Abstract
Objective:
To investigate the prognostic value of protocadherin 17 (PCDH17) promoter methylation in serum-derived DNA of patients with bladder cancer.
Methods:
DNA was isolated from serum of patients with bladder cancer and from age- and sex-matched controls. Methylation-specific polymerase chain reaction was used to examine the methylation status of the PCDH17 promoter. The correlations between methylation status and clinicopathological characteristics and overall survival were examined.
Results:
PCDH17 promoter methylation was detected in 79/151 (52.3%) of patients with bladder cancer, and none of the 43 control subjects. Methylation was significantly associated with larger tumour diameter (>3 cm), high grade (G3) and advanced stage (T2–T4). Patients with PCDH17 promoter methylation had significantly shorter overall survival than those with unmethylated PCDH17 promoter. Methylation was an independent predictor of overall survival.
Conclusions:
PCDH17 promoter methylation was significantly associated with malignant behaviour and poor prognosis of bladder cancer. The detection of PCDH17 promoter methylation in serum-derived DNA may be a convenient and noninvasive predictive biomarker in routine clinical practice.
Introduction
Bladder cancer is the second most common genitourinary neoplasm, with an estimated 72 570 new cases and 15 210 deaths expected to occur in the USA in 2013. 1 Due to the aging population, the incidence of bladder cancer morbidity is likely to increase in the future and to remain a global healthcare problem. 2 The initial presentation is superficial in 80% of cases, but superficial bladder cancer recurs in 70% of patients in spite of treatment standardization, with 10–30% of these patients developing invasive carcinoma. 3 Survival is negatively correlated with tumour progression. 3 Bladder cancers are histopathologically, morphologically and behaviourally heterogeneous, and it is therefore important to identify which tumours are likely to progress and which patients will need aggressive postoperative adjuvant therapy to improve their prognosis. 4 Conventional prognostic factors such as tumour stage and grade are clinically useful, but their accuracy remains unsatisfactory. 5 Novel, precise prognostic methods are therefore required.
The initiation and progression of bladder cancer is characterized by the gradual accumulation of genetic and epigenetic changes that lead to the activation of proto-oncogenes or inactivation of tumour suppressor genes. 6 The main epigenetic modification in human cancers is DNA methylation. 7 This is a potential prognostic biomarker, especially in cases where the aberrant methylation inactivates tumour suppressor genes. 8 DNA methylation can be detected in both tumour tissue-derived DNA and cell-free circulating DNA in serum. 9 Methylation of circulating DNA can usually be detected if the tumour DNA is methylated, since these DNA fragments show the same characteristics as the primary tumour DNA. 10 Analysis of DNA methylation in serum is sensitive, noninvasive, relatively cheap, and suitable for routine clinical use. 11
The protocadherin 17 (PCDH17) gene is located on chromosome 13q21.2 in humans, and functions as a tumour suppressor. Its TATA-less promoter contains CG-rich sequences that are susceptible to DNA methylation. PCDH17 methylation is found in several human carcinomas, including bladder, gastric and colorectal cancers.12–14 Methylation-specific polymerase chain reaction (MSP) is a simple, sensitive, specific and cost-effective method for detection of DNA methylation. 11 The present study used MSP to investigate PCDH17 promoter methylation in serum samples from patients with bladder cancer, in order to evaluate its prognostic significance.
Patients and methods
Study population
The study recruited patients with primary bladder cancer undergoing treatment at the Department of Urology, Third Hospital of Hebei Medical University, Shijiazhuang, China, between May 2003 and May 2008. Inclusion criteria were: histopathological diagnosis of transitional cell carcinoma of the bladder; no anticancer treatment before blood sampling; no other prior tumour; availability of complete follow-up data. Tumour diagnosis, staging, therapy and follow-up were completed according to international standards. 15 Overall survival was defined as the time from the date of diagnosis to the date of death from any cause, or last contact if the patient was still alive. Age- and sex-matched control subjects were recruited from healthy individuals with no history of any tumour, who were attending the same hospital for routine health checks.
Written informed consent was obtained from each participant. The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of the Third Hospital of Hebei Medical University.
Methylation-specific PCR
Peripheral blood samples were obtained from all the participants before any therapy using standard methods as described previously. 16 Serum was isolated and stored at −80℃ until use. DNA was extracted from 500 µl of serum using a QIAamp® DNA Blood mini kit (Qiagen, Valencia, CA, USA) then treated with bisulphite to convert any unmethylated (but not methylated) cytosine residues to uracil (EpiTect Bisulfite Kit; Qiagen). PCDH17 methylation status was examined using MSP as described previously. 12 Primer sequences were: PCDH17 unmethylated sense, 5′-AGATTATTGGGTGTTGTA GTTT-3′ and antisense, 5′-AACCCTAA CACAACATACACA-3′ (90-base pair product); and PCDH17 methylated sense, 5′-GATTATCGGGTGTCGTAGTTC-3′ and antisense, 5′-CCCTAACGCAACGTA CGCG-3′ (87-base pair product). The PCR cycling conditions were as follows: denaturation at 95℃ for 5 min, followed by 40 cycles of denaturation at 94℃ for 30 s, annealing at 60℃ for 1 min and extension at 72℃ for 1 min, followed by a final extension step at 72℃ for 5 min. In vitro methylated DNA and unmethylated DNA (Millipore, Billerica, MA USA) were used as positive controls for methylation and unmethylation, respectively. Water was used as a negative control in each assay. PCR products were separated in 2% agarose gels, stained with ethidium bromide, and visualized under ultraviolet illumination. Samples were scored as methylation positive when methylated alleles were present in the methylated DNA lane and methylation negative when bands were present only in the unmethylated DNA lane. 16 PCR was performed a minimum of three times per sample.
Statistical analyses
Data were presented as n (%). Fisher’s exact test was used to evaluate the difference in PCDH17 methylation status between patients and controls, and χ2-test was used to assess the relationship between PCDH17 methylation and clinicopathological parameters. Kaplan–Meier survival analysis and log-rank test were used to assess the difference in overall survival between methylated and unmethylated PCDH17. Multivariate Cox proportional hazard analysis was used to estimate the independent prognostic effect of PCDH17 methylation, controlling for classic risk factors (tumour size, stage and grade).17–19 Statistical analyses were carried out using SAS version 8.0 (SAS Institute, Cary, NC, USA). P-values <0.05 (two-sided) were considered statistically significant.
Results
The study included 151 patients with bladder cancer (107 male/44 female; median age 68 years, age range 21–89 years), and 43 age-and sex-matched control subjects (31 male/12 female; median age 67 years, age range 23–87 years). There were no statistically significant between-group differences in age or sex. PCDH17 promoter methylation was detected in 79/151 (52.3%) of patients and none of the controls (P < 0.0001). Representative MSP results are shown in Figure 1.
Representative methylation-specific polymerase chain reaction results for protocadherin 17 (PCDH17) promoter methylation in serum-derived DNA of seven patients with bladder cancer. M: methylated; U: unmethylated; bp: base pairs. Cases 71, 72 and 73 exhibited PCDH17 promoter methylation.
Relationship between protocadherin 17 (PCDH17) promoter methylation in serum-derived DNA and clinicopathological features of patients with bladder cancer (n = 151).
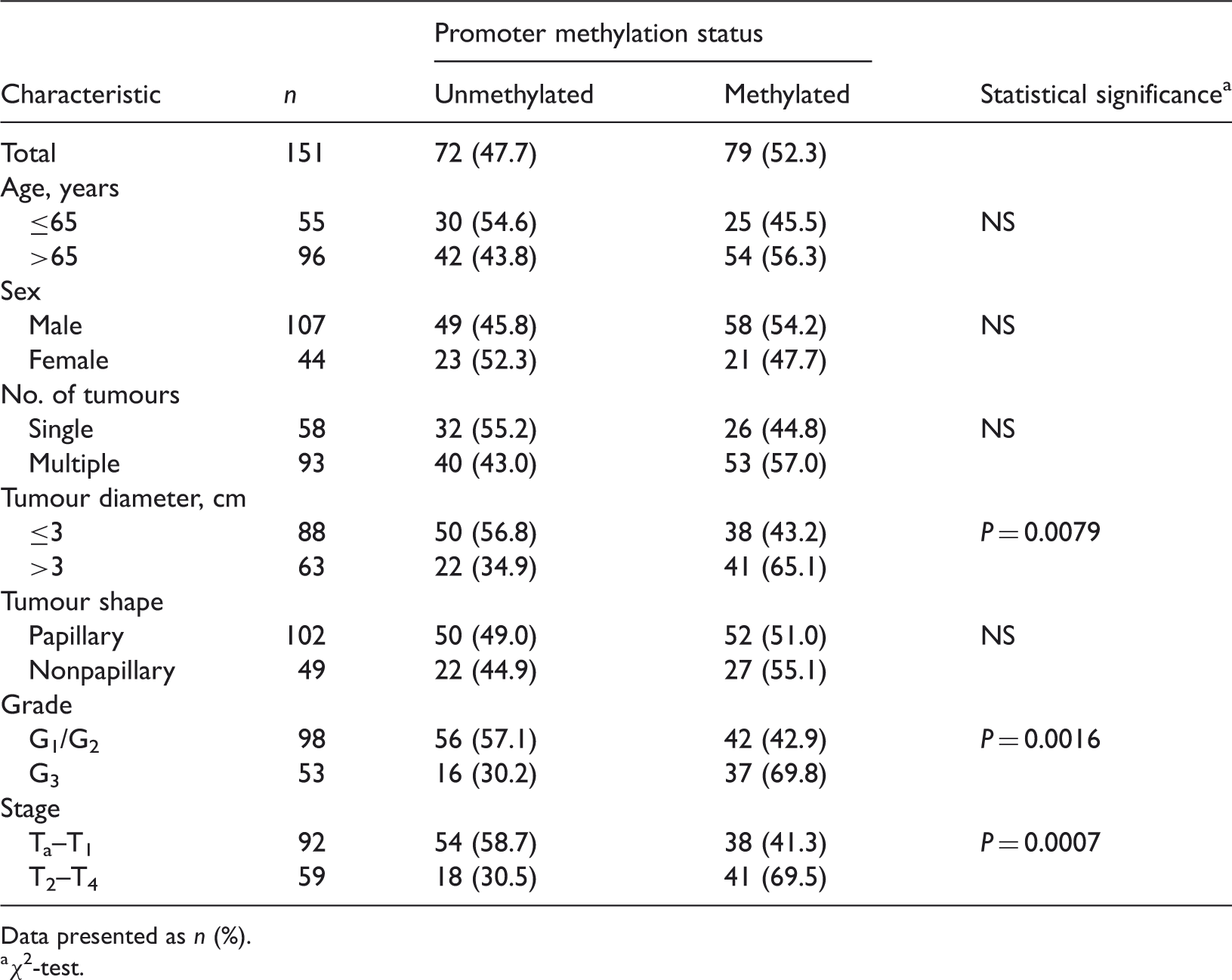
Data presented as n (%).
χ2-test.
Patients with unmethylated PCDH17 had significantly longer duration of overall survival than those with methylated PCDH17 (P < 0.0001; Figure 2). Cox proportional hazard analysis revealed that PCDH17 promoter methylation status, tumour stage, tumour grade and tumour diameter were independent predictors of overall survival (Table 2).
Kaplan–Meier survival analysis of patients with bladder cancer (n = 151), stratified according to protocadherin 17 (PCDH17) promoter methylation status assessed in serum-derived DNA. P < 0.0001; log-rank test. Multivariate Cox proportional hazard analysis of independent predictors of overall survival in patients with bladder cancer (n = 151), controlling for classic risk factors (tumour size, stage and grade). PCDH17, protocadherin-17; HR, hazard ratio.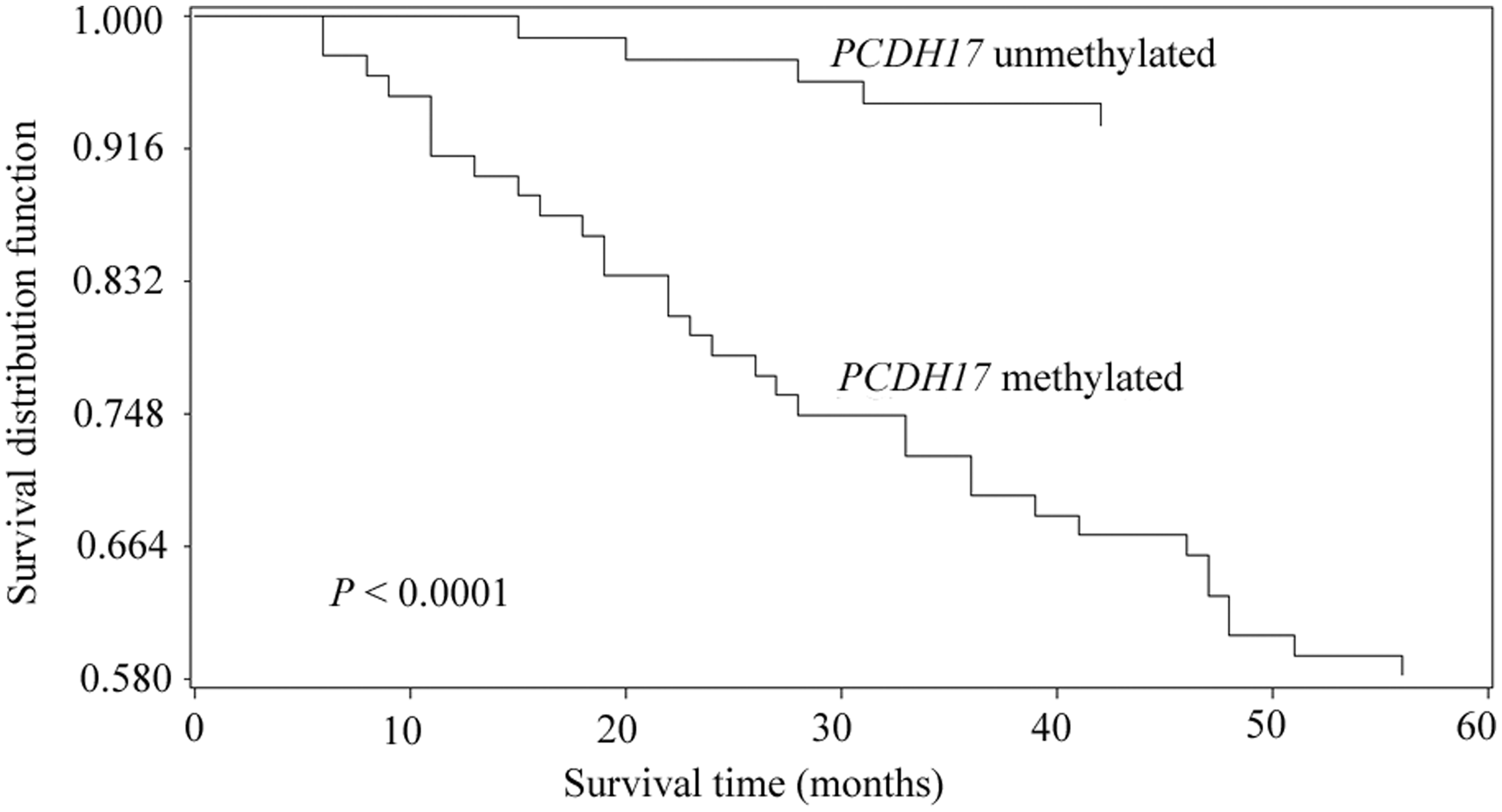

Discussion
DNA methylation is common in human cancers and plays a critical role in the regulation of gene expression, differentiation and tumour development. 20 DNA methylation occurs mainly in cytosine–guanine dinucleotide-rich areas (CpG islands) in gene promoter regions. 20 Promoter methylation may be used as a biomarker in cancer risk evaluation, diagnosis and prognosis, and in determining sensitivity to chemo- or radiotherapy.21–24 DNA methylation can be detected in body fluids as well as tumour tissue, since double-stranded DNA fragments are present in the serum of patients with cancer,9,11,25,26 and this DNA has a similar methylation status to that found in tumours. 4 Circulating tumour DNA may derive from tumour cells or result from DNA leakage caused by tumour necrosis or apoptosis.9,20 The current study investigated the relationship between PCDH17 promoter methylation detected in serum-derived DNA and clinicopathological characteristics and overall survival of patients with bladder cancer.
It has been shown that PCDH17 functions as a tumour suppressor and is downregulated by promoter methylation in human cancers.12,13 PCDH17 promoter methylation is common in urological cancers including bladder cancer, renal cell carcinoma and prostate cancer. 14 PCDH17 promoter methylation was detected in serum-derived DNA of 79/151 (52.3%) of patients but none of the control subjects in the present study, indicating that it is tumour-specific. In the current study, two cases were shown as being both unmethylated and methylated, which can be termed partial methylation. It is well known that serum-derived DNA originates from tumour tissue DNA; DNA methylation is a gradual process in which some DNA is methylated and some is not, thus resulting in partial methylation. In addition, PCDH17 methylation was significantly associated with larger tumour size, high grade and advanced stage in the current study, all of which are classic risk factors for unfavourable outcomes in bladder cancer.27,28 We found that patients with methylated PCDH17 had significantly shorter overall survival than those with unmethylated PCDH17, and PCDH17 methylation was an independent prognostic indicator. These findings suggest that PCDH17 promoter methylation detected in serum-derived DNA is a potential prognostic biomarker in bladder cancer. This study was limited by its small sample size. In addition, we did not use a positive control (a tissue-specific gene). Further larger-scale studies are required to confirm our findings.
In conclusion, this study found that PCDH17 promoter methylation was significantly associated with malignant behaviour and poor prognosis of bladder cancer. The detection of PCDH17 promoter methylation in serum-derived DNA may be a convenient and noninvasive predictive biomarker in routine clinical practice. Patients with PCDH17 promoter methylation may require systemic adjuvant therapy after initial surgery in order to achieve better outcomes.
Footnotes
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
