Abstract
Objectives
Docking Protein 3 (DOK3) is an adapter protein that has been implicated in various cellular processes relevant to diseases, such as cancer. In this study, we aimed to evaluate the role of DOK3 in kidney renal clear cell carcinoma (KIRC) by examining how its expression levels are correlated with patient characteristics and prognosis.
Methods
We analyzed KIRC-related data from The Cancer Genome Atlas and used several bioinformatics tools, such as LinkedOmics and Oncomine, to evaluate DOK3 mRNA expression in KIRC. DOK3 protein expression was examined in 150 clinical KIRC samples and 100 non-cancerous renal tissues with immunohistochemistry assays. The prognostic value of DOK3 mRNA expression on patient overall survival was analyzed retrospectively using Kaplan–Meier survival and Cox regression analyses.
Results
DOK3 mRNA expression was notably higher in KIRC samples compared with normal tissues. Significant correlations were found between DOK3 mRNA expression levels and tumor size, lymph node metastasis, distant metastasis, and pathological grade using the bioinformatics data. This was confirmed at the protein level with immunohistochemistry data. Survival analyses indicated that elevated DOK3 expression is linked to a lower overall survival rate in KIRC patients.
Conclusions
DOK3 is a potential biomarker for determining KIRC patient clinical prognosis.
Keywords
Introduction
Kidney cancer is the second most common malignant tumor of the urinary system, with an incidence only behind bladder cancer according to epidemiological surveys.1,2 Renal clear cell carcinoma was the most common pathological type, accounting for about 70% to 80% of kidney cancers.3,4 The incidence of kidney renal clear cell carcinoma (KIRC) has still been increasing in recent years. 5 Data suggest that 30% of KIRC patients had already developed metastases before diagnosis. 6 KIRC is insensitive to both radiotherapy and chemotherapy, but the introduction of immune checkpoint inhibitors (ICIs) has allowed various combinations of ICIs and drugs to become the first-line treatment modality for this disease. Cabozantinib and sunitinib are commonly used in combinations and have had satisfactory results in improving early mortality in metastatic renal clear cell carcinoma patients, significantly increasing their 5-year survival rate.7–9 Overall, KIRC remains a major challenge because of its poor prognosis and high early mortality. 10 Therefore, identifying reliable biomarkers for KIRC is essential to ensure early diagnosis and accurate prognostic assessment.
Docking Protein 3 (DOK3), which is encoded by the DOK3 gene, is associated with the immune response and is involved in regulating various signaling pathways downstream of immune receptors.11,12 For example, it participates in the formation of the tumor necrosis factor receptor-associated factor 3/TANK-binding kinase 1 complex in macrophages, which induces interferon-β production. 13 DOK3 also has an important role in the differentiation of plasma cells. 14 Additionally, this protein has significant involvement in a variety of cancers and has been shown to inhibit the proliferation and invasion rates of tumor cells.15,16 Berger et al. found that DOK3 expression levels were lower in lung adenocarcinoma samples than in the normal tissue adjacent to the tumor. Furthermore, knocking out the DOK3 gene accelerated the proliferation and survival of cancer cells. 17 Collectively, these previous studies demonstrate the importance of the DOK3 gene in tumorigenesis and assessing prognosis.
In this study, we aimed to investigate DOK3 expression in KIRC, determine its relationship with clinicopathological factors, and focus on its prognostic impact. First, we assessed DOK3 mRNA expression in KIRC by bioinformatics analysis using public databases. We then confirmed its role in 150 KIRC cases using immunohistochemistry (IHC) assays. Our results suggest that DOK3 may be a valuable biomarker for determining KIRC patient prognosis and assessing their treatment options.
Materials and methods
Oncomine
We examined data in the Oncomine database (https://www.oncomine.org) and filtered the results for the DOK3 gene in KIRC using the following criteria: 1) “Cancer Type: renal cell carcinoma”; 2) “Gene: DOK3”; 3) “Data Type: mRNA”; 4) “Analysis Type: Cancer vs Normal”; 5) Others: P-value <0.0001, fold change >2, gene rank = top 10%.
Statistical analysis
The Statistical Package for the Social Sciences (Version 26.0; IBM Corp., Armonk, NY, USA) was used to perform all statistical analyses. Categorical data were analyzed by χ2 or Fisher’s exact tests. Survival analysis was performed using the Kaplan–Meier (KM) method combined with the log-rank test. All tests were two-tailed statistical tests, with P-values less than 0.05 considered statistically significant.
Patient samples
Renal clear cell carcinoma tissue samples were collected from patients who underwent surgical resection at the Zhejiang Provincial People’s Hospital, Hangzhou, China, between February 2008 and March 2010. None of the patients had undergone radiotherapy or chemotherapy prior to the operation. Non-cancerous renal tissues were also collected from patients. The samples were used to prepare tissue microarrays (TMAs), which were constructed by Shanghai Xinchao Biotechnology Co. Ltd. (Shanghai, China). The project was approved by the Ethics Committee of Zhejiang Provincial People’s Hospital (approval no.: QT2022424) and written informed consent was obtained from all participants. All patient details were removed to follow protocol and protect patient privacy throughout the process. The study was performed in accordance with the Declaration of Helsinki and is reported in accordance with STROBE guidelines. 18
Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses
LinkedOmics (http://www.linkedomics.org/login.php) uses data from The Cancer Genome Atlas (TCGA) database. Thirty-two cancer species were included for each type of data, which can be used to analyze the relationships between gene expression at the mRNA level and features such as DNA methylation status and mutation sites. In this study, we used LinkedOmics to obtain information on DOK3 in KIRC with the following filter conditions: 1) “Gene: DOK3”; 2) “Analysis Type: Cancer vs Normal analysis”; 3) “Data Type: mRNA”; 4) “Cancer Type: Renal cell carcinoma”; 5) “Gene Summary: P-value <0.05, fold change = all, gene rank = top 10%”; 6) “Statistical Method: Pearson correlation test”. We then selected the “Gene Set Enrichment Analysis (GSEA)” tools with the following conditions: “GO analysis (biological process)”; “GO analysis (cellular component)”; “GO analysis (molecular function)”; “KEGG pathway”.
Protein–protein interaction (PPI) networks
The proteins associated with DOK3 and their interaction network were analyzed using the Interacting Gene Search database search tool (http://string-db.org/). The necessary data were obtained by searching for protein names, species, and other necessary information.
IHC
IHC analysis was performed on 150 paraffin-embedded specimens with standard methods. The samples were first heated to 70°C for 2 hours, dewaxed with xylene, and dehydrated with different concentrations of ethanol. Each TMA was then placed in TE buffer and boiled for 3 minutes at 120°C in an autoclave for antigen retrieval. Endogenous peroxidase activity in the samples was blocked by incubation in 3% H2O2 at room temperature for 15 to 20 minutes. Samples were incubated with normal goat serum as a blocking agent for 15 to 20 minutes at room temperature to reduce non-specific reactions. The samples were incubated with a rabbit anti-DOK3 polyclonal antibody (ab236609, Abcam, Cambridge, UK) diluted to 1:1000 in phosphate-buffered saline at 4°C overnight. The next day, the sections were incubated at room temperature for 15 minutes with a horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG secondary antibody (A0208, Beyotime, Shanghai, China). The sections were stained with 3,3-diaminobenzidine (DAB) and the nuclei were counterstained with hematoxylin. The slides were then mounted under resin-covered glass for drying.
The stained samples were semi-quantitatively assessed by two pathologists without knowledge of clinical information. Experts independently scored the samples according to their staining intensity and the proportion of tumor cells positively stained. The staining intensity was scored as 0 for no staining, 1 for weak staining, 2 for moderate staining, and 3 for strong staining. The cell staining fraction was scored as 0 for no cell staining, 1 for 1% to 25% cell staining, 2 for 25% to 50% cell staining, 3 for 51% to 75% cell staining, and 4 for >75% cell staining. From these results, the color intensity was multiplied by the scale score to obtain a composite score for coloring, with a score ranging from 0 to 12. In this study, a score of ≤6 denoted low DOK3 expression levels, while a score of ≥7 denoted strong expression.
Results
KIRC samples had significantly higher DOK3 expression levels than normal tissues
The TCGA data suggested that DOK3 mRNA is expressed at different levels between tumor samples and normal tissues and was assessed using the UALCAN database (Figure 1a). The results indicate that DOK3 was expressed in samples of many cancer types, with a significant difference between tumor and normal tissues in diffuse large B-cell lymphoma (DLBC) and KIRC. We further used Oncomine to extract the necessary information and assess DOK3 expression in normal and KIRC tissues. The DOK3 mRNA expression levels were significantly higher in the Yusenko Renal cohort (three normal tissues, 26 cancer tissues) compared with renal clear cell carcinoma (P < 0.05, Figure 1b). In the Gumz Renal cohort (10 normal tissues, 10 cancerous tissues), DOK3 mRNA was expressed at significantly higher levels than in renal clear cell carcinoma (P < 0.05, Figure 1c). In addition, we analyzed DOK3 expression in 72 normal kidney tissues and 533 renal clear cell carcinoma tissues using UALCAN. The results revealed that DOK3 expression was higher in KIRC patients with advanced TNM than in KIRC patients with early TNM staging (Figure 1d–g).

Docking Protein 3 (DOK3) mRNA expression in kidney renal clear cell carcinoma (KIRC) and normal tissues. (a) DOK3 mRNA expression in various cancers using The Cancer Genome Atlas (TCGA) database (UALCAN). (b–c) Boxplots showing DOK3 mRNA levels in the (b) Yusenko Renal and (c) Gumz Renal datasets (Oncomine). (d–g) DOK3 transcription in subgroups of KIRC patients stratified by grade, stage, and other criteria (UALCAN). (d) Boxplot showing relative expression of DOK3 in KIRC and normal tissue samples. (e–g) Association between DOK3 expression and KIRC histological grade, including (e) nodal metastasis status, (f) TNM stage, and (g) tumor grade.
Overall, 150 renal clear cell carcinoma tissue samples were collected from patients, which included 108 male patients and 42 female patients, aged 24 to 83 years (mean age of 56.98 years). Moreover, 100 non-cancerous renal tissue samples were collected from patients, including 60 cases of renal lesion tissues and 40 cases of precancerous tissues (Table 1).
Docking Protein 3 (DOK3) mRNA expression in kidney renal clear cell carcinoma (KIRC) and non-carcinomatous renal tissues.

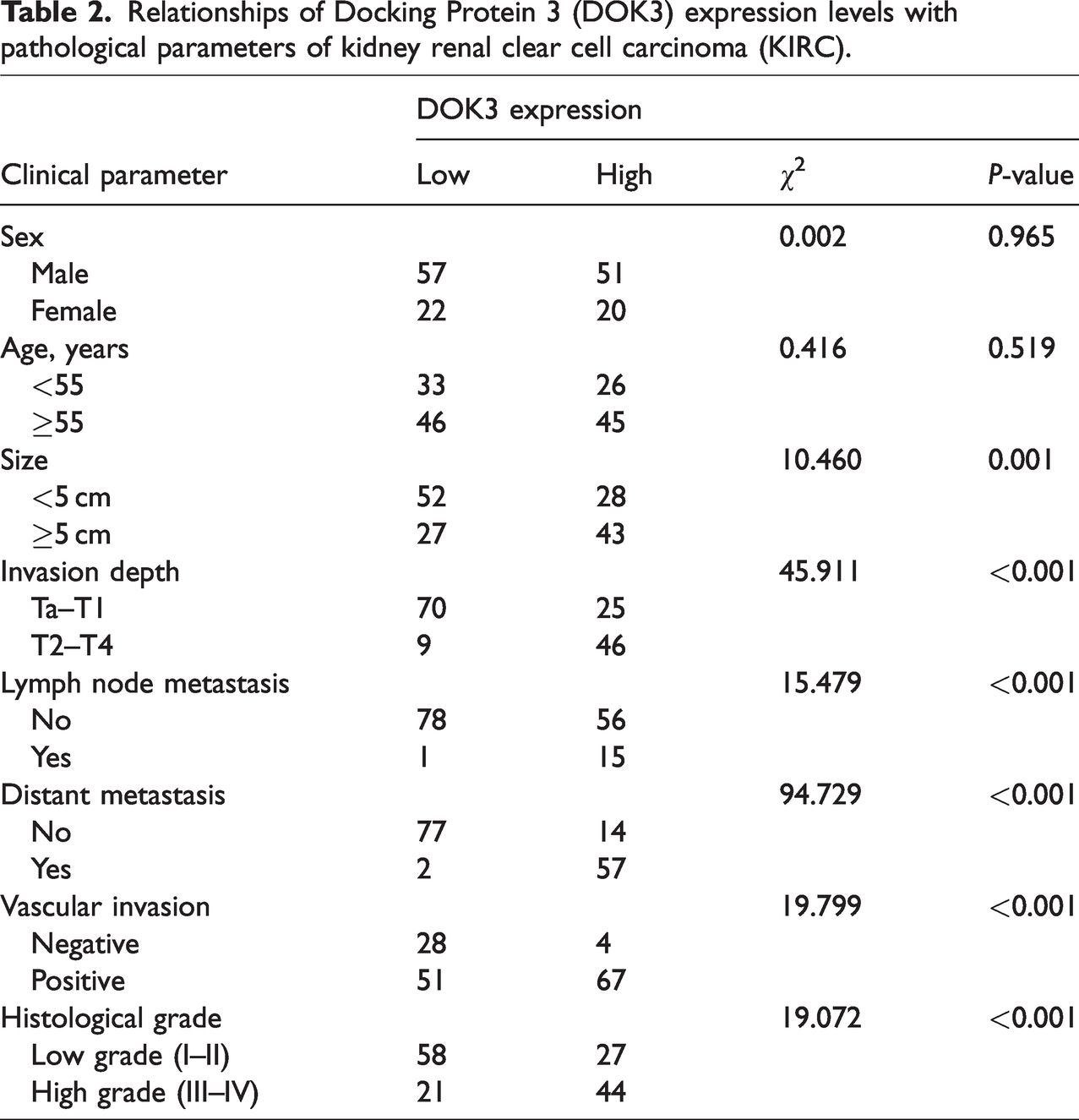
IHC staining confirmed that DOK3 protein was expressed in KIRC samples (Figure 2a–c). High DOK3 protein expression levels were detected in 71 cases (47.3%) in 150 kidney clear cell carcinoma tissue samples. After examining the correlations between DOK3 protein levels and the clinicopathological features of KIRC, we found that high DOK3 expression was associated with tumor size (P = 0.001), lymph node metastasis (P < 0.001), vascular invasion (P < 0.001), survival time (P = 0.033), distant metastasis (P < 0.001), and pathological grade (P < 0.001). However, there was no significant correlation between DOK3 expression and the other parameters examined, such as sex (P = 0.965) and age (P = 0.519) (Table 2).

(a) Low levels of Docking Protein 3 (DOK3) protein expression in kidney renal clear cell carcinoma (KIRC). (b) Immunohistochemistry staining of DOK3 in normal kidney tissue and (c) High DOK3 protein expression levels in KIRC. Magnification: 40× (A1–C1) and 400× (A2–C2).
Relationships of Docking Protein 3 (DOK3) expression levels with pathological parameters of kidney renal clear cell carcinoma (KIRC).
The clinical significance of DOK3 expression in KIRC prognosis
To examine the clinical importance of DOK3 expression levels in KIRC prognosis, the KM Plotter tool was employed. The data suggest that the high expression group had a significantly lower overall survival time than the low expression group (P < 0.05, Figure 3). The median survival time of patients with low DOK3 protein expression (55.77 ± 1.44 months) was significantly longer than that of patients with high DOK3 expression (46.35 ± 2.09 months). KM analyses of data from three databases all indicated that higher DOK3 mRNA expression levels were linked to a shorter overall survival time (P < 0.05, Figure 4). Moreover, we used univariate and multifactorial analyses to assess the prognostic factors affecting KIRC. The univariate analysis revealed that survival was associated with tumor size (hazard ratio (HR) = 3.918; 95% confidence interval (CI) = 1.665–9.221; P < 0.01), lymph node metastasis (HR = 4.575; 95% CI = 1.813–11.542; P < 0.01), distant metastasis (HR = 2.340; 95% CI = 1.108–4.941; P < 0.01), TNM stage (HR = 5.948; 95% CI = 2.607–13.569; P < 0.01), and pathological grade (HR = 4.152; 95% CI = 1.824–9.449; P < 0.01) (Table 3). Using multifactorial analysis to exclude confounding factors, the results showed that lymph node metastasis (HR = 4.892; 95% CI = 1.767–13.548; P = 0.002) and TNM stage (HR = 4.609; 95% CI = 1.546–13.743; P = 0.006) were independent prognostic factors affecting overall survival in patients with KIRC. Although DOK3 expression was not identified as an independent prognostic factor, its high expression plays a negative role in KIRC development and may be a factor affecting prognosis (Table 4).

Kaplan–Meier survival curves showing that kidney renal clear cell carcinoma (KIRC) patients with high expression levels of Docking Protein 3 (DOK3) have a poorer prognosis than those with low DOK3 expression levels (P < 0.05).

The prognostic value of Docking Protein 3 (DOK3) mRNA levels in kidney renal clear cell carcinoma (KIRC) patients. (a–c) Survival curves were plotted based on the (a) GEPIA, (b) UALCAN, and (c) LINKOMIC databases.
Univariate analysis of the correlations between clinicopathological parameters and survival time of kidney renal clear cell carcinoma (KIRC) patients (n = 150).
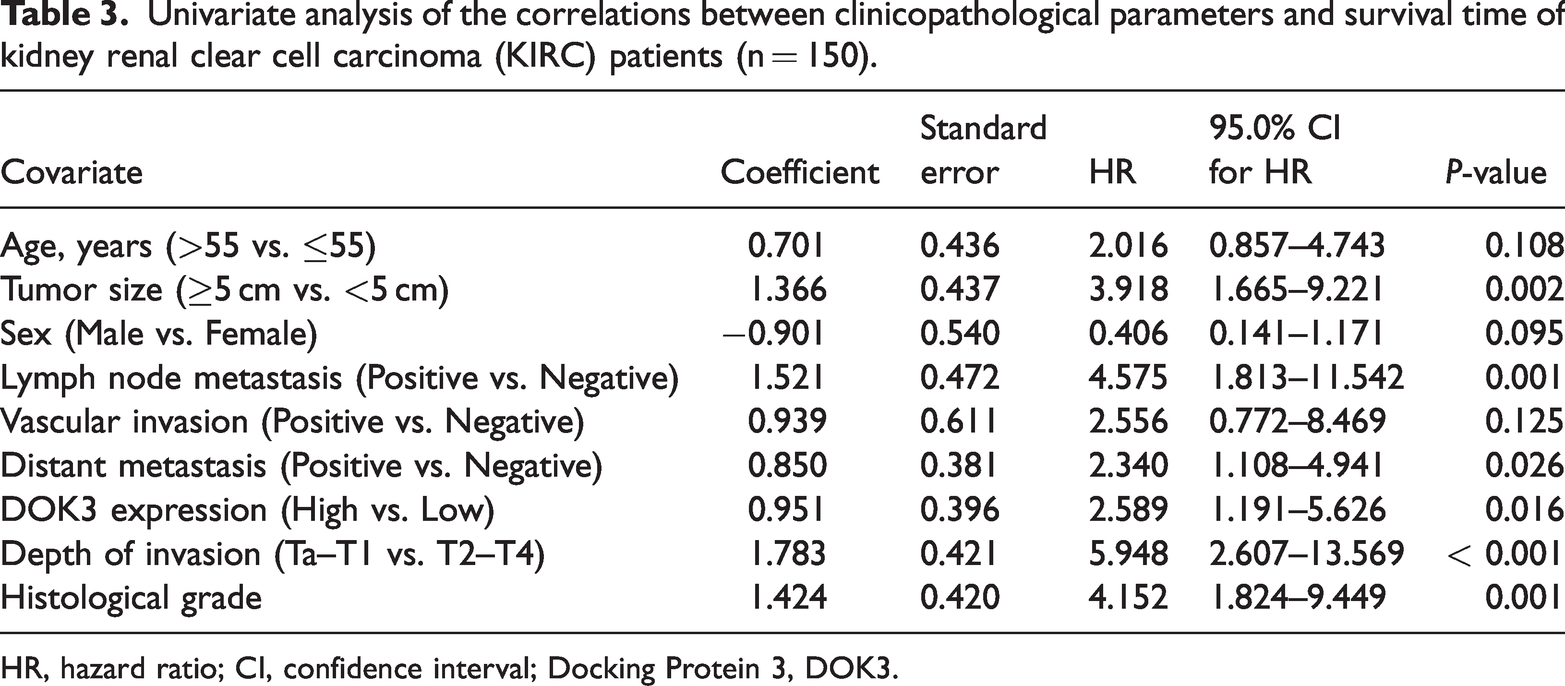
HR, hazard ratio; CI, confidence interval; Docking Protein 3, DOK3.
Multivariate analysis of the correlations between clinicopathological parameters and survival time of kidney renal clear cell carcinoma (KIRC) patients.
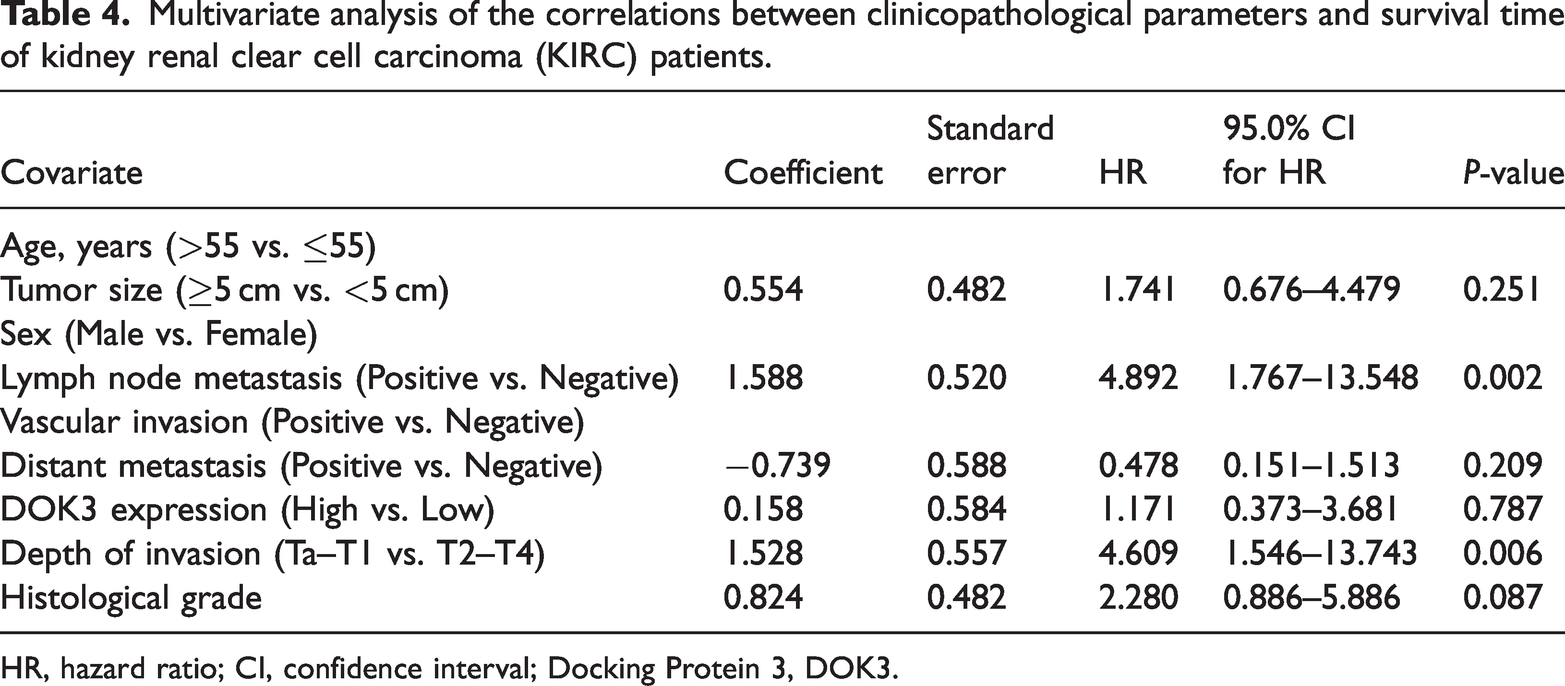
HR, hazard ratio; CI, confidence interval; Docking Protein 3, DOK3.
Genes co-expressed with DOK3 in KIRC: GO and KEGG pathway analyses
After analyzing the genes that are co-expressed with DOK3 in 533 KIRC samples from TCGA using LinkedOmics, we found 4903 genes to be negatively correlated (green spots; false discovery rate (FDR) <0.05) and 9220 genes to be positively correlated (red spots; FDR <0.05) with DOK3 according to the volcano plot (Figure 5a). The top 50 genes positively and negatively correlated with DOK3 are shown in the heat map (Figure 5b–c). WAS, SPI1, and MYO1F were the top three positively associated genes (Figure 5d–f), while SYPL1, MED21, and SUCLA2 were the top three negatively associated genes (Figure 5g–i). Moreover, we investigated the biological functions and pathways of action of DOK3-related proteins with GO enrichment analysis using LinkedOmics, which revealed that the genes co-expressed with DOK3 were mainly located in the immunological synapse, tertiary granule, and endocytic vesicle. They are involved in immunoglobulin binding, MHC protein binding, cytokine binding, and oxidoreductase activity, acting on the CH-OH group of donor molecules (Figure 6a–c). The KEGG pathway analysis suggested that the genes co-expressed with DOK3 are mostly involved in T helper 17 (Th17) cell differentiation, cytokine-cytokine receptor interactions, and cytokine receptor interactions. These results demonstrate that DOK3 broadly impacts the transcriptome (Figure 6d).

Genes co-expressed with Docking Protein 3 (DOK3) in kidney renal clear cell carcinoma (KIRC) (LinkedOmics). (a) Pearson’s test was used to analyze correlations between DOK3 and differentially expressed genes in KIRC. (b–c) Heat maps showing the top 50 genes positively and negatively correlated with DOK3 in KIRC. Red indicates positively correlated genes and green indicates negatively correlated genes. (d–f) Positive correlations between DOK3 expression and expression of the top three genes: (d) WAS, (e) SPI1, and (f) MYO1F. (g–i) Negative correlations between DOK3 expression and expression of the top three genes: (g) SYPL1, (h) MED21, and (i) SUCLA2.

Significantly enriched gene ontology (GO) annotations and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of Docking Protein 3 (DOK3) in kidney renal clear cell carcinoma (KIRC) (LinkedOmics). The significantly enriched GO annotations and KEGG pathways of genes co-expressed with DOK3 in KIRC were analyzed using gene set enrichment analysis. These included (a) cellular components, (b) biological processes, and (c) molecular functions. (d) KEGG pathway analysis results, with the x-axis representing the normalized enrichment score and the y-axis representing the GO term. Protein–protein interaction (PPI) network of DOK3 showing the interactions between DOK3 and other genes. These were obtained from the (e) STRING and (f) GeneMANIA web portals.
PPI network analysis
We constructed a PPI network using the STRING database to analyze the interactions between DOK3 and predicted protein molecules. The results revealed that DOK3 can interact with BAZ1A, PLEK, TYROBP, BTK, BLNK, GRB2, INPP5D, ABL1, CSK, and RASA1 (Figure 6e). Additionally, we analyzed the protein interactions using the GeneMANIA tool, with the data showing that ABL1, DOK4, INPP5D, DOK2, LCK, FRS3, FRS2, UHRF1BP1L, DOK1, DOK5, BTK, DOK7, IRS4, LRRK1, DOK6, GRAP, GRB2, IRS2, TRAF3, and MFNG all have an active relationship with each other (Figure 6f). Further analysis of the DOK3 PPI network showed that the genes closely related to DOK3 also have key functions in tumors.
Discussion
Renal tumors have the second highest incidence rate among urological tumors, only after bladder tumors, and account for approximately 2% to 3% of malignant tumors in adults. 19 The vast majority of these are renal clear cell carcinomas, accounting for 70% to 80% of kidney cancer cases. 20 The diagnosis rate of early KIRC is low, with asymptomatic kidney cancer accounting for 33% to 50% of cases. 21 Symptoms of hematuria, abdominal pain, and abdominal masses are not obvious and are often detected through physical examination. Early diagnosis of renal clear cell carcinoma is of great significance for improving patient prognosis. Therefore, it is important to identify effective KIRC tumor biomarkers.
The downstream of the kinase (DOK) adapter family consists of seven structurally related proteins that are not catalytically active. They primarily act as adapters to facilitate protein–protein interactions and possibly as scaffolds for nucleoprotein complexes in signal transduction pathways. 22 DOK3 is an articulator that binds molecules and thus participates in various intracellular signal transduction processes, particularly in B cells, where DOK3 acts as both an activator and inhibitor. 23 The function of DOK3 in tumors remains controversial. Paradoxically, DOK3 can be a tumor suppressor or a tumor promoter. The tumor-promoting role of DOK3 depends on activation of the NF-κB pathway by increasing cell proliferation and decreasing apoptosis. 24 In some cancer types, DOK3 is overexpressed and promotes cell proliferation and malignant transformation. However, there are also relevant studies showing that the upregulation of DOK3 plays an inhibitory role in lung epithelial cells and gliomas. 12 Taken together, DOK3 plays an important regulatory role in the biology of tumor cell growth and may represent a potential therapeutic target for human malignancies. However, no relevant exploration of DOK3 in KIRC has been previously reported. Therefore, we aimed to investigate the role of DOK3 in KIRC progression.
In this study, our data showed that DOK3 mRNA expression is positively correlated with tumor stage and is at considerably higher levels in KIRC samples compared with healthy kidney tissues. Our IHC assay findings also confirmed that KIRC samples had higher DOK3 protein levels than normal tissues. The bioinformatics analysis results of DOK3 mRNA from public databases were consistent with the IHC data for DOK3 protein levels. Overall, these data suggest that DOK3 has a tumor-promoting role in KIRC.
The prognostic value of DOK3 in KIRC was evaluated by univariate survival analysis, which revealed that tumor size, lymph node metastasis, distant metastasis, TNM stage, and pathological grading may affect KIRC patient prognosis. However, the multifactorial survival analysis showed that only lymph node metastasis and TNM stage were independent prognostic factors. This suggests that the mechanism of KIRC development is complex, with non-independent prognostic factors potentially still affecting its prognosis by interacting with other factors, such as tumor size and distant metastasis. Furthermore, we found that KIRC patients with high DOK3 mRNA expression levels had a significantly poorer prognosis compared with than those with low DOK3 mRNA expression. These results were consistent with KM Plotter database analysis. Therefore, we conclude that high DOK3 levels can promote tumor cell proliferation and can possibly be utilized to help predict KIRC tumor progression and metastasis in patients.
We also generated a DOK3 PPI network, finding that DOK3-related genes, including TYROBP, BTK, and RASA1, play important regulatory roles in tumor cell growth. The TYROBP gene is upregulated in KIRC cells and may promote tumor progression by interacting with immune cells to regulate the tumor microenvironment. 19 The BTK gene is also highly expressed in B cell malignant tumors, while it was also shown to play a role in tumor suppression through a strong correlation with P53. 25 RASA1, which is expressed at lower levels in KIRC tissues, plays a role in KIRC development and progression by suppressing miR-223-3p expression and enhancing FBXW7 expression. 26
Our study does have some limitations. First, the small patient sample size and RNA-sequencing data in public databases may not be representative of the entire KIRC patient population. Second, we used TMAs rather than whole tissue sections, which may not adequately reflect the full heterogeneity of primary KIRC. Finally, we assessed IHC-based DOK3 protein expression, which remains a semi-quantitative method. Therefore, more quantitative examinations are needed.
In summary, by combining information from public databases with IHC results from clinical samples, we have improved our understanding of the mechanisms of DOK3 in KIRC. This work validates the critical function that DOK3 plays in KIRC. Because elevated DOK3 expression in KIRC is linked to worse survival rates, our work may lead to new directions for the prevention and treatment of KIRC. Understanding DOK3 expression patterns in this disease will help identify patients with high metastatic potential. Thus, DOK3 expression levels may be a useful predictor of invasion and prognosis in KIRC patients.
Research Data
sj-pdf-1-imr-10.1177_03000605231174974 - Research Data for Expression and clinical significance of DOK3 in renal clear cell carcinoma
Research Data, sj-pdf-1-imr-10.1177_03000605231174974 for Expression and clinical significance of DOK3 in renal clear cell carcinoma by Yi-Fan Wang, Yu-Qi Hu, Yu-Ning Hu, Yu-Chen Bai, Heng Wang and Qi Zhang in Journal of International Medical Research
Footnotes
Author contributions
Data curation: HW, YFW; Writing-original draft: YFW, HW; Writing-review and editing: all authors; Visualization: YFW, YQH, YNH; Supervision: QZ; Project administration: QZ.
Availability of data and materials
The datasets generated and analyzed in the current study are available in the TCGA repository (https://portal.gdc.cancer.gov/), Oncomine database (https://www.oncomine.org), LinkedOmics (http://www.linkedomics.org/login.php), String (http://string-db.org/), UALCAN database (Ualcan.path.uab.edu/analysis), GEPIA (http://gepia.cancer-pku.cn/), and GeneMANIA database (![]() ). All patient data are available from the electronic medical records of Zhejiang Provincial People’s Hospital.
). All patient data are available from the electronic medical records of Zhejiang Provincial People’s Hospital.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was financially supported by the Zhejiang Provincial Science and Technology Department Public Welfare Technology Research Program (Grant No. LGF19H160025), Zhejiang Provincial Science and Technology Department Public Welfare Technology Research Program (Grant No. LBY21H050001), and Zhejiang Provincial Health Department Project (Grant No. 2023KY049).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
