Abstract
Single-cell RNA sequencing (scRNA-seq) is a high-throughput transcriptomic approach with the power to identify rare cells, discover new cellular subclusters, and describe novel genes. scRNA-seq can simultaneously reveal dynamic shifts in cellular phenotypes and heterogeneities in cellular subtypes. Since the publication of the first protocol on scRNA-seq in 2009, this evolving technology has continued to improve, through the use of cell-specific barcodes, adoption of droplet-based systems, and development of advanced computational methods. Despite induction of the cellular stress response during the tissue dissociation process, scRNA-seq remains a popular technology, and commercially available scRNA-seq methods have been applied to the brain. Recent advances in spatial transcriptomics now allow the researcher to capture the positional context of transcriptional activity, strengthening our knowledge of cellular organization and cell-cell interactions in spatially intact tissues. A combination of spatial transcriptomic data with proteomic, metabolomic, or chromatin accessibility data is a promising direction for future research. Herein, we provide an overview of the workflow, data analyses methods, and pros and cons of scRNA-seq technology. We also summarize the latest achievements of scRNA-seq in stroke and acute traumatic brain injury, and describe future applications of scRNA-seq and spatial transcriptomics.
Overview of next-generation sequencing techniques
Bulk-RNA sequencing
With the advent of high-throughput DNA sequencing technologies, including next-generation sequencing and third-generation sequencing, transcriptomic profiling of RNA molecules has emerged as a major force in biomedical research. RNA sequencing (RNA-seq) can help pinpoint key molecular pathways for mechanistic studies and possible diagnostic use, ushering in a new era of biological discovery. Indeed, bulk RNA-seq has become the most valuable and extensively used tool in understanding cancer biology from when the first expressed sequence tag (EST) library was developed using the Roche 454 Sequencer in 2007. 1 Bulk RNA-seq has shaped our understanding of RNA biology, improved disease classification methods, identified novel biomarkers, and accelerated gene fusion discoveries. 2 While highly effective and robust, the limitations of bulk RNA-seq in resolving the complexities of heterogeneous cell populations have also become evident. For instance, commonly used deconvolution approaches are mainly based on deep learning-based algorithms 3 or pseudo-bulk simulation strategies 4 that are available to mimic biological variance across cells in bulk RNA-seq. However, the deconvolution methods assume cell homogeneity and depend heavily on the quality and completeness of the reference data, which refer to known datasets that infer the composition of cell types, usually consisting of gene expression profiles from individual cell types. Additionally, given the differences in cell size or library size, those approaches are more susceptible to bias. Thus, bulk RNA-seq cannot distinguish individual cell populations, as it only measures the average gene expression profiles in the tissue or cell extracts. 5 Because of an intense demand to capture individual cell variations more precisely and identify rare cell types with potential roles in disease progression or responses to treatment, advancements in the RNA-seq techniques, such as single-cell RNA-seq (scRNA-Seq) and single-nucleus RNA-seq (snRNA-Seq), have profoundly altered the biomedical research landscape.
Single-cell RNA-sequencing
In 2009, the first single-cell transcriptome analysis based on next-generation sequencing was published. Tang et al. successfully detected 75% more genes than prior methods, using a single-cell gene expression profiling assay from a single mouse blastomere. 6 Since then, scRNA-seq technologies have enabled groundbreaking studies of gene expression at the level of individual cells, providing fresh insights into cellular heterogeneity and cell-to-cell variation that were masked in bulk RNA-seq analyses.6,7 The first applications of single-cell RNA sequencing to study brain and retina cells were published in 2015.8,9 Zeisel et al. used scRNA-seq to profile gene expression in individual cells from the mouse cortex and hippocampus, uncovering the molecular diversity of neural cell populations in these telencephalic regions. 8 Concurrently, Drop-Seq was first introduced to analyze mRNA transcripts and created a molecular atlas of gene expression of retinal cells by Macosko et al. 9 By analyzing thousands of single cells, researchers were now able to identify distinct cell types, characterize their transcriptional profiles, and better understand the functional specialization of neuronal and non-neuronal cell populations within the brain. Subsequently, single-cell RNA transcriptomic techniques have been leveraged for comprehensive investigation of embryonic and adult mouse brains. 10
Single-nucleus RNA-sequencing
The introduction of “Div-Seq” on individual, recently-divided cells in 2016 rendered single-nucleus RNA-sequencing (snRNA-seq) a more versatile method compared to scRNA-seq.11,12 For instance, snRNA-seq provides the flexibility to choose among fresh, frozen, or fixed samples while preserving partial spatial information. This method helped identify interconnected, rare, or unstable cells that could be lost during the tissue dissociation process required by scRNA-seq. Therefore, the transcriptomic information collected from snRNA-seq is potentially more reliable for the interpretation of normal and pathological conditions. 13 On the other hand, snRNA-seq has much lower throughput than scRNA-seq, as it exclusively harvests transcripts from the nucleus, where RNA is less abundant and may not be fully processed or mature. Nuclear RNA also may not represent the entire cellular transcriptome. Thus, as snRNA-seq is unable to provide sequence data on cytoplasmic RNA, the transcriptomic heterogeneity revealed by snRNA-seq is lower than scRNA-seq.13,14 Additionally, several studies comparing the two methods indicate that they share similar data quality and resolution but capture different cell types within the same tissues.14–16
Spatial transcriptomics
In 2016, Ståhl et al. introduced the concept of “spatial transcriptomics”, which allows researchers to visualize and map the location of RNA transcripts within intact tissues using positional molecular barcodes. 17 Incorporating spatial information in snRNA or scRNA analysis now allows for the direct determination of spatial patterns of recognized ligand-receptor interactions. 18 Two broad categories of spatial analysis include fluorescence in situ hybridization (FISH)-based (Seq-FISH 19 and MERFISH 20 ) and sequencing-based upon scRNA-seq.17,21 Spatial transcriptomics at single-cell resolution has recently been introduced with advanced features offered by CosMx 22 and Xenium. 23 These platforms now enable researchers to directly resolve gene expression profiles at the single-cell level in an intact spatial context, with the potential of uncovering specialized cellular roles of individual cell types and revealing molecular and cellular interactions.
While recent achievements in high-resolution sequencing technologies have provided insights into the spatial organization of cell populations and the localized networks of intercellular communication in situ, application of these methods to stroke or acute brain injury is still under development. The objectives of this review are to streamline a workflow for scRNA-seq data analyses and to examine how these analyses have advanced our understanding of the pathogenesis of stroke, acute traumatic brain injury, and neural recovery—through the lens of diversity in cellular subtypes within the central nervous system. We will also discuss limitations and challenges associated with single cell sequencing techniques and offer our perspective on the future applications of these technologies in stroke and brain injury.
ScRNA-seq workflow
Somatic alterations, transcriptional regulations, or epigenetic modifications result in cellular heterogeneity within an organism, as reflected at the transcriptomic level. The development of scRNA-seq technologies allows researchers to explore changing gene expression patterns at the level of individual cells. 24 After the initial documentation of this method in 2009, the field has witnessed concerted efforts aimed at refining and scaling up this technology (Figure 1). Noteworthy advancements include the incorporation of unique molecular identifiers to ensure precise RNA molecule quantification, 25 the implementation of cell barcoding techniques for enhanced sample multiplexing, 26 the adaptation of microfluidics for improved cell capture efficiency, 27 the optimization of experimental protocols to enhance data quality, and the streamlining of computational analysis workflows to facilitate data interpretation and insight generation.
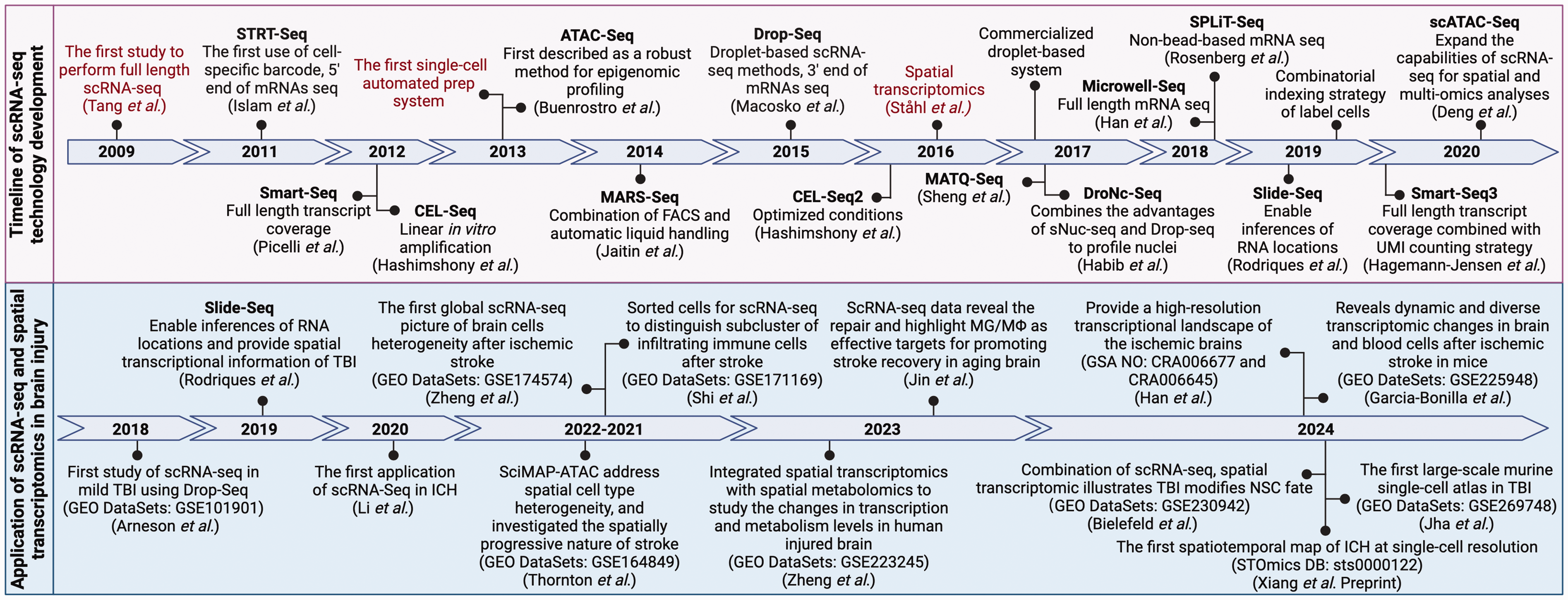
Timeline of scRNA-seq technology development and its applications in stroke and brain injury. A comprehensive timeline of the development of single-cell RNA sequencing (scRNA-seq) technology since 2009 (top panel). Key milestones in technological advancements are highlighted in red. The significant applications of scRNA-seq and spatial transcriptomics for revealing cellular heterogeneity in stroke and brain injury are listed in the bottom panel.
The main workflow of scRNA-seq shares a similar foundation as the protocols of bulk RNA-sequencing. A typical scRNA-seq experiment and analysis pipeline starts with sample preparation, followed by single cell/nucleus capture, RNA capture, reverse transcription, library preparation, sequencing, raw data processing, and downstream bioinformatic analyses. The use of unique cell barcodes enables accurate quantification of gene expression at the single-cell level and addresses some of the challenges inherent in conventional bulk RNA-seq methodologies. Additionally, sample preparation is crucial to ensure the accuracy and reliability of single-cell transcriptome data, and the protocols vary depending on the cell type being analyzed. For instance, extracting cells from peripheral blood is relatively easier, compared to extracting cells from solid tissues, as the former are already suspended as single cells in a liquid matrix. 28 Details of a typical dissociation workflow and technical overviews have been described previously. 29
scRNA-seq data analysis
The output of scRNA-seq undergoes a rigorous and multifaceted bioinformatics analysis pipeline, encompassing a series of intricate steps to extract meaningful biological insights from the massive amount of sequencing data generated (Figure 2). Currently, there is no universally accepted standard pipeline for conducting such analyses. In general, the workflow encompasses several main processing stages, including library preparation and template amplification, base calling, sequence mapping and alignment, and dataset quality control (QC) and processing, cell type annotation, followed by downstream biological characterization, which deciphers the underlying functions and mechanisms.
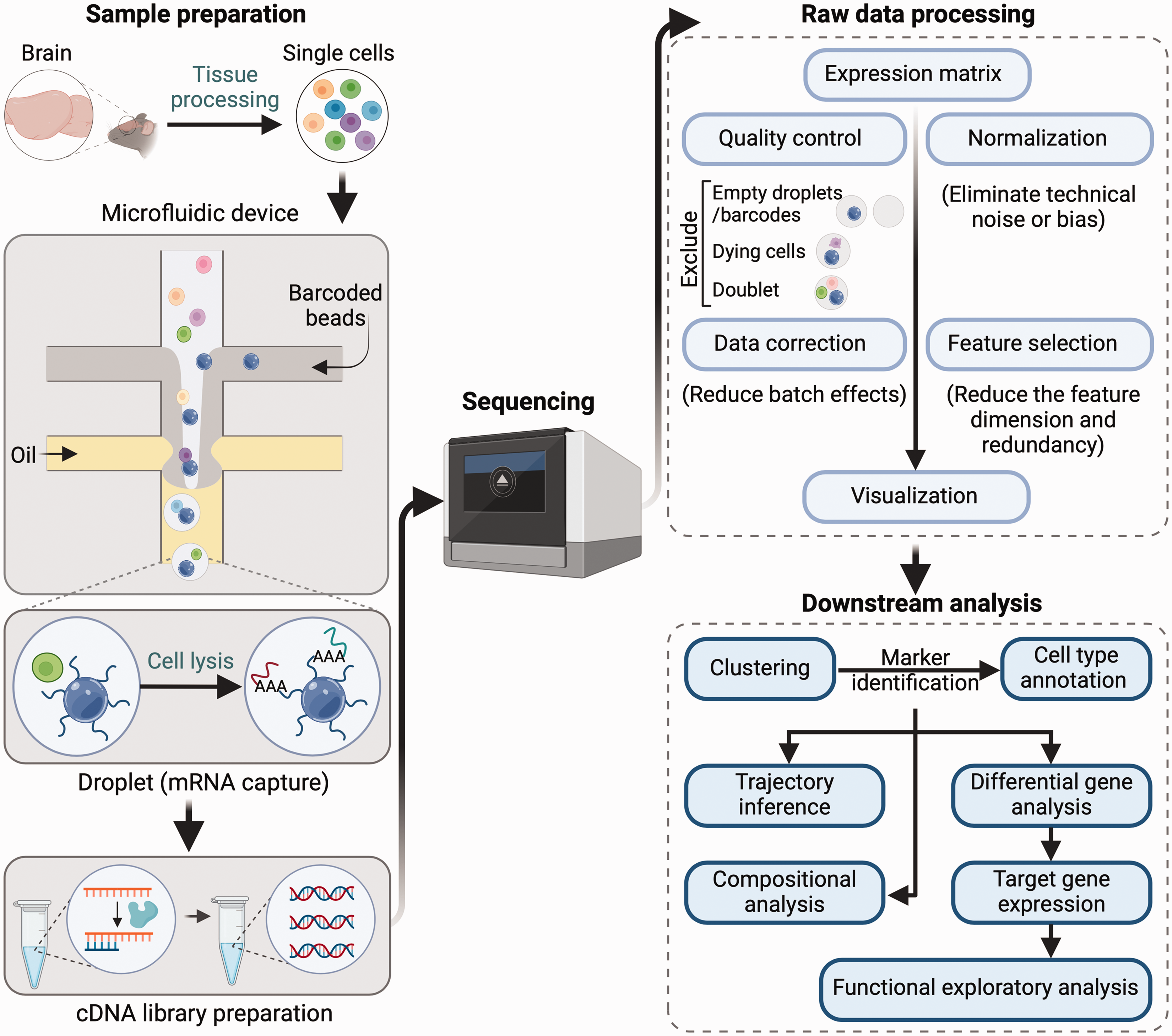
A typical single-cell RNA sequencing analysis workflow. A general workflow for single-cell RNA sequencing (scRNA-seq) includes several key steps: 1) Isolation of qualified single cells (viability >85%) from brain tissue; 2) Single-cell barcoding; 3) mRNA capture and cDNA amplification; 4) cDNA library preparation by reverse transcribing primed RNA into complementary DNA for sequencing; 5) Raw data processing, including conversion of sequence libraries into analysis-ready expression matrices, quality checks, data correction, normalization, and feature selection using bioinformatic tools; 6) Downstream data analysis and interpretation using advanced and specialized tools tailored to specific research interests. This process includes clustering, cell type annotation, trajectory inference, differential gene analysis, and pathway analysis etc.
During the primary base calling stage, raw BCL format data files are converted into FASTQ files. If samples were multiplexed before sequencing, an additional data demultiplexing step may be necessary before sequence alignment. Following the mapping and alignment of reads to the reference genome, collapsing of unique molecular identifiers (UMIs) is required to accurately measure gene expression levels at single-cell resolution. A common tool used for sequencing data processing pipeline is 10x Genomics Cell Ranger, which produces a filtered formatted matrix containing gene data from cells with valid barcodes, serving as the starting point for downstream analyses. 30 Quality control steps are applied at both the cell and gene levels, with advanced options of doublet detection and removal of ambient RNAs. Subsequent data scaling and normalization is essential to mitigate technical variations or sample biases, and feature selection techniques are employed to identify genes that distinguish different cell types based on their expression profiles. 31 As this technology has matured, more analytic tools have been developed (Table 1, Supplementary Material). R and Python are two widely used programming languages, with streamlined pipelines implemented as general guides, such as Seurat built with R, and SCANPY built with Python, along with other web-based tools. Comprehensive tutorials describing the full analytic process of single-cell RNA sequencing are also available and can be practiced with publicly available data.32–34
Available packages in basic scRNA-seq data processing developed after 2015.
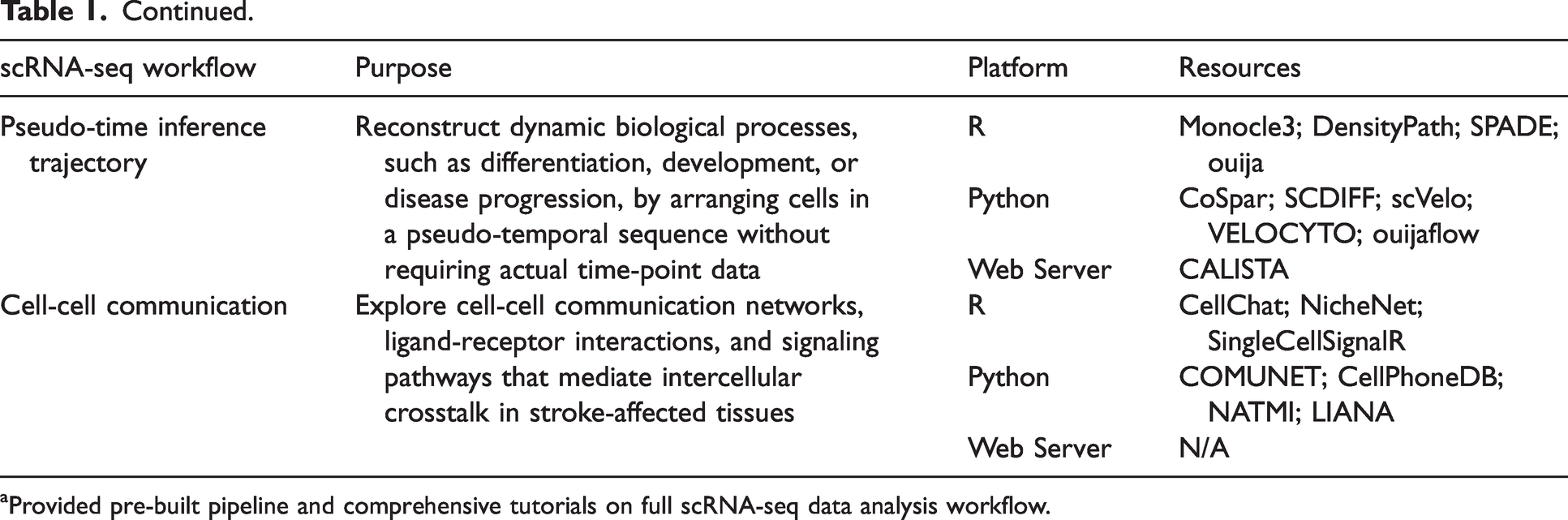
Provided pre-built pipeline and comprehensive tutorials on full scRNA-seq data analysis workflow.
Following the initial phases of data processing, researchers transition towards characterization of the biological and functional aspects of the data, to understand molecular mechanisms underlying cellular heterogeneity. As a high-throughput method, single-cell transcriptomics results in sparse gene expression across cells. Thus, effective dimension reduction and visualization algorithms are crucial for downstream analysis. Among these algorithms, Uniform Manifold Approximation and Projection (UMAP) and t-distributed Stochastic Neighbor Embedding (t-SNE) are the most popular techniques for illustrating meaningful cell clusters with high reproducibility. UMAP is designed to be more scalable than t-SNE and can handle large-scale datasets more efficiently, while t-SNE offers the advantage of preserving local structures.35,36 The next steps involve selecting feature genes for each cell group, focusing on differentially expressed genes. Various feature gene selection algorithms exist, including those based on Bayesian methods generalized linear models,37,38 and other machine learning approaches. 39 A single toolkit may incorporate multiple algorithms for researchers to choose based on their specific needs. Once meaningful clusters or cell types are identified, the researcher can probe more deeply and compare specific cells across treatment or injury groups. Further exploration includes functional enrichment analysis, cell trajectory inference, and cell communication prediction. Gene-based enrichment analyses aim to decipher gene functions by mapping genes to clusters with known functions. For example, the ability to obtain dynamic information on thousands of cells from a single system over time allows the researcher to capture diverse cell types involved in developmental processes and to infer developmental trajectories. In addition, by leveraging ligand-receptor databases and single-cell gene expression data, potential interactions between different cell types can be computed and predicted. With advanced computational tools, researchers can gain a deeper understanding of gene expression patterns, functional pathways, transcriptional regulation, cell-cell interactions, and developmental trajectories at the single-cell level (Table 1).
ScRNA-seq applications in acute brain injuries
Stroke injury is initiated by acute disruption of cerebral blood flow (ischemic stroke) or sudden bleeding of brain vessels (hemorrhagic stroke), which eventually results in brain damage and a wide range of long-term neurological deficits, including motor impairments, loss of sensory function, cognitive impairments, and speech difficulties. 40 According to the latest Global Burden of Disease (GBD) report, ischemic stroke accounts for approximately 62.4% of all strokes, and the remaining are ascribed to hemorrhagic strokes, including 27.9% intracranial hemorrhages (ICH) and 9.7% subarachnoid hemorrhages (SAHs). 41 Ischemic and hemorrhagic strokes share many risk factors, such as hypertension, diabetes, hyperlipidemia, smoking, obesity, and a sedentary lifestyle. In addition to lifestyle, aging and sex are also significant risk factors. 41 In contrast to multifaceted risk factors for stroke, traumatic brain injury (TBI) is mainly caused by falls and road injuries. 42 On the other hand, lifestyle, age, and sex also partly impact functional recovery following mechanical brain trauma. Thus, it is important to understand the molecular mechanisms underlying brain damage, explore potential therapeutic approaches against brain injury, and find effective means of promoting functional recovery after acute brain damage.
Over the past decade, scRNA-seq has rapidly emerged as a new technology for uncovering brain complexity. Following the work of Zeisel et al. 8 and Tasic et al., 43 scRNA transcriptomics has been largely focused on embryonic and adult mouse brains. Recently, scRNA-seq has begun to deepen our understanding of the molecular mechanisms of neurological diseases, including stroke and traumatic brain injury, by refining traditional cell groups into more subtypes, discovering novel or rare cell subsets or transitional cells, and through spatial transcriptomic approaches, providing a complete transcriptomic profile of cell subsets at a high spatial resolution (Figure 1).44–46
Heterogeneity of the neurovascular unit, cells in perivascular spaces, and infiltrated immune cells in stroke and traumatic brain injury
The neurovascular unit (NVU) is an essential structural and functional unit in the brain parenchyma, which incorporates vascular cells (endothelial cells, pericytes, smooth muscle cells), glial cells (astrocytes, microglia, oligodendrocytes), neurons, and the extracellular matrix surrounding endothelial cells. The functional coordination of NVU cells, also called neurovascular coupling, matches blood flow to adjacent neuronal activity and regulates the integrity of the blood-brain barrier (BBB). NVU dysfunction is widely investigated in acute brain injury, although interactions between the NVU and peripheral-derived immune cells are also important mediators of neurological outcomes. Breakdown of the BBB after acute brain injuries and the release of chemokines by microglia and endothelial cells result in extravasation of patrolling immune cells, including monocytes/macrophages, neutrophils, T cells, B cells, and dendritic cells, into the CNS. The infiltration of peripheral immune cells is attributed to both BBB and blood-cerebrospinal fluid barrier (BCSFB) leakage. The permeability of the BCSFB under physiological conditions is regulated by tight junctions between a layer of choroidal epithelial cells with higher permeability than the BBB. Peripheral immune cells, such as CD4+ T cells, CD8+ T cells, NK cells, dendritic cells, and monocytes are rarely detected in brain parenchyma but have been detected in the cerebrospinal fluid (CSF) of normal brains. 47 Thus, the BCSFB is more permeable to circulation-derived neurotoxic substances and invasion of immune cells, especially after brain injury.
The CSF is predominantly synthesized by the choroid plexus and circulates in the ventricular system and the subarachnoid space to provide buoyancy and cushioning of the entire CNS. 48 In addition to basic cellular components (such as fibroblast cells, endothelial cells, and ependymal cells), CNS-associated macrophages (CAMs), also named border-associated macrophages (BAMs), inhabit the perivascular spaces surrounding vessels, meninges, and the choroid plexus where the BCSFB separates the CSF from blood. 49 The function of these specific CNS resident macrophages has not been investigated as intensely as the professional phagocyte of the CNS, the microglial cell. 50 The BCSFB contacts the brain parenchyma, and its components function as an interface between neurons and glial cells in the CNS and immune cells from the peripheral circulation, suggesting that the BCSFB plays a role in brain injury. 49 In the next section, we will discuss the heterogeneity of the NVU, BAMs and ependymal cells distributed in perivascular spaces, and infiltrated immune cells, as revealed by scRNA-seq analysis after stroke and brain injury.
Brain resident cells
Neurons
scRNA-seq and snRNA-seq have been used to distinguish multiple neuron types in the injured brain, such as GABAergic neurons (Rgs9, Rgrb, Adarb2, Gad1, Gad2), glutamatergic neurons (Satb2, Sv2b, Slc17a7), pyramidal neurons (Cux2), and interneurons (Sv2c, Sst, Vip, Chat).51–53 One study investigating neuron proportions at different time points after stroke injury found that the proportion of ipsilateral neurons decreased to a minimum at 3 days, and then gradually rose to normal levels. Contralateral neurons also tended to show proportions higher than those observed in the sham group, except at day 1. Two subpopulations of injury-related neurons were observed: one transient cluster present mainly at 4 hours post-stroke on the ipsilateral brain and another persistent cluster found in both hemispheres. Functional analyses showed that the transient neuron subpopulation expressed genes associated with inflammatory and immune responses. 54 Another scRNA-seq study also reported high levels of chemokines such as Ccl4 in post-stroke neurons. 51 However, neurons not only contribute to inflammatory and immune responses but are also actively involved in post-injury repair processes. Nakamura et al., investigated neurons by snRNA-seq in the peri-infarct region post-stroke and found that glutamatergic neurons upregulated genes involved in nervous tissue reconstruction, synaptic organization, and remyelination at 4 days post-injury. 53
Arneson et al. were the first to systematically analyze TBI-induced hippocampal cell damage, utilizing Drop-seq to identify known cell types such as GABAergic interneurons (Gad1, Gad2), DG granule cells (Dsp, Sema5a), four subtypes of CA pyramidal neurons (Wfs1, Ptn, Atp2b1, Ly6e, Dcn), and two novel neuronal subpopulations. The latter novel subpopulations do not express established neuronal subtype markers but do express genes related to differentiation or self-renewal (Tshz2, Ddit4l, Nrep, Ndnf, Marcks). 44 It is unclear whether brain injury affects the balance between neurogenesis and astrogliogenesis, as neural stem cells can differentiate into both neurons and astrocytes. Bielefeld et al. reclustered neural stem cells, radial glia-like cells, neuron-like cells, and astrocyte-like cells in hippocampal tissue during the chronic phase (15 days) after TBI, categorizing them into neuronal versus astrocytic lineages. Their scRNA-seq analysis revealed that brain injury does not significantly affect the differentiation pathway of neural stem cells into neuronal or astrocytic lineages. Rather, the differentiation trajectories remain unchanged. However, the balance between neurogenesis and astrogliogenesis is disrupted, favoring neurogenesis at the expense of astrogliogenesis. Using molecular cartography spatial transcriptomics, Bielefeld et al. also found that neural stem cell-derived neuronal and astrocytic lineages are anatomically misaligned within the injured hippocampus. Specifically, TBI causes a lower percentage of neuronal lineage (N-stage 3) cells in the subgranular zone (SGZ) and a higher percentage in the granule cell layer (GCL). Similarly, there is a lower percentage of astrocytic lineage (A-stage 1) cells in the SGZ and a higher percentage in the hilus region of the dentate gyrus (DG). 55 Shi et al. conducted snRNA-seq on amygdala tissues extracted after TBI and discovered that brain injury amplifies decorin (Dcn) expression in amygdala neurons. Furthermore, stereotactic injections of AAV-hSyn-Cre into the amygdaloid complex of Dcnflox/flox mice to knock down Dcn expression markedly alleviated the TBI-associated fear conditioning response. 56 Due to technical limitations of sample preparation for scRNA-seq, snRNA-seq has so far been more popular in revealing neuron-related molecular mechanisms after injury.
Astrocytes
Astrocytes are an abundant type of neural cell, ranging from 20% to ∼40% of cells in different regions of the CNS. In the adult CNS, astrocytes are classified into protoplasmic astroglia of the gray matter and fibrous astroglia of the white matter. In the healthy brain, astrocytes primarily express markers such as Aldoc and Mfge8. However, astrocytes undergo significant phenotypic changes following stroke or traumatic brain injury. A new cluster of reactive astrocytes emerges upon ischemic stroke and is characterized by high expression of markers such as Gfap, Serpina3n, and Cd44.51,57–59 The functional heterogeneity of astrocytes has been reported previously as diversification into at least two astrocyte types: A1 (neurotoxic/pro-inflammatory) and A2 (neuroprotective/anti-inflammatory), akin to the M1/M2 classifications in macrophages and microglia. 60 However, recent single-cell analyses in the stroke brain suggest that astrocyte heterogeneity is more complex than a binary categorization. 57 Spatial transcriptomics shows that the A1 and A2 markers are not region-specific but highly expressed around the lesion at 7 and 21 days after hemorrhagic stroke. 61 Bormann et al. further categorized injury-associated astrocytes into 3 clusters (AS_3 to AS_5) at 48 hours after stroke. Both AS_3 and AS_4 clusters are involved in maintaining the BBB and are characterized by high expression of tight junction proteins such as Tjp1 and Tjp2. AS_3 is implicated in interactions with peripherally infiltrated cells, involving chemokine signaling (e.g., Ccl2, Shc3, Shc4) and myeloid leukocyte migration/adhesion. On the other hand, AS_4 is associated with the VEGF response, characterized by the high expression of genes such as Nrp2, Rock2, and Hspb1. One subgroup is associated with a VEGF response (e.g., Nrp2, Rock2, Hspb1), while the other is implicated in interactions with infiltrating peripheral cells, involving chemokine signaling (e.g., Ccl2, Shc3, Shc4) and myeloid leukocyte migration/adhesion. Notably, a subset of reactive astrocytes AS_5 appears in both the ipsilateral and contralateral hemispheres, associated with cilium function and axonemal assembly and movement (expressing Rfx3, Dnah9, Cfap46). These data suggest that focal injuries such as stroke may cause the activation of astrocytes in the non-injured hemisphere. 59 Kim et al. used snRNA-seq to identify 3 clusters almost exclusively present in the stroke brain (cluster_8: Lcn2, Serpina3n, S1pr3, Ptx3; cluster_12: Mki67, Top2a; cluster_13: Oasl2, Gbp2, Gbp5, Gbp6, Gbp7). Cluster_8 displays functional shifts over time. In the acute (within 3 days) phase, Cluster_8 upregulates genes related to signaling molecule secretion, immunomodulation, and synaptic structure, but in the chronic (14 days) phase, Cluster_8 primarily transitions to functions related to neuronal support and tissue remodeling. 62 Additionally, Shi et al. compared chronic astrocytes from different types of stroke injuries and found that hemorrhagic stroke astrocytes included a lower proportion of subpopulations involved in synaptic pruning compared to ischemic stroke astrocytes, indicating heterogeneity of stroke subtype-specific astrocytes. 63
In TBI, Koupourtidou et al. identified three clusters of injury-associated astrocytes: AG5 (Gfap, Vim, S100a6), AG6 (Id3, Ckb, Fth1), and AG7 (Hmox1, Cdkn1a, Gfap, Vim). AG6 and AG7 are predominantly present at 3 days post-insult, whereas AG5 is mainly observed at 5 days post-TBI. 64 AG5 shares transcriptional similarities with certain microglial and oligodendrocyte precursor cells (OPC) subpopulations and is primarily involved in cell proliferation, innate immunity, and type I interferon signaling pathways. 64 Qiu et al. also found that astrocytes upregulate several non-canonical neurotrophic factors during the subacute phase of TBI, including midkine (MDK), pleiotrophin (PTN), and prosaposin (PSAP). 65
In addition to transcriptional heterogeneity, reactive astrocytes also show spatial heterogeneity. Astrocytes closest to the injury core express high levels of Clcf1, Fosl1, Tnfrsf12a, Flnc, and Tuba1c, which are associated with myeloid leukocyte migration/adhesion. Those situated slightly further from the core express high levels of Lcn2, Gfap, Serpina3n, Timp1, Ifitm3, Pclaf, Top2a, Rrm2, Tk1, and Birc5, which are related to inflammatory responses and cell proliferation. 58 These reactive and proliferative astrocytes are transiently predominant at 3 days post stroke. Transcription factor analyses further suggest that this group of astrocytes may be primarily regulated by Brca1. 62
The advantage of single-cell transcriptomics lies in its high resolution, while spatial transcriptomics technologies such as 10X Visium retain spatial information. Although these methods complement each other, they still rely on various algorithms to infer spatial cellular heterogeneity, falling short of achieving true single-cell resolution within a spatial context. To address this, Scott et al. developed a platform named tDISCO (digital microfluidic isolation of single cells for -Omics), enabling spatially resolved single-cell transcriptomics and proteomics to more faithfully examine the phenotypic heterogeneity of astrocytes. 66 In subacute stroke, researchers divided the infarct area into three zones based on proximity to the injury: Zone A (0–200 µm), Zone B (200–400 µm), Zone C (400–600 µm), and more distal areas (Zone D). Zone A astrocytes express high levels of Fkbp11, Sdhb, Gltp, Ltc4s, Dffb, and Zfp799. Zone B astrocytes express genes related to the MAPK pathway and neurotrophic signaling, such as Dnahb2, Egin2, Mapk10, and Ntrk2. Zone C astrocytes show no significant differences compared to Zone D. These findings suggest that astrocytes accumulate around the injury site, with those closer to the core primarily inducing immune cell infiltration and migration, while those closer to the perilesional area focus on cell proliferation and neurotrophic support. However, two spatial transcriptomics studies on hemorrhagic stroke found that astrocytes primarily exhibit a reparative phenotype in the acute phase, while their pro-inflammatory phenotype becomes more prominent in the subacute and chronic phases, further emphasizing the disease-specific heterogeneity of astrocytes. 61 In summary, astrocytes are remarkably heterogeneous across the dimensions of space and time, and their dynamic transcriptomes also show disease-specific signatures.
Microglia
Microglia, the resident immune cells of the brain, undergo substantial transcriptional changes following stroke, as evidenced by a large number of differentially expressed genes after stroke. Various studies have identified multiple subpopulations of microglia, commonly categorized into homeostatic microglia (P2ry12, Tmem119), interferon-associated microglia (Ifit3, Isg15), proliferative microglia (Stmn1, Top2a, Mki67), inflammation-associated microglia (Cxcl10, Ccl2, IL1β), and reparative microglia (Igf1, Lpl, Gpnmb).46,67–76 Single-cell sequencing and spatial transcriptomics both indicate that proliferative microglia predominantly appear in the acute phase (≤3 days post-stroke) and are primarily located at the periphery of the lesion. 77 Zhang et al. discovered that proliferative microglia share signatures with interferon-associated microglia and reparative microglia at 3 days after stroke. After accounting for cell cycle effects, the distribution differences between proliferative microglia and these two microglia clusters disappeared on UMAP, suggesting that proliferative Mki67+ microglia could differentiate into interferon-associated microglia and reparative microglia. 78 While reparative microglia dominate in the chronic phase (14 days), the proportion of inflammation-associated microglia is higher in the chronic phase (14 days) than in the acute phase (2 days). 68 This spatiotemporal transition also highlights the dynamic nature of microglial responses following brain injury. 68 Reducing the prolonged pathological activation of microglia and promoting their return to homeostasis is crucial for the prevention of long-term neurological dysfunction following brain injury. The fate of disease-induced microglial subpopulations, such as the Spp1-positive subset, may vary with developmental stage and age in the context of stroke. In neonates, microglia return to a homeostatic state as stroke resolves, whereas in juveniles, microglia undergo irreversible cell death. 79 Microglia repopulation or transplantation of young mouse microglia effectively improves stroke outcomes in aged mice. 80 Although studies examining the impact of aging on post-stroke microglia at the single-cell level are limited, aging appears to lead to the upregulation of interferon-stimulated genes in microglia, both before and after stroke.68,81 In addition, spatial transcriptomics reveals that proliferative microglia around stroke lesions are gradually replaced by microglia with functions related to inflammatory signaling molecules and lipid metabolism genes in the subacute phase (7 days). 77
In TBI, snRNA-seq studies reveal significant upregulation of IFN-I signaling and the inflammasome pathway in microglia at subacute phases (7 days) after the insult. 82 Zambusi et al. used scRNA-seq on a zebrafish brain injury model to identify an injury-induced microglial subpopulation. Over time, this subpopulation gradually transitions back to homeostatic microglia and is characterized by enhanced lipid droplet metabolism and increased expression of granulin genes (grna, grnb, grn1-2). Further investigations using double knockout of grna and grnb revealed a failure of microglia to revert to a homeostatic state. Instead, the grna and grnb knockout microglia exhibited a sustained pro-inflammatory phenotype and increased lipid metabolism. 83 Recently, Jha et al. created for the first time a large-scale murine single-cell atlas to systematically reveal signature variations according to sex, brain region, and time in TBI. Certain microglial subpopulations (1, 2, and 6) decrease 24 hours post-TBI, with partial repopulation (1 and 6) by 6 months, while others (3, 4, and 5) emerge post-TBI with varying temporal dynamics. Microglia-3 are elevated at 24 hours, peak at 7 days post-TBI, and return to baseline by 6 months. The highest proportions of microglia-4 are noted at 24 hours, decline at 7 days, and reach baseline levels by 6 months. Unlike their temporal dynamics, differential gene expression in microglia-4 is most extensive at 7 days and persists for 6 months, with nine genes (Spp1, Slpi, Celc4a, Hp, G0s2, S100a6, S100a8, S100a9, Il1b) consistently upregulated post-TBI. Functional enrichment analysis shows that microglia-4 shifts from innate immune response towards RNA splicing/processing and synapse pruning by 7 days, and towards immune response and regulation of peptide secretion by 6 months. The microglia-5 subpopulation shows a persistent increase at 7 days and returns to baseline by 6 months. This study also highlights biological sex differences, with females showing lower proportions of most microglia subtypes, except for Microglia-3. The microglia-4 subpopulation shows significant sex-based transcriptional heterogeneity, with an enhanced neuroinflammatory response in females. Additionally, another subset in females upregulated genes related to increased synapse organization and formation (Atp1a2, Slc1a2, Ntrk2, Tubb2b, Ptn, Akap12, Plat, Itpr1, Pcp4, Abat, Mt3, Ncam1). 84
Notably, several studies have identified subpopulations of cells expressing both microglia/macrophage and other cell markers, such as neutrophil, 85 oligodendrocyte, 59 T cell,86,87 and B cell88,89 markers. These cells are almost exclusively present in the injured brain. Through single-cell data analysis of various diseases, Wang et al. found that cells expressing both B cell and microglia/macrophage markers are present in conditions such as stroke, TBI, Alzheimer's disease (AD), and glioblastoma (GBM), indicating the universality of this cell subtype. 89 Additionally, studies have shown that microglia upregulate oligodendrocytic transcripts such as Plp1 and Mbp and downregulate homeostatic microglia markers like P2ry12 after myelin phagocytosis, suggesting that cells expressing both oligodendrocyte and microglia/macrophage markers may be phagocytic microglia/macrophages rather than oligodendrocytes. 90 The reason for the existence of cells with double cell identity is not fully understood, but could arise from technical artifacts, such as doublets introduced during single-cell processing, phagocytosis, or other causes. Although it is not proven that “double cell type identities” can be created by cell engulfment and heterophagy, the increase in cells with double cell type identities in response to acute brain damage supports the upregulation of phagocytic functions in cells such as microglia/macrophages. Higher-resolution spatial transcriptomics may shed light on the spatial distribution of these unique cells and extend our knowledge of cell-cell interactions and debris clearance mechanisms after acute brain injury.
Oligodendrocytes and their precursor cells
Recent scRNA-seq studies have characterized the heterogeneity of oligodendrocyte (OL) lineage cells in the healthy brain. In young rodents, the proportion of the homeostatic OPCs (Pdgfra, Hes5) significantly decreases following stroke, while a proliferative subpopulation (OPC2), marked by high expression of cell cycle genes such as Mki67, becomes dominant in acute stroke.51,59,80 Enrichment analyses show that this OPC subpopulation primarily functions in migration, survival, and differentiation. 59 Using cell trajectory analyses, a new subset of OPCs has been discovered in the stroke brain. The latter OPC subset expresses markers associated with committed oligodendrocyte precursors (COPs) such as Bmp4 but acts as a transitional state between OPCs and OLs. By day 14 post-stroke, substantial numbers of Bmp4+Sox10+ OPCs were observed in the peri-infarct regions in immunostaining experiments, suggesting ongoing OPC differentiation processes. 80 Spatial transcriptomics reveals the presence of OPCs around the lesion. 77 The transition of OPCs to mature OLs is not fully understood.
Like OPCs, OLs also diverge into new subpopulations during the acute phase (≤3 days) and subacute phase (14 days) post-stroke. The acute phase subpopulation shows low expression of mature OL genes (e.g., Dlg1, Lamc1, Psem4) and high expression of immature OL markers such as Bcas1. Gene enrichment analysis reveals that myelination processes are downregulated in the acute phase subpopulation but show partial recovery in the chronic phase (14 days) subpopulation, and are accompanied by sustained enhancement in phospholipid transport processes. 80 Different recovery patterns in myelination have been observed between young and aged mice. 80 Interestingly, young mice (10–12 weeks) show weaker myelination at 14 days post-stroke compared to sham-injured mice, while aged mice (19-20 months) exhibit myelination levels comparable to sham-injured mice at the same time point. Recent studies have identified a group of disease-associated OLs (DAOs) in neurodegenerative and demyelinating conditions, characterized by high expression of genes like Serpina3n, C4b, Cdkn1a, Tnfrsf12a, Irf9, and Ifit1. 91 However, comparing the marker genes of the new post-stroke OL subpopulations with DAO signature reveals limited overlap in the acute phase. 59
Notably, beyond their well-known function of forming myelin sheaths around axons, recent research has revealed that OLs and OPCs can also activate the expression of immune-related genes and regulate immune function and inflammation under disease conditions. Garza et al. analyzed snRNA-seq data from brain tissues of TBI patients and found that OLs exhibit a robust innate immune response under brain injury conditions. The innate immune response is characterized by the activation of interferon pathways and the transcriptional activation of major histocompatibility complex (MHC) classes I and II. 92 Spatial transcriptomics and scRNA-seq have also revealed that a subpopulation of OPCs upregulates genes such as Cxcl10, Oasl2, and Ifi27l2a, which are involved in innate immunity and the type I interferon signaling pathway. 64 These findings highlight the multifaceted roles of OLs in brain injury and repair processes, suggesting that they not only support myelination but also contribute to the immune response following brain injury. 92
Endothelial cells
As a major component of the NVU, highly specialized cerebrovascular endothelial cells are essential elements of the BBB, which shields the CNS from circulating blood, bacteria, and neurotoxins. scRNA-seq studies have identified several endothelial cell subclusters, categorized in a way that reflects anatomical differences: venous capillaries (e.g., Car4, Tfrc), large veins (e.g., Vcam1, Cfh, Scl38a5), arterial capillaries (e.g., Fos, Fosb), arteries (e.g., Gkn3, Mgp, Stmn2, Bmx, Clu, Cdh13), venules (e.g., Icam1, Vcam1), fenestrated brain vascular (e.g., Plvap, Plpp3, Igfbp3, Plpp1, Cd24a, Ldb2), and proliferation (e.g., Pimreg, Pclaf). 68 ScRNA-seq analysis revealed a novel subset of endothelial cells (EC3: Lrg1, Mgp, Vwf) that emerges transiently in the acute phase of injury (≤3 days post-stroke).68,80 This subset gradually diminishes over the following two weeks. Gene Ontology Biological Process enrichment analysis has shown transcriptional perturbations in numerous angiogenesis-related processes within this EC3 population, affecting both pro-angiogenic and anti-angiogenic activities. For example, processes such as vasculature development, blood vessel development, and angiogenesis were significantly altered. Consistently, spatial transcriptomics confirmed the presence of a large number of endothelial cells in the infarct core during the acute phase of stroke. 77 In the aged sham-injured brain, the predominant endothelial cell subset was EC2, which showed increased expression of apoptosis-related genes (Depp1, Cdkn1a) and decreased expression of circadian genes (Dbp, Nr1d1) compared to the young brain. After a stroke, aged brains primarily exhibited EC3 and EC4 subsets, with EC4 displaying a strong interferon response at 3 days after insult. Despite retaining some angiogenic markers (Lrg1, Tmem252, Litaf), aged EC3 cells had inhibited angiogenesis-related processes, with significant downregulation of angiogenesis genes (Aplnr, Nr4a1, Jun, Sparc). 80
Jha et al. identified two groups of endothelial cells (endothelial-1 and endothelial-2) following TBI. Endothelial-1 cells exhibited greater changes in transcriptional activity during the acute phase (24 hours after TBI), and both groups showed significant sex differences in their transcriptional responses. 84 Surprisingly, endothelial-1 cells of the contralateral hemisphere exhibited activation of hemoglobin/haptoglobin, oxygen binding, and neuroinflammation, including innate immunity, leukocyte chemotaxis, and regulation of apoptosis. The mechanisms underlying these unexpected changes in the contralateral hemisphere are unclear but may relate to enhanced angiogenesis for functional recovery. 84
Pericytes
In the acute phase of stroke injury (≤24 h), scRNA-seq studies have identified a subpopulation of pericytes (Il11, Il6, Ccl2, and Ccl11) emerging as early as 12 hours post-stroke. This subpopulation is characterized by high expression of Il11, Il6, Ccl2, and Ccl11. Functional enrichment analysis shows that these cells are primarily involved in inflammatory responses, apoptosis, TNFα signaling, and cytokine-mediated signaling pathways, suggesting they may mediate detrimental aspects of the pericyte response in the acute injury phase. 93 In the subacute injury phase (7 days post-stroke), pericytes in the infarct core display the strongest pro-fibrotic activity and downregulate genes associated with pericyte-endothelium junction and vascular integrity. In contrast, peri-infarct pericytes exhibit more diversity, upregulating both regenerative pathways (e.g., regeneration and response to wounding) and inflammatory pathways (e.g., immune response and response to oxidative/endoplasmic reticulum stress). 94 These findings emphasize the importance of pericytes, alongside astrocytes and myeloid cells in the post-stroke inflammation and recovery process.95,96 Pericyte heterogeneity is rarely addressed in scRNA-seq studies of the aging brain after stroke. Considering the essential role of pericytes in the NVU, this cell type deserves further investigation during the brain injury and recovery process, across neurodevelopmental stage, age, and sex.
Brain border-associated cells
Border-associated macrophages
Brain-resident macrophages (BAMs) reside within the brain barriers, such as meningeal macrophages distributed at the brain surface, perivascular macrophages at the blood-brain barrier, and choroid plexus (ChP) macrophages found at the blood-cerebrospinal fluid barrier, to help maintain homeostasis. 49 The current understanding of the roles of BAMs lags behind that of microglia. BAMs are a highly heterogenous population with multiple subtypes, similar to microglia, and are also activated following stroke insults. 68 Using sc-RNA sequencing techniques, Garcia-Bonilla et al. reported the heterogeneity of BAMs at different phases after stroke and across ages. In the latter study, BAMs were classified into five clusters, with BAM1 and BAM2 being the most abundant. BAM1 cells were characterized by Cd209f, Ccl24, and Slc40a1 expression, while BAM2 cells were characterized by choroid plexus (ChP) macrophages major histocompatibility complex class II (MHC class II) markers (e.g., H2-Ab1, H2-Eb1, H2-Aa, H2-DMb1) and perivascular macrophage marker Lyve1. BAM3 cells (Wwp1, Abhd12, Dhrs3, Hpgd, Trf) were detected less frequently at the subacute phase (14 days) after stroke, while BAM4 cells (Spp1, Ccl8, Cstb, Lgals1) appeared mainly at the acute phase (2 days) after stroke. BAM5 cells were characterized by damage-associated molecular pattern-recognition molecules (S100A4, Lgals3, Clec4d, Clec4e). 68 A recent study reported alterations in the BAM transcriptomic phenotype during aging. The age-related upregulation of MHC class II components such as H2-Aa, H2-Ab1, H2-Eb1, and Cd74, adhesion molecules such as Vcam1, Ceacam1, and Selplg, and chemokines such as Ccl5 and Ccl12 may be related the trafficking of peripheral immune cells. BAMs acquire immunoregulatory functions during aging, as the loss of BAMs exacerbates neurological dysfunction after ischemic stroke in aged but not young brains, suggesting an essential role of BAMs in older animals. 97
Ependymal cells
Ependymal cells are a type of glial-like support cell that line the ventricles of the brain and central canal of the spinal cord. Ependymal cells produce and help circulate CSF and play a role in cellular regeneration in the CNS. In the acute phase of TBI, ependymal cells become more abundant, 44 and five ependymal subpopulations have been identified. 84 Ependymal subtypes 1-3 shared gene-expression constituents but varied in expression levels. The Ependymal-4 subpopulation was the only one to express Il1b, while Ependymal-5 showed high expression of genes related to the inflammatory response (Ly6a, Arg1). 84 Ependymal-4 is a newly found contributor to cytokine-mediated inflammation, as the transcriptional signature for ependymal-4 leads to dramatic changes in biological processes, such as the upregulation of cytokine activity, changes in the extracellular matrix, and decreased leukocyte proliferation, activation, and antigen-presentation. As the primary recipient (and key source) of IL-1, ependymal-4 communicates with other cell types, such as neutrophils, B cells, and microglia, which may contribute to the progression of inflammation after TBI.
Ependymal cells exhibit stem cell-like plasticity post-stroke and are capable of generating neuroblasts and astrocytes through Notch signaling. However, this notion is controversial and may be attributed to type B neural stem cells in the ventricular-subventricular zone. Thus, Shah et al. developed αSMACreERT2 reporter mice to trace ependymal cells while excluding neural stem cells. Their scRNA-seq and in vivo fate mapping indicated that, despite sharing the expression of some genes with quiescent neural stem cells, ependymal cells express other genes (e.g., Rarres2, Ccdc153, Nnat, Tmem212, Mia, etc.) that are not detected in neural stem cells. In addition, ependymal cells do not exhibit mitosis, loss, or migration from their niche before or after injury. 98 Spatial transcriptomics using deconvolution methods reveal that, although ependymal cells generally reside in the ventricular lining before injury, a small number of ependymal cells are present within the injury region during the acute phase post-stroke. 77 This observation differs from the findings of Shah et al., indicating that further investigation is needed.
CNS-infiltrating peripheral cells
Monocytes/macrophages
Upon stroke, circulation-derived monocytes/macrophages infiltrate the brain parenchyma, accumulate in the affected brain regions, and undergo significant phenotypic changes, transitioning from classical inflammatory monocytes (Ly6C+/CCR2+) in the early injury phase to patrolling monocytes (CX3CR1+) in later injury stages.99,100 Recent studies have shown that classical inflammatory monocytes (Ly6C+/CCR2+) originate from two distinct sources: granulocyte-macrophage progenitors (GMPs) and monocyte-dendritic progenitors (MDPs). GMP-derived monocytes primarily express Sell, Chil3, and Cd177, while MDP-derived monocytes express MHCII molecules.101,102 Although not explicitly stated in previous studies, differential gene expression analyses confirm that both GMP- and MDP-derived monocytes are present in peripheral blood and brain tissue post-stroke.68,86,103,104 GMP-derived monocyte numbers increase significantly during the acute phase and slightly decrease during the chronic phase after stroke, whereas MDP-derived monocytes increase during the acute phase and continue to rise during the chronic phase. 68 The specific roles of these two monocyte populations in stroke pathology and recovery have not been defined. GMP-derived monocytes exhibit greater heterogeneity than MDP-derived monocytes. During the acute injury phase, GMP-derived monocytes are characterized by the expression of chemokines and inflammatory cytokines (Cxcl2, Cxcl3, Il1b) as well as M2-like markers (Arg1, Pf4, Fabp5). In the chronic phase, GMP-derived monocytes predominantly express phagocytic and reparative markers such as Igf1, C1qa, and Gpnmb. 68 The partial depletion of circulating monocytes/macrophages with clodronate has been leveraged in studies of experimental TBI. Subsequent scRNA-seq work during the acute phase of TBI reveals that the surviving peripheral blood monocytes upregulate neutrophil-related genes (Mmp8, Lyz2, Lcn2, Elane, Prtn3, Ctsg, Mpo) in the clodronate-treated group. Upon further analysis of subpopulations, it was found that the proportion of classical inflammatory monocytes increased significantly with clodronate application. Additionally, the authors identified and named a new cellular subpopulation (neutrophil-like monocytes). The neutrophil-like monocytes display high expression of neutrophil markers (Elane, Prtn3, Ctsg, Mpo) and low expression of MHC II molecules, suggesting that their transcriptional profile may resemble GMP-derived monocytes. Transfer of monocytes derived from clodronate-treated TBI donor mice into post-TBI recipients significantly reduced lesion volumes, indicating that the monocytes surviving clodronate application have neuroprotective properties. 105 MDP-derived monocytes, which primarily express MHCII molecules, may play a role in antigen presentation during stroke. In aged mice, the presence of MDP-derived monocytes is significantly higher in both peripheral blood and brain tissues compared to younger mice, suggesting that these monocytes might have a detrimental effect on stroke recovery. 103
The burden of stroke extends beyond the immediate brain injury and is heavily influenced by chronic comorbidities that develop as secondary conditions. However, the chronic effects of stroke on systemic immunity have been underexplored. Recent studies have begun to shed light on these mechanisms. By performing single-cell sequencing on multiple peripheral organs (blood, bone marrow, spleen, heart, lung, liver) and combining these findings with single nucleus ATAC-seq, researchers revealed the role of myeloid innate immune memory as a significant contributor to remote organ dysfunction following stroke. Specifically, stroke-induced elevation of IL-1β influences epigenetic modifications in innate immune cells (particularly monocytes/macrophages) via H3K4me3, primarily affecting pro-inflammatory pathways. Additionally, IL-1β promotes the recruitment of classical inflammatory monocytes to the heart. By blocking IL-1β or monocyte recruitment, post-stroke cardiac dysfunction may be prevented. 106
Multiple spatial transcriptomics studies have shown that peripherally infiltrated monocytes/macrophages appear around the injury site during the acute phase and gradually fill the core area after stroke.58,77,107 However, it remains unclear whether there is spatial heterogeneity in these two sources of monocytes/macrophages.
Neutrophils
The functional diversity of neutrophils has been recently recognized. In the acute phase of stroke, the levels of peripheral blood neutrophils, predominantly consisting of immature neutrophils (Mmp8, Retnlg, and Lcn2) remain constant. However, in the chronic phase, mature neutrophils (Cd101+) begin to predominate, expressing genes associated with tissue infiltration (Ninj1, Cd300c2) and growth inhibition (Lst1, Creg1). In the stroke brain, immature (Retnlg, Mmp8, Wfdc21) and early mature neutrophils (Camp, Ltf, Chil3, Lcn2) gradually decrease, while interferon-related neutrophils (Isg15, Ifit3) and cytokine-related neutrophils (Ccl3, Ccl4, Csf1) increase. Cell trajectory analyses indicate that brain neutrophils at different stages resemble their peripheral neutrophil counterparts, suggesting that neutrophils primarily migrate into the CNS from the periphery without undergoing significant differentiation. 68
In the aged brain, two major clusters of infiltrated neutrophils display high levels of Ptprc, Itgam, and Cxcr2. In contrast to the high expression of homeostatic genes in Cluster 2, such as ltf, Camp, and Ngp, Cluster 1 neutrophils exhibit high levels of mt-Nd1-4, Aif1, Ly86, Il1b, and protein synthesis-related genes rather than homeostatic genes. 85 Compared to young mice, aged mice exhibit higher proportions of neutrophils in both peripheral blood and brain, with a greater degree of maturity (Cxcr4 upregulation, Cd62l/Cxcr2 downregulation), stronger inflammatory responses (Il1b, Mmp9 upregulation), and enhanced antigen presentation capabilities (MHCII molecule expression upregulation). 103
Dendritic cells (DC)
Circulation-derived DCs have been detected in brain parenchyma after brain injury. Recently, spatial transcriptomic studies also confirmed the distribution of DCs around the injury site during the acute phase of stroke. 77 scRNA-seq work has identified five DC subpopulations: conventional cDC1 (Xrc1, Clec9a) and cDC2 (Cd209a, Cd172a), plasmacytoid DCs (Siglech, Ccr9, Bst2), migratory DCs (Ccr7, Fscn1), and monocyte-derived DCs (Ms4a7, Lyz2). cDC2 is the most abundant DC subtype in the brain and can be further divided into four subpopulations. Among these, the subpopulation associated with interferon, stress, and inflammation (Ifitm1, Ddit4, Il1r2) predominates during the acute phase, while the subpopulation related to antigen presentation (H2-Oa, H2-DMb2) is more prevalent in the chronic phase. 68 Surprisingly, scRNA-seq has also identified a proliferative subpopulation of cDC1 cells that increase in proportion during the subacute injury phase, 68 suggesting that at least part of the subacute cDC1 population relies on local proliferation. In the aged brain, Li et al. reported four clusters of DCs after stroke, Cd209a (DC1), Xcr1 (DC2), Ccr7 (DC3), and Ccr9 (DC4). Among them, DC1 cells were characterized by upregulated expression of MHC class II-related genes, such as H2-Aa, H2-Ab1, and Cd74, indicating antigen-presenting functions of DC1 after ischemic stroke. 85 These findings highlight the complexity and dynamic nature of DC responses following stroke, emphasizing the need for further research to elucidate the roles and therapeutic potential of these distinct DC subpopulations in stroke recovery.
Lymphocytes
ScRNA-seq has identified various lymphocyte subpopulations, including CD4 T cell subpopulations (CD4 naïve: Sell, Ccr7; CD4 memory: Cd44; Th1: Ifng, Ccr5, Cxcr6; Th17: Ccr6, Ccr4; CD4 Tregs: Foxp3, Il2ra; IFN-associated CD4: Ifit3, Isg15), CD8 T cell subpopulations (CD8 naïve: Ccr7, Sell; CD8 central memory: Ccr7, Cd44; CD8 effector memory: Ccr7low, Cd44; IFN-associated CD8: Ifit3, Isg15; exhausted-like CD8: Pdcd1, Lag3, Ctla4; CD8 effector: Klrg1, Gzma, Gzmb; CD8 Treg: Klra1, Klra6, Klra7), and γδT cell (Trdc, Trdv4, Tcrg-C1).68,103 CD4 Tregs express TNF receptors (Tnfrsf4, Tnfrsf9, Tnfrsf18), similar to those found in non-lymphoid-tissue-derived Treg cells. 68 CD4 Tregs can be classified into naive Tregs (Sell, Ccr7) and effector Tregs (Itgae, Lgals1). Effector Tregs exhibit enhanced adhesion, chemotaxis, and secretion. 103 IL-2 can effectively expand CD4 Treg cells. By engineering astrocytes to serve as a source of IL-2 within the brain, CD4 Treg cells can be significantly expanded. Single-cell analysis revealed that the transcriptome of the expanded Treg cells remained largely conserved, with notable upregulation of IL-2R components and the anti-apoptotic gene Bcl2. 108 Most peripheral immune cells exhibit a rapid increase shortly after stroke, followed by a decrease. In contrast, scRNA sequencing revealed proliferative T cell subpopulations of lymphocytes, characterized with Ccna2, Ccnb2, and Mki67 expression as well as Ki67 immunostaining, suggestive of in situ proliferation of brain-resident lymphocytes. 109 Hence, the sustained presence of brain lymphocytes may be at least partly due to their proliferation in the CNS after injury.
The number of effector Tregs is higher in aged mice, with effector-related genes being more highly expressed in naive Tregs compared to in young mice. 103 Additionally, aged mice exhibit a significant increase in CD8 effector memory and CD8 exhausted-like T cells, which express higher levels of inflammatory chemokines (Ccl3, Ccl4, Il1b). The increase in CD8 effector memory cells due to aging is also evident post-TBI. 110 The number of CD8 Tregs is significantly reduced in aged mice, and they upregulate genes related to cytotoxicity (Fasl, Gzmm), suggesting a shift towards a cytotoxic and pro-inflammatory phenotype in the aging brain. 103
Chen et al. conducted scRNA-seq analysis to investigate the heterogeneity of brain cells in young (14 weeks) and aged (20 months) mice four months post-TBI. They discovered that the proportion of T cells was significantly higher in aged mice compared to young mice before injury, and this proportion increases further post-TBI. Further classification of T cells reveals distinct subpopulations: CD8 naïve-like (Ccr7, Sell), CD8 early active (Tox, Ccl5, Gzmk), CD8 effector memory (Tox, Pdcd1, Gzmb, Gzmk), and Th1 (Tcf7, Toxlow, Gzmklow). Differential gene expression analysis showed that CD8 T cells in aged TBI mice upregulated the expression of inflammatory mediators S100a8/a9, Nr4a2, and Cd28 (T cell activation). Conversely, genes related to CD8 T cell function, such as Fgl2 (immunosuppressive activity) and Cst3 (cytotoxicity), are downregulated. Th1 cells upregulate Gpr183 (positioning of immune cells) and Igf1r (T cell expansion), while downregulating genes associated with homing and migration (Ccl5). Functional enrichment analysis revealed that CD8 T cells are involved in macromolecule biosynthesis processes, inactivation of MAPK activity, and negative regulation of apoptosis. Th1 cells are implicated in the regulation of dendritic cell differentiation, negative regulation of translation, complement activation, and cellular response to IL-4. 110
Natural killer (NK) cells
NK cells are a type of innate lymphoid cell, initially classified into two major subpopulations based on CD56 expression: CD56high and CD56dim. However, under certain conditions, a group of adaptive NK cells has also been identified, indicating NK cell heterogeneity. 111 Recent studies have shown that NK cells can infiltrate the brain during the acute phase of stroke and exacerbate brain injury.45,112 NK cells infiltrating the brain undergo reprogramming towards enhanced cell activation, cytotoxicity, and the production of cytokines and chemokines. Similar changes are observed in peripheral blood NK cells post-stroke, supporting their neurotoxic potential. 45 Cho et al. found comparable results in immune cells from the peripheral blood of stroke patients via scRNA-seq. 113 Another scRNA-seq study reported that the proportion of NK cells increased from the acute stroke phase (2 days) to the chronic phase (14 days) and identified two subpopulations: NK1 and NK2. NK1 cells were transcriptionally similar to peripheral blood NK cells, suggesting that NK2 might be the reprogrammed NK subpopulation. 68
Cell-cell interactions
Following stroke, dramatic increases are predicted in the interactions between microglia and other brain-resident cells such as astrocytes, endothelial cells, pericytes, and oligodendrocytes/OPCs, as well as peripheral immune cells. In the acute phase, there is a notable rise in interactions involving chemokines, cytokines, and TNF family members. These include molecules such as CCL2, CCL5, IL-1α, IL-1RΑ, CD300a, and TNFα, which bind to their corresponding receptors, 46 further facilitating inflammatory responses. Additionally, microglia/macrophages and astrocytes express various reparative factors that may contribute to brain repair and functional recovery from stroke. These factors include VEGF, GDF15, IGF1, MIF, Galectin-9, FGF1, Prosaposin (PSAP), and Pleiotrophin (PTN), which interact with receptors on endothelial cells or oligodendrocyte lineage cells such as VEGFR-2, TGFBR2, IGF1R, CXCR7, CD44, FGFR-2 (CD332), GPR37, and Nucleolin. These interactions might promote angiogenesis and OPC proliferation and differentiation.58,68,80,114–116 Microglia/macrophages also increase the expression of CXCL10, which may activate the CXCR3 receptor on CD8 T cells 14 days after stroke. Spatial transcriptomics studies have revealed that peripherally infiltrated monocyte/macrophages accumulate in the injury core post-stroke, while activated microglia predominantly localize to the outer regions.58,77 The emergence of these unique niches may facilitate interactions between these two cell types in the injured brain. Cell ligand-receptor inference analysis has identified numerous potential interactions between microglia and monocytes, with the interaction between microglial SPP1 and monocyte CD44 being the most significantly upregulated post-stroke. 117 Recent scRNA-seq studies have found extensive transcriptomic changes in microglia when peripheral infiltrating monocyte/macrophages are absent.118,119 One study reported a significant reduction in microglial subpopulations associated with leukocyte migration and the response to IFN. These subpopulations of microglia shared markers with disease-associated microglia (DAM) (e.g., Ccl6, Clec7a, Cd63, Ctsb, Cst7, Apoe). Additionally, other microglial subpopulations downregulated expression of IFN-stimulated genes (ISGs), suggesting a role for peripheral monocyte/macrophages in modulating microglial interferon responses. 118 Another study, based on biological experiments and scRNA-seq data, found that infiltrating monocytes promote microglial reprogramming to support brain vasculature repair by secreting IL-6, which binds to receptors on microglia. 119
In addition to intracerebral cell interactions, stroke also impacts peripheral signaling. Xu et al. performed scRNA-seq on the ipsilateral and contralateral skull bone marrow in the acute phase post-stroke, revealing significant differences in signaling pathways. Notably, Icam, Spp1, and Hgf were significantly upregulated in the ipsilateral skull, while L1cam and Thbs were significantly downregulated. 120 Wang et al. combined scRNA-seq and spatial transcriptomics to show that the interaction between Thbs1 and Cd47 was significantly activated in the meninges during the acute phase of hemorrhagic stroke. Evidence favoring this pairing was also provided by spatial transcriptomics work. 121
Aging significantly impacts the nature of cell-cell communication. In pre-stroke conditions, the binding of Tnf secreted by microglia/macrophages to Tnfrsf1a receptors on endothelial cells and oligodendrocytes/OPCs is notably stronger in aged mice compared to young mice. In the post-stroke phases, there is a noticeable reduction or delay in the interactions involving reparative factors and their receptors in aged mice. Interestingly, aging does not appear to affect the binding of these reparative factors secreted by astrocytes to their receptors on endothelial cells or oligodendrocytes/OPCs, indirectly highlighting a more critical role of microglia/macrophages in stroke recovery in the context of aging. Additional experiments supporting this view have shown that transplanting microglia from young mice into aged mice significantly promotes angiogenesis, oligodendrogenesis, and motor and memory functions. 80
Sex differences significantly influence cell-communication networks following brain injury. In females, overall cell-cell communication is less frequent and weaker compared to males at the acute injury stage, with notable exceptions in NK cells and specific microglia subpopulations. Males exhibited enrichment in signaling pathways (MHC-1, Bst2, Thy, Lair1, Cd86, Cd29, Cspg4) related to immune signaling and endothelial basement membrane stabilization. In contrast, females showed enrichment in pathways (Selplg, Sell, Ngl, Cdh) associated with brain development and leukocyte adhesion. 84
These findings highlight the importance of understanding cell-cell interactions in stroke recovery, particularly in the context of aging. As current studies on cell-cell interactions mainly rely on single-cell sequencing and overlook physical distances, they may lead to false positive and false negative findings. Thus, additional research will be needed to verify these interactions at the spatial level and leverage cell-cell interactions as therapeutic targets for brain repair.
Limitations, challenges, and future directions
ScRNA-seq has revolutionized the field of genomics by enabling researchers to analyze gene expression at the single-cell level. However, scRNA-seq has limitations that researchers must consider when designing experiments and interpreting data.
Inherent limitations of scRNA-seq
One of the primary limitations lies in cell isolation methods for scRNA-seq, especially in studies of the CNS. Due to the complexity of biological samples and the heterogeneity of cell populations, isolating and capturing single cells can be a technically challenging and labor-intensive process. However, to date, no standardized cell isolation protocol is compatible with all tissue preparations in scRNA-seq. Most studies in the stroke and TBI fields use enzymatic dissociation, others use mechanical dissociation.85,104,119 Among the enzymatic methods, commercially available neural tissue dissociation kits for adult brains (Miltenyi Biotec) are popular,55,58,63,68,80 while others use custom cocktails containing enzymes such as trypsin, collagenase, DNase, and papain.73,79,122,123 Different dissociation methods may influence the types and proportions of cells identified, which can impact data interpretations. Thus, researchers have optimized dissociation methods according to the specific cell or tissue type of interest. 123 For example, Buizza et al. used a combination of collagenase IV, dispase I, and DNase I to enrich non-neuronal cells, 93 and Wu et al. added pronase to their tissue dissociation reagent specific for mouse cortical tissue. 54 Additionally, the intrinsic cellular stress responses caused by the tissue dissociation process may also alter the transcriptional profile of the sample, and result in disproportionate mRNA degradation as well as cell death. 124 To reduce the cellular stress and preserve the transcriptional state during single-cell preparation, cocktails containing RNA polymerase inhibitor (actinomycin D, triptolide), NMDA receptor blocker (AP-V), and/or sodium channel inhibitor (TTX), have been added to the chilled dissociation solution.68,122,123 To acquire reliable data, cell viability values at or above ∼90% are recommended. Rare cells that make up less than 1% of the total cell population are particularly difficult to harvest using current scRNA-seq techniques, leading to underrepresentation of certain cell types or phenotypic states in the final analysis. 24
With snRNA-seq, there is a lower probability of a cellular stress response and loss of viability from tissue dissociation compared to scRNA-seq.68,122,123 In addition, the structural complexity of some cell types may pose another caveat in scRNA-seq. For instance, neurons in the brain have extensive neurites that are difficult to keep intact during cell dissociation processes. Considering the advantages of sn-RNA seq in retaining neuronal heterogeneity, the combination of snRNA-seq and scRNA-seq offers a reasonable technical compromise.14,16 Additionally, to understand the connection between transcriptomic profiles and dynamic neuronal functions, Patch-seq has been developed as a novel multimodal approach that combines electrophysiological recordings of brain slices with collection of the mRNA from neurons in various brain regions.125–128
Besides sample dissociation-induced cellular stress, scRNA-seq only focuses on a few thousand unique transcripts from a single cell—far less than the whole transcriptomic profile captured by bulk RNA-seq. 129 This limited coverage of the transcriptome can lead to incomplete or biased representation of gene expression within individual cells, potentially obscuring important biological insights. Additionally, scRNA-seq typically offers a shallow read depth compared to bulk RNA-seq, in that the number of sequencing reads obtained per cell is lower. The reduced read depth negatively impacts the accuracy and sensitivity of gene expression measurements, especially for genes or transcripts expressed at low levels. 130 Researchers must consider these limitations carefully when interpreting scRNA-seq data and be wary of drawing conclusions based on incomplete or noisy gene expression profiles.
The relatively high cost of scRNA-seq compared to bulk-RNA or flow cytometry also restricts its widespread application and is a major limiting factor in obtaining solid conclusions with a reasonable sample size. Thus, data-sharing approaches and publicly available repositories, like the NCBI’s Gene Expression Omnibus (GEO, with GSE prefix in DataSets) serve as invaluable resources for researchers to expand their findings or discover new possibilities. For instance, with the limited studies on stroke and brain injury, there are publicly available resources with the following GSE IDs (Supplementary material).
Although scRNA-seq has the potential to answer many research questions, the details of the answers can depend on the protocols used, how many genes can be detected, if the genes of interest are expressed, and if the cells of interest are involved.129,131 The RNA sequencing technique commences with single-cell dissociation or single-nucleus isolation and is followed by a single-cell barcoding step. Most available scRNA-seq techniques, including 10× Chromium, 30 Drop-seq, 9 inDrop-seq, 132 Smart-Seq,133,134 and CEL-Seq2, rely on beads with oligos that capture poly-A tails on the 3’ ends of mRNA transcripts and then add cell-specific barcodes. 135 Targeting the poly-A tails on the 3’ end is conventional, but this approach excludes a sizable fraction of the non-polyadenylated RNAs, such as long non-coding RNA (lncRNA), ribosomal RNA (rRNA), and small non-regulatory non-coding RNAs. Rosenberg et al. developed a non-bead-based sc/snRNA sequencing platform named split-pool ligation-based transcriptome sequencing (SPLiT-seq), which labels the cellular origin of RNA via combinatorial barcoding without any special instrumentation. 136 Moreover, most histone modification-related mRNAs do not have the polyA structure. 137 The importance of including non-poly-A RNAs has become evident in recent years, and investigators can use Small-seq to capture microRNAs, tRNAs fragments or small nucleolar RNAs, 138 RamDA-Seq to capture lncRNA, histone mRNA, or enhancer RNA, 139 and VASA-Seq to generally capture both polyadenylated and non-polyadenylated transcripts. 140 However, those single-cell techniques do not rely on poly-A tails and are not widely applied in the field of stroke, which limits the findings.
Overcoming the limitations of scRNA-seq in stroke and TBI
Incorporating the inclusion of biological variables
Since the first application of scRNA-seq in focal brain injuries, such as stroke and TBI, cellular heterogeneity has been mainly investigated in young male animals at the acute and subacute phases (within 14 days). Recently, the impact of aging and sex on heterogeneity has been illustrated in experimental stroke and TBI.68,84,106 Garcia-Bonilla et al. highlight changes in immune cells from brain and blood during stroke, comparing the differences in transcriptomic profile between young (8–12 weeks old) and aged (17–18 months old) male mice within 14 days after focal ischemic stroke. 68 Our recent study revealed the differences in non-immune cells (EC, OPC/OL) at different time points (acute and chronic phases) and ages (young and aged) in the stroke brain, and demonstrated the therapeutic effect of myeloid cells from young donors in the aged stroke brain. 80 Simats et al. reported single-cell changes in peripheral organs (blood, spleen, liver, lung, heart, bone marrow) up to one month after stroke, except the brain. 106 Although microglia heterogeneity has been illustrated in neonatal and juvenile brains up to 28 days after stroke, the heterogeneity of the NVU in chronic injury phases after stroke is still poorly understood. 79
Jha et al. systemically investigated the variation of single-cell signatures among different TBI models, including repetitive-closed-head-injury TBI (rCHI), controlled cortical impact (CCI), and CCI with hemorrhagic shock, and evaluated the impact of sex and brain region at several time points. 84 This was the first study to investigate gene expression changes up to 6 months after TBI; it provided a single-cell atlas after TBI and emphasized the impact of immune cell heterogeneity on brain recovery during the chronic injury stage. Brain injury is accompanied by brain repair, and neurogenesis is one of the critical steps in this process. Leveraging scRNA sequencing, Bielefeld et al. found that TBI modified the fate of neural stem cells (NSCs) to promote neurogenesis without affecting astrogliogenesis in the adult hippocampal dentate gyrus 15 days post-injury. 55 However, brain recovery in the NVU is complex, involving revascularization, angiogenesis, neurogenesis, synaptogenesis, axonal sprouting, oligodendrogenesis, and remyelination, among other processes. Key components involved in NVU recovery, such as neurons, vascular-associated cells (endothelial cells, pericytes), ependymal cells, and choroid plexus cells, are often overlooked. Furthermore, the impact of different variables including the interventions on brain recovery may vary across different stages. Zhang et al. found that thyroid hormone T4 administered as an intervention at 1 and 6 hours after TBI affected a greater variety of cell types and genes in the subacute (7 days after TBI) and chronic phases (21 days after TBI), compared to the acute phase (24 hours after TBI). 141 This finding further underscores the significance of time as an independent variable and the inclusion of long-term timepoints. Thus, comprehensive RNAseq studies that account for spatial and temporal differences remain sparse, and few studies of brain injury and recovery have simultaneously incorporated biological variables such as age and sex or compared systemic and local changes.
Improving clinical relevance and translatability
Recent studies on clinical samples from patients are gradually being undertaken, marking a promising start. However, combining single-cell data with clinical metrics, such as patient history, comorbidities, and lifestyle factors, and expanding studies to include diverse human populations, accounting for differences in age, sex, ethnicity, genetic backgrounds, and lifestyle factors, remains a significant challenge.
The ideal approach to address these challenges is to conduct a single comprehensive study that simultaneously includes all these variables, which would minimize batch effects and other sources of bias. However, considering that it is nearly impossible for a single team to achieve this, integrating high-quality data from different research teams might be a viable strategy to maximize resource utilization. Given the current limited number of multivariable studies, it is essential to first conduct single-cell research that accounts for biological variables and includes long-term studies.
Facilitating single-cell transcriptomics in spatially intact tissues
Spatial information is crucial for unraveling the complexities of tissue architecture, cell signaling pathways, and developmental processes, particularly in diffuse injuries such as blast TBI. Before the widespread adoption of spatial transcriptomics, researchers primarily used several methods to obtain spatial information: inferring single-cell spatial location through algorithms, 142 and segmenting different anatomical regions (e.g., cortex, hippocampus, peri-lesion, core) for single-cell sequencing. Currently, there are two main types of spatial transcriptomics: sequencing-based methods and imaging-based methods. Sequencing-based methods include Visium (10× Genomics) etc., while imaging-based approaches include Xenium (10× Genomics), CosMx (NanoString), and MERSCOPE (Vizgen). To date, Visium is the most used technology to collect spatial information in focal brain injury.55,58,66,121 However, due to Visium’s resolution of 55 µm, each spot contains 1–10 cells (Figure 3(a)). Therefore, it often requires complementary scRNA-seq data and several computational approaches, such as cell2location, 143 CARD, 144 and SpatialScope, 145 to infer the cellular composition of each spot (Supplementary file). These methods allow researchers to link cellular phenotypes with anatomical locations in the tissue microenvironment, offering many insights into tissue architecture and pathophysiology.
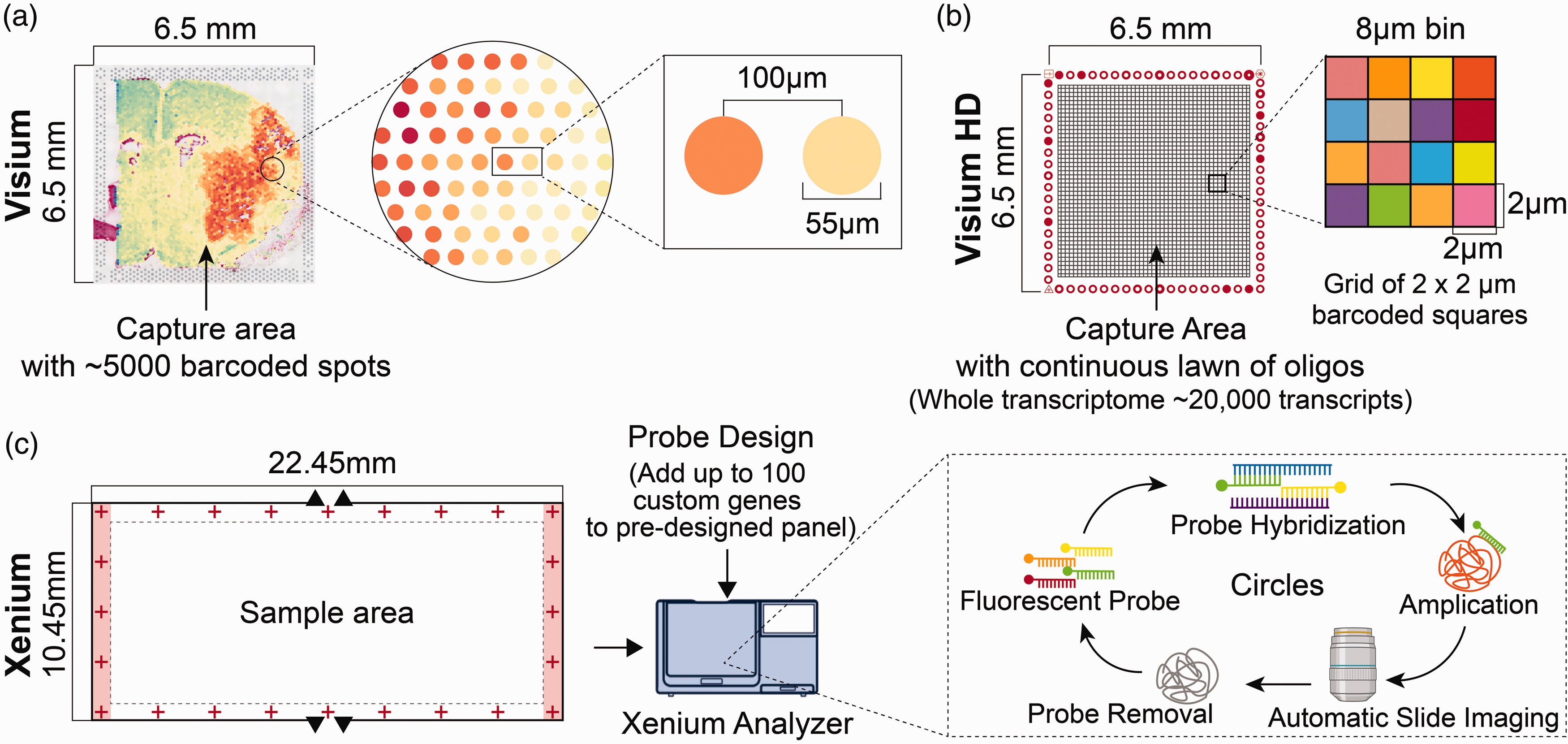
Illustration of current spatial transcriptomics tools. (a) Visium v2 is the most widely used technology to collect spatial information in focal brain injury. However, Visium’s resolution is 55 µm with a 45 μm gap between adjacent spots, each spot contains 1–10 cells. (b) The newly released 10× Genomics Visium HD achieves a resolution of 2 µm without gaps. 8 μm x 8 μm bin is a recommended starting point for data analysis. Visium HD spatial gene expression platform enables whole genome transcriptome analysis at the single-cell level with spatial context. It is a powerful tool for discovery-based research and generating hypotheses and (c) Xenium for in situ analysis was first introduced by 10× Genomics in 2022. An updated version, Xenium Prime 5 K Mouse Pan Tissue & Pathways Panel was released in 2024, with a higher throughput than the first generation, containing 5000 genes and allowing up to 100 more custom genes. Xenium is an efficient tool for targeted analysis and focused insights attributed to the availability of customized probes.
The field of stroke and TBI research increasingly demands higher-resolution spatial transcriptomics technologies. In 2024, 10× Genomics launched Visium HD, achieving a resolution of 2 µm without gaps (Figure 3(b)). The 10× Genomics Visium HD Spatial Gene Expression platform enables whole genome transcriptome analysis at the single-cell level of resolution with spatial context on formalin-fixed, paraffin-embedded (FFPE) tissue sections. Additionally, 10× Genomics introduced Xenium for in situ analysis as early as 2022. 23 Xenium offers single-cell spatial imaging across whole tissue sections with exceptional specificity, sensitivity, and speed (Figure 3(c)). Initially, it provides a 247-gene panel for the neuroscience field, allowing researchers to add up to 100 custom genes or use entirely custom panels (up to 480 genes), significantly increasing throughput. An updated version, Xenium Prime 5 K Mouse Pan Tissue & Pathways Panel has been released by 10× Genomics in 2024, containing 5006 genes and allowing up to 100 custom genes, with a higher throughput than the first generation. (Figure 3) We have recently applied the newly released, powerful Xenium In Situ platform to TBI brain injury, using the 247-gene mouse brain probe panel, as shown in representative images (Figure 4(a)–(c)). These results confirm the distinctive expression patterns of Wfs1, Slits, and Neurod6 in hippocampal pyramidal neurons, as reported recently. 146 Immunofluorescent staining of cell markers such as NeuN on post-Xenium slides can also be leveraged to identify the expression of target genes in specific cell types (Figure 4(d)). Considering the potential whole genome spectrum of Visium HD and the spatial precision of Xenium, particularly with cell segmentation enhancing resolution at the cellular and subcellular levels to visualize cell outlines or intracellular organelles, the two technologies can be used complementarily to better address the relevant research questions. A fair comparison of transcript detection efficiency between the two platforms may be of interest. This will involve comparing the number of transcripts (read counts) of a particular gene by Xenium to those captured by Visium HD on a post-Xenium slide. Other commercial platforms, such as CosMx from NanoString 22 and MERSCOPE from Vizgen, also enable single-cell level transcriptomics analysis with intact spatial information. Comprehensive comparisons have concluded that Xenium outperforms CosMx in sensitivity, accuracy, reproducibility, and spatial resolution, while MERSCOPE demonstrated the best performance on a large panel of 483 genes, given the lower rates of incorrect molecular assignments.147–149 In the future, an ideal strategy may involve sequencing-based methods that provide non-targeted capture of transcriptomes, while imaging-based approaches could offer high-throughput, accurate, and sensitive validation. Recently, spatial transcriptomics advanced further with the development of Open-ST by Schott et al., offering high-resolution spatial transcriptomics in three dimensions. 150 This technology is expected to be applied to stroke and TBI research in the future.
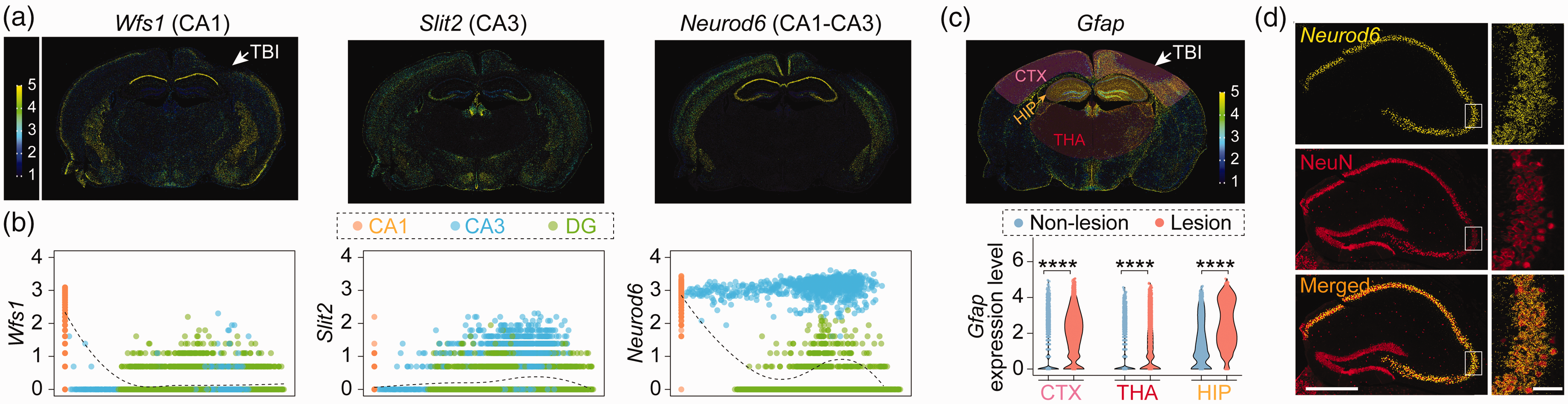
Spatial transcriptomics visualization and analysis of gene expression in TBI mouse brain using 10X Genomics Xenium. (a) Representative Xenium images showing spatial expression of Wfs1, Slit2, and Neurod6 at 3 days after TBI; higher expression is shown in yellow and lower expression is shown in blue. (b) Quantification of Wfs1, Slit2, and Neurod6 gene expression in hippocampus. (c) Representative Xenium image showing Gfap expression 3 days after TBI; higher expression is shown in yellow and lower expression is shown in blue (top panel). The bottom panel is the quantification of Gfap expression in three labeled ROIs, including CTX (pink), hippocampus (orange), and thalamus (red) from both lesion and non-lesion hemispheres. ****p ≤ 0.0001 vs. indicated group analyzed by Wilcoxon rank-sum test and (d) Post-Xenium immunofluorescent staining of NeuN. Representative images showing the expression of Neurod6 (yellow) in Xenium and NeuN (red) in the lesioned hemisphere 3 days after TBI. Merged image shows the co-localization of Neurod6 and NeuN in the hippocampus. Scale bar = 500 μm. The right column shows the enlarged images of ROI in CA3 (white box). Scale bar = 50 μm.
Superior integration with other omics methods
The combination of transcriptomic data with proteomic, metabolomic, or chromatin accessibility data is a valuable multi-omics approach. In the fields of stroke and TBI, integration with proteomic data is particularly important. Our research has shown that stroke involves persistent protein synthesis inhibition, suggesting that changes at the RNA level may not accurately reflect protein levels. 151 CITE-seq 152 and REAP-seq 153 enable the integration of transcriptomic data with proteomic data to improve our understanding of downstream biological responses. For example, studies have demonstrated the potential of CITE-seq in differentiating ambiguous NK cell subgroups in COVID-19 samples. 154 Additionally, single-cell ATAC-seq also facilitates investigations of the association between transcriptional and epigenetic variations.155,156
Conclusion
Single-cell technologies continue to evolve, with spatial transcriptomics gradually being integrated into the study of stroke and TBI. The breadth and depth of single-cell studies have increased with incorporation of key biological variables (e.g., age, sex) and the extension of observations beyond the acute injury phase to longer timepoints. Additionally, single-cell research is no longer limited to cellular heterogeneities and functional changes caused by diseases. Additional emphasis is now placed on understanding cellular diversity and functions in healthy populations. The launch of the large cell atlas has significantly enhanced the basic understanding of major cell functions across various tissues, with novel targets available specifically for the brain. 157 This approach also extends to exploring cellular fate, gene/drug effects, and other factors, thereby driving forward our understanding of cellular heterogeneity and functional diversity.
Future single-cell research is advancing into the era of spatial multi-omics, with improvements in spatial transcriptomics technologies, including higher resolution, sensitivity, and throughput, as well as the integration of omics data from genomics, proteomics, metabolomics, etc. Vandereyken et al. have summarized a detailed list of single-cell modalities, including genomic, transcriptomic, epigenomic, and proteomic information. 158 These advancing technologies hold great promise for the field of brain injury. For example, scRNA-seq is poised to pinpoint specific cell types and their roles in neurological disorders, while spatial transcriptomics adds another dimension by mapping gene expression patterns within the spatial context of brain tissue, which is crucial for understanding complex brain architecture and the interplay among various cell types. Multi-omics approaches will further provide a comprehensive view of the molecular mechanisms underlying brain function and pathology.
Before fully incorporating all types of multi-omics data, researchers are proceeding within a three-dimensional framework of spatial-cellular-molecular integration. This framework can be further validated through mechanistic in vivo and in vitro experiments, facilitating a comprehensive portrait of diverse cellular processes perturbed by stroke and TBI in both the brain and peripheral tissues. For instance, a recent study introduced a multi-omics platform integrating scRNA-seq, spatial transcriptomics, and proteomics, thereby advancing the resolution of our knowledge on gene expression in astrocytes in stroke injury and revealing their potential roles in stroke recovery. 66 By incorporating additional omics perspectives and considering variables such as sex, neurodevelopmental stage, age, brain region, and time course, novel biomarkers and therapeutic targets may be identified for stroke or TBI, and ultimately provide new therapeutic opportunities in the clinic.
Supplemental Material
sj-pdf-1-jcb-10.1177_0271678X241305914 - Supplemental material for Single-cell RNA sequencing in stroke and traumatic brain injury: Current achievements, challenges, and future perspectives on transcriptomic profiling
Supplemental material, sj-pdf-1-jcb-10.1177_0271678X241305914 for Single-cell RNA sequencing in stroke and traumatic brain injury: Current achievements, challenges, and future perspectives on transcriptomic profiling by Ruyu Shi, Huaijun Chen, Wenting Zhang, Rehana K Leak, Dequan Lou, Kong Chen and Jun Chen in Journal of Cerebral Blood Flow & Metabolism
Footnotes
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: J.C. is the Richard King Mellon Professor of Neurology and a recipient of a VA Senior Research Career Scientist Award and is supported by the US Department of Veterans Affairs Merit Review I01BX002495 and I01BX003377 and NIH grant NS124673. RKL is grateful for support by R15 NS130532-01 and 1R21 NS141002-01 from NINDS and the School of Pharmacy at Duquesne University. We thank Patricia Strickler for administrative support. BioRender was used to create figures.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplementary material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
