Abstract
Purpose
Osteosarcoma is the most common primary tumor of bone tissue. To date, the expression of transmembrane and tetratricopeptide repeat containing 4 (TMTC4) in osteosarcoma has not been reported. This study aimed to investigate the significance of TMTC4 in the prognosis of osteosarcoma.
Methods
We screened the expression of downstream genes after interleukin 11 receptor subunit alpha knockout in osteosarcoma using the Gene Expression Omnibus database. Data from The Cancer Genome Atlas database and a cloud-based platform were used to further analyze the expression levels and functional impact of TMTC4 in osteosarcoma, along with subgroup analysis. Polymerase chain reaction was used to detect TMTC4 expression in osteoblast and osteosarcoma cells, and a cell counting kit-8 assay was employed to assess cell proliferation.
Results
Eleven downregulated genes, including TMTC4, were identified using the GSE191215 database. Kaplan–Meier survival curve analysis showed that osteosarcoma cells with low TMTC4 expression were associated with a higher survival rate. Enrichment analysis suggested that low TMTC4 expression might be related to immunoregulatory pathways. Methylation analysis revealed that TMTC4 methylation was significantly increased in osteosarcoma cells. In addition, TMTC4 expression was significantly higher in osteosarcoma cells than in osteoblast cells, and inhibition of TMTC4 expression hindered osteosarcoma cell proliferation.
Conclusion
TMTC4 is an important biomarker in patients with osteosarcoma, and its low expression is associated with improved prognosis.
Keywords
Introduction
Soft tissue sarcoma and bone tissue sarcoma are heterogeneous malignant tumors of mesenchymal origin. 1 Primary bone sarcomas are rare tumors that include osteosarcoma, Ewing’s sarcoma, and chondrosarcoma. 2 Among these, osteosarcoma is one of the most common types of bone tumor worldwide. 3 It is the most common primary malignant bone tumor, with an annual incidence rate of 4.4–5 cases per million in the general population.4,5 Osteosarcoma is an invasive primary bone tumor that predominantly affects children and adolescents, 6 particularly those with rapidly growing bones, 7 and a second incidence peak is observed in older adults. 8 Currently, there is no clear and effective strategy for the prevention and treatment of osteosarcoma. Therefore, this study focuses on identifying a potential molecular biomarker for osteosarcoma to support early detection, intervention, and rehabilitation.
In medicine, one of the primary applications of bioinformatics is the identification of biomarkers. 9 Interest in using bioinformatics methods to examine disease-associated biomarkers has increased. 10 Conducting bioinformatics analyses of diseases typically requires access to multiple public databases. Currently, more than 1 million microarrays and high-throughput sequencing datasets are stored in public repositories such as Gene Expression Omnibus (GEO) and ArrayExpress. 11 Cancer genome sequencing projects, such as The Cancer Genome Atlas (TCGA), have generated large amounts of data and enabled the development of bioinformatics algorithms to identify cancer-driving genes. 12 Currently, TCGA includes histological data for more than 20 cancer types. 13 Its data portal provides a platform for researchers to search, download, and analyze datasets. 14 The SMART application is an interactive web tool for comprehensive DNA methylation analysis and visualization. 15 UALCAN is a portal that facilitates gene expression and survival analysis of tumor subgroups.16,17 MethSurv is a network tool for multivariate survival analysis based on DNA methylation data. 18
This study focuses on the expression of transmembrane and tetratricopeptide repeat containing 4 (TMTC4), a gene previously unreported in the context of osteosarcoma, and its potential implications for patient prognosis. Although several molecular markers associated with osteosarcoma have been identified, the specific role of TMTC4 remains unclear, representing an important gap in current knowledge. Our findings indicate that low TMTC4 expression is associated with improved survival outcomes in patients with osteosarcoma, suggesting a protective effect that requires further investigation. Additionally, the observed relationship between TMTC4 expression and immunoregulatory pathways highlights its potential relevance as a therapeutic target. The significantly increased methylation of TMTC4 in osteosarcoma further underscores its involvement in disease pathophysiology. Collectively, these associations suggest that TMTC4 has functional significance in osteosarcoma and could serve as a useful biomarker for guiding clinical decisions and therapeutic strategies.
This study employs a multifaceted approach to investigate the role of TMTC4 in osteosarcoma, using a combination of bioinformatics analysis and experimental validation. Integrating data from the GEO and TCGA enables a comprehensive assessment of TMTC4 expression and its potential prognostic significance in osteosarcoma. This methodology leverages large-scale genomic datasets for initial screening, followed by targeted laboratory experiments to validate findings in a controlled environment. The primary objective of this research is to elucidate the relationship between TMTC4 expression levels and patient outcomes, thereby contributing to the understanding of its role as a prognostic biomarker. By correlating TMTC4 expression with survival rates and examining its involvement in immunoregulatory pathways, this study aimed to provide insights that may inform future therapeutic strategies and improve clinical management of osteosarcoma.
Materials and methods
Dataset analysis
The GEO database (http://www.ncbi.nlm.nih.gov/geo) is a publicly accessible data platform maintained by the National Center for Biotechnology Information. The dataset related to osteosarcoma (GSE191215) was selected for this study. GSE191215 includes 3 control groups and 12 treatment groups, which comprise 4 different short hairpin RNA (shRNA) sequences targeting enhancer of zeste homolog 2 (EZH2), with 3 groups for each sequence.
Differentially expressed gene (DEG) analysis
We used the default GEO2R tool (https://www.ncbi.nlm.nih.gov/geo/geo2r/) to analyze the GSE191215 dataset. As the treatment group included four different shRNA sequences targeting EZH2, we analyzed each sequence separately against the control group. We set the threshold at p < 0.05 to screen for DEGs. For each sequence, we constructed a volcano plot to visualize differential expression and classified the DEGs as upregulated or downregulated based on log fold change (logFC) values. Subsequently, we integrated and analyzed the upregulated and downregulated DEGs from the four groups separately and visualized the results using a Venn diagram.
Functional and pathway enrichment analysis
The clusterProfiler package was used to conduct Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses on the common upregulated and downregulated DEGs across the four groups. For GO analysis, biological processes (BP), molecular functions (MF), and cellular components (CC) were included.
Construction of Kaplan–Meier curve and predictive nomogram
We used the survival package to perform the proportional hazards assumption test and to fit survival regression models. The results were visualized using survival and ggplot2 packages. Additionally, the survival package was used for Cox regression analysis, and the rms package was employed to construct and visualize the nomogram-related model.
Gene set enrichment analysis (GSEA)
To investigate potential molecular mechanisms by which TMTC4 expression affects osteosarcoma prognosis, we conducted GSEA. Patients were divided into two groups based on the median TMTC4 expression level: low TMTC4 expression group and high TMTC4 expression group. Subsequently, we identified DEGs between the two groups and performed GSEA. The results of enrichment analysis were visualized using the ggplot2 package.
Evaluation of immune infiltration
To examine the correlation between TMTC4 expression and infiltration of 24 immune cell types in osteosarcoma, we used Spearman correlation analysis and presented the results using a lollipop plot. Differences in dendritic cell (DC) enrichment levels between the low TMTC4 expression group and high TMTC4 expression group were analyzed using scatter plots and grouped comparison plots. Using data from the TIMER database (http://timer.cistrome.org/), we performed Kaplan–Meier analysis and plotted Kaplan–Meier curves accordingly.
Evaluation of methylation level
The SMART online tool (http://www.bioinfozs.com/smartapp/) was used to visualize the chromosomal distribution of methylation probes associated with TMTC4 and to obtain detailed genomic information on TMTC4. The UALCAN platform (http://ualcan.path.uab.edu/) was used to analyze differences in TMTC4 methylation levels between osteosarcoma and normal tissues. The MethSurv platform (https://biit.cs.ut.ee/methsurv/) was employed to evaluate the prognostic value of TMTC4 methylation levels in patients with osteosarcoma.
RNA extraction and polymerase chain reaction (PCR)
Initially, we washed the prepared human osteoblast (hFOB1.19) and human osteosarcoma (MG-63, U2OS, and Saos-2) cells with phosphate-buffered saline (PBS) three times. Then, we extracted total RNA using Trizol reagent. Complementary DNA (cDNA) was synthesized from the extracted RNA using the Takara RR047A reverse transcription kit. Real-time quantitative PCR was performed on the synthesized cDNA using the Takara RR820A reagent kit, and the results were statistically analyzed. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as an internal reference. The primer sequences were as follows: (a) TMTC4 forward primer, GCCGTGTTGGACACTGATTTTG; (b) TMTC4 reverse primer, TCCCACTAACTTAGCCCAG; (c) GAPDH forward primer, GGAGCGAGATCCCTCCAAAAT; and (d) GAPDH reverse primer, GGCTGTGTCATACTTTCTCATGG.
Cell transfection
We obtained the TMTC4 overexpression plasmid, sh-TMTC4, and a nontargeting control sequence as a negative control (NC) for TMTC4 from GenePharma Co., Ltd. (Shanghai, China. According to the manufacturer’s instructions, we transfected MG-63, U2OS, and Saos-2 cells with the plasmids using Lipofectamine 2000 (Invitrogen, Inc.).
Cell counting kit-8 (CCK-8) assay
We seeded the transfected MG-63, U2OS, and Saos-2 cells into 96-well plates. After 24 h, the CCK-8 reagent (Dojindo Molecular Technologies, Inc.) was added to each well and incubated at 37°C for 60 min. Cell viability was assessed according to the manufacturer’s instructions. The cells were subjected to enzyme-linked immunosorbent assay (ELx808, BioTek, Inc.), and the optical density (OD) was measured at 450 nm, with the activity expressed as the OD value.
Immunohistochemistry (IHC)
Tissue specimens from 10 patients with osteosarcoma and 10 patients with osteochondroma were collected at the General Hospital of Northern Theater Command between March 2023 and January 2024. This study was approved by the Ethics Committee of the General Hospital of Northern Theater Command (approval no. 2025-108). Tissue specimens were fixed in 4% formaldehyde, dehydrated overnight, and embedded in paraffin. Finally, we prepared 3-µm thick tissue sections and deparaffinized them in xylene. After rehydration, antigen retrieval was performed, and endogenous peroxidase activity was blocked with 3% hydrogen peroxide (H2O2) for 10 min. Sections were washed with PBS and incubated with 5% bovine serum albumin (BSA) at room temperature for 30 min. Sections were then incubated overnight at 4°C with primary rabbit polyclonal anti-TMTC4 antibody (ProteinTech Group, 25374-1-AP), followed by incubation with a goat anti-rabbit secondary antibody at room temperature for 90 min. The sections were subsequently treated with ABC working solution (Zsbio) at room temperature for 20 min and visualized using 3,3-diaminobenzidine (DAB; Zsbio). Brown staining indicated positive expression, which was observed under a Leica microscope.
Statistical analysis
All data were analyzed using Statistical Package for the Social Sciences (SPSS) version 23.0 and GraphPad Prism version 5.0. Statistical significance was set at p < 0.05. During the DEG analysis, genes were classified as upregulated (logFC ≥ 1) or downregulated (logFC ≤ −1). Each assay was performed independently three times, and measurement data were presented as mean ± SD. Differences between the two groups were assessed using an unpaired Student’s t-test. The experimental procedures of this study are summarized in a flow chart (Figure 1).
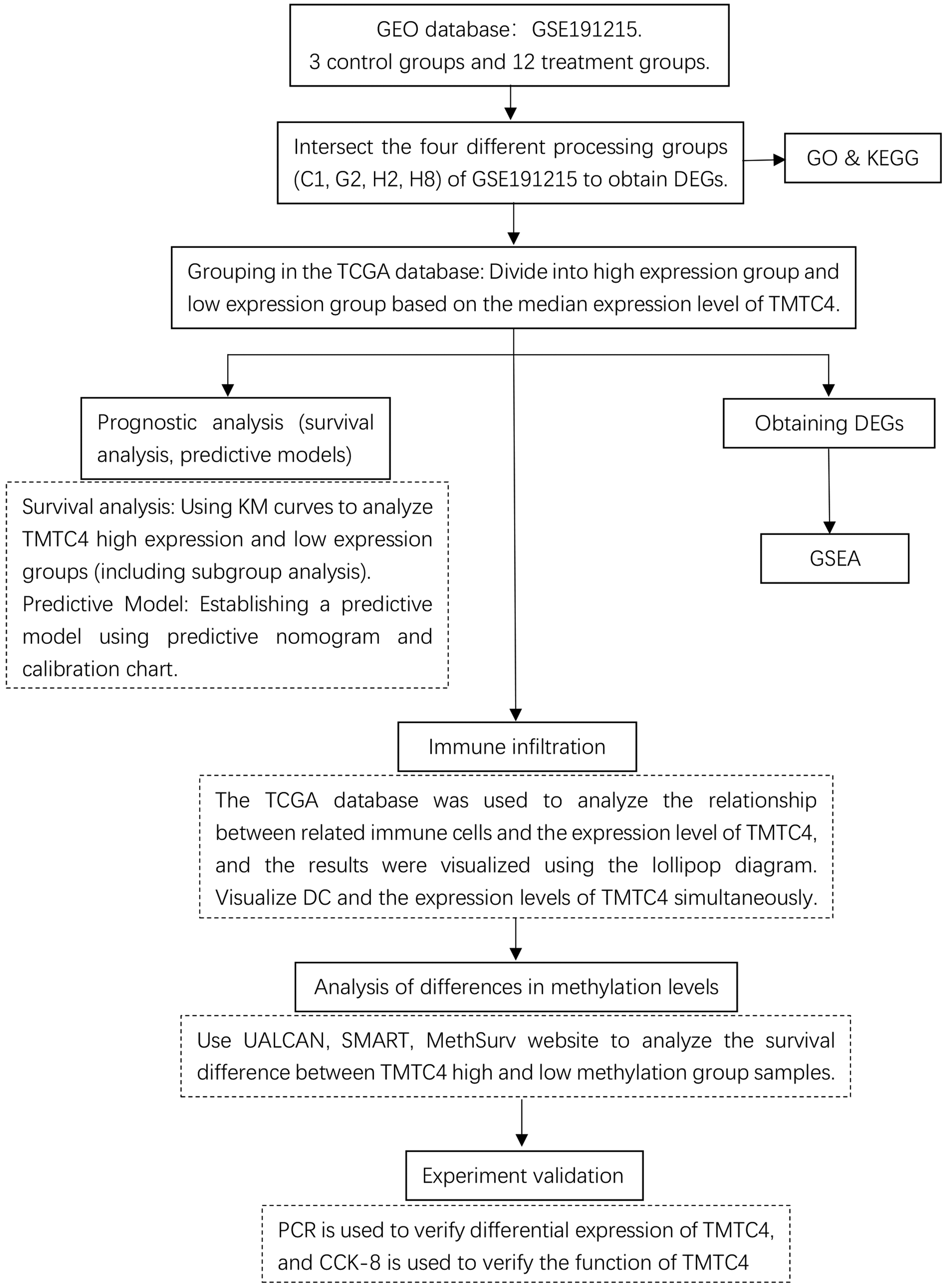
Flow chart of the study. The study workflow is illustrated as follows: first, the GSE191215 dataset was analyzed to identify TMTC4; next, prognostic analysis of TMTC4 in osteosarcoma was performed; subsequently, immune infiltration and methylation levels of TMTC4 in osteosarcoma were evaluated; finally, cell experiments were conducted to confirm that TMTC4 can serve as a prognostic factor in osteosarcoma.
Results
DEG analysis
The GSE191215 dataset includes three control samples and 12 treatment samples representing 4 different shRNA sequences targeting EZH2 (3 samples per sequence). Each treatment group (C1, G2, H2, and H8) was compared with the control group using the default GEO2R tool. Volcano plots were generated to visualize DEGs for each group (Figure 2(a) to (d)). The intersection between upregulated and downregulated DEGs from the four groups was presented using a Venn diagram (Figure 2(e) to (f)). A total of 42 DEGs were identified with p-values <0.05, of which 31 were upregulated and 11 were downregulated.

DEG analysis of GSE191215. (a–d) Analysis of DEGs in GSE191215 using GEO2R for C1, G2, H2, and H8 compared with control, respectively. (e) Overlap of upregulated DEGs across the four groups and (f) overlap of downregulated DEGs across the four groups. DEGs: differentially expressed genes; GEO2R: Gene Expression Omnibus 2R.
Functional and pathway enrichment and least absolute shrinkage and selection operator (LASSO) analysis
GO analysis showed that the DEGs were enriched in the following processes (only the top five were reported when more than five results were obtained). The downregulated genes were enriched in the following processes:
BP. Nuclear-transcribed mRNA catabolic process, signal transduction in response to DNA damage, mRNA catabolic process, neuron apoptotic process, regulation of response to DNA damage stimulus; CC. Axonemal dynein particle, RNA-induced silencing (RISC) complex, RNA interference (RNAi) effector complex, chaperone complex, integral component of the mitochondrial membrane; MF. 5′–3′ exonuclease activity, death receptor binding, telomerase RNA binding, mannosyltransferase activity, receptor-regulated SMAD (R-SMAD) binding (Figure 3(a), Table 1).

Enrichment and LASSO regression analyses of downregulated DEGs. (a) GO term enrichment analysis of downregulated DEGs. (b) KEGG pathway enrichment analysis of downregulated DEGs. (c) Coefficient variation of downregulated DEGs based on LASSO regression and (d) selection of the optimal λ value in the LASSO regression model using cross-validation. GO: Gene Ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes; DEGs: differentially expressed genes; λ: lambda (regularization parameter); LASSO: least absolute shrinkage and selection operator.
GO term enrichment analysis of downregulated DEGs.

GO: Gene Ontology; DEGs: differentially expressed genes; BP: biological process; CC: cellular component; MF: molecular function; mRNA: messenger RNA; RISC: RNA-induced silencing complex; RNAi: RNA interference; R-SMAD: receptor-regulated SMAD.
KEGG pathway analysis indicated enrichment in the following pathways for downregulated genes (top 5 reported): apoptosis—multiple species, viral myocarditis, platinum drug resistance, p53 signaling pathway, and RNA degradation (Figure 3(b), Table 2).
KEGG pathway enrichment analysis of downregulated DEGs.

KEGG: Kyoto Encyclopedia of Genes and Genomes; DEGs: differentially expressed genes; RNA: ribonucleic acid.
LASSO regression analysis was performed to construct a prognostic model based on the prognostic value of the DEGs (Figure 3(c)). A LASSO variable trajectory plot illustrating the regression results is shown in Figure 3(d).
Among the upregulated genes, no DEGs were found to be significantly enriched.
Clinical features
A total of 263 patients with osteosarcoma were included in this study, comprising 144 women (54.8%) and 119 men (45.2%). Among them, 130 patients (49.5%) were aged <60 years. Overall, 199 patients (83.2%) had no multifocal tumor, while 40 (16.7%) had multifocal tumors. Furthermore, 21 patients (10.1%) had superficial tumor depth, while 188 patients (90%) had deep tumor depth. Additional clinicopathological characteristics of the patients are summarized in Table 3.
Clinicopathological data of patients.

TMTC4: transmembrane and tetratricopeptide repeat containing 4.
Expression level of TMTC4 and prognosis of osteosarcoma
Analysis of the GSE191215 dataset showed that TMTC4 expression in KRIB cells was significantly reduced following targeted knockout of interleukin 11 receptor subunit alpha (IL11RA) compared with the control group. These results suggest that low TMTC4 expression is closely associated with improved prognosis in osteosarcoma.
Kaplan–Meier curve survival analysis indicated that patients with low TMTC4 expression had better overall survival (OS) and disease-specific survival (DSS) than those with high TMTC4 expression (p = 0.025 and p = 0.016, respectively; Figure 4(a) and (b)).

Prognostic value of TMTC4 expression in osteosarcoma. (a) Kaplan–Meier curves showing the correlation of TMTC4 expression with OS probability; (b) Kaplan–Meier curves showing the correlation of TMTC4 expression with DSS probability and (c) forest plot of univariate Cox regression analysis for OS. TMTC4: transmembrane and tetratricopeptide repeat containing 4; OS: overall survival; DSS: disease-specific survival; Cox: Cox proportional hazards model.
Univariate Cox survival analysis showed a significant difference in survival between the high and low TMTC4 expression groups (hazard ratio (HR): 1.579; 95% confidence interval (CI): 1.058–2.359; p = 0.025; Figure 4(c)). In addition to TMTC4 expression, metastasis, residual tumor, tumor multifocality, and margin status were identified as independent prognostic factors in osteosarcoma.
Kaplan–Meier subgroup analysis revealed that among women, those with lower TMTC4 expression had improved OS (p = 0.046), whereas this association was not significant in men (p = 0.143) (Figure 5(a) and (b)). Lower TMTC4 expression was associated with higher OS in patients with deep tumor involvement (p = 0.042), but not in patients with superficial tumor depth (p = 0.469; Figure 5(c) and (d)). In patients with tumor metastasis, lower TMTC4 expression was correlated with improved OS (p = 0.019), whereas no significant difference was observed in patients without metastasis (p = 0.592; Figure 5(e) and (f)). Among patients with negative tumor margins, lower TMTC4 expression was associated with higher OS (p = 0.025), while no significant difference was observed in patients with positive tumor margins (p = 0.947; Figure 5(g) and (h)).

Kaplan–Meier subgroup survival analysis of TMTC4 expression. (a, b) Subgroup analysis by sex; (c, d) subgroup analysis by tumor depth; (e, f) subgroup analysis by metastasis and (g, h) subgroup analysis by margin status. TMTC4: transmembrane and tetratricopeptide repeat containing 4.
Construction of predictive nomogram
A nomogram was constructed using patient characteristics, including sex, race, age, tumor multifocality, tumor necrosis, tumor depth, and TMTC4 expression, to predict the 1-, 2-, and 3-year OS probabilities of patients with osteosarcoma (Figure 6(a)). The calibration plots indicated that the nomogram-predicted 1-, 2-, and 3-year OS probabilities closely corresponded to the observed outcomes (Figure 6(b) to (d)). These results demonstrate that the nomogram can accurately predict OS in patients with osteosarcoma and may provide useful information for individualized clinical evaluation and treatment planning.

Nomogram construction and OS prediction for osteosarcoma patients. (a) Nomogram integrating TMTC4 and other prognostic factors to predict 1-, 2-, and 3-year OS and (b–d) calibration plots comparing nomogram-predicted OS with observed survival rates at 1-, 2-, and 3-year intervals. TMTC4: transmembrane and tetratricopeptide repeat containing 4; OS: overall survival.
GSEA
To explore the potential molecular mechanisms underlying the prognostic value of TMTC4 in osteosarcoma, GSEA was conducted using transcriptome data from TCGA. The results indicated that patients with low TMTC4 expression showed enrichment in immunoregulatory pathways and antigen activation processes (Figure 7(a) and (b)).

GSEA of DEGs in high- and low-TMTC4 expression samples. (a) Enrichment plot showing two representative pathways and (b) Ridge plot depicting the density distribution for the top 5 gene sets. GSEA: gene set enrichment analysis; DEGs: differentially expressed genes; TMTC4: transmembrane and tetratricopeptide repeat containing 4.
Evaluation of immune infiltration
The correlation between TMTC4 expression and 24 immune cell types in osteosarcoma patients was analyzed and visualized using a lollipop plot. The strongest negative correlation was observed with DCs (p < 0.001; Figure 8(a)). Further analysis focused on DCs to compare enrichment fractions between the high and low TMTC4 expression groups (Figure 8(b) and (c)). Kaplan–Meier survival curves were generated using the TIMER database (Figure 8(d)). The results showed that among patients with low TMTC4 expression, those with high DC infiltration had significantly higher survival (p < 0.001). In contrast, among patients with high TMTC4 expression, there was no significant difference in survival between the high and low DC infiltration groups (p = 0.145).

Correlation of TMTC4 expression with immune infiltration in osteosarcoma. (a) Correlation between TMTC4 expression and immune cell infiltration; (b) enrichment scores of DCs in low- versus high-TMTC4 expression samples; (c) negative correlation between TMTC4 expression and DC infiltration levels and (d) Kaplan–Meier curves showing the association of DC infiltration levels with survival probability. TMTC4: transmembrane and tetratricopeptide repeat containing 4; DC: dendritic cell.
Evaluation of methylation level of TMTC4 in osteosarcoma
Analysis using the UALCAN platform indicated that TMTC4 promoter methylation levels were significantly higher in tumor tissues than in adjacent normal tissues (p < 0.001; Figure 9(a)). Further analysis using the SMART platform provided the chromosomal distribution of methylation probes associated with TMTC4 (Figure 9(b)) and detailed genetic information for TMTC4 (Figure 9(c)). MethSurv analysis revealed significant survival differences between patients with high and low TMTC4 methylation levels, with patients exhibiting high methylation showing higher OS (Figure 10(a)). In addition, methylation status stratified by maxstat (Figure 10(b)) as well as methylation profiles based on age (Figure 10(c)) and sex (Figure 10(d)) were obtained.

Methylation analysis of TMTC4 in osteosarcoma. (a) Promoter methylation levels of TMTC4 in osteosarcoma versus normal tissue; (b) chromosomal distribution of methylation probes associated with TMTC4 and (c) detailed genomic information of TMTC4. TMTC4: transmembrane and tetratricopeptide repeat containing 4.

Correlation of TMTC4 methylation level with survival probability. (a) Kaplan–Meier curves showing the association between TMTC4 methylation level and survival probability; (b) splitting of methylation level by maxstat; (c) methylation profiles stratified by age; (d) methylation profiles stratified by sex. TMTC4: transmembrane and tetratricopeptide repeat containing 4; maxstat: maximum statistic.
Expression level of TMTC4 in osteosarcoma tissue specimens and cells
To examine TMTC4 expression in osteosarcoma tissue specimens, osteochondroma tissues were used as controls. IHC analysis showed TMTC4 protein expression was significantly higher in osteosarcoma tissues compared with that in osteochondroma tissues (control group) (Figure 11 (a)), validating the findings from the database analysis.
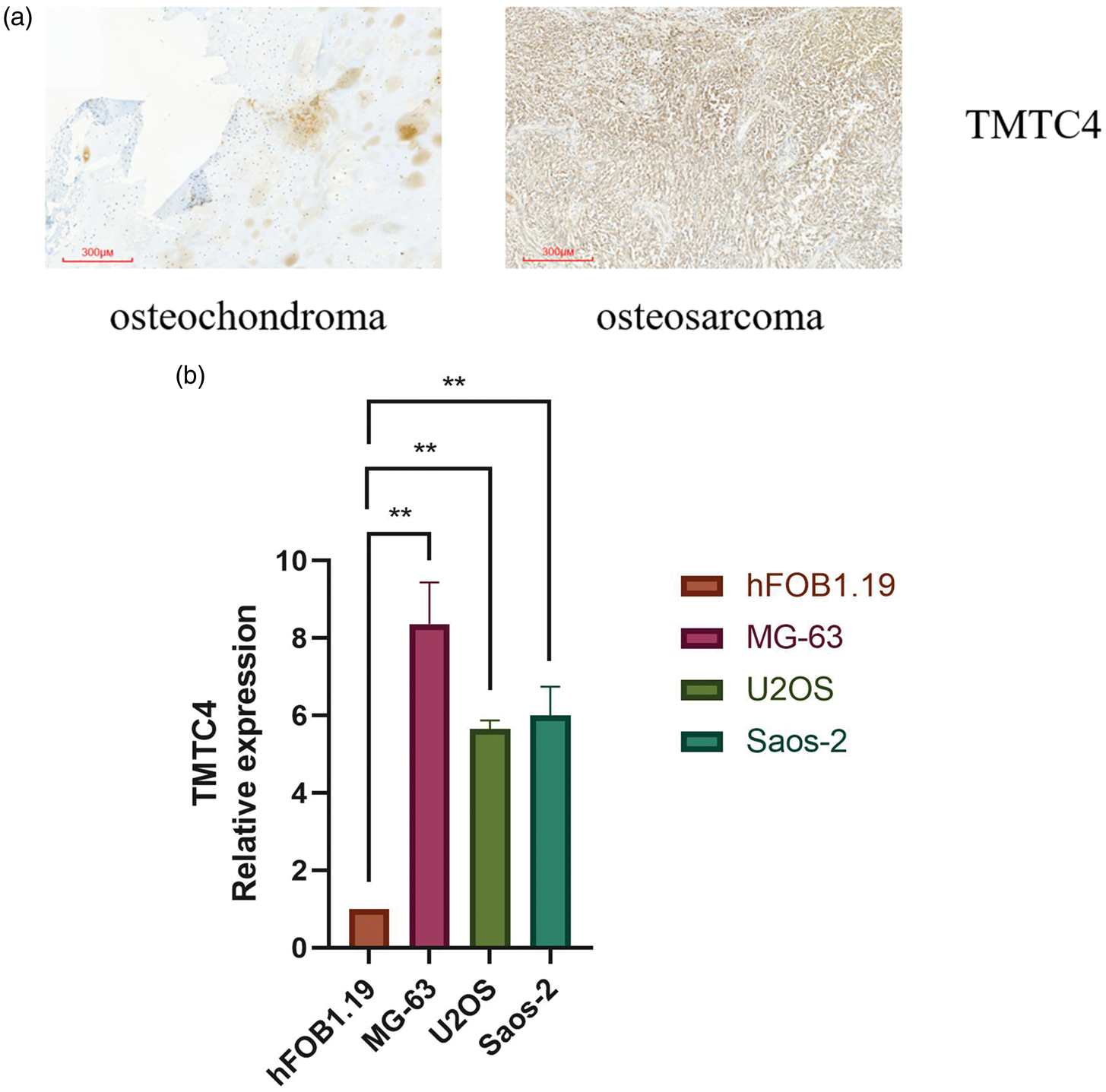
TMTC4 expression in osteosarcoma tissue specimens and osteosarcoma cells. (a) TMTC4 protein expression in osteochondroma and osteosarcoma tissue specimens and (b) TMTC4 mRNA expression in osteoblasts (hFOB1.19) and osteosarcoma cells (MG-63, U2OS, and Saos-2). TMTC4: transmembrane and tetratricopeptide repeat containing 4; mRNA: messenger RNA.
To assess TMTC4 expression in osteosarcoma cells, hFOB1.19 cells were used as controls. Quantitative PCR analysis revealed that TMTC4 mRNA expression was significantly higher in MG-63, U2OS, and Saos-2 cells compared with that in hFOB1.19 cells (p < 0.01; Figure 11(b)), confirming the database-based observations.
Inhibition of TMTC4 expression impedes cell proliferation
To further investigate the role of TMTC4 in osteosarcoma cells, we transfected TMTC4 overexpression plasmids, sh-TMTC4, and the corresponding NCs into osteosarcoma cells. Successful transfection was confirmed (Figure 12(a)). Cell proliferation was then assessed using the CCK-8 assay. The results showed that knockout of TMTC4 inhibited cell proliferation, whereas overexpression of TMTC4 promoted proliferation (Figure 12(b)).

Inhibition of TMTC4 expression impedes cell proliferation. (a) Transfection efficiency of TMTC4 overexpression plasmids, sh-TMTC4 and corresponding NCs and (b) viability of MG-63, U2OS, and Saos-2 cells after transfection with TMTC4 overexpression plasmids, sh-TMTC4 and corresponding NCs. TMTC4: transmembrane and tetratricopeptide repeat containing 4; sh-TMTC4: short hairpin RNA targeting TMTC4; NC: negative control; OD: optical density.
Discussion
TMTC4 encodes a transmembrane protein that belongs to a protein family containing an N-terminal transmembrane domain and a C-terminal tetratricopeptide repeat (TPR) domain. The TPR domain mediates protein–protein interactions in various cellular processes, including synaptic vesicle fusion, protein folding, and protein translocation. A pseudogene of TMTC4 has been identified on chromosome 5 (RefSeq, Apr 2017). Previous studies have shown that TMTC4 deletion activates the unfolded protein response (UPR) and leads to postnatal hearing loss. 19 The proteins encoded by human TMTC1, TMTC2, TMTC3, and TMTC4 are newly characterized O-mannosyltransferases of the GT-C/PMT family, with ligand-binding sites. 20 TMTC4 has also been identified as a novel diagnostic marker for prostate cancer with high specificity and sensitivity. 21 However, to date, no studies have explored the biological role of TMTC4 in osteosarcoma.
The methodological rigor of this study highlights its strengths in elucidating the role of TMTC4 in osteosarcoma. By integrating multiple high-throughput datasets, including GEO and TCGA, we performed a comprehensive analysis of TMTC4 expression and its prognostic significance across diverse osteosarcoma cohorts. The GSE191215 dataset was initially used to screen DEGs related to osteosarcoma, and subsequent enrichment analysis suggested that changes in TMTC4 expression may be associated with osteosarcoma pathogenesis. Kaplan–Meier survival and enrichment analyses further supported the association between TMTC4 expression and patient outcomes. Analysis of TCGA data confirmed that low TMTC4 expression correlates with improved prognosis in osteosarcoma, underscoring its potential utility as a prognostic biomarker. In addition, we analyzed TMTC4 expression in relation to immune infiltration and methylation status in osteosarcoma and found that these factors are closely associated with patient prognosis. Enrichment analysis indicated that TMTC4 is involved in immunoregulatory pathways, with its downregulation potentially promoting a more immunogenic tumor microenvironment, thereby enhancing responses to immunotherapy. Furthermore, the observed hypermethylation of TMTC4 in osteosarcoma highlights the contribution of epigenetic alterations to tumor aggressiveness. Our in vitro experiments corroborated these findings: quantitative PCR confirmed TMTC4 expression in osteoblasts and osteosarcoma cells, and functional assays, including the CCK-8 assay, demonstrated that inhibition of TMTC4 reduced osteosarcoma cell proliferation. These results underscore the functional relevance of TMTC4 in osteosarcoma and support its potential as a therapeutic target, providing a foundation for future studies exploring its mechanistic role in disease biology.
This study provides important insights into the role of TMTC4 in osteosarcoma; however, several limitations should be noted. First, the reliance on publicly available databases such as GEO and TCGA may introduce biases due to variations in sample collection, processing, and annotation, which could affect the generalizability of our findings. Second, although in vitro experiments offered functional insights into TMTC4, they do not fully replicate the complex tumor microenvironment in vivo. Third, the lack of longitudinal clinical data limits our ability to establish a direct causal relationship between TMTC4 expression and patient outcomes. Finally, the mechanisms underlying the observed association between TMTC4 methylation and expression remain unclear, warranting further investigation. These limitations underscore the need for caution in interpreting our results and highlight the importance of complementary studies to validate the prognostic and therapeutic significance of TMTC4 in osteosarcoma.
Conclusions
This study identifies TMTC4 as a crucial prognostic biomarker in osteosarcoma, with its low expression levels associated with improved patient survival. These findings suggest that TMTC4 not only serves as a prognostic marker but also plays an active role in the pathophysiology of osteosarcoma. The inverse correlation between TMTC4 expression and survival highlights its potential clinical relevance.
Moreover, TMTC4’s involvement in immunoregulatory pathways and its elevated methylation status in osteosarcoma provide insights into the molecular mechanisms contributing to tumor progression and immune evasion. Experimental evidence demonstrated that silencing TMTC4 inhibits osteosarcoma cell proliferation, reinforcing its functional importance and potential as a therapeutic target.
These findings collectively support incorporating TMTC4 expression analysis into the prognostic evaluation of osteosarcoma, which may guide therapeutic decision-making and improve patient outcomes. Future research should focus on elucidating the mechanistic pathways by which TMTC4 influences tumor behavior and evaluating its potential as a therapeutic target. Such studies could facilitate the development of innovative strategies for tumor prevention and treatment, establishing TMTC4 as a key biomarker in clinical oncology.
Footnotes
Acknowledgments
We thank the researchers who made their experimental data publicly available and all members of our team for their hard work and contributions.
Authors’ contributions
Yihao Tian contributed to the study conception and design. Zihan Li performed language editing, tabulation, and figure preparation. All authors read and approved the final version of the manuscript.
Availability of data and materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Declaration of conflicting interests
The authors declare that the study was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethics approval and consent to participate
All procedures involving human participants were conducted in accordance with the ethical standards of the institutional research committee and the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Institutional Review Board of the General Hospital of the Northern Theater Command (approval no. 2025-108).
Funding
Not applicable.
