Abstract
Objectives
Mitofusion-2 (Mfn2) may have a role in mitochondrial oxidative stress and insulin resistance that can promote the development of metabolic dysfunction associated fatty liver disease (MAFLD). This retrospective and case control study aimed to explore the relationships between common Mfn2 single nucleotide polymorphisms (SNPs) and MAFLD in a northern Han Chinese population.
Methods
Six Mfn2 SNPs (rs2336384, rs873458, rs873457, rs4846085, rs2878677, and rs2236057) were genotyped using the ligase detection reaction in 466 MAFLD patients and 423 healthy controls. Genotype and allele frequencies were calculated, along with haplotype analysis and pairwise linkage disequilibrium.
Results
The genotype distribution of rs2336384, rs2878677, and rs2236057 among the MAFLD patients showed a significantly different pattern from that of healthy controls. The data showed that an increased risk of MAFLD was significantly correlated with patients carrying the GG genotype of rs2336384, CC genotype of rs873457, TT genotype of rs4846085, TT genotype of rs2878677, and the AA genotype of rs2236057. Moreover, The
Conclusion
The current findings suggest a strong link between certain Mfn2 gene polymorphisms and MAFLD.
Keywords
Introduction
Previously known as non-alcoholic fatty liver disease (NAFLD), metabolic dysfunction associated fatty liver disease (MAFLD) is a chronic condition characterized by hepatic fat accumulation combined with underlying metabolic dysregulation. 1 It is the most common chronic liver disease worldwide. 2 A quarter of the adult population worldwide is affected by MAFLD, resulting in a significant clinical and financial burden.3,4 The high prevalence of this disease may be at least partly caused by increasingly sedentary lifestyles with little physical exercise, overnutrition, and unhealthy diets. 5 However, there is evidence suggesting that MAFLD is a multifactorial disease influenced not only by environmental determinants, but also by genetic predisposition, which promotes geographical demarcation of the disorder. 4 Thus, there is a need to more precisely define patient phenotypes and genetic backgrounds to develop effective treatment strategies.6,7 Many studies have aimed to identify the specific susceptibility loci for fatty liver disease. Patatin-like phospholipase domain-containing protein 3 (PNPLA3) and transmembrane 6 superfamily member 2 (TM6SF2) have been associated with MAFLD. 8 There are some indications that Mitofusion-2 (MFN2) is associated with MAFLD, however, information concerning the connection between Mfn2 gene polymorphisms and MAFLD is limited.
The Mfn2 gene maps to chromosome 1p36.22 and encodes a protein with GTPase activity in the mitochondrial outer membrane. Mitochondria produce most of the ATP required for metabolism, and the aerobic nature of ATP generation also produces reactive oxygen species (ROS). Thus, mitochondrial dysfunction may result in damage to liver cells. 9 The Mfn2 gene plays roles in mitochondrial fusion, division, energy generation, and thereby cell metabolism. Therefore, aberrant expression of Mfn2 may promote oxidative and endoplasmic reticulum (ER) stress, resulting in liver cell damage. 10 Several studies have linked Mfn2 to insulin resistance (IR), 11 mitophagy, 12 and the proliferation of vascular smooth muscle cells,13,14 all of which are associated with metabolic disorders. We speculated that Mfn2 may have an impact on MAFLD via the pathological processes outlined above. Previous studies have examined the associations between Mfn2 gene polymorphisms and metabolic syndrome diseases, such as type 2 diabetes 15 and essential hypertension.16–18 However, there have been few reported indications of any correlation between Mfn2 single nucleotide polymorphisms (SNPs) and MAFLD. In this study, we aimed to address these deficiencies by performing an in-depth investigation of the relationship between Mfn2 polymorphisms and MAFLD in a northern Han Chinese population.
Materials and methods
Subjects
Subjects presenting at the physical examination center of Third Hospital of Hebei Medical University, Shijiazhuang, China between October and December 2020 were enrolled in the study. All individuals were of northern Han descent and there was no evidence of intermarriage within the cohort. The new diagnostic criteria for MAFLD were applied, with hepatic steatosis being identified by two clinical ultrasound experts. 2 All subjects were over 18 years of age. Subjects who had undergone liver surgery, had forms of liver disease other than MAFLD, or had any other disease likely to cause fatty liver were excluded. The retrospective study was approved and supervised by the Ethics Committee of the Third Hospital of Hebei Medical University (approval number: 2019-005-1; date: 2019-4-23) and complied with the Principles of the Helsinki Declaration. All participants provided written informed consent for genetic testing. We have de-identified all patient details in this report. The reporting of this study conforms to STROBE guidelines. 19
Baseline information, including age, sex, medical history, and family history of MAFLD, was collected by questionnaire. Patient height and body weight were measured using an electronic instrument (Omron, Shenzhen, China) and the body mass index (BMI) was calculated according to the formula: body weight (kg)/height 2 (m2). Electronic sphygmomanometers were used to measure blood pressure (Citizen, Jiangmen, Japan). Venous blood was collected in the morning after fasting for 8 to 12 hours. An automated blood cell analyzer was used to produce a white blood cell (WBC) count and an automated biochemical analyzer was used to generate liver biochemical indices (Beckman Coulter, Brea, CA, USA).
SNP identification and genotyping
Six Mfn2 SNPs (rs2336384, rs873458, rs873457, rs4846085, rs2878677, and rs2236057) were selected from previous reports of variants associated with metabolic dysfunction.15–19 Rs2336384, rs873458, rs873457, and rs4846085 are located within intron 2, rs2878677 within intron 3, and rs2236057 within intron 11 of Mfn2. Genomic DNA was extracted from peripheral blood leukocytes using a genomic DNA kit (GENEray Biotechnology, Shanghai, China). Mfn2 gene polymorphisms were detected using the ligase detection reaction (LDR). Briefly, after extracting the DNA from the peripheral blood leukocytes, PCR amplification was performed using the following conditions: an initial denaturation of 5 minutes at 95°C, followed by 35 cycles of denaturation at 94°C for 20 s, annealing at 55°C for 20 s, and extension at 72°C for 40 s, with a final extension at 72°C for 10 minutes. For the LDR, the probe was added to each PCR reaction well, mixed, spun by centrifugation, and amplified for 30 cycles at 94°C for 20 s and 58°C for 90 s. Then, 1 µL of that product was added to 9 µL of highly deionized formamide, desaturated at 95°C for 3 minutes, plunged into an ice bath, and tested immediately.
Primer Premier 5.0 software (Premier Biosoft Intl., San Francisco, CA, USA) was used to design primers for amplification based on the Mfn2 Genbank reference sequence (NG_007945.1; Supplementary Table 1). Probes were designed using GENEray Biotechnology (Shanghai, China; Supplementary Table 2). Genotypes were evaluated using DNA sequencing on an ABI 3730 genetic analyzer (Applied Biosystems, Waltham, MA, USA) following the manufacturer’s instructions, and sequenced with GeneMarker V2.2.0 (SoftGenetics, LLC., State College, PA, USA).
Statistical analyses
Statistical analyses were performed using IBM SPSS Statistics software, version 26.0 (IBM Corp., Armonk, NY, USA). Continuous variables are presented as the mean ± standard deviation (SD), skewed variables as the median (interquartile range), and categorical variables as percentage. Student’s t-tests, Mann–Whitney U tests, and chi-squared tests were used, respectively, to compare paired groups. Risk estimates for MAFLD with respect to the six Mfn2 SNPs for different genetic models (additive, recessive, dominant, and co-dominant models) were determined by binary logistic regression. Hardy–Weinberg equilibrium (HWE), linkage disequilibrium (LD), and haplotype blocks were assessed using Haploview software 4.2 (r2 was set to 0.85) (http://www.broad.mit.edu/mpg/haploview/). A two-tailed P-value < 0.05 was considered statistically significant.
Results
Characteristics of the subjects
A total of 889 unrelated participants were enrolled in the study, including 466 MAFLD cases (208 men and 258 women) and 423 healthy controls (162 men and 261 women). A flow chart of the study protocol is shown in Figure 1. The clinical characteristics and laboratory data for all participants are shown in Table 1. There was adequate sex matching between MAFLD subjects and controls. The mean age of the MAFLD group was higher than that of controls (P < 0.001). The mean values of the following variables were significantly higher among the MAFLD patients compared with the controls: BMI, diastolic blood pressure (DBP), systolic blood pressure (SBP), WBC count, and levels of alanine transaminase (ALT), aspartate transaminase (AST), total cholesterol (TC), triglyceride (TG), low-density lipoprotein cholesterol (LDL-C), fasting plasma glucose (FPG), and uric acid (UA) (P < 0.001; Table 1). The mean values for high-density lipoprotein cholesterol (HDL-C) levels were lower in the MAFLD group (P < 0.001). The mean plasma albumin concentrations were significantly lower among women and in the entire MAFLD cohort (P < 0.001), but not among male MAFLD patients.

Flowchart showing the patient enrollment process in this study.
Demographic and biochemical characteristics of the healthy controls and metabolic dysfunction associated fatty liver disease (MAFLD) patients.

Continuous variables are expressed as the mean ± standard deviation when normally distributed and as the median (interquartile range) when non-normally distributed. MAFLD, metabolic associated fatty liver disease; BMI, body mass index; DBP, diastolic blood pressure; SBP, systolic blood pressure; WBC, white blood cell count; ALB, albumin; ALT, alanine transaminase; AST, aspartate transaminase; TC, total cholesterol; TG, triglyceride; HDL-C, high-density lipoprotein; LDL-C, low-density lipoprotein; FPG, fasting plasma glucose; UA, uric acid.
Detection and distribution of SNPs
All six Mfn2 SNPs were successfully genotyped for all participants (Table 2). All SNPs showed genotype frequencies in accordance with HWE in the control group (Table 2). Chi-square analyses indicated significant differences in the distributions of the rs2336384, rs2878677, and rs2236057 genotypes in MAFLD patients compared with healthy controls (P < 0.05). For rs2878677, the C allele prevalence was significantly higher among MAFLD patients (P < 0.05), whereas no differences were found for the distributions of rs2336384 and rs2236057. Analysis by sex revealed a different genotype distribution for rs2878677 between female MAFLD patients and controls. No sex-based differences in the allele ratio was found.
Genotype distribution and allele frequency of the Mfn2 gene in healthy controls and metabolic dysfunction associated fatty liver disease (MAFLD) patients.

SNP, single nucleotide polymorphism.
Correlation analyses
Different genetic models, additive, recessive, dominant, and co-dominant, were assessed by logistic regression analyses. The rs2336384 polymorphism was inversely correlated with MAFLD when analyzed by both the recessive genetic model (GG vs. GT + TT: P = 0.006; odds ratio (OR) = 1.585; 95% confidence interval (CI) = 1.138–2.207) and co-dominant genetic model (GG + TT vs. GT: P = 0.022; OR = 1.365; 95% CI = 1.047–1.781), suggesting that having the G allele of rs2336384 carries a higher risk of MAFLD (Table 3). Furthermore, analysis by the additive genetic model (TT vs. CT vs. CC: P = 0.025; OR = 1.247; 95% CI = 1.028–1.513) and the recessive genetic model (TT vs. CT + CC: P = 0.002; OR = 1.722; 95% CI = 1.229–2.415) demonstrated an increased risk of MAFLD for subjects carrying the TT genotype of the rs2878677 polymorphism. SNPs of rs873457, rs4846085, and rs2236057 were inversely correlated with MAFLD in recessive genetic models (rs873457: CC vs. CG + GG: P = 0.017; OR = 1.485; 95% CI = 1.073–2.056; rs4846085: TT vs. CT +CC: P = 0.048; OR = 1.388; 95% CI = 1.003–1.921; rs2236057: AA vs. AG + GG: P = 0.012; OR = 1.560; 95% CI = 1.102–2.208). No significant association between the rs873458 polymorphism and MAFLD risk was found (Table 3). Sex-based subgroup analyses revealed a different finding in recessive modelling of rs2336384 in men (GG vs. GT + TT: P = 0.040; OR = 1.772; 95% CI = 1.026–3.060) and recessive and co-dominant modelling of rs2878677 in women (TT vs. CT + CC: P = 0.009; OR = 1.791; 95% CI = 1.159–2.768; TT + CC vs. CT: P = 0.039; OR = 1.439; 95% CI = 1.018–2.033; Table 3).
Association of Mfn2 gene polymorphisms with metabolic dysfunction associated fatty liver disease (MAFLD) interpreted by different genetic models.

SNP, single nucleotide polymorphism; OR, odds ratio; CI, confidence interval.
Haplotype analysis
Calculations of r2 were made for all SNPs under investigation. The Haploview program showed that all six Mfn2 polymorphisms were located in a strong LD block and were interlinked with each other (Figure 2). The full haplotype analyses of all six Mfn2 polymorphisms in MAFLD patients and controls are presented in Table 4. The haplotype, GGCTTA (rs2336384 - rs873457 - rs873458 - rs4846085 - rs2878677 - rs2236057), was lower among MAFLD patients (39.7%) than in the controls (45.2%; P = 0.029). No significant association was found between other haplotypes and MAFLD risk.
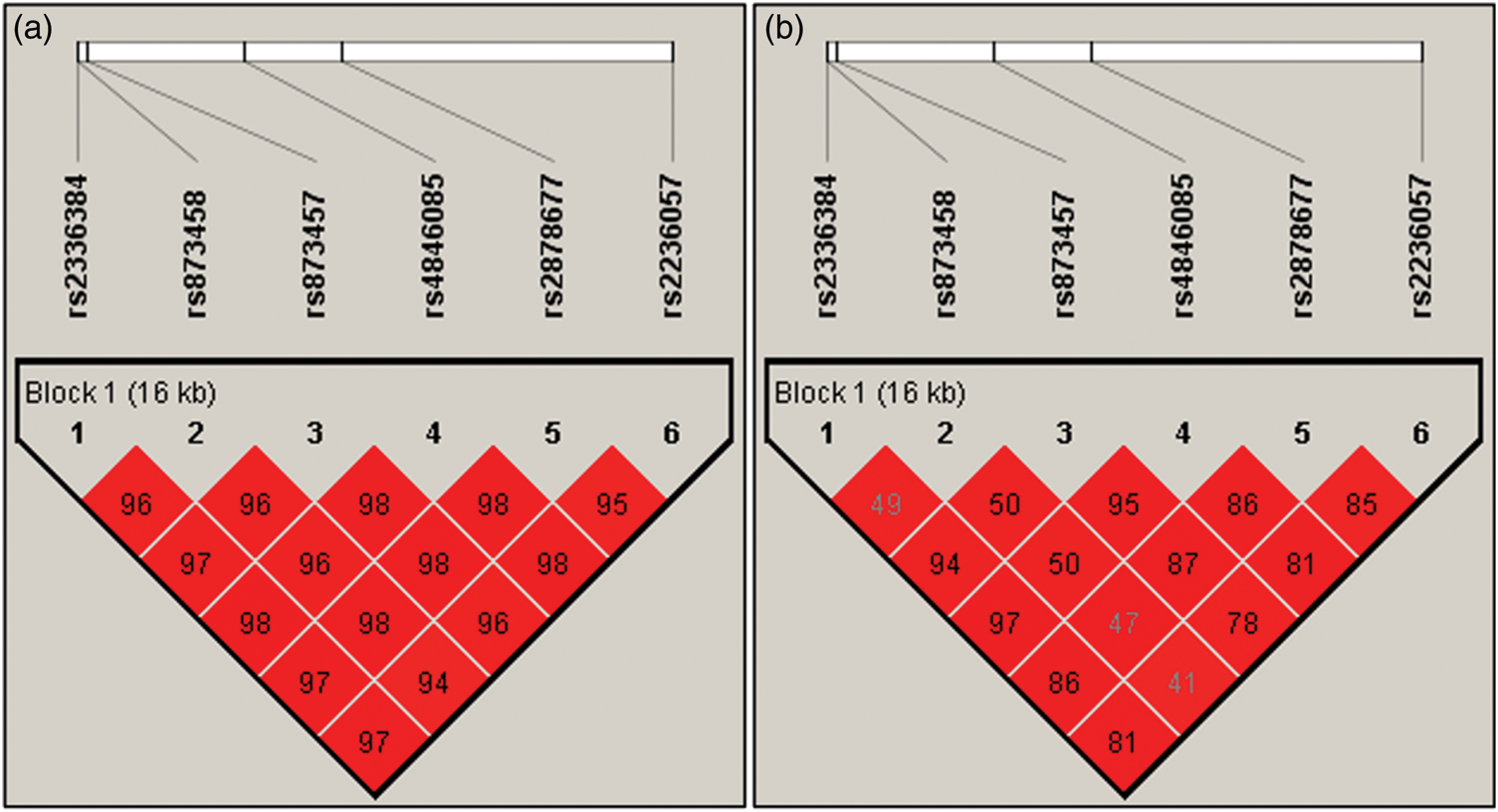
Linkage disequilibrium (LD) block defined by the Haploview program using the solid spine of LD method. (a) LD of D′ and (b) LD of r2.
Haplotype analyses of the Mfn2 polymorphisms in healthy controls and metabolic dysfunction associated fatty liver disease (MAFLD) patients

M1: rs2336384, M2: rs873458, M3: rs873457, M4: 4846085, M5: 2878677, M6: 2236057.
Discussion
In this study, we identified and genotyped six SNPs of the Mfn2 gene using Haploview software and SPSS statistical analysis to assess the risk factors for MAFLD by binary logistic regression analyses. Our findings show that Mfn2 polymorphisms rs2336384, rs2878677, and rs2236057 are strongly correlated with MAFLD risk in the Han Chinese population. Moreover, the haplotype, GGCTTA, appears to be a protective factor for MAFLD. Sex-specific subgroup analysis demonstrated the relationship between the rs2236384 and rs2878677 polymorphisms and risk of MAFLD in men and women, respectively. To our knowledge, the current study is the first to focus on Mfn2 gene polymorphisms and MAFLD in the Chinese population before or because MAFLD has been newly defined.
Several previous studies have investigated associations between Mfn2 gene polymorphisms and diseases related to metabolic dysfunction. In 2011, Wang et al. 16 and Li et al. 17 found that Mfn2 gene polymorphisms (rs873457, rs2336384, rs147486, rs4846085, rs2236055 16 and rs2336384, rs2295281, rs17037564, rs2236057, rs2236058, rs3766741, 17 respectively) were associated with the sub-groups of MAFLD patients who have essential hypertension (EH) in Chinese subjects. In addition, Liu et al. 15 established a link between Mfn2 polymorphisms (rs873458, rs2878677, rs2236058, rs3766742, and rs3766741) and type 2 diabetes in Han Chinese people. In combination, these findings suggest that Mfn2 gene polymorphisms play a major role in the occurrence and development of metabolism-related diseases. However, the studies listed above do not demonstrate a direct association between SNPs and MAFLD. To bridge this gap in knowledge, we selected six representative SNPs in the Mfn2 gene and explored the links between them and MAFLD in patients from a northern Han Chinese population. It was considered that the population being studied shows consistent characteristics of geographical stability, environmental exposure, and genetic homogeneity.20,21 It is vital to exclude the influence of such factors when investigating the genetics of a metabolic disorder, such as MAFLD, and our findings were consistent with those of previous studies on metabolic dysfunction.15–18,22 Our current findings suggest that Mfn2 gene polymorphisms can be correlated with the risk of MAFLD.
Subgroup analysis revealed a degree of sex-specificity regarding the impact of Mfn2 polymorphisms. We found an association between rs2336384 and MAFLD risk in men and a similar association for rs2878677 in women. There are several possible reasons for these sex-specific differences: 1) Differing genetic structure between women and men 23 may influence MAFLD susceptibility genes; 2) Regulation of Mfn2 gene expression may be different in men compared with women because of varying rates of smoking, drinking, mental stress, low estrogen levels, and other reasons; 24 3) Some receptors relevant to the current study may have sex-specific activities, such as peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC-1α), peroxisome proliferator-activated receptor alpha (PPARα), farnesoid X receptor (FXR), and liver X receptor (LXR), all of which may contribute to the steatosis that characterizes MAFLD.24,25 Any or all of these risk factors could be hypothesized to interact with the Mfn2 gene variation and influence the regulation of fatty acid metabolism.
While a single SNP has a vanishingly small influence on phenotype, multiple SNPs, especially when closely linked, may have a large collective impact. Hence, haplotype analysis can contribute more to genetic studies of complex diseases, such as MAFLD, than studies of single SNPs. The six SNPs examined here were located in one block and displayed strong LD with each other. Haplotype analysis indicated an association of GGCTTA (rs2336384 - rs873458 - rs873457 - rs4846085 - rs2878677 - rs2236057) with a decreased risk of MAFLD. We believe that the current report is the first to identify these six Mfn2 polymorphisms (rs2336384, rs873458, rs873457, rs4846085, rs2878677, and rs2236057) as a protective haplotype block for MAFLD. Moreover, we found that the haplotypes, TAGCCG, TGGCCG, and GGCTCG, are more likely to be associated with an increased risk of MAFLD, albeit there was no difference between the MAFLD and control groups.
The complex and multifactorial development of MAFLD is characterized by pathophysiological mechanisms, including disordered lipid metabolism, oxidative stress, and IR. Mfn2 has a prominent regulatory role in this process13,26 and shows reduced activity in obesity and type 2 diabetes.27,28 Indeed, Gan et al. demonstrated an impact of Mfn2 protein deficiency on disordered mitochondrial architecture, decreased oxidative capacity, and glucose-derived pyruvate in mice. These phenotypes could be reversed by overexpressing Mfn2. The insulin signaling pathway appeared to be the target of these effects.11,26 Insulin may block the Mfn2-dependent mitogen-activated protein kinase kinase 1 (MEK) cascade and activate the phosphoinositol 3-kinase/protein kinase B (PI3K/Akt) signaling pathway, which stimulates mitochondrial activity and promotes mitochondrial fusion. 29 Conversely, downregulation of Mfn2 can decrease mitochondrial β-oxidation of fatty acids, enhancing TG accumulation and impairing the respiratory chain. This ultimately leads to ROS generation. Lower Mfn2 activity can also result in activation of the JNK and p38MAPK pathways and inhibition of the Akt pathway. The cumulative effect of these alterations can promote IR in both skeletal muscle and liver.30,31 Ablation of Mfn2 in hypothalamic proopiomelanocortin neurons has also been shown to lead to ER stress, leptin resistance, and consequentially hyperphagia, reduced energy expenditure, and obesity. All of these effects could be reversed by relieving hypothalamic ER stress. 32 Furthermore, lipotoxin damage induced by saturated lipids reduces Mfn2 expression, stimulating the ER stress response and IR in hypothalamic-derived cells. 33 Taken together, these findings confirm the role for Mfn2 and its impact on ER stress in local and systemic energy balance.
To summarize, fatty acids and glucose-derived pyruvate are metabolized in mitochondria through the tricarboxylic acid cycle. An increase in glucose metabolism, along with reduced fatty acid oxidation, stimulates ROS production and causes decreased Mfn2 expression. Downstream consequences include activation of mitochondrial fission, suppression of mitochondrial calcium release, impaired mitochondrial metabolism, and reduced mitochondrial membrane potential. The resulting oxidative stress and ER stress promote mitochondrial apoptosis, thereby enhancing the metabolic disorders that lead to MAFLD.34–38 The current genetic analysis, in combination with evidence from previous studies, supports the conclusion that Mfn2 influences the occurrence of MAFLD and is a potential therapeutic target for this disorder.
Introns are crucial for the regulation of eukaryotic gene expression by interacting with enhancers or alternative promoters of transcriptional regulatory elements, or through the intron-mediated enhancement that increases the processivity of the transcription machinery at the elongation stage. 39 There are many reports of genes with complete promoters that fail to be expressed without introns. 40 The current study identified three positive SNPs in Mfn2 gene introns that were significantly associated with MAFLD. It is likely that these polymorphisms have an impact on the transcriptional regulation of Mfn2. Further mechanistic studies should be performed in cultured cells using transfection to introduce different polymorphisms of the Mfn2 introns to assess their impact on gene expression.
This study has some limitations. The subjects were exclusively of northern Han origin, and it is unclear how our findings may apply to other populations. Only relatively severe cases of MAFLD were enrolled and matched with control participants who were not overweight or obese. Some selection bias may have been introduced because the study participants were more health-conscious than the general population. Moreover, there was some deficiency in data collection, such as detailed alcohol consumption. Any correlations between Mfn2 gene polymorphisms and MAFLD biomarkers were not examined. We intend to focus on this aspect in future work to identify any potential associations.
Conclusions
The current study revealed that SNPs of the Mfn2 gene (rs2336384, rs2878677, and rs2236057) and the haplotype,
Supplemental Material
sj-pdf-1-imr-10.1177_03000605231187953 - Supplemental material for Mitofusin-2 gene polymorphisms and metabolic dysfunction associated fatty liver disease: a case-control study in a Chinese population
Supplemental material, sj-pdf-1-imr-10.1177_03000605231187953 for Mitofusin-2 gene polymorphisms and metabolic dysfunction associated fatty liver disease: a case-control study in a Chinese population by Xiwei Yuan, Mengmeng Hou, Yiqi Wang, Siyu Zhang, Lu Li, Yingjun Mi, Huijuan Du, Songhao Yu and Yuemin Nan in Journal of International Medical Research
Footnotes
Acknowledgements
Author contributions
YN contributed to the study design and revised the manuscript draft. XY performed statistical analysis and wrote the manuscript draft. MH, YW, and SY collected the data and performed the study. SZ and LL verified the statistical analysis. YM and HD provided technical support. All authors have read and approved the final manuscript.
Data availability statement
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
The study was supported by National Natural Science Foundation of China (Grant No. 81970504), Natural Science Foundation of Hebei Province (Grant No. H2018206326), and Key Research and Development Program of Hebei Province (Grant No. 19277779D).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
