Abstract
Objective
To identify differentially expressed long noncoding RNAs (lncRNAs) in nasopharyngeal carcinoma (NPC) compared with chronic nasopharyngitis (CNP) tissues.
Methods
This prospective cohort study enrolled patients with NPC and CNP. The levels of lncRNAs in NPC and CNP tissues was detected by deep sequencing and quantitative polymerase chain reaction. Kyoto Encyclopedia of Genes and Genomes pathway analysis and antisense prediction were performed to reveal the function of differentially expressed lncRNAs (DELs). Receiver operating characteristic (ROC) curve and correlation analyses were used to evaluate the clinical and prognostic value of lncRNAs.
Results
A total of 30 NPC and 27 CNP tissues were analysed. A total of 296 DELs were identified. LncRNAs ENSG00000227084 and ENSG00000230489 might play important roles in the development of NPC by interfering with the Rap1 signalling pathway and natural killer cell-mediated cytotoxicity, respectively. Antisense prediction showed that lncRNA ENSG00000230489 was paired with VAV3 mRNA. LncRNAs ENSG00000230489 (area under the ROC curve = 0.9138) and ENSG00000227084 (area under the ROC curve = 0.8037) may be diagnostic markers for NPC. Furthermore, low levels of ENSG00000230489 was positively associated with distant metastasis.
Conclusion
LncRNAs ENSG00000230489 and ENSG00000227084 may be potential diagnostic markers and therapeutic targets for NPC.
Introduction
Nasopharyngeal carcinoma (NPC) is an epithelial malignancy characterized by a unique set of geographical, aetiological and biological features distinct from those of other head and neck cancers. 1 The incidence is high in the south-eastern region of China but rare in the western world. 2 Many patients with NPC are missed by the current diagnostic approaches in the early stages, leading to treatment failure. 3 In addition, the high incidence of distant metastasis in NPC is an important risk factor of mortality. 4 To date, radiotherapy and chemotherapy are the major strategies to improve the survival rate of patients. 4 However, the molecular mechanism of NPC remains unknown.
Long noncoding RNAs (lncRNAs) are noncoding RNAs that are usually longer than 200 nucleotides and exert biological functions through their interaction with DNA and proteins. 5 LncRNAs play important roles in transcription control, post-transcriptional processing, chromatin remodelling and protein metabolism. 6 However, little is known about their associations with NPC, although they have been reported as biomarkers specifically expressed in the progression of various tumours. 6
Deep sequencing can be used to explore a large lncRNA pool and has obvious advantages for the identification of lncRNA sequence variations and the discovery of novel lncRNAs. 7 A previous study identified 2127 differentially expressed lncRNAs (DELs) in patients with bladder urothelial cancer, four of which had potential value in the treatment and diagnosis of tumours. 8 However, knowledge of the role of lncRNAs in NPC remains limited.
The aim of the current study was to identify DELs in NPC tissues compared with chronic nasopharyngitis (CNP) tissues. The study then confirmed a subgroup of DELs using quantitative polymerase chain reaction (qPCR) in order to determine a DEL-based risk score in order to investigate the potential regulatory network of these lncRNAs.
Patients and methods
Patient samples
This prospective cohort study enrolled patients with NPC and patients with CNP in the Department of Otorhinolaryngology, Head and Neck Surgery, Shenzhen Baoan Hospital, Southern Medical University, Shenzhen, China between January 2016 and December 2017. None of the patients had received therapy before their biopsy. Fresh tissues were stored in liquid nitrogen for transportation and at –70°C afterwards. The nature of all specimens was confirmed by pathological examination. The Tumor-Node-Metastasis classification was used according to the criteria of the 7th edition of the International Union for Cancer Control and the American Joint Committee on Cancer. 9 Samples of NPC and CNP were used for deep sequencing.
The study was approved by the Ethics Committee of the Affiliated Shenzhen Baoan Hospital of Southern Medical University (no. 2016010631). All patients providing samples for the study were required to provide written informed consent.
RNA extraction and RT–qPCR
Total RNA, containing lncRNAs, was extracted from freshly frozen tissues using a QIAGEN miRNeasy Mini Kit (Qiagen, Hilden, Germany). Sample quality and integrity were assessed using an Agilent 2100 Bioanalyzer system (Agilent Technologies, Santa Clara, CA, USA). A reverse transcription (RT)–qPCR assay was carried out using a PrimeScript™ RT reagent kit (Takara Bio, Kusatsu, Japan) according to the manufacturer’s instructions. The primers were synthesized by Kangcheng (Shanghai, China) and the sequences are listed in Table 1. The cycling programme involved preliminary denaturation at 95 °C for 30 s, followed by 40 cycles of denaturation at 95 °C for 5 s and annealing at 62 °C for 34 s. Relative lncRNA levels were normalized relative to the expression of glyceraldehyde 3-phosphate dehydrogenase and calculated using the 2−ΔΔCt method. Considering the observed fold changes,
Primer sequences used in reverse transcription–quantitative polymerase chain reaction analysis of long noncoding RNAs.

Library preparation and lncRNA RNA sequencing
Isolated total RNA was subjected to ribosomal RNA depletion using a RiboZero Kit (Epicentre, Madison, WI, USA), cut randomly into short fragments and then reverse-transcribed into cDNA (Illumina, San Diego, CA, USA). Sequencing adapters were ligated to both ends of the cDNA fragments. After amplification of the adapter-ligated fragment by PCR, fragments with inserts size between 200 and 400 base pairs (bp) were selected and sequenced using an Illumina HiSeqTM 2500 sequencer (Illumina) according to the standard Illumina protocol. Analysis of the quality and filtering of the raw RNA-sequencing (seq) data to remove low quality reads, adaptor sequences, contaminant DNA and PCR duplicates was performed using the Trimmomatic software version 0.32 (Illumina). The trimmed RNA-seq reads were mapped to the human reference genome hg 19 using TopHat2 software version 2.0.13. 10 Cufflinks software version 1.0.3 was used to process the reads and calculate the transcription level of each gene (University of Washington, Seattle, WA, USA). To obtain the differential expression profiles, the fragment per kilobase of transcript per million mapped reads (FPKM) values were calculated for each lncRNA.
KEGG enrichment analysis
The Kyoto Encyclopedia of Genes and Genomes (KEGG) was used to understand the biological significance of the differentially expressed genes. Fisher’s exact test and χ2-test were used as statistical tests and the threshold of significance was defined based on the false discovery rate and
Statistical analyses
All statistical analyses were performed using IBM SPSS Statistics for Windows, Version 22.0 (IBM Corp., Armonk, NY, USA) and GraphPad Prism (GraphPad, La Jolla, CA, USA). Heat maps were obtained using the R package.
11
Student’s
Results
A cohort of patients, 30 with NPC (7 females and 23 males; mean ± SD age 38.5 ± 14.5 years) and 27 with CNP (8 females and 19 males; mean ± SD age 41.0 ± 16.0 years), were enrolled in the study.
Four patients with NPC and four with CNP were randomly selected and their RNA was extracted for deep sequencing. The sequencing data have been deposited in the Sequence Read Archive database of the National Center for Biotechnology Information, US National Library of Medicine (BioProject: PRJNA451367; SRA: SRP142570). The Cufflinks method was used to identify significantly deregulated genes in NPC (tumour, T) and CNP (inflammation, I) samples. The gene expression level was normalized as FPKM. A correlation analysis of the gene expression among samples was performed using the Pearson’s coefficient (

Long noncoding RNA (lncRNA) expression profile in tissue samples. (a) LncRNA expression profiles in patients with nasopharyngeal carcinoma (NPC-T in the graph) and chronic nasopharyngitis (CNP-I in the graph) using deep sequencing technology. (b) Heat map showing distinguishable lncRNA expression profiles between patients with NPC (T1–T4) and chronic nasopharyngitis CNP (I1–I4). The colour version of this figure is available at: http://imr.sagepub.com.
The top 20 differentially expressed long noncoding RNAs (lncRNAs) in nasopharyngeal carcinoma compared with chronic nasopharyngitis.

To visualize the DELs and further explore the difference between the NPC and CNP samples, hierarchical cluster analysis was undertaken. The results of this analysis showed that lncRNAs could be clearly classified into two groups: the tumour and chronic inflammation groups (Figure 1b).
The KEGG pathway analysis showed that the DELs annotated to the Rap1 signalling pathway, natural killer cell-mediated cytotoxicity and the thyroid hormone signalling pathway were statistically significant (Table 3) (
The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway of enriched differentially expressed target genes.

aHypergeometric distribution.
Rap1, Ras-related protein 1; FoxO, Forkhead box protein O.
The antisense prediction method is recognized as an important strategy for the prediction of the association between lncRNAs and mRNAs. 12 Based on the antisense prediction method, 18 lncRNAs were found to be complementarily paired with mRNAs (Table 4). LncRNA ENSG00000230489 was in the top 20 deregulated lncRNAs and was paired with VAV3 mRNA.
Antisense prediction of long noncoding RNAs (lncRNAs) and mRNA associations.

Reverse transcription–qPCR assays were used to confirm the deep sequencing results for the following six lncRNAs: ENSG00000250451, ENSG00000243479, ENST00000230798, ENST00000228630 (upregulated), ENSG00000230489 and ENSG00000227084 (downregulated). Their expression was consistent with the sequencing results (Figure 2a). LncRNAs ENSG00000230489 and ENSG00000227084 were decreased in NPC tissues compared with their levels in CNP tissues (Figures 2b and 2c).

Validation of the RNA sequencing results. (a) The levels of six long noncoding RNAs (lncRNAs) were evaluated by quantitative polymerase chain reaction in samples from patients with nasopharyngeal carcinoma (NPC;
The lncRNAs ENSG00000227084 and ENSG00000230489 were in the top 20 downregulated lncRNAs in NPC (Table 2). Moreover, lncRNAs ENSG00000227084 and ENSG00000230489 might play essential roles in the development of NPC by interfering with the Rap1 signalling pathway and natural killer cell-mediated cytotoxicity, respectively. The potential diagnostic value of lncRNAs ENSG00000230489 and ENSG00000227084 for NPC was evaluated with CNP patients as the control subjects. The areas under the ROC curves for lncRNAs ENSG00000230489 and ENSG00000227084 were 0.9138 and 0.8037, respectively. These results suggested that they have potential diagnostic value for NPC (Figure 3).

Receiver operating characteristic (ROC) curves for the nasopharyngeal carcinoma samples. (a) ROC curve of ENSG00000230489 (area under the ROC curve = 0.9138); (b) ROC curve of ENSG00000227084 (area under the ROC curve = 0.8037).
The potential diagnostic value of the two lncRNAs was evaluated by a correlation analysis between clinical characteristics and the expression of lncRNAs ENSG00000230489 and ENSG00000227084. This analysis found that a low level of lncRNA ENSG00000230489 but not lncRNA ENSG00000227084 was positively associated with distant metastasis (
Relationship between the levels of two long noncoding RNAs (lncRNAs) and the clinicopathological characteristics of patients with nasopharyngeal carcinoma (NPC;

Data presented as mean ± SD, interquartile range and
aχ2-test and Fisher’s exact test; NS, no significant association (
Since the biological function and clinical value of these two lncRNAs has never been studied, further research is required to elucidate the role of these two lncRNAs in NPC, for example, by determining an lncRNA-mRNA co-expression network (Figure 4).
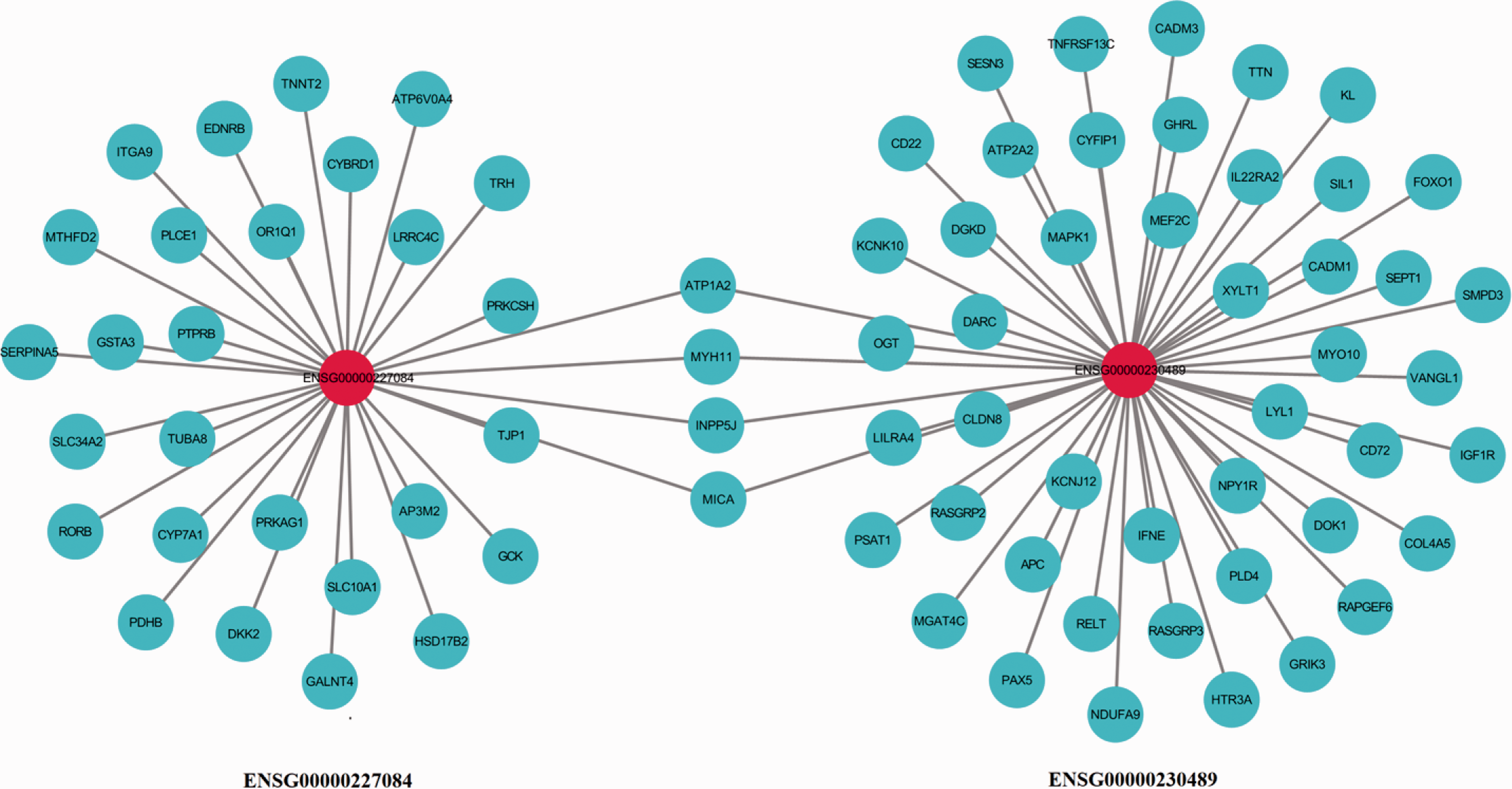
Gene network of the correlative genes of long noncoding RNAs (lncRNAs) ENSG00000230489 and ENSG00000227084. The LncRNAs are shown as red nodes while mRNAs are shown as green nodes.
Discussion
Nasopharyngeal carcinoma is one of the most common carcinomas of the head and neck in China. 2 The molecular features underlying the pathogenesis of NPC remain largely unknown. In addition, early detection of NPC is not easy due the lack of obvious clinical features in the early stages. 13 Increasing evidence suggests that lncRNAs play an important role in the progression of cancer by affecting chromatin remodelling and gene expression. 14 At the same time, lncRNAs have been reported to be biomarkers for predicting tumorigenesis and development in the diagnosis of multiple diseases. 15 Previous studies have demonstrated that a few dysregulated lncRNAs, such as AFAP1-AS, ANRIL and LET, are linked to NPC.16–18 However, the expression signatures and the diagnostic value of lncRNAs in NPC development remain largely unknown.
Deep sequencing has been widely used for clinical detection and has greatly accelerated the discovery and characterization of novel ncRNAs. For example, a previous study demonstrated the expression profiles of microRNAs between NPC and CNP patients by deep sequencing and identified miR-34c as an important gene in NPC tumorigenesis. 19 However, to the best of our knowledge, this current study is the first to use deep sequencing technology to evaluate the genome-wide expression of lncRNAs in tumours and inflammatory tissues derived from the nasopharynx.
The current deep sequencing results were consistent with those of previous studies on the known NPC-related lncRNA ENSG00000228630 (HOTAIR), 20 which is aberrantly upregulated in various types of cancer, including breast, hepatocellular and bladder cancer.21–23 Moreover, lncRNAs ENSG00000250451 (HOXC-AS1), ENSG00000230798 (FOXD3-AS1) and ENSG00000243479 (MNX1-AS1) have been reported to contribute to tumorigenesis in several tumors;24–26 and this current study is the first to confirm their expression in NPC, further supporting their potential diagnostic value in tumorigenesis.
The lncRNAs ENSG00000227084 and ENSG00000230489 (VAV3-AS1) are related to the Rap1 signalling pathway and natural killer cell-mediated cytotoxicity, respectively. A previous study showed that increased and aberrant activation of Rap1 signalling can lead to tumour formation and progression in human breast epithelial cells. 27 Since Rap1 signalling has been increasingly recognized as an important pathway linked to cancer biogenesis and metastasis, 28 it is possible to speculate that lncRNA ENSG00000227084 may be a key factor promoting the development of NPC.
Natural killer cell-mediated cytotoxicity plays an important role in the natural immune defence against tumours. 29 According to KEGG pathway analysis, lncRNA ENSG00000230489 might play important roles in the development of NPC by interfering with natural killer cell-mediated cytotoxicity. In addition, antisense prediction showed that lncRNA ENSG00000230489 was associated with VAV3. A previous study reported that some specific noncoding RNAs can regulate the proliferation and metastasis of tumour cells by targeting VAV3. 30 Furthermore, this current study found that NPC patients with distant metastases had lower ENSG00000230489 levels than those without metastases, indicating that lncRNA ENSG00000230489 may function as a potential upstream regulator of VAV3 as a tumour suppressor.
Using ROC curve analysis to predict the potential diagnostic value of the lncRNAs ENSG00000227084 and ENSG00000230489, the current study found that both of these have diagnostic value for NPC. Moreover, the diagnostic value of ENSG00000230489 was higher than that of ENSG00000227084. Whether lncRNAs ENSG00000230489 and ENSG00000227084 could be potential targets for the treatment of NPC requires further investigation.
In conclusion, this current study provides, for the first time, a global lncRNA expression profile in NPC by deep sequencing. Furthermore, this study suggests that lncRNAs ENSG00000230489 and ENSG00000227084 may be potential diagnostic and treatment biomarkers of NPC.
Footnotes
Declaration of conflicting interest
The authors declare that there are no conflicts of interest.
Funding
The study was supported by grants from the National Natural Science Foundation of China (no. 81470071 and no. 81171652), the Jiangsu Natural Science Youth Fund Project (no. BK20180293) and the Shenzhen Baoan Technology Projects (no. 2016CX173).
