PAAD) primary tumors and normal tissues.
Associations between immune cell infiltration and EphA2 expression in PAAD patients
Pancreatic cancer tumors have low immunogenicity and high immunosuppression, leading to poor clinical immunotherapy effects. 6 Therefore, we conducted a systematic analysis of the associations between EphA2 expression and immune cell infiltrates using TIMER. The Gene module was used to explore these correlations. A multivariant Cox proportional risk model was constructed to evaluate the prognostic factors of pancreatic cancer. We found that EphA2 expression was significantly and positively associated with CD4+ T cell infiltration (P = 0.0431; Figure 6). Additionally, 165 patients were included to construct a multivariate Cox proportional risk model, of which 89 patients died. Prognostic factors for pancreatic cancer included age, CD4+ T cell infiltration, and EphA2 gene expression (P < 0.05; Table 1).

Correlations of immune cell infiltration and EphA2 expression in pancreatic adenocarcinoma (PAAD) patients.
Multivariate Cox proportional risk model of pancreatic adenocarcinoma (PAAD).

Coef, coefficient; HR, hazard ratio; CI, confidence interval; EphA2, ephrin receptor A2.
P < 0.05 indicates statistical significance.
PPI analysis of EphA2 in PAAD patients
To investigate the interactions among EphA2 and various proteins of interest, we used the STRING tool for PPI network analysis. We identified 11 nodes and 29 edges in the PPI network (Figure 7a). The top 10 proteins in Score ranking were EFNA2 (Score = 0.999), EFNA5 (Score =0.999), EFNA1 (Score = 0.999), INPPL1 (Score = 0.998), EFNA3 (Score = 0.997), EFNA4 (Score = 0.997), EPHA1 (Score =0.976), EFNB2 (Score = 0.973), ABL1 (Score = 0.9751), and EFNB3 (Score =0.944). Additionally, we applied the GeneMANIA webtool to further elucidate the potential functions of these proteins. The size of the gene node indicates the strength of the interaction. The results showed that the top three associated functional intensities of interactions with EphA2 included ENFNA1, ACP1, and CDC42 (Figure 7b). PPI enrichment analysis indicated that the top three associated functional changes were EPH-Ephrin signaling, the ephrin receptor signaling pathway, and EPH-ephrin mediated repulsion of cells (Figure 7c).

Protein-protein interaction (PPI) analysis of EphA2 in pancreatic adenocarcinoma (PAAD) patients. (a) PPI network of EphA2. (b) Potential functions of the proteins of interest and (c) Molecular complex detection components identified in the gene lists.
Functional enrichment analysis of EphA2 in PAAD patients
To investigate the role of EphA2 and its associated proteins, the online webtool Metascape was subsequently used to annotate their functions. The gene ontology (GO)/Kyoto Encyclopedia of Genes and Genomes (KEGG) terms suggested that morphogenesis of an epithelium, regulation of cell size, regulation of epithelial cell proliferation, cell-cell adhesion, and spermatogenesis were associated with PAAD carcinogenesis and progression (Figure 8).

Functional enrichment analysis of EphA2 in pancreatic adenocarcinoma (PAAD) patients.
IHC analysis of EphA2 expression in PAAD patient samples
IHC assays showed high expression of EphA2 protein in 80% of the pancreatic cancer tissues examined (12/15), which was higher than in paracancer tissues (2/15; 13.34%). This difference was statistically significant (χ2 = 13.393, P < 0.001). These results revealed increased EphA2 protein expression in pancreatic cancer samples compared with paracancer tissues. Representative IHC staining images are shown in Figure 9.
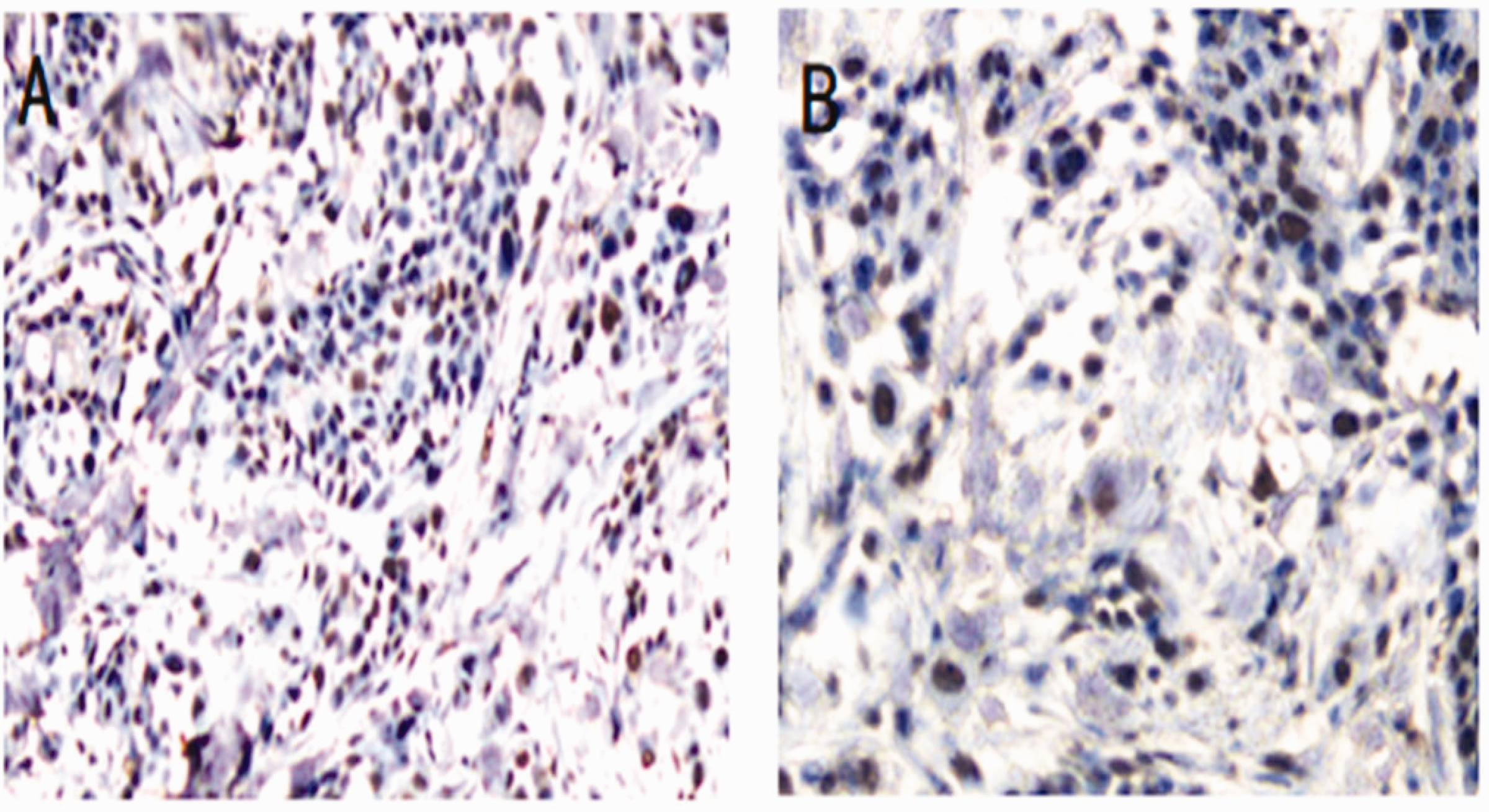
Analysis of EphA2 protein expression using immunohistochemistry (IHC) in tissue samples from pancreatic adenocarcinoma (PAAD) patients. Representative images are shown at 100x magnification. (a) High EphA2 protein expression in pancreatic cancer tissue and (b) Low EphA2 protein expression in paracancerous tissue.
Discussion
Pancreatic cancer is the 12th most common malignant tumor worldwide. It ranks seventh for cancer-related deaths, with global morbidity and mortality rates increasing each year. 7 It is necessary to identify new tumor biomarkers with strong sensitivity and specificity to improve pancreatic cancer patient survival rates. 8 High EphA2 expression levels have been observed in glioblastoma multiforme, prostate cancer, ovarian cancer, uterine cancer, gastric cancer, melanoma, and breast cancer.3,9–13 EphA2 expression was shown to be closely related to tumor cell proliferation, invasion, and metastasis. In certain cancers, EphA2 was linked to poor prognosis and decreased patient survival. 4 However, there are few reports on the function and mechanism of EphA2 in pancreatic cancer. In this study, we analyzed EphA2 expression in pancreatic cancer tissues using bioinformatics methods, then systematically evaluated its prognostic value in this disease.
Here, we examined the expression patterns of EphA2 in PAAD. The results show that PAAD tumor tissues have aberrantly high EphA2 expression levels compared with normal tissues. We also observed a significant correlation between EphA2 expression and PAAD pathological stage. PAAD patients with high EphA2 expression had a poor prognosis, as shown by both OS and DFS rates. These data suggest that EphA2 has an essential role and clinical prognostic value in PAAD. Brantley-Sieders et al. reported that EphA2 expression is correlated with poor survival in basal-like breast cancers. 14 Moreover, the EphA2 gene methylation levels showed a higher trend in PAAD samples compared with normal tissues, but this difference was not statistically significant. These results demonstrated that EphA2 methylation may not be the relevant mechanism in PAAD.
Pancreatic tumors have been reported to have low immunogenicity and high immunosuppression, leading to poor clinical immunotherapy effects. This is related to the immunosuppression and immune deficiency of the pancreatic cancer microenvironment. 15 Therefore, we evaluated any potential correlation between immune cell infiltration and EphA2 expression in PAAD. In this study, we found a significantly positive association between EphA2 expression and CD4+ T cell infiltration, suggesting that EphA2 may be a promising biomarker for immune therapy. Furthermore, age, CD4+ T cell infiltration, and EphA2 gene expression were positively correlated with PAAD prognosis. Lymph node metastasis and late stage have been previously identified as factors affecting the prognosis of pancreatic cancer patients.16,17 The sample size may lead to bias in the statistical results, which would be inconsistent with those of relevant reports in the literature. Our results identified EphA2 expression as an independent prognostic factor of PAAD, which was consistent with related literature reports. 12 Additionally, IHC assays showed high protein expression of EphA2 in pancreatic cancer tissues. Thus, EphA2 is not only a potential prognostic biomarker, but may also participate in immune regulation in PAAD.
The function of EphA2 in pancreatic cancer is unclear. Huang et al. showed that EphA2 can induce chemotherapy resistance in tumor cell lines by increasing yes-associated protein stability and nuclear localization. 18 Recent studies have associated EphA2 protein in gastric tumors with the yes-associated protein, a transcriptional coactivator that can promote cell proliferation, stem cell maintenance, and tissue homeostasis. 19 Our results confirmed that EphA2 functions through interactions with various proteins, including ENFNA1, ACP1, ENFNA1, and ENFNA2. These data may provide new insights for studying EphA2 function in PAAD. We also conducted a functional enrichment analysis of EphA2 with its interacting proteins using GO/KEGG enrichment items. Our findings showed that these proteins were enriched in cancer development and progression-related functions, like the ephrin receptor signaling pathway and ribonucleoprotein complex biogenesis. The EphA receptors play key roles in multiple mechanisms of cancer biology and are also likely implicated in stem cell biology of the tumor microenvironment.4,20 Overall, these data suggest that EphA2 has a significant role in pancreatic cancer development and is a potential therapeutic target for this disease. This work provides new evidence and directions for better understanding the mechanism of EphA2 in pancreatic cancer.
Conclusions
Taken together, these data imply that EphA2 is highly expressed in PAAD tissues, which may promote the occurrence and development of this cancer. EphA2 expression is closely related to PAAD patient prognosis. EphA2 is therefore a potential novel target and biomarker for the diagnosis and treatment of PAAD. However, additional validation experiments are needed in the future.
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231218559 - Supplemental material for Identification of therapeutic targets and prognostic biomarkers of the ephrin receptor subfamily in pancreatic adenocarcinoma
Supplemental material, sj-pdf-1-imr-10.1177_03000605231218559 for Identification of therapeutic targets and prognostic biomarkers of the ephrin receptor subfamily in pancreatic adenocarcinoma by Jingjng Zou, Kun Zhang, Jinde Zhu, Chaoyong Tu and Jingqiang Guo in Journal of International Medical Research
Footnotes
Author contributions
CT and JG designed the study. J. Zou completed the first draft of the manuscript, performed the experiments, and collected the data. KZ and J. Zhu assisted with data collection. JG analyzed the data and revised the manuscript. All authors have read and approved the final version of the manuscript.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
This study was supported by the Natural Science Foundation of Zhejiang (#LY19H030004) and the Project of Lishui City Key Research and Development (2022ZDYF08).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
