Abstract
Objectives
Cancer is a threat to human health, and many molecules are involved in the transformation of malignant cells. Hepatocyte nuclear factor 1A (HNF1A) is an important transcription factor that regulates multiple biological processes. Our research focused on elucidating the expression and function of HNF1A in cancer through bioinformatic analysis.
Methods
UALCAN, Kaplan–Meier plotter, COSMIC, Tumor IMmune Estimation Resource, and Cancer Regulome were used to obtain relevant data for HNF1A.
Results
HNF1A was abnormally expressed in multiple cancers, and its expression was associated with differences in overall survival in patients with cancer. HNF1A mutations widely exist in tumors and interact with different genes involved in various processes. Additionally, we found that HNF1A was associated with the infiltration of immune cells, and it affected the prognostic value of these cells in some cancers.
Conclusions
HNF1A plays a crucial role in cancer, and it may represent a biomarker and target for future cancer immunotherapy.
Keywords
Introduction
Cancer is one of the main diseases that threaten human health. In recent years, the incidence and mortality of cancer have increased. 1 Cancer is caused by changes in several metabolic processes that affect cell proliferation, growth, replication, and death. 2 A basic challenge in cancer biology research is identifying the molecular mechanisms that lead to the transformation of malignant cells.3,4 Although cancer treatments are diverse, including surgery, radiotherapy, chemotherapeutic drugs, and biologic therapy, the overall survival rate of patients remains poor. 5 Therefore, elucidating the crucial molecular alterations in cancers is critical for understanding the mechanisms leading to disease and improving therapeutic approaches.
Hepatocyte nuclear factor 1A (HNF1A) is one of the most important transcription factors in the liver. 6 This molecule regulates the expression of various liver-specific genes and participates in multiple biological processes, such as protein synthesis, substance metabolism, and injury repair. 7 HNF1A also has an important role in maintaining pancreatic function and regulating insulin expression, glucose transporters, and other molecules. 8 HNF1A can control development and substance absorption in the intestine. 9 This transcription factor participates in the development and progression of many malignant tumors,10,11 and studies of the underlying mechanism will be conducive to the early detection and treatment of cancer, which are important for improving the survival rate of patients.
In this study, we first applied The Cancer Genome Atlas (TCGA) database to analyze the expression and mutations of HNF1A in different types of cancer. Furthermore, we evaluated the association of HNF1A with the infiltration of immune cells and their prognostic value in patients with cancer. Our findings revealed the important role of HNF1A in cancer and provided a potential mechanism related to immune infiltration.
Materials and methods
Expression analysis
The UALCAN database (http://ualcan.path.uab.edu) was used to determine the expression of HNF1A in cancer and normal tissues. This database is based on TCGA, and it contains RNA-seq data and clinical information for various cancers. 12
Mutation analysis
The COSMIC database (https://cancer.sanger.ac.uk/cosmic/) was used to investigate HNF1A mutations in human cancers. 13 This database includes large and comprehensive resources for researchers to rapidly and intuitively perform interactive analyses.
Genome-wide association analysis
The Cancer Regulome tool (http://explorer.cancerregulome.org/) was employed to discover the association of HNF1A expression levels with those of other genes in cancer. The strength of the association was represented by the Spearman correlation coefficient, 14 and the criteria were set as Abs = 0.5 and P < 0.01.
Survival analysis
The Kaplan–Meier plotter database (http://kmplot.com/analysis/) was applied to analyze the prognostic value of HNF1A in cancer. 15 The correlation between HNF1A expression and overall survival in different types of cancer was revealed using Kaplan–Meier survival plots. The log-rank test was used to evaluate the statistical significance of the correlation.
Immune association analysis
The Tumor IMmune Estimation Resource (TIMER) database (https://cistrome.shinyapps.io/timer/) was used to discover the association of HNF1A expression with immune cell infiltration in different cancers. 16 The TIMER algorithm was also adopted to conduct comprehensive prognostic analysis of HNF1A and immune cell infiltration.
Results
HNF1A expression levels in human cancers
To explore HNF1A levels in different types of cancer tissues compared with those in normal tissues, we first analyzed the UALCAN database. The analysis illustrated that HNF1A expression in various cancers was substantially different from that in normal tissues (Figure 1). HNF1A expression was significantly increased in cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), esophageal carcinoma (ESCA), head and neck squamous cell carcinoma (HNSC), kidney renal clear cell carcinoma (KIRC), liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), pheochromocytoma and paraganglioma (PCPG), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), stomach adenocarcinoma (STAD), and uterine corpus endometrial carcinoma (UCEC). Conversely, HNF1A expression was decreased in chromophobe renal cell carcinoma (KICH), thyroid carcinoma (THCA), and cervical squamous cell carcinoma (CESC).

HNF1A mRNA expression was obviously different between cancer tissues and normal tissues based on an analysis of the UALCAN database.
HNF1A mutations in cancer
We applied the COSMIC database to explore HNF1A mutations in different cancers. As presented in Figure 2, the mutation types of HNF1A included nonsense mutations, missense mutations, synonymous substitutions, in-frame insertions, frameshift insertions, in-frame deletions, frameshift deletions, complex mutations, and others. Missense mutation was the main mutation type in breast cancer (48.78%), endometrial cancer (51.52%), hematopoietic and lymphoid cancer (23.53%), kidney cancer (66.67%), cancer of the large intestine (41.91%), liver cancer (54.12%), lung cancer (67.12%), esophageal cancer (35.71%), prostate cancer (37.93%), skin cancer (62.22), soft tissue cancer (50.00%), and cancer of the urinary tract (68.42%). Additionally, C > T and G > A mutations were most commonly observed in the HNF1A coding strand. Other types of mutations were also found in different cancers.

The proportions of different HNF1A mutation types in human cancers based on the COSMIC database (only those with more than two types of mutations in one cancer are presented).
Genome-wide associations with HNF1A mRNA levels in cancer
Subsequently, we applied the Regulome Explorer to analyze the relevant human genome location and the associations between certain genes and HNF1A in human cancer. The associations of HNF1A with other genes were analyzed at multiple levels, including gene expression, DNA methylation, somatic copy-number alteration, somatic mutation, and protein expression. As presented in Figure 3, HNF1A was associated with other genes that could be detected in adrenocortical carcinoma, bladder urothelial carcinoma, breast invasive carcinoma, COAD/READ, colorectal carcinoma, ESCA, STAD, glioblastoma multiforme, HNSC, renal clear cell carcinoma, LIHC, LUAD, LUSC, cutaneous melanoma, and UCEC.
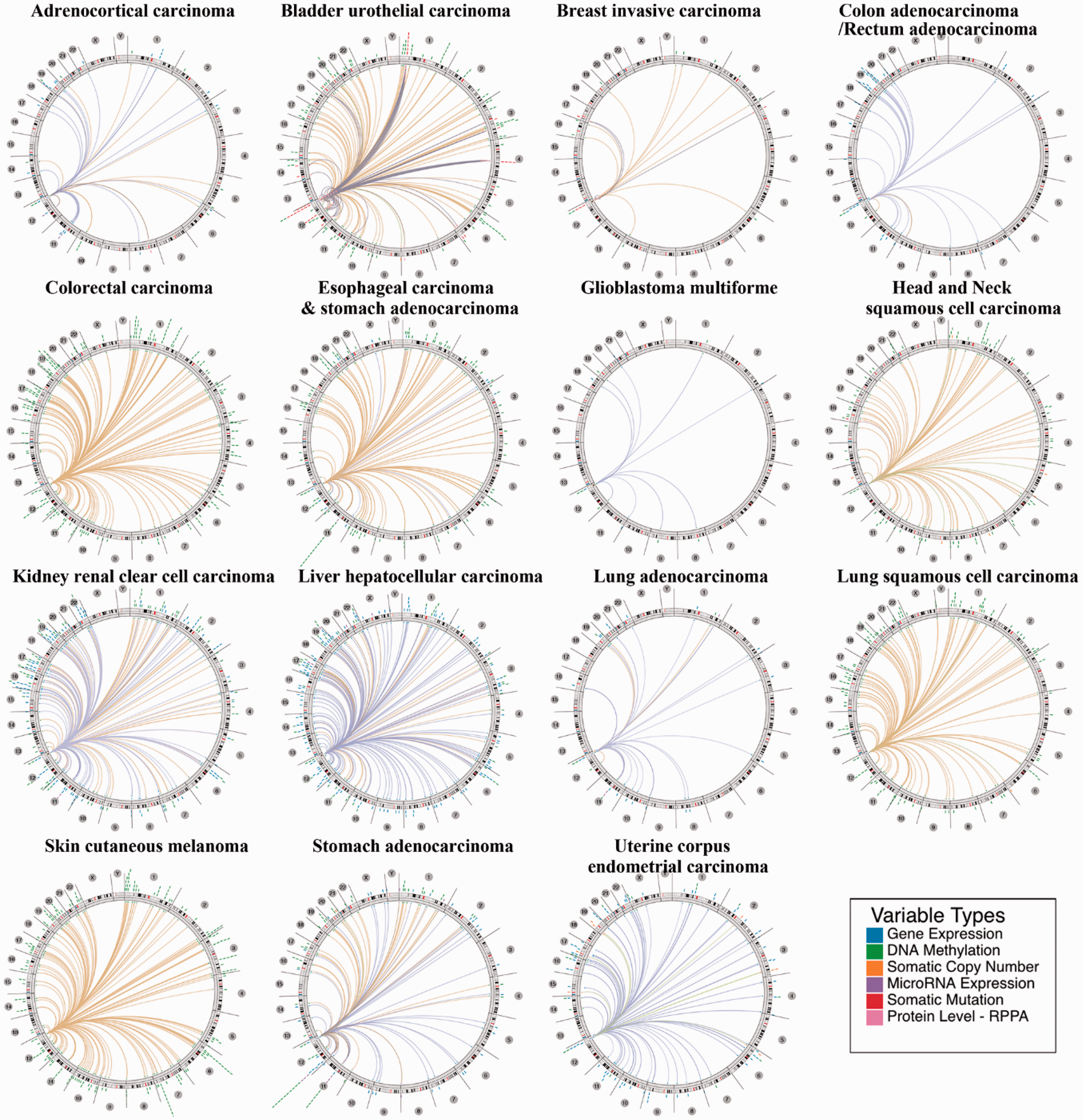
The associations of HNF1A with other genes based on the Cancer Regulome tool. The correlation criteria were set as Abs = 0.5 and P < 0.01, and only the 200 top results for each cancer are presented in the circus plots.
Prognostic value of HNF1A in cancer
Furthermore, we constructed Kaplan–Meier survival curves to explore the potential prognostic value of HNF1A in patients with cancer. As presented in Figure 4, HNF1A was positively correlated with overall survival in bladder carcinoma (hazard ratio [HR] = 0.67, P = 0.0078), breast cancer (HR = 0.52, P = 8.3e−05), HNSC (HR = 0.66, P = 0.0065), KIRC (HR = 0.45, P = 2.2e−07), pancreatic ductal adenocarcinoma (HR = 0.59, P = 0.039), READ (HR = 0.36, P = 0.011), THCA (HR = 0.22, P = 0.0043), and UCEC (HR = 0.56, P = 0.007). However, higher HNF1A expression portended worse overall survival in ESCA (HR = 2.74, P = 0.03), LUAD (HR = 1.39, P = 0.041), ovarian cancer (HR = 1.37, P = 0.019), and LUSC (HR = 1.49, p = 0.0067).
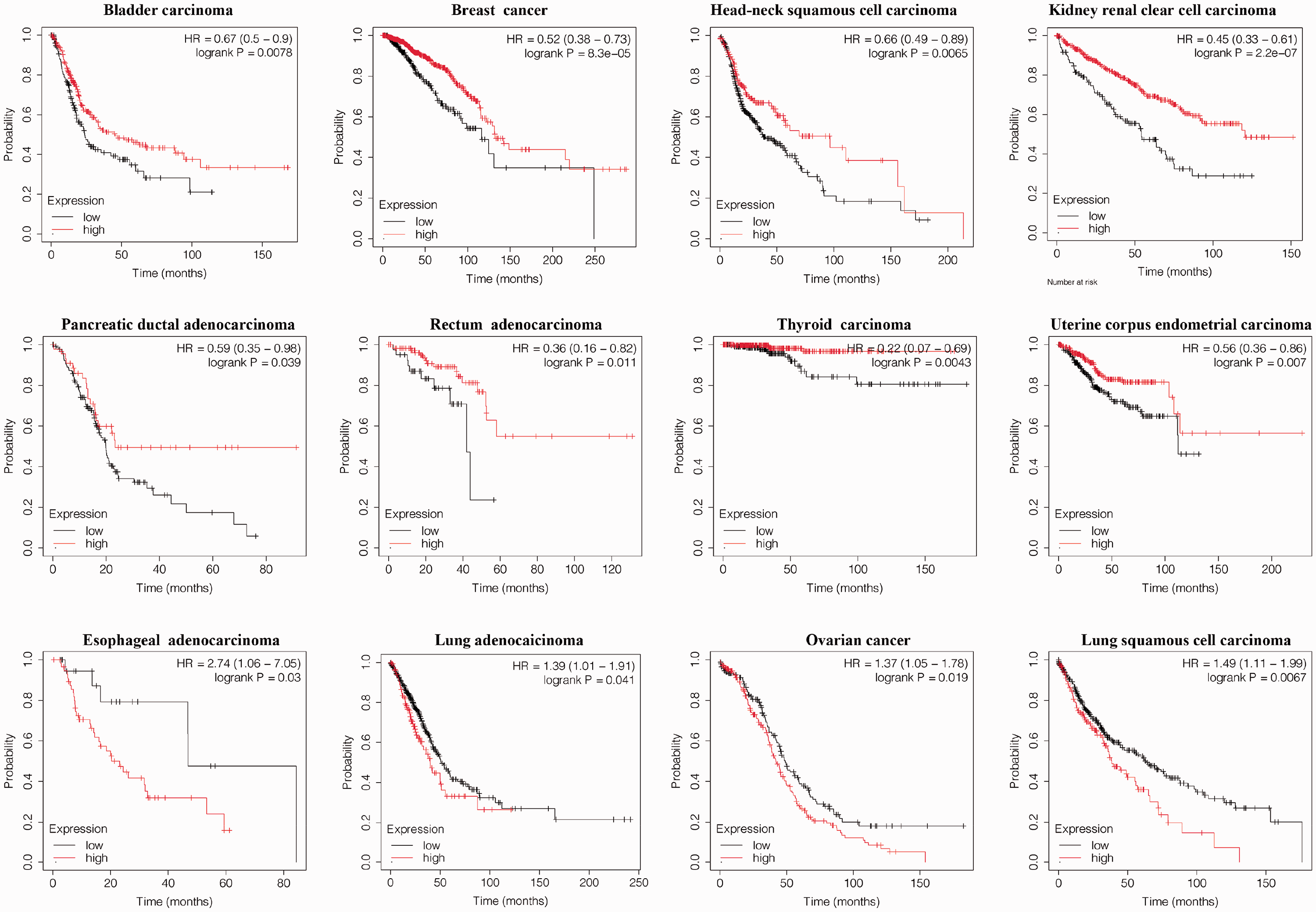
The potential value of HNF1A in predicting overall survival in patients with cancer based on Kaplan–Meier analysis.
Association of HNF1A expression with immune cell infiltration in cancer
Based on TCGA data and the TIMER algorithm, we analyzed the correlation of HNF1A expression with the levels of various infiltrating immune cells in cancer. As presented in Figure 5a, a positive correlation between HNF1A expression and CD8+ T cell infiltration levels was found in diffuse large B-cell lymphoma (r = 0.434, P = 4.62e−03), KICH (r = 0.389, p = 1.34e−03), lower-grade glioma (LGG, r = 0.16, P = 4.63e−04), primary skin cutaneous melanoma (SKCM, r = 0.222, P = 2.50e−02), and UCEC (r = 0.213, P = 4.68e–02), whereas HNF1A was negatively associated with CD8+ T cell infiltration levels in pancreatic adenocarcinoma (PAAD), testicular germ cell tumor, LumA− invasive breast carcinoma, glioblastoma multiforme, HNSC, KIRC, STAD, and thymoma. Moreover, the results indicated that HNF1A was significantly associated with macrophage infiltration in multiple cancers (Figure 5b).

HNF1A expression is associated with CD8+ T cell (a) and macrophage (b) infiltration levels in cancer (TIMER algorithm).
Comprehensive prognostic analysis of HNF1A and immune cell infiltration in cancer
Finally, we applied the TIMER algorithm to explore the comprehensive prognostic value of HNF1A expression and immune cell infiltration in different tumors. As illustrated in Figure 6a, CD8+ T cell counts were positively associated with overall survival in the low HNF1A expression group in SKCM (HR = 0.627, P = 0.0362). In uveal melanoma, CD8+ T cell infiltration was negatively associated with overall survival in the low HNF1A expression group (HR = 24.8, P = 3e−04). Moreover, HNF1A levels influenced the prognostic value of CD4+ T cell infiltration in LGG, PAAD, and sarcoma (Figure 6b). B cell infiltration was revealed to be negatively associated with overall survival in the high HNF1A expression group in LGG (HR = 1.92, P = 0.0115) and COAD (HR = 2.02, P = 0.043) (Figure 6c). High B cell infiltration indicated better overall survival in the high HNF1A expression group in LUAD (HR = 0.601, P = 0.012).

Comprehensive prognostic value of HNF1A expression and CD8+ T cell (a), CD4+ T cell (b), and B cell (c) infiltration levels based on the TIMER algorithm.
Discussion
HNF is a transcription factor involved in controlling liver development and differentiation and maintaining liver function by regulating the expression of downstream genes. 17 HNF1A is a member of the HNF family, which is mainly expressed in the liver, kidneys, and small intestine. 18 In recent years, an increasing number of studies have focused on the relationship between HNF1A and cancer. 19 For example, HNF1A is closely related to the occurrence and development of hepatocellular carcinoma, 20 and HNF1A and HNF4A cooperatively regulate the occurrence of renal cell carcinoma. 21 In our study, we found that HNF1A expression was upregulated at different levels in CHOL, COAD, ESCA, HNSC, KIRC, LIHC, LUAD, LUSC, PCPG, PRAD, READ, STAD, and UCEC and downregulated in KICH, THCA, and CESC. These results indicated that HNF1A may be a good potential diagnostic biomarker for different cancers.
The human HNF1A gene contains 10 exons, and its encoded protein consists of 631 amino acids. 22 Studies revealed that HNF1A gene mutations are closely related to maturity-onset diabetes of the young type 3, type 2 diabetes, pancreatic cancer, and obesity. 23 Our results revealed the wide existence of HNF1A mutations in human cancers. Missense mutations and synonymous mutations were common types. The analysis indicated that C > T and G > A mutations widely occur in tumors. These mutations may cause abnormal transcription and translation, eventually leading to changes in phenotype and function. 24 HNF1A was reported to interact with other genes and pathways in cancer,25,26 which is consistent with our results. These phenomena may explain the distinct prognostic value of HNF1A in different types of cancers. Many molecular targeted therapies have been approved by the Food and Drug Administration, 27 and HNF1A may be a potential target.
The immune microenvironment is an essential component of all tumors, playing a key role in tumor development. 28 In addition, the ability to escape immune destruction has been identified as one of the hallmarks of cancer. 29 Immune-associated therapy is a new type of treatment strategy for cancer with significant curative effects, and its types include nonspecific immunotherapy, cytokine therapy, adoptive cell therapy, and monoclonal antibody therapy.30,31 In this study, HNF1A was found to be significantly associated with CD8+ T cell and macrophage infiltration levels in different cancers. In addition, the prognostic value of immune cell infiltration differed according to the expression of HNF1A. Few studies have assessed the relationship between HNF1A and immune cells, and further research is needed. For patients treated with immunotherapy, the expression of HNF1A may be an important observation index.
In summary, our research revealed that the HNF1A mRNA levels are abnormal in various cancers, and these differences were associated with distinct overall survival rates in patients with cancer. HNF1A mutations are widely found in cancers, and this mutant gene interacts with different genes in different cancer types, which may affect the different prognostic value of HNF1A. In addition, HNF1A is associated with the level of immune cell infiltration and prognosis in some cancers. These results were mainly based on databases, and it is necessary to further confirm the result using clinical samples. We will collect clinical samples and conduct experiments for verification in the future.
Footnotes
Acknowledgments
The authors thank TIMER, UALCAN, COSMIC, and Kaplan–Meier plotter for the availability of the data.
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This study was supported by the Natural Science Foundation of Zhejiang Province (Project No. LQ21H160015) and the Medical and Health Science and Technology Project of Zhejiang Province (Project No. 2019KY070).
